Abstract
Purpose
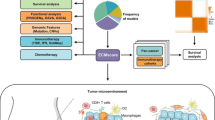
With the heterogeneous genetic background, prognosis prediction and therapeutic targets for testicular germ cell tumors (TGCTs) are still unclear. We defined the tumor immune microenvironment activation status (TIMEAS).
Methods
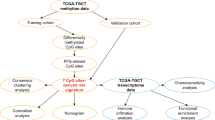
We collected a total of 314 TGCT patients from four cohorts, including a 48-case microarray. A nonnegative matrix factorization algorithm was applied to identify the “immune factor”, derived the top 150 weighted genes to divide patients into immune and non-immune classes, and further separated the immune class into activated and exhausted subgroups by nearest template prediction. Tumor mutant burden, gene mutation, and copy number alteration were compared with our recently developed package “MOVICS”. A random forest algorithm was performed to establish a prediction model with fewer genes. Immunohistochemistry staining was performed to identify TIMEAS in the microarray.
Results
We constructed the TIMEAS in the TCGA-TGCT cohort and further validated it in the GSE3218 and GSE99420 cohorts. The immune class contained the activated status of T-lymphocytes, B-lymphocytes, and macrophages, while Treg cells and the WNT/TGFβ signature were more activated in the immune-suppressed subgroup. Patients in the immune-exhausted subgroup had the worst prognosis, and 22.9% of patients in the immune-activated subgroup had KRAS mutations, which might stimulate the response of the immune system and lead to a favorable prognosis. The immune-exhausted group benefited more from chemotherapy, while the immune-activated subgroup responded well to anti-PD-1/PD-L1 therapy. FSCN1 was validated as the target of the immune-exhausted microenvironment by immunohistochemistry.
Conclusion
TIMEAS classification can separate TGCT patients; patients in the immune-activated subgroup could benefit more from anti-PD-L1 immunotherapy, and those in the immune-exhausted subgroup are more suitable for chemotherapy.







Similar content being viewed by others
Data availability
The raw data for this study were generated at the corresponding archives. Derived data supporting the findings are available from the corresponding author [LCZ] upon reasonable request.
References
K.A. McGlynn, M.B. Cook, Future Oncol. 5, 1389–1402 (2009). https://doi.org/10.2217/fon.09.116
A.A. Ghazarian, B. Trabert, S.S. Devesa, K.A. McGlynn, Andrology 3, 13–18 (2015). https://doi.org/10.1111/andr.288
P. Carrière, P. Baade, L. Fritschi, J. Urol. 178, 125–128 (2007)
J. Ferlay, I. Soerjomataram, R. Dikshit, S. Eser, C. Mathers, M. Rebelo, D.M. Parkin, D. Forman, F. Bray, Int. J. Cancer 136, E359–E386 (2015). https://doi.org/10.1002/ijc.29210
O. Khan, A. Protheroe, Postgrad. Med. J. 83, 624–632 (2007)
H.J. Schmoll, R. Souchon, S. Krege, P. Albers, J. Beyer, C. Kollmannsberger, S.D. Fossa, N.E. Skakkebaek, R. de Wit, K. Fizazi, J.P. Droz, G. Pizzocaro, G. Daugaard, P.H.M. de Mulder, A. Horwich, T. Oliver, R. Huddart, G. Rosti, L. Paz Ares, O. Pont, J.T. Hartmann, N. Aass, F. Algaba, M. Bamberg, I. Bodrogi, C. Bokemeyer, J. Classen, S. Clemm, S. Culine, M. de Wit, H.G. Derigs, K.P. Dieckmann, M. Flasshove, X. Garcia del Muro, A. Gerl, J.R. Germa-Lluch, M. Hartmann, A. Heidenreich, W. Hoeltl, J. Joffe, W. Jones, G. Kaiser, O. Klepp, S. Kliesch, L. Kisbenedek, K.U. Koehrmann, M. Kuczyk, M.P. Laguna, O. Leiva, V. Loy, M.D. Mason, G.M. Mead, R.P. Mueller, N. Nicolai, G.O.N. Oosterhof, T. Pottek, O. Rick, H. Schmidberger, F. Sedlmayer, W. Siegert, U. Studer, S. Tjulandin, Ann. Oncol. 15, 1377–1399 (2004) von der Maase, P. Walz, S. Weinknecht, L. Weissbach, E. Winter and C. Wittekind
F. Bremmer, C.L. Behnes, S. Schweyer, Pathologe 35, 238–244 (2014). https://doi.org/10.1007/s00292-014-1900-8
V. Seetharam, Z.B.M. Hameed, S.B. Talengala, J. Thomas, BMJ Case Rep. 2014 (2014) https://doi.org/10.1136/bcr-2013-203085
P. Chieffi, Intractable Rare Dis Res 6, 319–321 (2017). https://doi.org/10.5582/irdr.2017.01070
P. Chieffi, Mini Rev. Med. Chem. 11, 1075–1081 (2011)
J.S. Parker, M. Mullins, M.C.U. Cheang, S. Leung, D. Voduc, T. Vickery, S. Davies, C. Fauron, X. He, Z. Hu, J.F. Quackenbush, I.J. Stijleman, J. Palazzo, J.S. Marron, A.B. Nobel, E. Mardis, T.O. Nielsen, M.J. Ellis, C.M. Perou, P.S. Bernard, J. Clin. Oncol. 27, 1160–1167 (2009). https://doi.org/10.1200/JCO.2008.18.1370
W. Choi, S. Porten, S. Kim, D. Willis, E.R. Plimack, J. Hoffman-Censits, B. Roth, T. Cheng, M. Tran, I.L. Lee, J. Melquist, J. Bondaruk, T. Majewski, S. Zhang, S. Pretzsch, K. Baggerly, A. Siefker-Radtke, B. Czerniak, C.P.N. Dinney, D.J. McConkey, Cancer Cell. 25, 152–165 (2014). https://doi.org/10.1016/j.ccr.2014.01.009
A. Batool, N. Karimi, X.-N. Wu, S.-R. Chen, Y.-X. Liu, Cell. Mol. Life Sci. 76, 1713–1727 (2019). https://doi.org/10.1007/s00018-019-03022-7
D.D. Lee, H.S. Seung, Nature 401, 788–791 (1999)
R.A. Moffitt, R. Marayati, E.L. Flate, K.E. Volmar, S.G.H. Loeza, K.A. Hoadley, N.U. Rashid, L.A. Williams, S.C. Eaton, A.H. Chung, J.K. Smyla, J.M. Anderson, H.J. Kim, D.J. Bentrem, M.S. Talamonti, C.A. Iacobuzio-Donahue, M.A. Hollingsworth, J.J. Yeh, Nat. Genet. 47, 1168–1178 (2015). https://doi.org/10.1038/ng.3398
J. Meng, X. Lu, Y. Zhou, M. Zhang, Q. Ge, J. Zhou, Z. Hao, S. Gao, F. Yan, C. Liang, Mol. Ther. Oncolytics 20, 410–421 (2021). https://doi.org/10.1016/j.omto.2021.02.001
K. Yoshihara, M. Shahmoradgoli, E. Martínez, R. Vegesna, H. Kim, W. Torres-Garcia, V. Treviño, H. Shen, P.W. Laird, D.A. Levine, S.L. Carter, G. Getz, K. Stemke-Hale, G.B. Mills, R.G.W. Verhaak, Nat. Commun. 4, 2612 (2013). https://doi.org/10.1038/ncomms3612
X. Lu, J. Meng, Y. Zhou, L. Jiang, F. Yan, Bioinformatics (2020) https://doi.org/10.1093/bioinformatics/btaa1018
C.H. Mermel, S.E. Schumacher, B. Hill, M.L. Meyerson, R. Beroukhim, G. Getz, Genome Biol 12, R41 (2011) https://doi.org/10.1186/gb-2011-12-4-r41
Y. Yin, L. Xu, Y. Chang, T. Zeng, X. Chen, A. Wang, J. Groth, W.-C. Foo, C. Liang, H. Hu, J. Huang, Mol. Cancer 18, 11 (2019) https://doi.org/10.1186/s12943-019-0941-2
J. Chen, C. Zhan, L. Zhang, L. Zhang, Y. Liu, Y. Zhang, H. Du, C. Liang, X. Chen, Inflammation 42, 1705–1718 (2019). https://doi.org/10.1007/s10753-019-01030-0
A. Paschalis, B. Sheehan, R. Riisnaes, D.N. Rodrigues, B. Gurel, C. Bertan, A. Ferreira, M.B.K. Lambros, G. Seed, W. Yuan, D. Dolling, J.C. Welti, A. Neeb, S. Sumanasuriya, P. Rescigno, D. Bianchini, N. Tunariu, S. Carreira, A. Sharp, W. Oyen, J.S. de Bono, Eur. Urol. 76, 469–478 (2019). https://doi.org/10.1016/j.eururo.2019.06.030
M. Reich, T. Liefeld, J. Gould, J. Lerner, P. Tamayo, J.P. Mesirov, Nat. Genet. 38, 500–501 (2006)
J. Meng, Y. Zhou, X. Lu, Z. Bian, Y. Chen, J. Zhou, L. Zhang, Z. Hao, M. Zhang, C. Liang, Mol. Oncol. 15, 1358–1375 (2021). https://doi.org/10.1002/1878-0261.12887
K.N. Sivanathan, S. Gronthos, D. Rojas-Canales, B. Thierry, P.T. Coates, Stem Cell. Rev. Rep. 10, 351–375 (2014). https://doi.org/10.1007/s12015-014-9495-2
E. Batlle, J. Massagué, Immunity 50, 924–940 (2019). https://doi.org/10.1016/j.immuni.2019.03.024
Y. Dong, X. Li, L. Zhang, Q. Zhu, C. Chen, J. Bao, Y. Chen, BMC Immunol. 20, 27 (2019). https://doi.org/10.1186/s12865-019-0309-9
J.-F. Liu, L. Wu, L.-L. Yang, W.-W. Deng, L. Mao, H. Wu, W.-F. Zhang, Z.-J. Sun, J. Exp. Clin. Cancer Res. 37, 44 (2018). https://doi.org/10.1186/s13046-018-0713-7
B.A. Youngblood, J. Leung, R. Falahati, J. Williams, J. Schanin, E.C. Brock, B. Singh, A.T. Chang, J.A. O’Sullivan, R.P. Schleimer, N. Tomasevic, C.R. Bebbington, B.S. Bochner, Cells 10 (2020) https://doi.org/10.3390/cells10010019
E. Rajpert-De, Meyts, Andrology 7, 391–393 (2019). https://doi.org/10.1111/andr.12675
P. Chieffi, M. De Martino, F. Esposito, Expert Rev. Anticancer Ther. 20, 189–195 (2020). https://doi.org/10.1080/14737140.2020.1736566
J. Houldsworth, J.E. Korkola, G.J. Bosl, R.S.K. Chaganti, J. Clin. Oncol. 24, 5512–5518 (2006)
T. Fukawa, H.-O. Kanayama, Int. J. Urol. 25, 337–344 (2018). https://doi.org/10.1111/iju.13519
P.J. Siska, R.A.N. Johnpulle, A. Zhou, J. Bordeaux, J.Y. Kim, B. Dabbas, N. Dakappagari, J.C. Rathmell, W.K. Rathmell, A.K. Morgans, J.M. Balko, D.B. Johnson, Oncoimmunology 6, e1305535 (2017) https://doi.org/10.1080/2162402X.2017.1305535
N. Yu, M.-J. Wu, J.-X. Liu, C.-H. Zheng, Y. Xu, IEEE Trans. Cybern 51, 3952–3963 (2021). https://doi.org/10.1109/TCYB.2020.3000799
J. Pan, N. Gillis, IEEE Trans. Pattern Anal. Mach. Intell. 43, 1546–1561 (2021). https://doi.org/10.1109/TPAMI.2019.2956046
Y. Hoshida, PLoS One 5, e15543 (2010). https://doi.org/10.1371/journal.pone.0015543
G. Bindea, B. Mlecnik, M. Tosolini, A. Kirilovsky, M. Waldner, A.C. Obenauf, H. Angell, T. Fredriksen, L. Lafontaine, A. Berger, P. Bruneval, W.H. Fridman, C. Becker, F. Pagès, M.R. Speicher, Z. Trajanoski, J. Galon, Immunity 39, 782–795 (2013). https://doi.org/10.1016/j.immuni.2013.10.003
T. Strohmeyer, S. Peter, M. Hartmann, S. Munemitsu, R. Ackermann, A. Ullrich, D.J. Slamon, Cancer Res. 51, 1811–1816 (1991)
S.L. Woldu, J.F. Amatruda, A. Bagrodia, Curr. Opin. Urol. 27, 41–47 (2017)
M. Malumbres, M. Barbacid, Nat. Rev. Cancer 3, 459–465 (2003). https://doi.org/10.1038/nrc1097
C.M. Kelly, L. Gutierrez Sainz, P. Chi, J. Hematol. Oncol. 14, 2 (2021) https://doi.org/10.1186/s13045-020-01026-6
H. Joensuu, E. Wardelmann, H. Sihto, M. Eriksson, K. Sundby Hall, A. Reichardt, J.T. Hartmann, D. Pink, S. Cameron, P. Hohenberger, S.-E. Al-Batran, M. Schlemmer, S. Bauer, B. Nilsson, R. Kallio, J. Junnila, A. Vehtari, P. Reichardt, JAMA Oncol. 3, 602–609 (2017). https://doi.org/10.1001/jamaoncol.2016.5751
F.E. von Eyben, J. Parraga-Alava, Int. J. Mol. Sci. 21 (2020) https://doi.org/10.3390/ijms21124487
G. Bignell, R. Smith, C. Hunter, P. Stephens, H. Davies, C. Greenman, J. Teague, A. Butler, S. Edkins, C. Stevens, S. O’Meara, A. Parker, T. Avis, S. Barthorpe, L. Brackenbury, G. Buck, J. Clements, J. Cole, E. Dicks, K. Edwards, S. Forbes, M. Gorton, K. Gray, K. Halliday, R. Harrison, K. Hills, J. Hinton, D. Jones, V. Kosmidou, R. Laman, R. Lugg, A. Menzies, J. Perry, R. Petty, K. Raine, R. Shepherd, A. Small, H. Solomon, Y. Stephens, C. Tofts, J. Varian, A. Webb, S. West, S. Widaa, A. Yates, A.J.M. Gillis, H.J. Stoop, R.J.H.L.M. van Gurp, J.W. Oosterhuis, L.H.J. P.A. Looijenga, Futreal, R. Wooster and M.R. Stratton, Genes Chromosomes Cancer 45, 42–46 (2006)
M. Yeste-Velasco, T. Guo, X. Mao, E. Stankiewicz, G. Scandura, H. Li, C.S. Wang, S. Kudahetti, T. Oliver, D. Berney, J. Shamash, Y.-J. Lu, Am. J. Cancer Res. 9, 855–871 (2019)
R. Nakayama, J.P. Jagannathan, N. Ramaiya, M.L. Ferrone, C.P. Raut, J.E. Ready, J.L. Hornick, A.J. Wagner, BMC Cancer 18, 1296 (2018). https://doi.org/10.1186/s12885-018-5188-6
Y. Chen, T. Tian, Z.-Y. Li, C.-Y. Wang, R. Deng, W.-Y. Deng, A.-K. Yang, Y.-F. Chen, H. Li, Cell. Death Dis. 10, 356 (2019). https://doi.org/10.1038/s41419-019-1574-5
M. Zhang, Z. Zhao, X. Duan, P. Chen, Z. Peng, H. Qiu, J. Cell. Physiol. 233, 4748–4758 (2018). https://doi.org/10.1002/jcp.26264
C.-Q. Wang, C.-H. Tang, Y. Wang, L. Jin, Q. Wang, X. Li, G.-N. Hu, B.-F. Huang, Y.-M. Zhao, C.-M. Su, Sci. Rep. 7, 15887 (2017). https://doi.org/10.1038/s41598-017-16196-6
Acknowledgements
We greatly appreciate the patients and investigators who participated in the corresponding medical project for providing data.
Funding
This work was supported by the National Natural Science Foundation of China [grant numbers: 82170787, 82071637, 81973145]; the Supporting Project for Distinguished Young Scholars of Anhui Colleges [grant number: gxyqZD2019018]; the National Key R&D Program of China (2019YFC1711000), and the Key R&D Program of Jiangsu Province [Social Development] (BE2020694).
Author information
Authors and Affiliations
Contributions
Conceptualization, MJL, GJJ and LCZ; methodology, MJL, GR, LXF, and ZXS; formal analysis, MJL, LXF, YFR and WHT; writing the original draft, LX, LY and HZY; visualization, MJL, LX, LXF and WHT; funding acquisition, HZY, ZXS and LCZ; supervision, LCZ and ZXS.
Corresponding authors
Ethics declarations
Ethics approval and consent to participate
Ethical approval for the microarray was obtained from the Ethics Committee of the First Affiliated Hospital of Anhui Medical University (PJ-2022-06-36), and patient consent for the retrospective cohorts was waived. As the other data used in this study are publicly available, no ethical approval was needed.
Consent for publication
Not applicable.
Competing interests
The authors declare no competing interests.
Conflicts of interest
The authors have no conflicts of interest.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Below is the link to the electronic supplementary material.
ESM 1
(DOCX 636 KB)
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Meng, J., Gao, J., Li, X. et al. TIMEAS, a promising method for the stratification of testicular germ cell tumor patients with distinct immune microenvironment, clinical outcome and sensitivity to frontline therapies. Cell Oncol. 46, 745–759 (2023). https://doi.org/10.1007/s13402-023-00781-1
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13402-023-00781-1