Abstract
The agro-industrial sector generates diverse waste that requires effective handling to mitigate economic loss and pollution. Consequently, this study assessed bacterial isolates from poultry dumpsites for feather decomposition and keratinase production capacity. The protein hydrolysates generated from feather dismemberment by two bacteria identified as Chryseobacterium proteolyticum FGNn (accession no. MW165478) and Pseudomonas aeruginosa GNFx (accession no. MW165479) were profiled for amino acids composition. The biochemical properties of the bacteria-associated keratinases were determined. Initial keratinase production (with percentage feather degradation), demonstrated by the isolates, was 693.63 ± 62.99 U/mL (81%) and 619.09 ± 37.28 (76%) against FGNn and GNFx, respectively. At optimized process conditions, C. proteolyticum and P. aeruginosa extracellular keratinase production was 1756.36 ± 2.57 U/mL at 72 h and 2055.45 ± 50.14 U/mL at 96 h, respectively. Analysis of the feather hydrolysates showed a relatively high abundance of arginine (3.18%) and glycine (3.26%) for FGNn and glutamic acid (6.05%), serine (3.31%), aspartic acid (4.74%), glycine (3.71%), alanine (3.43%), valine (3.09%), and leucine (3.23%) for GNFx. The keratinolytic proteases showed pH and temperature optima of 8.0 and 50°C against FGNn, and 8.0 and 60°C against GNFx. GNFx keratinase was thermostable, displaying a half-life time of more than 60 min at 80°C. In addition, GNFx keratinolytic enzyme was chemical agent tolerant post-treatment. The findings underlined the significance of C. proteolyticum FGNn and P. aeruginosa GNFx as suitable in the valorisation of keratinous biomass. Also, the robust stability profile of GNFx keratinase highlights its prospects in green technology.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
The growth trajectory in the agricultural sector has been steeply upward and that translates into more food and food-associated products, which are crucial to sustaining the teeming global population [1]. Consequently, huge byproducts emanate from agro-processing and, some waste biomass have the potential for reused as feedstock in the innovation value chain in the bioeconomy. Considering that agro-wastes come in different forms, such as chitinous, lignocellulosic, and keratinous biomass [2], the molecular compositions and interactive forces in the agro-residues determine their degree of structural stability. Amongst these renewable resources, keratinous biomass is predominantly generated from livestock and poultry processing, of which avian feathers are the most abundant. Proper handling of feather waste incurs additional costs due to their exceptional structural stability orchestrated by high keratin content (90%); hence, most agro-industrialists dispose of the waste materials through landfilling and incineration [3].
Considering the protein content of feathers, the disposal approach is economically inefficient and pose environmental concerns including the increase on the carbon footprint and pollution associated problems [4]. To harness the economic values of the large keratinous feather biomass estimated at 102 million tonnes in 2021 [5], several techniques have been employed for the conversion of the waste biomass into high-value products [6]. In the same vein, keratin extraction using acid/base or hydrophobic ionic liquid hydrolysis yielded products considered crucial for application in green chemistry, pharmaceutical, biomedical, and cosmetic industries [7, 8]. Another method of feather recycling involves the thermo-mechanical conversion of feathers into feather meals with the feathers serving as protein supplements in feeds [9]. These value-addition approaches are not ecologically and economically sustainable due to the environmental concerns posed by chemical agents and high energy requirements, respectively [6]. In addition to intense energy involvement in thermo-mechanical processing, the feather meals produced are unsuitable for animal consumption because of the loss of heat-labile nutrients and poor digestibility of the bio-product in ruminants and monogastrics [3].
The efficient valorization of keratinous feathers into high quality products with versatile applicability has been successfully implemented via microbial and enzyme technology. Some bacteria within the class Gammaproteobacteria, Flavobacteriia, Actinomycetia, Bacilli, and Sphingobacteriia have demonstrated an excellent metabolic diversity with high expression of keratinolytic systems which directs a complete dismemberment of keratin. The extracellularly secreted keratinolytic enzymes dismember keratinous polymer by altering the various inherent linkages such as hydrophobic interactions, hydrogen, disulfide, and peptide bonds that stabilized the biopolymer [10]. Keratinolytic bacteria inductively secrete keratinases and other associated metabolites in response to the presence of keratin-rich biomass as main substrates for carbon and energy requirements [11]. Biodegradation of keratinous agro-residues produces hydrolysates rich in bioactive peptides, amino acids, plant growth factors, and non-protein nitrogenous compounds [10, 12,13,14]. The mechanistic approach for keratin degradation may vary from one bacterium to another; hence, it would be expedient to allude that the mechanism is not fully elucidated; however, synergistic actions of disulfide reductase and keratinolytic peptidase have also been advanced as one of the pathways of keratin decomposition into simpler protein units [15]. Gram-positive bacterial isolates have been extensively studied for their role in keratin degradation, with Bacillus spp. as the lead degrader of recalcitrant keratin biomass.
However, Gram-negative bacteria have been reported less compared to Gram-positive isolates; hence an emerging research hotspot for exploring potentially robust keratinase producers with efficient biosynthetic gene clusters. Gram-negative keratinolytic isolates have been implicated in colonizing many ecological niches due to their metabolic diversity [16], suggesting their plasticity to degrading and utilizing various recalcitrant biomass for their nutritional requirements. Another interesting property of Gram-negative bacteria, including Pseudomonas sp., Chryseobacterium sp., and Stenotrophomonas sp., among others, is their ability to demonstrate optimal keratinolytic activity at mesophilic conditions, making the bioprocess cost-effective from an economic perspective [17,18,19]. Keratinolytic bacteria isolation from waste dumpsites has been identified to have a stronger impact on biomass degradation owing to improved microbial acclimatization and colonization of the substrates that subsequently induce the biosynthesis and secretion of necessary metabolites to mediate polymer decomposition. Therefore, this study assessed the keratinolytic potential of bacterial isolates indigenous to a poultry waste dump site. The two most potent isolates were identified as Gram-negative bacteria – Pseudomonas aeruginosa and Chryseobacterium proteolyticum. Selected important process conditions influencing enzyme production by bacterial isolates were optimized while maintaining a cost-effective production medium. The protein hydrolysates generated through chicken feather disintegration by both isolates were analyzed for various amino acids. In addition, the properties and laundry detergent tolerance of the extracellularly produced bacterial keratinases were determined.
2 Materials and methods
2.1 Sample collection
Chicken feathers and soil samples were sourced from a local poultry slaughter facility located in the Eastern Cape (geographic coordinates 32.7450° S, 27.0268° E), Republic of South Africa. Poultry feathers and soil samples were collected in sterile sampling bags and then transported to the laboratory in a cooler box for processing within six hours of collection.
2.2 Keratinous substrate processing
At the laboratory, blood stains and other debris on the chicken feathers were removed by several rinsing with tap water. The feathers were air-dried and further dried in an oven set at 60°C for 48 h. Some portions of the dried feathers were milled into 2 mm particle size powder and stored under room temperature using sterile containers as previously reported [18].
2.3 Isolation of bacterial strains
The bacterial strains were isolated from the soil samples using minimal salt media (MSM) supplemented with chicken feathers [20]. MSM that comprised (g/L): K2HPO4, 0.3; KH2PO4, 0.4; MgCl2, 0.2; CaCl2, 0.22; NH4Cl, 0.5 (Merck, South Africa); milled chicken feather (MCF), 10 and pH 6.5 was prepared and dispensed in 250 mL Erlenmeyer flasks containing 50 mL working volume. The flasks were autoclaved at 121°C, 15 psi, and soil samples (2 g) were added into the sterile media. The flasks were incubated at 30°C for 5 days in a rotary shaker (Labotec, South Africa). A serial dilution of the samples was prepared post-incubation; about 10 μL of 10-5 dilution was inoculated on MCF agar plates by spread plate method. MCF agar plates comprised (g/L): K2HPO4, 0.3; KH2PO4, 0.4; MgCl2, 0.2; CaCl2, 0.22; MCF, 10 and bacteriological agar, 15 (Merck, South Africa). Also, 80 mg/L of nystatin (Merck, South Africa) was aseptically added to the media before pouring it into Petri dishes to prevent the potential growth of fungi. The inoculated plates were incubated at 30°C for 48 h and then monitored for observable bacterial colony growth. Distinct colonies were purified by sub-culturing on new MCF agar plates. Axenic colonies were kept on MCF agar slant at 4°C for subsequent inoculum preparation and in 25% glycerol at -80°C for long-term storage.
2.4 Inoculum preparation and bacteria screening for proteolytic potential
Axenic bacterial colonies grown on MCF agar plates were scooped with wire loop into micro-centrifuge tubes that contained 1-mL sterile saline. The bacterial suspensions’ optical density was standardized to read 0.1 at 600 nm. An aliquot of each standardized bacterial suspension (5 μL) was placed onto the skimmed milk agar. The plates were placed in an incubator set at 30°C, and the incubation was done for 48 h. The minimal salts, agar bacteriological (15 g/L) and skimmed milk (10 g/L) were all constituents of skimmed milk agar. Bacterial isolates that showed halo zones on skimmed milk agar plates were selected as positive proteolytic strains, and the measurement of the zones was taken in millimetre (mm).
2.5 Evaluation of the proteolytic bacteria for keratinolytic activity
Fresh inocula of proteolytic bacterial isolates were prepared and used to ferment chicken feathers in 100 mL bottles. In each bottle with 20 mL MSM, poultry feathers (10 g/L) were included, then sterilized by autoclaving. The pH of the medium was determined and subsequently adjusted to 6.0 in an aseptic condition. Each bottle was inoculated with 2% (v/v) of the freshly prepared starter culture and incubated for 72 h at 30°C and 150 rpm. The fermented broths were filtered with Whatman no. 1 filter paper to recover remnant feathers that were utilized to estimate the hydrolysis percentage [18]. The resulting filtrates were subjected to centrifugation (HERMLE Labortechnik GmbH, Wehingen, Germany) and the cell-free extracts were then used to study keratinase activity, medium pH drift, protein, and thiol concentrations.
2.6 Assay for keratinase activity
The enzyme assay was performed following the method described by Kuo et al. [21]. In microtubes, 10 g/L of keratin azure was added to 500 μL of 100 mM Tris-HCl buffer (pH 8). The crude enzyme (500 μL) was combined with the keratin azure solution, and the microtubes were incubated in a water bath set at 50°C. The reaction was stop after 1 h incubation by immersing the tubes in ice water. Subsequently, the tubes were centrifuged for 10 min at 12,500 rpm, and the absorbance (A595) of the resulting filtrates was determined using SYNERGYMX 96 well microtitre plate reader (BioTek, USA). A similar protocol was used for the control, except that a denatured enzyme solution was used instead. The enzyme’s amount raising the absorbance by 0.01 under the standard assay protocols was defined as one keratinase unit (KU).
2.7 Total protein and sulfhydryl (thiol) group quantification
Using bovine serum albumin (BSA) as a reference protein, the protocol developed by Bradford was used to determine the crude extract’s protein content [22], while the sulfhydryl group concentration in the extract samples was determined in accordance with Ellman’s method [23].
2.8 Identification of bacterial isolates with promising feather-degrading capacity
The Quick-DNA Fungal/Bacterial Miniprep Kit (Zymo Research Catalogue No D6005) was used to isolate the genomic DNA of the keratinolytic bacterial strains. OneTaq Quick-Load 2X Master Mix and a set of primers 27F- AGAGTTTGATCMTGGCTCAG and 1492R- CGGTTTACCTTGTTACGACTT were used to amplify the target 16S rRNA gene sequence of the isolates by polymerase chain reaction (PCR) under standard thermal cycling conditions [24]. The PCR products were analysed on 1% (w/v) agarose gel, and Zymoclean Gel DNA Recovery Kit (Zymo Research, Catalogue No. D44001) was used to recover the DNA fragments. The isolated fragments were bi-directionally sequenced (Nimagen, BrilliantDye Terminator Cycle Sequencing Kit V3.1, BRD3- 100/1000) and purified using Zymo Research, ZR-96 DNA Sequencing Clean-up Kit (Catalogue No. D4050). ABI 3500xl Genetic Analyser (Applied Biosystems, ThermoFisher Scientific) was used to sequence the pure DNA fragments. In order to access the .ab1 files generated by the ABI 3500XL Genetic Analyzer, CLC Bio Main Workbench version 7.6 was utilized, and the obtained sequences were compared with other reference sequences following the basic local alignment search tool hits on the National Centre for Biotechnology Information (NCBI) database to infer the evolutionary relationships.
2.9 Construction of the optimal fermentation variables for improved keratinase production
The effect of significant parameters, including inoculum size, chicken feather concentration, initial medium pH, and incubation temperature on keratinase production by bacterial isolates was studied using one-variable-at-a-time method. The parameters were varied: starter culture (1–7% v/v; at 1 unit interval), chicken feather (0.5–3.0% w/v; at 0.5 unit interval), initial medium pH (pH 3.0–9.0; at 1 unit interval) and incubation temperature (25–45°C; at 5 unit interval).
For the study of keratinase production kinetics, fermentation was performed at optimized process conditions. Then, 5 mL was withdrawn at 24 h intervals post-fermentation to ascertain keratinase titre, pH change, protein, and free sulfhydryl groups concentration. The Kinetics of bacterial cell growth was determined by the plate count method [25].
2.10 Feather hydrolysate preparation and amino acids analysis
The feather hydrolysates were prepared by filtering fermented medium through muslin. The filtrate was centrifuged for 15 min at 15,000 rpm before freeze-drying with a vacuum concentrator (Martin Christ Gefriertrocknungsanlagen GmbH, Germany). For the amino acid analysis, the methods used involved acid hydrolysis, pre-column derivatization, high-performance liquid chromatography (HPLC) separation, and fluorescence detection, as previously described [26].
2.11 Determination of optimal pH and temperature for keratinase activity
The pH and temperature optima for keratinase activity were determined with a crude extract from the bacterial fermentation without further purification. For pH, buffer solutions, including sodium citrate (pH 5), potassium phosphate (pH 6–7), Tris-HCl (pH 8–pH 9), and Glycine-NaOH (pH 10–11), were used. The temperature optima for keratinase activity were determined by carrying out the enzyme activity assay from 30 to 80°C (at 10°C intervals). The thermostability profile was further assessed by pre-heating the enzyme solution for 2 h at temperatures ranging from 50 to 80 (°C). Then, the residual keratinase activity was determined under the standard assay protocol. The keratinase activity of the crude extract evaluated without pre-heating served as the control and was set at 100%.
2.12 Effect of metal ions and other chemical agents on enzyme stability
To investigate how metal ions, such as Al3+, K+, Na+, Ba2+, Ca2+, Co2+, Cu2+, Fe2+, Mg2+, Fe2+, and Zn2+ affect the stability of the bacterial keratinases under investigation, the solutions of the various metal ions (5 mM final concentration) were pre-incubated with the enzymes for 1 h at 30°C. Subsequently, the remaining keratinase activity was then assessed under the established assay protocols. Similarly, the impact of ethylenediaminetetraacetic acid (EDTA), sodium dodecyl sulfate (SDS), 1,10-phenanthroline, phenylmethylsulfonyl fluoride (PMSF), and dithiothreitol (DTT) on enzyme performance was evaluated at 5 mM. In addition, the impact of 1% (v/v) of acetonitrile, triton X-100, tween-80, hydrogen peroxides (H2O2), and dimethyl sulfoxide (DMSO) on enzyme catalytic performance was studied. The crude extract pre-incubated with simply distilled water was regarded as 100% effective and was taken as the control.
2.13 Effect of laundry detergents on keratinase stability
The protocol described by Paul et al. [27] was used to assess the impact of Omo, Sunlight, Surf, Ariel, Skyy, Prowash, Fresh Wave, and Evaklin on the stability of the keratinolytic enzyme. Briefly, various detergent powders were solubilized in tap water. The detergent solutions (7 mg/mL) were heated at 100°C for 30 min to inactivate any pre-existing endogenous proteolytic enzymes. After the detergent solutions cooled to 50°C, crude keratinase was added and incubated; then, remaining keratinase activity was ascertained at 30 min and 60 min post-incubation. The control experiment that was set at 100% was crude extract pre-incubated with tap water.
2.14 Analysis of statistics
All the experimental runs were performed in triplicate unless otherwise stated. The datasets generated were analyzed using SigmaPlot v11.0, and the mean significant difference was compared at P < 0.05.
3 Results
3.1 Bacterial isolation and keratinolytic potential evaluation
Based on distinct morphological characteristics (result not shown), twenty-two (22) axenic bacteria were isolated from the soil samples collected from the poultry dumpsite. Eighteen (18) bacterial isolates showed variable zones of hydrolysis on skimmed milk agar plates ranging from 5 ± 0.71 mm for PSS-03 and 25.5 ± 0.71 mm for DSS-03 (Supplementary Fig. 1; Supplementary Table 1), while 4 isolates (DSS-04, PSS-01, PSS-05, PSS-11) showed no clearance zone around the bacterial colonies on skimmed milk agar plates. Out of the 18 proteolytic bacteria, 9 isolates showed different abilities to degrade whole chicken feathers in MSM (Supplementary Fig. 2), with percentage feather hydrolysis ranging from 24% (PSS-10) to 81% (DSS-02) as shown in Table 1. At the same time, the extracellular keratinase titre quantified using the crude extract of the fermented media indicated that the lowest enzyme activity was recorded by PSS-10 (63.63 ± 41.14 U/mL), while DSS-02 displayed the maximum extracellular keratinase production of 693.63 ± 62.99 U/mL within the incubation timeline used (Table 1).
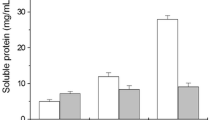
Expectedly, the medium inoculated with isolate PSS-10 recorded the least total protein (0.02 ± 0.01 mg/mL), whereas the maximum protein content of 0.18 ± 0.01 mg/mL was determined using samples from DSS-01 fermentation media (Table 1). The free sulfhydryl groups liberated during feathers degradation by the isolates showed marginal differences except for isolates PSS-10 and DSS-03, which had 0.03 ± 0 and 0.05 ± 0 (mM), respectively (Table 1). The medium pH drifted from the initial pH of 6.5 toward the spectrum's neutral/alkaline (pH 7.2–8.2) range (Table 1).
3.2 Identification of potent keratinolytic isolates and process conditions optimization
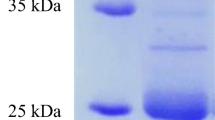
The performance of the bacterial isolates during whole feather hydrolysis was an index for selecting four potent isolates (DSS-02, PSS-04, PSS-12, PSS-14) for identification study. Polymerase chain reaction amplification of the 16S rRNA gene sequence of the isolates, sequencing of the amplicons, and phylogenetic analysis against the NCBI database showed that the isolates DSS-02, PSS-04, PSS-12, and PSS-14 are closely related to reference strains Chryseobacterium proteolyticum (NCBI: AB039830), Chryseobacterium lactis (NCBI: KT945035), Chryseobacterium aquifrigidense (NCBI: MN099388) and Pseudomonas aeruginosa (NCBI: MT633047), with percentage similarities of 100%, 99.28%, 99.67%, and 100%, respectively. Accordingly, the isolates were identified as Chryseobacterium proteolyticum FGNn, Chryseobacterium lactis GNNcx, Chryseobacterium aquifrigidense NGFc and Pseudomonas aeruginosa GNFx; and their respective nucleotide sequences were deposited in the GenBank with the accession numbers MW165478, MW165480, MW165481, and MW165479 (Supplementary Table 2).
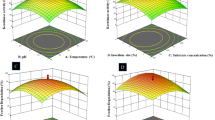
Two (Chryseobacterium proteolyticum FGNn and Pseudomonas aeruginosa GNFx) of the four identified keratinolytic bacteria were carefully chosen based on their keratinolytic potential for further optimization study. The results of the optimization study showed that the enzyme production by C. proteolyticum FGNn was best at pH (4), incubation temperature (30°C), inoculum size (2%, v/v), and chicken feather concentration (15 g/L) (Supplementary Fig. 3). Meanwhile, P. aeruginosa GNFx optimally secreted keratinase at pH, incubation temperature, inoculum size, and chicken feather concentration of 7, 30°C, 5% v/v, and 25 g/L, respectively (Supplementary Fig. 3).
3.3 Time course study of keratinolytic activities
Keratinase production by C. proteolyticum FHNn showed an enzyme titre of 653.64 ± 37.28 U/mL at 24 h and then decreased at 48 h of incubation. Further incubation engendered a spike in the extracellular keratinase with enzyme activity of 1756.36 ± 2.57 U/mL (a 2.5-fold increase compared to unoptimized condition (Table 1)) after 72 h coinciding with the late exponential/stationary phase of the bacterial growth curve (Fig. 1a). Beyond 72 h, keratinase titre decreased steadily with increasing fermentation period, having 488.18 ± 34.71 U/mL at 168 h. Similarly, free sulfhydryls released during the decomposition of the keratinous polymer showed a low concentration after the first 24 h of fermentation (Fig. 1a). Incubating the production medium further saw a sharp free sulfhydryls increment, achieving a maximum concentration (7.19 ± 0.13 mM) at 72 h similar to optimal keratinase production. Free sulfhydryl groups decreased consistently after 72 h of incubation, with only 1.01 ± 0.14 mM recorded after 168 h. In addition, protein concentration consistently increased within the first 96 h of fermentation and suddenly showed a spike at 120 h with a maximum concentration of 530.31 ± 13.15 μg/mL (Fig. 1a). The pH evaluation indicated that the fermentation medium experienced constant alkalinization following feather degradation which caused the production medium pH to change from the initial value of 4.0 to 8.14 after 168 h (Fig. 1a).
A similar pattern of keratinase production was shown by P. aeruginosa GNFx, except that the maximum extracellular keratinase titre (2055.45 ± 50.14 U/mL) was obtained at 96 h of fermentation, coinciding with the death phase of the microbial growth curve (Fig. 1b). The enzyme production indicated a 3.3-fold increment compared to that achieved during initial evaluation (Table 1). Free sulfhydryl concentration increased from 24 to 48 h of fermentation, and subsequently, decreased slightly at 72 h. After 72 h, free sulfhydryls liberation in the medium increased consistently, with a maximum concentration of 7.84 ± 0.08 mM recorded after 168 h (Fig. 1b). A significant amount of protein consistently built-up in the fermentation medium as the incubation time was increasing, and then peaked at 144 h with a total concentration of 986.22 ± 14.55 μg/mL (Fig. 1b). The initial medium pH (7.0) used to start bacterial fermentation marginally decreased to 6.86 ± 0.14 after 24 h of incubation. Beyond 24 h, the pH drifted towards alkalinity, with the highest value of 9.14 ± 0.17 recorded at 168 h (Fig. 1b).
3.4 Amino acids profile of the generated protein hydrolysates
Amino acid analysis showed that different amino acids were liberated in various proportions during the bacteria growth on the chicken feathers. The total protein contents and amino acids concentration of 58.23% and 34.67% for C. proteolyticum FGNn and 50.42% and 49.13% for P. aeruginosa GNFx, respectively, were determined using lyophilised feather hydrolysates. Arginine (3.18%) and glycine (3.26%) were more abundant than other amino acids, including serine (2.80%), glutamic acid (2.04%), threonine (1.98%), proline (2.15%), valine (2.39%), phenylalanine (2.03%), leucine (2.49%), histidine (2.09%), and lysine (2.78%) in FGNn hydrolysate (Fig. 2). Strain GNFx feather hydrolysate showed a preponderant presence of glutamic acid (6.05%), and significant concentrations of serine (3.31%), aspartic acid (4.74%), glycine (3.71%), alanine (3.43%), valine (3.09%), and leucine (3.23%). However, both hydrolysates had relatively low cysteine, tryptophan, and tyrosine concentrations, while HO-proline was not detected (Fig. 2).
3.5 Effect of pH and temperature on keratinase activity and stability
The activity of both bacterial keratinases was optimal at pH 8.0 and decreased as pH tended towards higher alkalinity (Fig. 3a). FGNn keratinase showed maximum enzyme activity at 50°C, and beyond this temperature, the activity declined drastically (Fig. 3b). Whereas GNFx keratinase displayed optimal activity at 60°C, with significant relative activity of 84.49 ± 1.16% and 65.20 ± 1.87% at 70°C and 80°C, respectively (Fig. 3b).
Optimal (a) pH and (b) temperature determination for maximum activity of keratinases from C. proteolyticum FGNn and P. aeruginosa GNFx. (c) Thermostability study of P. aeruginosa GNFx keratinase for 120 min of heating at different temperatures. Each point on the line graphs indicates the mean and standard deviation of replicate experiments
The thermostability study indicated that at 50°C, GNFx keratinase activity was progressively enhanced following the thermal pretreatment with residual activity of 132.78 ± 6.38% at 120 min (Fig. 3c). Also, the catalytic property of the enzyme was not significantly perturbed at 60°C and 70°C; but rather showed noticeable recovery of catalytic efficiency after 60 min at 60°C and 90 min at 70°C. The final residual activity recorded after 120 min of heating was 109.86 ± 6.34% and 105.28 ± 4.26% at 60°C and 70°C, respectively. At 80°C heat treatment, the keratinolytic protease maintained 93.66 ± 4.98% residual activity after 30 min. The enzyme stability declined as the thermal treatment time increased to 60 min but showed more than a half-life time. Further heating at 80°C engendered more disruption of the catalytic function as the residual activity dropped to 29.57 ± 5.98% at 120 min (Fig. 3c).
3.6 Chemical agents’ effect on keratinase stability
The presence of serine protease inhibitor – PMSF did not affect the enzyme activity significantly as the keratinases from FGNn, and GNFx showed residual activity of 97.93 ± 4.65% and 96.87 ± 3.93%, respectively (Table 2). Similarly, GNFx keratinase had a marginal decrease in residual activity against metalloprotease inhibitors, EDTA, and 1,10-phenanthroline. However, FGNn keratinolytic protease was significantly inhibited by EDTA and 1,10-phenanthroline with respective residual activity of 29.48 ± 1.12% and 28.29 ± 0.48%. FGNn keratinase demonstrated high stability post-treatment with DTT (90.07 ± 5.45%), H2O2 (103.41 ± 3.85%), acetonitrile (100.74 ± 2.89%), triton-X100 (118.52 ± 1.92%), tween-80 (130.67 ± 3.53%), SDS (108.59 ± 1.12%). Likewise, GNFx keratinase was highly stable in the presence of DDT (99.33 ± 2.95%); while showing enhanced residual activity of 119.34 ± 1.32%, 140.40 ± 3.94%, 150.93 ± 3.28%, and 151.28 ± 4.27% against H2O2, triton-X100, tween-80 and SDS, respectively (Table 1). Contrarily, acetonitrile negatively impacted GNFx keratinase; hence it retained only 74.76 ± 3.60% of the original activity after 1 h pretreatment. C. proteolyticum FGNn keratinase showed remarkable stability against K+ with residual keratinase activity of 99.49 ± 1.08%. However, the rest of the metal ions tested perturbed the catalytic efficiency of the protease as it maintained residual activity ranging from 47.95 ± 1.09% (Zn2+) to 80.89 ± 1.99% (Mg2+). Metal ions pretreatment positively influenced the biocatalysis of GNFx keratinase compared to the control (Table 2). Among the metal ions tested, Fe3+ and K+ caused significantly improved enzymatic activity, with residual activity of 144.91 ± 2.44% and 137.12 ± 0.77%, respectively, while the least residual activity of 94.06 ± 0.73% was recorded against Co2+ (Table 2).
3.7 Assessment of keratinase stability in the presence of laundry detergent
Keratinase stability assessment against selected laundry detergents showed that P. aeruginosa GNFx keratinolytic protease demonstrated robust structural stability and enhanced catalytic activity with residual activity of 153.85% (Ariel), 119.78% (Omo), 141.76% (Sunlight), 128.57% (Surf), 146.15% (Sky), 93.52% (FreshWave), 126.48% (Eva), and 107.69% (PROWASH) as shown in Fig. 4. Contrariwise, C. proteolyticum FGNn keratinase generally demonstrated a decrease in stability compared to the control experiment, with residual activity of 80.49%, 78.64%, 81.55%, 95.73%, 69.35%, 76.78%, 98.59%, and 88.55% for Ariel, Omo, Sunlight, Surf, Sky, FreshWave, Eva, and PROWASH, respectively (Fig. 4).
4 Discussion
In recent time, microorganisms have become game-changers of sustainable development in the various sectors of the bioeconomy due to their genetic diversity. Also, the growing transition from classical methods to sustainable means of processing, production, and recycling has brought microbial technology to the fore of the circular bioeconomy. Poultry processing has significantly contributed to the stream of waste materials emanating from the agroindustrial sector. Proper handling and management of poultry feathers using a biobased approach have become imperative for quality protein recovery and eco-restoration. Depending on the intended applicability, new microbes are continually sought from various habitats for unique traits that would revolutionize the landscape of biotechnological processes. In light of this, keratinolytic bacteria from poultry dumpsites were evaluated for chicken feather degradation and keratinase production in minimal salt media. Different poultry or tannery dumpsites have been explored for robust keratinolytic bacteria as it is believed that their survival in this habitat is an indication of excellent adaptability [28,29,30]. A few isolates recovered from the same site showed neither proteolytic nor keratinolytic activity. At the same time, some bacteria with proteolytic activity could not degrade keratinous chicken feathers in basal media. The varied capacities exhibited by these isolates may suggest a symbiotic relationship among the microbial community in that particular niche, especially regarding macro- and micro-nutrient accessibility for survival. The preliminary evaluation showed that bacteria’s feather degradation and keratinase production capacities differed significantly due to the strain dissimilarity and genetic diversity [16]. Principally, keratinolysis releases appreciable concentrations of ammonia that perturb the prevailing pH of the fermentation medium by causing a drift toward the alkaline spectrum [31]. A similar observation was noted for two keratinolytic bacteria – Aquamicrobium defluvii FH 20 and Bacillus safensis LAU 13 during their growth on keratinous feathers where the cultivation medium pH changed from 6.0 to 8.3 [32]. The alkalization of the media is caused by the deamination of amino-containing compounds emanating from feather keratin decomposition [33].
Based on the pecking order of the isolates’ keratinolytic propensity, the identity of the four most potent bacteria was confirmed as C. proteolyticum FGNn, C. lactis GNNcx, C. aquifrigidense NGFc, and P. aeruginosa GNFx. These bacteria identity further corroborated the Gram-negative bacterial species previously reported to dismember keratinous materials into bio-accessible products through extracellular keratinase production [16, 33, 34]. The absence of Bacillus spp. among these top degraders shows that keratin colonisation and decomposition diversified into several bacterial strains; hence continuous exploration is imperative to discovering more novel isolates.
A time course study showed that strain GNFx had maximum extracellular keratinase titre at a longer incubation time than strain FGNn, even though the latter had better keratinolytic propensity in unoptimized conditions. These findings suggest that bacterial species' peculiarity and their microenvironment contribute significantly to their optimal productivity. C. proteolyticum FGNn optimal keratinase production at a shorter incubation time than P. aeruginosa GNFx is desirable, especially if the enzyme commercialisation is in perspective. Contrary to our findings, keratinase production by P. aeruginosa YK17 [19] and Pseudomonas sp. LM19 [17] peaked at 72 h post-incubation with lower enzyme titres. Similar to the present report, keratinase production by Chryseobacterium gleum climaxed at 72 h [35], whereas Chryseobacterium sp. RBT had a maximum keratinase yield of 89.12 U/mL at 48 h [36]. Generally, because of genetic diversity and strain variations, proteolytic enzyme production by wild bacterial strains has suffered many shortfalls, including but not limited to low yield and prolonged production time. Therefore, gene cloning, heterologous expression, and molecular optimization are currently being explored to upscale productivity. The decrease in keratinase activity after 48 h of fermentation among keratinolytic bacteria could be ascribed to the fluctuation in the cellular processes during the biosynthesis of primary metabolites that support bacteria growth and reproduction [37, 38]. The detection of a high concentration of free sulfhydryls during the fermentation process is a clear indication of efficient hydrolysis of disulfide bonds which subsequently promotes proteolytic catabolism of feather keratin into simpler and soluble products with multifaceted application potential [39]. Protein quantitation further corroborated the keratinolytic prowess of the bacterial isolates under investigation as a high amount of protein amassed following feather biodegradation. Protein accumulation in fermentation media is one of the indicators of keratinolysis, which might suggest the efficiency of the studied strains' keratinolytic system towards keratin utilization. A similar finding was recorded for other bacterial isolates growing on feather meal [40].
Furthermore, the amino acid analysis indicated that though C. proteolyticum FGNn generated feather hydrolysate, which was marginally higher in protein content, P. aeruginosa GNFx associated hydrolysates had more abundance of various amino acids than strain FGNn hydrolysate. The litany of amino acids present in the analyzed feather hydrolysates was in consonant with that generated by similar keratinolytic bacteria – Chryseobacterium sediminis RCM-SSR-7 [41] and Pseudomonas sp. LM19 [17] growing on keratinous biomass. The results also highlighted the dismemberment of keratinous feathers into simpler protein units by strain GNFx with a unique pattern of amino acid cleavage from feather keratin compared to isolate FGNn. This observation further suggested the peculiarity of keratinolytic bacterial strains to degrading keratinous biomass. Therefore, based on the nature of catalytic tendencies displayed by the isolates, the use of microbial consortium could improve the valorisation prospects of these isolates from the nutritional value point of view. Metagenomic profiling of the bacterial community involved in chicken feathers bioconversion indicated that more than 90% of the consortium belonged to the genera – Chryseobacterium, Stenotrophomonas, and Pseudomonas [33], which further substantiates the present report. Also, Kang and colleagues report on genomic analysis of keratinolytic Chryseobacterium sp. KMC2 provided an understanding of metabolic pathways that participate in keratinolysis [42].
Most of the keratinolytic proteases already characterized are optimally active at alkaline conditions. This characteristic behaviour suggests that proper alignment of the catalytic cleft and other essential residues that promote biocatalysis are favoured in alkaline conditions. Temperature rise positively influences the performance of biocatalysts until the optimal temperature is reached. Beyond optimum temperature, the enzyme's active site tends to lose its catalytic conformation due to the perturbation of the protein's tertiary structure, decreasing the efficiency of substrate binding and product formation [43]. The keratinases under investigation showed different optimal temperatures attributable to their bacterial sources. Several reports on Chryseobacterium spp. keratinases indicate that their optimal temperature ranges between 30°C and 50°C [44,45,46]. Therefore, protein engineering could offer beneficial attributes if keratinases from Chryseobacterium spp. are targeted for processes that involve elevated temperature conditions.
On the other hand, P. aeruginosa GNFx keratinase exhibited a relatively higher optimal temperature compared to C. proteolyticum FGNn keratinase. In line with the present findings, keratinases secreted by P. aeruginosa S-04 [20], P. aeruginosa 4-3 [47], and P. aeruginosa KS-1 [48] were optimally active at 60°C. The thermostability profile of GNFx keratinase indicates the ability of the enzyme to retain viable catalytic orientation after heating the enzyme solution for a reasonable period, which was also evident at 80°C as the keratinolytic protease maintained a half-life time of more than 60 min. This attribute further highlights the industrial and biotechnological significance of P. aeruginosa keratinase.
Proteases are generally classified following their inhibition patterns to different protease inhibitors. Repression of enzyme activity of FGNn keratinase by metal ion chelators, EDTA, and 1,10-phenanthroline suggests that it belongs to metallopeptidases [46]. Metallokeratinases are widely distributed among various bacterial strains, including, Pedobacter sp. 3.14.7 [2], Chryseobacterium indologenes TKU014 [46], Bacillus subtilis NRC 3 [49], Microbacterium sp. kr10 [50], P. aeruginosa S-04 [20], Acinetobacter sp. R-1 [51], Lysobacter NCIMB 9497 [52] among others, where the enzymes have been isolated and characterized for their value addition potential. Contrariwise, the biocatalytic efficiency of GNFx keratinase was not drastically affected by the presence of serine- or metallo-protease inhibitors tested; this suggests that it may belong to another class of protease that was not assayed for in this study. Furthermore, P. aeruginosa GNFx keratinase was catalytically enhanced after pretreatment with various chemical agents, including reducing agents, organic solvents, surfactants, and metal ions. Similarly, keratinase from P. aeruginosa S-04 demonstrated catalytic improvement post-treatment with DMSO, tween 20, triton X-100, Mn2+, Fe2+, ethanol, and isopropyl alcohol [20]. The behaviour of GNFx keratinase towards chemical agents underscores its multifaceted industrial application prospects. In addition, the enhanced activity observed after treatment with surfactants and hydrogen peroxide suggests promising candidacy of the protease for detergent production. Likewise, C. proteolyticum FGNn keratinase showed remarkable stability in the presence of reducing agents, hydrogen peroxide, organic solvent, and surfactants, supporting its potential candidacy for an industrial sector. However, various metal ions tested perturbed the catalytic conformation of FGNn keratinase. It can be suggested that the interaction or association of the enzyme with the tested metal ions negatively impacted the catalytic integrity of the biomolecule, and similar stability reduction was observed among other bacterial keratinases previously characterized [53].
The bacterial keratinases demonstrated different stability patterns when tested against selected laundry detergents, reflecting their bacterial origin and peculiar robust tendencies. C. proteolyticum keratinase showed lesser stability compared to the P. aeruginosa counterpart. The variable stability trend observed among various detergents might be attributed to the discrepancy in the ingredients used to formulate the detergents [25]. Contrarily, the activity of GNFx keratinase was remarkably enhanced following pretreatment with the various detergents. This is suggestive of a proper alignment of catalytic residues and binding sites following the enzyme’s interaction with the components of the detergents [54]. The pronounced stability profile exhibited by GNFx keratinase against FGNn keratinase might suggest that the bacterial enzymes’ variable side chains reacted differently following incubation with the various detergents. The activity enhancement of P. aeruginosa keratinolytic protease after detergent treatment underlines its compatibility and industrial prospects for detergent formulation [55] and green processes, respectively, if further explored.
5 Conclusions
In conclusion, the keratinolytic capacity of bacterial species recovered from poultry dumpsite was determined following halo zone formation on skimmed milk agar and intact chicken feathers degradation in basal media. By secreting higher titres of keratinolytic protease, isolates identified as C. proteolyticum FGNn and P. aeruginosa GNFx successfully disassembled keratinous chicken feathers. These isolates may be useful candidates for recycling feather wastes since the protein hydrolysates generated by hydrolysing keratinous chicken feathers were abundant in various amino acids. The study of the bacterial enzymes showed that GNFx keratinase exhibited superior catalytic properties vis-à-vis thermoactivity, thermostability, chemical agents’ tolerance, and laundry detergent stability compared to FGNn keratinase. The GNFx keratinase’s outstanding stability against the laundry detergents evaluated underlines its potential as a detergent bioadditive. The attributes of P. aeruginosa GNFx and associated keratinase highlight its crucial place in green technological processes that promote a circular bioeconomy. Consequently, future investigations on mining the bacterial genome for keratinolytic determinants are crucial to upscaling the product yield.
Data availability
The data that support the findings of this study are available from the corresponding author upon reasonable request.
References
Ossai IC, Hamid FS, Hassan A (2022) Valorisation of keratinous wastes: A sustainable approach towards a circular economy. Waste Manage 151:81–104. https://doi.org/10.1016/j.wasman.2022.07.021
Rios P, Bezus B, Cavalitto S, Cavello I (2022) Production and characterisation of a new detergent-stable keratinase expressed by Pedobacter sp. 3.14. 7, a novel Antarctic psychrotolerant keratin-degrading bacterium. J Genet Eng Biotechnol 20:81. https://doi.org/10.1186/s43141-022-00356-x
Siddharthan N, Balagurunathan R, Hemalatha N (2022) A novel feather-degrading bacterial isolate Geobacillus thermodenitrificans PS41 isolated from poultry farm soil. Arch Microbiol 204:565. https://doi.org/10.1007/s00203-022-03179-z
Reddy CC, Khilji IA, Gupta A, Bhuyar P, Mahmood S, KAS AL-J, Chua GK (2021) Valorisation of keratin waste biomass and its potential applications. J Water Process Eng 40:101707. https://doi.org/10.1016/j.jwpe.2020.101707
Mishra A, Jung D, Kim NK, Bhattacharyya D (2023) Influence of chicken feather fibre processing technique on mechanical and fire performances of flame-retardant polypropylene composites. Compos Part A Appl Sci Manuf 165:107338. https://doi.org/10.1016/j.compositesa.2022.107338
Zhou L, Xie X, Wu T, Chen M, Yao Q, Zhu H, Zou W (2020) Compound enzymatic hydrolysis of feather waste to improve the nutritional value. Biomass Conv Bioref 12:287–298. https://doi.org/10.1007/s13399-020-00643-y
Wang YX, Cao XJ (2012) Extracting keratin from chicken feathers by using a hydrophobic ionic liquid. Process Biochem 47:896–899. https://doi.org/10.1016/j.procbio.2012.02.013
Pourjavaheri F, Pour SO, Jones OA, Smooker PM, Brkljača R, Sherkat F, Blanch EW, Gupta A, Shanks RA (2019) Extraction of keratin from waste chicken feathers using sodium sulfide and l-cysteine. Process Biochem 82:205–214. https://doi.org/10.1016/j.procbio.2019.04.010
Yadav S, Bumbra P, Laura JS, Khosla B (2022) Optimisation of nutritional and physical parameters for enhancing the keratinase activity of Bacillus cereus isolated from soil of poultry dump site in Gurugram, Haryana. Bioresour Technol Rep 18:101108. https://doi.org/10.1016/j.biteb.2022.101108
Liya SM, Umesh M (2023) Bioconversion of chicken feather waste into feather hydrolysate by multifaceted keratinolytic Bacillus tropicus LS27 and new insights into its antioxidant and plant growth-promoting properties. Biomass Conv Bioref. https://doi.org/10.1007/s13399-023-04664-1
Sypka M, Jodłowska I, Białkowska AM (2021) Keratinases as versatile enzymatic tools for sustainable development. Biomolecules 11:1900. https://doi.org/10.3390/biom11121900
Gurav R, Nalavade V, Aware C, Vyavahare G, Bhatia SK, Yang YH, Bapat V, Jadhav J (2020) Microbial degradation of poultry feather biomass in a constructed bioreactor and application of hydrolysate as bioenhancer to vegetable crops. Environ Sci Pollut Res 27:2027–2035. https://doi.org/10.1007/s11356-019-06536-6
Kshetri P, Singh PL, Chanu SB, Singh TS, Rajiv C, Tamreihao K, Singh HN, Chongtham T, Devi AK, Sharma SK, Chongtham S (2022) Biological activity of peptides isolated from feather keratin waste through microbial and enzymatic hydrolysis. Electron J Biotechnol 60:11–18. https://doi.org/10.1016/j.ejbt.2022.08.001
Qin X, Xu X, Guo Y, Shen Q, Liu J, Yang C, Scott E, Bitter H, Zhang C (2022) A sustainable and efficient recycling strategy of feather waste into keratin peptides with antimicrobial activity. Waste Manage 144:421–430. https://doi.org/10.1016/j.wasman.2022.04.017
Shen N, Yang M, Xie C, Pan J, Pang K, Zhang H, Wang Y, Jiang M (2022) Isolation and identification of a feather degrading Bacillus tropicus strain Gxun-17 from marine environment and its enzyme characteristics. BMC Biotechnol 22:11. https://doi.org/10.1186/s12896-022-00742-w
Riffel A, Brandelli A (2006) Keratinolytic bacteria isolated from feather waste. Braz J Microbiol 37:395–399. https://doi.org/10.1590/S1517-83822006000300036
Mohamad N, Phang LY, Abd-Aziz S (2017) Optimisation of metallo-keratinase production by Pseudomonas sp. LM19 as a potential enzyme for feather waste conversion. Biocatal Biotransform 35:41–50. https://doi.org/10.1080/10242422.2017.1280031
Bokveld A, Nnolim NE, Digban TO, Okoh AI, Nwodo UU (2023) Chryseobacterium aquifrigidense keratinase liberated essential and nonessential amino acids from chicken feather degradation. Environ Technol 44:293–303. https://doi.org/10.1080/09593330.2021.1969597
Moonnee YA, Foysal MJ, Hashem A, Miah MF (2021) Keratinolytic protease from Pseudomonas aeruginosa for leather skin processing. J Genet Eng Biotechnol 19:53. https://doi.org/10.1186/s43141-021-00149-8
Ramalingum N, Bhagwat P, Permaul K, Pillai S (2022) Production, characterisation, and application of Pseudomonas aeruginosa S-04 keratinase for feather utilisation. Biomass Conv Bioref. https://doi.org/10.1007/s13399-022-03218-1
Kuo JM, Yang JI, Chen WM, Pan MH, Tsai ML, Lai YJ, Hwang A, Pan BS, Lin CY (2012) Purification and characterization of a thermostable keratinase from Meiothermus sp. I40. Int Biodeterior Biodegrad 70:111–116. https://doi.org/10.1016/j.ibiod.2012.02.006
Bradford MM (1976) A rapid and sensitive method for the quantitation of microgram quantities of protein utilising the principle of protein-dye binding. Anal Biochem 72:248–254. https://doi.org/10.1016/0003-2697(76)90527-3
Ellman GL (1959) Tissue sulfhydryl groups. Arch Biochem Biophys 82:70–77
Lorenz TC (2012) Polymerase chain reaction: basic protocol plus troubleshooting and optimisation strategies. J Vis Exp 63:e3998. https://doi.org/10.3791/3998
Jeong JH, Park KH, Oh DJ, Hwang DY, Kim HS, Lee CY, Son HJ (2010) Keratinolytic enzyme-mediated biodegradation of recalcitrant feather by a newly isolated Xanthomonas sp. P5. Polym Degrad Stab 95:1969–1977. https://doi.org/10.1016/j.polymdegradstab.2010.07.020
Einarsson S, Josefsson B, Lagerkvist S (1983) Determination of amino acids with 9-fluorenylmethyl chloroformate and reversed-phase high-performance liquid chromatography. J Chromatogr A 282:609–618. https://doi.org/10.1016/S0021-9673(00)91638-8
Paul T, Das A, Mandal A, Halder SK, Jana A, Maity C, DasMohapatra PK, Pati BR, Mondal KC (2014) An efficient cloth cleaning properties of a crude keratinase combined with detergent: towards industrial viewpoint. J Clean Prod 66:672–684. https://doi.org/10.1016/j.jclepro.2013.10.054
Riffel A, Ortolan S, Brandelli A (2003) De-hairing activity of extracellular proteases produced by keratinolytic bacteria. J Chem Technol Biotechnol Inter Res Process Environ Clean Technol 78:855–859
Sekar V, Kannan M, Ganesan R, Dheeba B, Sivakumar N, Kannan K (2016) Isolation and screening of keratinolytic bacteria from feather dumping soil in and around Cuddalore and Villupuram, Tamil Nadu. Proc Natl Acad Sci India Sect B Biol Sci 86:567–575. https://doi.org/10.1007/s40011-014-0483-8
Barman NC, Zohora FT, Das KC, Mowla MG, Banu NA, Salimullah M, Hashem A (2017) Production, partial optimisation and characterisation of keratinase enzyme by Arthrobacter sp. NFH5 isolated from soil samples. AMB Express 7:181. https://doi.org/10.1186/s13568-017-0462-6
He Z, Sun R, Tang Z, Bu T, Wu Q, Li C, Chen H (2018) Biodegradation of feather waste keratin by the keratin-degrading strain Bacillus subtilis 8. J Microbiol Biotechnol 28:314–322. https://doi.org/10.4014/jmb.1708.08077
Adelere IA, Lateef A (2022) Valorisation of feather by Bacillus safensis and Aquamicrobium defluvii for growth promotion in leafy vegetables. Waste Biomass Valori 14:723–737. https://doi.org/10.1007/s12649-022-01904-9
Kang D, Huang Y, Nesme J, Herschend J, Jacquiod S, Kot W, Hansen LH, Lange L, Sørensen SJ (2021a) Metagenomic analysis of a keratin-degrading bacterial consortium provides insight into the keratinolytic mechanisms. Sci Total Environ 761:143281. https://doi.org/10.1016/j.scitotenv.2020.143281
Kang D, Herschend J, Al-Soud WA, Mortensen MS, Gonzalo M, Jacquiod S, Sørensen SJ (2018) Enrichment and characterisation of an environmental microbial consortium displaying efficient keratinolytic activity. Bioresour Technol 270:303–310. https://doi.org/10.1016/j.biortech.2018.09.006
Chaudhari PN, Chaudhari BL, Chincholkar SB (2013) Iron containing keratinolytic metallo-protease produced by Chryseobacterium gleum. Process Biochem 48:144–151. https://doi.org/10.1016/j.procbio.2012.11.009
Gurav RG, Tang J, Jadhav JP (2016) Sulfitolytic and keratinolytic potential of Chryseobacterium sp. RBT revealed hydrolysis of melanin containing feathers. 3 Biotech 6:145. https://doi.org/10.1007/s13205-016-0464-0
Levine E, Hwa T (2007) Stochastic fluctuations in metabolic pathways. Proc Natl Acad Sci 104:9224–9229. https://doi.org/10.1073/pnas.0610987104
Horak I, Engelbrecht G, van Rensburg PJ, Claassens S (2019) Microbial metabolomics: essential definitions and the importance of cultivation conditions for utilising Bacillus species as bionematicides. J Appl Microbiol 127:326–343. https://doi.org/10.1111/jam.14218
Ramnani P, Singh R, Gupta R (2005) Keratinolytic potential of Bacillus licheniformis RG1: structural and biochemical mechanism of feather degradation. Can J Microbiol 51:191–196. https://doi.org/10.1139/w04-123
Pereira JQ, Lopes FC, Petry MV, da Costa Medina LF, Brandelli A (2014) Isolation of three novel Antarctic psychrotolerant feather-degrading bacteria and partial purification of keratinolytic enzyme from Lysobacter sp. A03. Inter Biodeterior Biodegrad 88:1–7. https://doi.org/10.1016/j.ibiod.2013.11.012
Kshetri P, Roy SS, Sharma SK, Singh TS, Ansari MA, Prakash N, Ngachan SV (2019) Transforming chicken feather waste into feather protein hydrolysate using a newly isolated multifaceted keratinolytic bacterium Chryseobacterium sediminis RCM-SSR-7. Waste Biomass Valor 10:1–11. https://doi.org/10.1007/s12649-017-0037-4
Kang D, Shoaie S, Jacquiod S, Sørensen SJ, Ledesma-Amaro R (2021b) Comparative genomics analysis of keratin-degrading Chryseobacterium species reveals their keratinolytic potential for secondary metabolite production. Microorganisms 9:1042. https://doi.org/10.3390/microorganisms9051042
Akram F, Haq IU, Hayat AK, Ahmed Z, Jabbar Z, Baig IM, Akram R (2021) Keratinolytic enzyme from a thermotolerant isolate Bacillus sp. NDS-10: an efficient Green biocatalyst for poultry waste management, laundry and hide-dehairing applications. Waste Biomass Valor 12:5001–5018. https://doi.org/10.1007/s12649-021-01369-2
Lv LX, Sim MH, Li YD, Min J, Feng WH, Guan WJ, Li YQ (2010) Production, characterisation and application of a keratinase from Chryseobacterium L99 sp. nov. Process Biochem 45:1236–1244. https://doi.org/10.1016/j.procbio.2010.03.011
Hong SJ, Park GS, Jung BK, Khan AR, Park YJ, Lee CH, Shin JH (2015) Isolation, identification, and characterization of a keratin-degrading bacterium Chryseobacterium sp. P1-3. J Appl Biol Chem 58:247–251. https://doi.org/10.3839/jabc.2015.039
Wang SL, Hsu WT, Liang TW, Yen YH, Wang CL (2008) Purification and characterization of three novel keratinolytic metalloproteases produced by Chryseobacterium indologenes TKU014 in a shrimp shell powder medium. Bioresour Technol 99:5679–5686. https://doi.org/10.1016/j.biortech.2007.10.024
Pei XD, Li F, Yue SY, Huang XN, Gao TT, Jiao DQ, Wang CH (2022) Production and characterisation of novel thermo-and organic solvent–stable keratinase and aminopeptidase from Pseudomonas aeruginosa 4–3 for effective poultry feather degradation. Environ Sci Pollut Res 30:2480–2493. https://doi.org/10.1007/s11356-022-22367-4
Sharma R, Gupta R (2010) Substrate specificity characterisation of a thermostable keratinase from Pseudomonas aeruginosa KS-1. J Ind Microbiol Biotechnol 37:785–792. https://doi.org/10.1007/s10295-010-0723-8
Tork SE, Shahein YE, El-Hakim AE, Abdel-Aty AM, Aly MM (2013) Production and characterisation of thermostable metallo-keratinase from newly isolated Bacillus subtilis NRC 3. Inter J BiolMacromol 55:169–175. https://doi.org/10.1016/j.ijbiomac.2013.01.002
Thys RCS, Brandelli A (2006) Purification and properties of a keratinolytic metalloprotease from Microbacterium sp. J Appl Microbiol 101:1259–1268. https://doi.org/10.1111/j.1365-2672.2006.03050.x
Zhang RX, Gong JS, Zhang DD, Su C, Hou YS, Li H, Shi JS, Xu ZH (2016) A metallo-keratinase from a newly isolated Acinetobacter sp. R-1 with low collagenase activity and its biotechnological application potential in leather industry. Bioprocess Biosyst Eng 39:193–204. https://doi.org/10.1007/s00449-015-1503-7
Allpress JD, Mountain G, Gowland PC (2002) Production, purification and characterization of an extracellular keratinase from Lysobacter NCIMB 9497. Lett Appl Microbiol 34:337–342. https://doi.org/10.1046/j.1472-765X.2002.01093.x
Gegeckas A, Šimkutė A, Gudiukaitė R, Čitavičius DJ (2018) Characterization and application of keratinolytic paptidases from Bacillus spp. Int J Biol Macromol 113:1206–1213. https://doi.org/10.1016/j.ijbiomac.2018.03.046
Rai SK, Mukherjee AK (2011) Optimization of production of an oxidant and detergent-stable alkaline β-keratinase from Brevibacillus sp. strain AS-S10-II: application of enzyme in laundry detergent formulations and in leather industry. Biochem Eng J 54:47–56. https://doi.org/10.1016/j.bej.2011.01.007
Benkiar A, Nadia ZJ, Badis A, Rebzani F, Soraya BT, Rekik H, Naili B, Ferradji FZ, Bejar S, Jaouadi B (2013) Biochemical and molecular characterization of a thermo-and detergent-stable alkaline serine keratinolytic protease from Bacillus circulans strain DZ100 for detergent formulations and feather-biodegradation process. Int Biodeterior Biodegrad 83:129–138. https://doi.org/10.1016/j.ibiod.2013.05.014
Acknowledgements
This work was supported by the Industrial Biocatalysis Hub, funded by the Department of Science and Innovation and the Technology Innovation Agency.
Funding
Open access funding provided by University of Fort Hare. This work was supported by the Industrial Biocatalysis Hub, funded by the Department of Science and Innovation and the Technology Innovation Agency.
Author information
Authors and Affiliations
Contributions
Conceptualization: Nnolim Nonso, Uchechukwu Nwodo; Methodology: Nonkonzo Giwu, Nonso Nnolim; Formal analysis and investigation: Nonso Nnolim; Writing - original draft preparation: Nonkonzo Giwu; Writing - review and editing: Nonso Nnolim, Uchechukwu Nwodo; Funding acquisition: Uchechukwu Nwodo; Resources: Uchechukwu Nwodo; Supervision: Nonso Nnolim, Uchechukwu Nwodo.
Corresponding author
Ethics declarations
Ethical approval
Not applicable.
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Giwu, N., Nnolim, N.E. & Nwodo, U.U. Keratinases produced by Chryseobacterium proteolyticum FGNn and Pseudomonas aeruginosa GNFx liberated amino acids from poultry feathers. Biomass Conv. Bioref. (2023). https://doi.org/10.1007/s13399-023-05035-6
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s13399-023-05035-6