Abstract
Lipids exert substantial influences on vertebrate embryogenesis, but their metabolic dynamics at detailed structural levels remains elusive, primarily owing to the lack of a tool capable of resolving their huge structural diversity. Herein, we present the first large-scale and spatiotemporal monitoring of unsaturated lipids with C=C specificity in single developing zebrafish embryos enabled by photochemical derivatization and tandem mass spectrometry (MS). The lipid isomer composition was found extremely stable in yolk throughout embryogenesis, while notable differences in ratios of C=C location (e.g., PC 16:0_16:1 (7) vs. 16:0_16:1 (9)) and fatty acyl composition isomers (e.g., PC 16:1_18:1 vs. 16:0_18:2) were unveiled between blastomeres and yolk from zygote to 4 h post fertilization (hpf). From 24 hpf onwards, lipid isomer compositions in embryo head and tail evolved distinctively with development, suggesting a meticulously regulated lipid remodeling essential for cell division and differentiation. This work has laid the foundation for functional studies of structurally defined lipids in vertebrate embryology.

Similar content being viewed by others
Introduction
Lipids, typically abundant in oocytes and embryos, are of profound importance to vertebrate embryonic development in aspects of cell membrane construction [1], energy provision [2], and signaling [3, 4]. Although efforts have been made on global lipid profiling of oocytes and embryos using mass spectrometry (MS) [5,6,7,8], our knowledge of lipids in the dynamic context of vertebrate embryogenesis is considerably limited and sketchy compared with that of nucleic acids and proteins. This situation is further complicated by the vast structural complexity of lipids owing to the five layers of lipid moieties, namely lipid class, fatty acyl identity, fatty acyl sn-position, carbon-carbon double bond (C=C) location, and C=C geometry (cis/trans), wherein the latter three present great challenges to conventional MS workflows [9,10,11,12,13]. Consequently, the metabolic dynamics and developmental significances of a large variety of structurally defined lipids remain largely overlooked and poorly understood in vertebrate embryology.
The zebrafish (Danio rerio) is a model vertebrate which has contributed substantially to developmental biology [14], drug discovery [15], human genetic [16,17,18], and cardiovascular disease studies [19, 20]. While recent advances have enabled the tracing of developmental trajectories of zebrafish embryonic cells at the single-cell transcriptomic level [21, 22] and a large-scale embryonic proteome analysis [23], the distribution, transport, and metabolism of lipids during zebrafish development have recently attracted increasing research interests [24,25,26,27,28,29]. Fraher et al. performed lipid profiling of the body and yolk of zebrafish embryos from 0 to 120 h post fertilization (hpf) and revealed that the two sections had characteristic lipid compositions which were dynamic with development [30]. However, the question of how lipids are processed in the first several hours of embryogenesis, during which cells undergo rapid division and differentiation, remains unanswered. Dueñas et al. applied high-resolution 3D mass spectrometric imaging (MSI) to visualize lipid distributions in the early-stage (up to 16-cell) zebrafish embryos and found lipids of major classes had symmetrical distributions in the blastomeres but each had its own specific location [31]. Pirro et al. developed an integrated desorption electrospray ionization (DESI) imaging and nanoESI-MS platform to study lipid dynamics in developing zebrafish embryos [32]. Nonetheless, most of these studies failed to resolve the detailed lipid structural moieties, which has great impacts on the biological functions of lipids.
Zebrafish embryos develop independently using yolk nutrients for the first 5 days post fertilization (dpf) before they are able to commence feeding (Figure 1a). Lipids, along with other cytoplasmic components, are initially stored in the yolk and transported to the animal pole upon fertilization to form the blastodisc. The embryo body and yolk are separated by the yolk syncytial layer (YSL), wherein many crucial genes for lipid transport and lipoprotein production are expressed [33, 34]. It is well established that the YSL is responsible for the processing and delivery of yolk lipids to the embryo body; however, a key question remains as to how this lipid remodeling process affects lipid compositions in the embryo body at deep structural levels. An enhanced lipidomics workflow that offers comprehensive lipid structural characterization at a high detection sensitivity should be able to provide more insights in this regard.
(a) Developmental stages of zebrafish embryos that were monitored in this work. (b) Experimental workflow for shotgun lipidomic analysis of one zebrafish embryo with C=C specificity. The blastomeres and yolk were manually dissected for separate lipid extraction. Lipid extracts were re-dissolved for online photochemical C=C derivatization and nanoESI-MS/MS analysis. (c) Scheme for locating the C=Cs in unsaturated lipids
In this study, we performed for the first time an in-depth and dynamic lipid profiling with C=C specificity of single developing vertebrate embryos. Zebrafish embryos were chosen owing to transparency in their early developmental stages, which simplified observation and handling. The photochemical Paternò–Büchi (PB) reaction was coupled with tandem MS [35, 36] (Figure 1b) to pinpoint C=C locations in glycerophospholipids (GPs), which were invisible to conventional lipid analysis methods. MS/MS fragmentation of the PB reaction product released two diagnostic ions distinctive to the C=C location, spaced 26 Da from each other (Figure 1c). The relative amount of a lipid C=C isomer could be represented by the summed abundance of its C=C-specific diagnostic ions [37,38,39], eliminating the use of internal standards. Recent improvements on the PB reaction-based workflow included the coupling with chromatographic separations [39,40,41], selection of new PB reagents [40, 42, 43], and development of custom software and databases [39, 44]. Moreover, the PB reaction, along with other tools for deeper lipid structural analysis, has been coupled with MSI for spatial profiling of lipids with isomeric resolutions [12, 38, 45]. These features have opened an avenue for the large-scale and time-resolved lipid profiling down to the C=C location level for developmental studies.
To investigate the possible lipid alterations between lipid source (i.e., the yolk) and destination (i.e., the blastomeres or embryo body), the blastomeres and yolk were dissected with their lipid contents separately extracted and analyzed by MS. Specifically, for early embryogenesis, lipid dynamics was monitored at a high temporal resolution as the embryo developed from 1-cell (zygote), 4-cell, 16-cell (cleavage period), 1k-cell until 4 hpf (blastula period). Later on, lipids in different parts of the embryo body, i.e., head and tail, were profiled at 24 hpf (end of segmentation period), 72 hpf (end of hatching period), and 5 dpf (early larval period).
Experimental
Reagents
Methanol (HPLC grade), acetonitrile (HPLC grade), and tricaine mesylate (MS-222) were purchased from Sigma-Aldrich (St. Louis, MO, USA), acetone (analytical grade) was purchased from Beijing Chemical Works (Beijing, China), chloroform was purchased from Beijing Tongguang Fine Chemicals Company (Beijing, China), and ammonium acetate was purchased from Aladdin Chemical Co., Ltd. (Shanghai, China). Deionized water was prepared from a Milli-Q water purifier (Millipore, Belford, MA, USA).
Lipid Nomenclature
Unless otherwise noted, ΔX was used to annotate C=C locations in fatty acyls, in which X indicated the lower numbered carbon in each C=C. For glycerophospholipids, the sn-positions of fatty acyls were not determined; therefore, “_” was used to denote unspecified sn-positions, e.g., PC 16:0_18:1.
Collection of Zebrafish Embryos and Lipid Extraction
In this study, only wild-type zebrafish embryos of Tübingen (TU) strain were used. Fish maintenance and embryo raising followed the ethical guidelines of the Tsinghua University Animal Care and Use Committee. Embryos were raised at 28.5 °C and staged according to Kimmel et al. [46]. When reaching to desired stages, embryos were transferred into a new Petri dish containing deionized water and dechorionated manually under a microscope. For stages of 1-cell, 4-cell, 16-cell, 1k-cell, and 4 hpf, separation of blastomeres and yolk was achieved by manual dissection using two fine needle tips of syringes under a microscope. For stages of 24 hpf, 72 hpf, and 5 dpf, embryos were transferred to a new Petri dish containing deionized water and MS-222 (ca. 100 mg/L) for anesthesia, and then the yolk was first dissected from the embryo body, after which the body was cut into head and tail (Figure S1). Each part from a dissected embryo was transferred into a 0.6-mL centrifuge tube, with 20-μL methanol immediately added to quench metabolic activities. All samples were stored at − 20 °C prior to lipid extraction.
Lipids were extracted using a modified Bligh-Dyer method [47]. Briefly, 20-μL deionized water and 40-μL chloroform were added into each tube which already contained the sample and 20-μL methanol. For head and tail samples of 24, 72 hpf, and 5 dpf stages, 2–5 min of sonication was applied for complete pulverization. The mixture was vortexed for 2 min and subject to centrifugation at 14,000 rpm for 5 min, and then the organic phase was transferred to a new centrifuge tube. Twenty microliters of chloroform was added into the remaining aqueous phase to repeat the vortex and centrifugation steps once, and then the organic phases were combined and dried. Lipid extracts were re-dissolved into the spray solvent, i.e., a mixture of acetone, acetonitrile, and 10 mM aqueous ammonium acetate (40/40/20, v/v/v). For each blastomere/head/tail sample, 5-μL solvent was added. For each yolk sample at stages from 1-cell to 24 hpf, 10-μL solvent was added, while only 5-μL solvent was used for each yolk sample at 72 hpf due to its shrinking size. The yolk of a single embryo at 5 dpf stage was too small for analysis, and only the head and tail parts were analyzed.
Mass Spectrometry
All experiments were performed on a Thermo Q-Exactive Orbitrap mass spectrometer (Thermo Fisher Scientific, San Jose, CA) unless otherwise noted. Major parameters were set as follows: resolution, 70,000; microscans, 1; maximum inject time, 50 ms; capillary temperature, 320 °C; and S-lens RF level, 50. For MS2, the high collision dissociation (HCD) energy was set as 20 (normalized collision energy, NCE), and the mass isolation window was 0.8 Da. The commercial ESI ion source was removed and replaced by a homemade nanoESI source prior to all experiments. The homemade nanoESI source was constructed on a lifting platform where a pulled emitter was fixed. The high voltage (+ 1700 V and − 1800 V for positive and negative mode analyses, respectively) was applied via a copper wire inserted from the back of the emitter to be in touch with the solution. NanoESI emitters with a tip diameter of 3 μm were fabricated by pulling borosilicate glass capillaries (o.d., 1.50 mm; i.d., 1.10 mm, Sutter Instrument) using a P-1000 micropipette puller (Sutter Instrument, Novato, CA, USA). Parameters for capillary pulling were set as follows: heat, 500; pull, 0; vel., 30; time, 250; and pressure, 500.
Online Photochemical Derivatization for Locating the C=C
Acetone was used as the photochemical reagent for PB reaction towards the C=Cs in unsaturated lipids. An ultraviolet lamp with a major emission at 254 nm was placed beside the nanoESI source. The distance between the lamp and nanoESI capillary was 2.5 cm. PB reaction products were identified by a 58-Da mass shift relative to intact lipids, which corresponded to the addition of one acetone molecule. For lipids containing polyunsaturated fatty acyls, only the first-stage (with one acetone addition) products were used for C=C localization. Diagnostic ions corresponding to all C=C locations were detected since PB reaction was not specific to C=Cs at different locations. Although we could not perform relative quantitation of C=C location isomers due to the lack of commercially available internal standards for translating the abundance ratio of diagnostic ions to concentration ratio, the term “relative amount” is used herein to indicate the isomer quantity.
Results and Discussion
Lipid Diversity in Zebrafish Embryos Revealed Through Deep Structural Characterization with C=C Specificity
As a proof of principle, MS analysis of the lipid extracts of the blastomeres and yolk dissected from a single zebrafish embryo revealed similar lipid compositions (Figure 2a, b) dominated by unsaturated phosphatidylcholines (PCs). The relative amounts of major lipids varied between the blastomeres and yolk. Generally, polyunsaturated lipids, such as PC 38:6 and PC 40:7, were more concentrated in the yolk. This suggested that lipids stored in the yolk were remodeled in response to developmental needs. Despite these differences, the blastomeres and yolk shared the same set of dominant species. The fatty acyl composition of each PC was then acquired via –MS/MS analysis of their acetate adducts ([PC + CH3COO]−). Using this strategy, we found that PC 38:5 was a mixture of PC 16:0_22:5, PC 18:1_20:4, and PC 18:0_20:5 (Figure 2e). Similarly, PC 38:6 was not pure PC 16:0_22:6 either [30]. Rather, it was composed of PC 16:0_22:6 and PC 18:1_20:5 (SI, page 31).
Lipid profiling with C=C specificity in a developing zebrafish embryo. Mass spectra of the lipid extracts from (a) blastomeres and (b) yolk of a single zebrafish embryo at 4-cell stage. (c, d) After photochemical derivatization, PB reaction products of unsaturated lipids were identified via a 58-Da mass shift. (e) Fragmentation of the acetate adduct of PC 38:5 released fatty acyls of C16:0, C18:1, C18:0, C20:5, C20:4, and C22:5; asterisk denotes 44 Da (CO2) loss from C20:5. (f) Tandem mass spectrum of PB products of PC 38:5 at m/z 866.6. Several sets of diagnostic ions specific to C20:5 (5, 8, 11, 14, 17), C22:5 (7, 10, 13, 16, 19), C20:4 (5, 8, 11, 14), C18:1 (9), and C18:1 (11) were detected. (g) PC 38:5 was a mixture of four fatty acyl composition and C=C location isomers. The sn-position of fatty acyls and C=C cis/trans geometry could not be determined
A more detailed layer of lipid structural information was acquired by locating the C=Cs in fatty acyls. Following rapid (< 2 min) photochemical PB derivatization, unsaturated lipids shifted to higher mass range after a 58-Da mass increase due to acetone addition (Figure 2c, d). For instance, several pairs of C=C-specific diagnostic ions were generated from PB products of PC 38:5 (PBPC 38:5) at m/z 866.6 (Figure 2f). Diagnostic ions at m/z 622/648, 662/688, 702/728, 742/768, and 782/808 were assigned to the five C=Cs at Δ5, 8, 11, 14, and 17 in C20:5 and those at Δ7, 10, 13, 16, and 19 in C22:5, while those at m/z 620/646, 660/686, 700/726, and 740/766 were assigned to Δ5, 8, 11, and 14 within C20:4. All these polyunsaturated fatty acyls (C20:4, C20:5, and C22:5) consisted of a single isomer. By contrast, consistent with previous studies of human and rat samples [48, 49], C18:1 consisted of Δ9 and Δ11 isomers, with C=C-specific ions at m/z 698/724 and 726/752, respectively. It is worth noting that the ions at m/z 726.47 from the Δ11 C=C in C18:1 and m/z 726.50 from the Δ11 C=C in C20:4 were well resolved by the Orbitrap MS at a mass resolution of 70,000 (Figure S2), which enabled accurate calculation of abundance ratios of diagnostic ions. Therefore, PC 38:5 in the zebrafish embryo was a mixture of PC 16:0_22:5 (7, 10, 13, 16, 19), PC 18:1 (9)_20:4 (5, 8, 11, 14), PC 18:1 (11)_20:4 (5, 8, 11, 14), and PC 18:0_20:5 (5, 8, 11, 14, 17) (Figure 2g).
In this way, over fifty lysoPCs (LPCs) and PCs were identified with fatty acyl compositions and C=C locations from a single embryo (Table S1). Monounsaturated fatty acyls of C16:1 and C18:1 were each found to contain two C=C location isomers, i.e., C16:1 (7)/C16:1 (9) and C18:1 (9)/C18:1 (11). Polyunsaturated fatty acyls existed in only one isomeric form: For C18:3, C20:5, C22:5, and C22:6, only the ω-3 isomers were detected, while C18:2 and C20:4 solely existed as ω-6 forms. A set of phosphatidylethanolamine (PE) and phosphatidylinositol (PI) species were also detected and characterized (Figure S3 and Table S2). Specifically, the C=C localization of PEs and PIs was achieved using the LC-PB-MS/MS platform (see Supplemental Information for details) [39].
Collectively, the structural characterization of GPs with C=C specificity was achieved by identifying fatty acyls and C=C locations within them, presenting a significant improvement in lipid structural elucidation. Our results thus revealed a remarkable structural diversity of embryo lipids in terms of fatty acyl composition and C=C location. The intrinsic diversity of maternally deposited yolk lipids was pivotal to support myriad developmental needs of the increasingly specified embryonic cells.
Alterations in Lipid C=C Location Isomers Unveiled during Zebrafish Embryogenesis
The relative amounts of C=C location isomers for C16:1- and C18:1-containing lipids were monitored during embryonic development. As mentioned before, relative amount of a structurally defined lipid isomer was achieved by calculating the intensity ratio between C=C-specific diagnostic ions. For instance, diagnostic ions at m/z 622/648 and 650/676 (Figure 3a, b) were observed for PBPC 16:0_16:1 (m/z 790.6), indicative of the presence of PC 16:0_16:1 (7) and PC 16:0_16:1 (9) (Figure S4A). The relative amount of PC 16:0_16:1 (7) was calculated as the intensity ratio between its C=C-specific ions and C=C-specific ions to all C=C location isomers, i.e., (I622 + I648)/(I622 + I648 + I650 + I676).
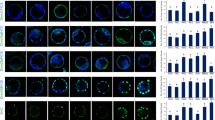
PB-MS/MS analysis of PC 32:1 in the (a) blastomeres and (b) yolk gave rise to two pairs of diagnostic ions, which led to the identification of PC 16:0_16:1 (7) and PC 16:0_16:1 (9). PB-MS/MS analysis of PC 34:1 in the (c) blastomeres and (d) yolk also produced two pairs of diagnostic ions indicative of the presence of PC 16:0_18:1 (9) and PC 16:0_18:1 (11). Differences in the relative amounts of (e) PC 16:0_16:1 (7) and (f) PC 16:0_18:1 (9) were revealed in blastomeres/yolk from 1-cell to 4 hpf and head/tail/yolk from 24 hpf onwards. N = 5 for stages from 1-cell to 4 hpf and N = 6 for 24 hpf, 72 hpf, and 5 dpf stages; *p < 0.05, **p < 0.01, ***p < 0.001 (Student’s two-tailed t test)
Notable differences in the C=C location isomer compositions of PC 16:0_16:1 were found between the blastomeres and yolk. In the blastomeres, the level of PC 16:0_16:1 (7) was significantly elevated than in the yolk from 10–15% to 26–34% (Figure 3e). The presence of C16:1 (7), most likely derived from β-oxidation of C18:1 (9) [50], was rarely reported and its function was not understood well [51, 52]. Conventionally, mitochondrial fatty acid β-oxidation (FAO) pathway is important to maintain energy homoeostasis in organs including heart, liver, and skeletal muscle [53]. However, the shortening of C18:1 (9) to C16:1 (7) via β-oxidation should not be simply considered as an energy production process but a pathway for lipid remodeling, and C16:1 (7) was not merely an intermediate of C18:1 degradation. An additional evidence to support a specific biological role of C16:1 (7) was that the composition of C16:1 C=C location isomers was fairly stable in the blastomeres and yolk during the first 4 h of development, indicating a tight regulation on its production. Even at 24 hpf, separate lipid profiling of the head and tail revealed that the C=C isomer compositions of PC 16:0_16:1 were basically unchanged (30% in the head vs. 31% in the tail) compared with that in the blastomeres prior to 4 hpf. Only at 72 hpf and 5 dpf, a notable decrease of the Δ7 isomer to 15% was observed in the tail, while little changed in that of the head (Figure 3e).
Similar analysis was carried out for the much more abundant PC 34:1, which contained almost exclusively PC 16:0_18:1 (SI, page 19). As aforementioned, C18:1 consisted of Δ9 and Δ11 isomers (Figure 3c, d and Figure S4B). The relative amount of C18:1 (9) was, however, only slightly higher in the blastomeres (85–87%) than in the yolk (79–85%). From 24 hpf to 5dpf, the relative amount of C18:1 (9) in the tail kept decreasing from 83 to 75%, which was notably lower than that in the head (82% at 5 dpf, Figure 3f). Consequently, using the developed method, we have achieved the monitoring of temporal changes of structurally defined lipids during zebrafish embryogenesis, which were meticulously regulated during different developmental stages.
Alterations in Both Fatty Acyl Composition and C=C Location Isomers for Polyunsaturated Lipids
Besides C=C location isomers, we also found distinctive compositions of fatty acyl isomers. For instance, PC 34:2 in the embryo was revealed to be a mixture of PC 16:0_18:2 and the more abundant PC 16:1_18:1 (SI, page 20). We found that the level of PC 16:1_18:1 in PC 34:2 was significantly upregulated in the blastomeres than in the yolk (79–88% vs. 63–70%) prior to 4 hpf (Figure S5A-B). Afterwards, its amount decreased in the tail (69% at 5 dpf) and more obviously in the head (57% at 5 dpf) (Figure 4a). Likewise, PC 36:2 was a mixture of PC 18:0_18:2 and PC 18:1_18:1 (SI, page 27), with the latter consistently more concentrated in the blastomeres and embryo body (95–99%) than in the yolk (ca. 90%, Figure 4c and S5C-D). Interestingly, the distributions of fatty acyl composition isomers for PC 34:2 and PC 36:2 had the same trend; viz., both PC 16:1_18:1 and PC 18:1_18:1 were more concentrated in the blastomeres and embryo body. The underlying molecular mechanism and biological implications of these fatty acyl isomer differences remain unclear. It is speculated that such a lipid remodeling of yolk lipids ensures appropriate lipid content required for embryonic cell division and membrane construction, although verification of this hypothesis requires extensive research.
Variations in the distributions of fatty acyl composition and C=C location isomers of (a, b) PC 34:2 and (c, d) PC 36:2. N = 5 for stages from 1-cell to 4 hpf and N = 6 for 24 hpf, 72 hpf, and 5 dpf stages; *p < 0.05, **p < 0.01, ***p < 0.001 (Student’s two-tailed t test). (e) Relative amounts of C18:1 (9) in eight C18:1-containing lipids in the yolk monitored from 1-cell to 72 hpf stages (N = 5). Isomeric compositions of C18:1 in most species were basically unchanged
Furthermore, variations were also uncovered in the C=C location isomer compositions of PC 16:1_18:1 and PC 18:1_18:1. For PC 16:1_18:1, diagnostic ions for Δ7 and Δ9 isomers of C16:1 were identical with those for Δ9 and Δ11 isomers of C18:1 (Figure S5A-B and Figure S6). As C16:1 (7) was the β-oxidation product of C18:1 (9), we chose to sum their C=C-specific ion abundances to represent their total amount. The relative amounts of C16:1 (7) and C18:1 (9) were notably higher in the blastomeres (55–62%) than in the yolk (41–45%). From 24 to 72 hpf, the relative amounts of C16:1 (7) and C18:1 (9) decreased in both embryo head and tail, but they were still higher than those of the yolk (Figure 4b). For PC 16:0_16:1, C16:1 (7) was found to be more concentrated in blastomeres, while for PC 16:0_18:1, the relative amounts of C18:1 (9) were quite similar in the blastomeres and yolk. The higher relative amount of C16:1 (7) and C18:1 (9) of PC 16:1_18:1 in the blastomeres, therefore, was also consistent with the above observations. As for PC 18:1_18:1, C=C location isomer profiles were close between the blastomeres and yolk (66–70% vs. 63–65%). Even after 24 hpf, very little changes in isomer ratios were observed among the head, tail, and yolk (Figure 4d).
Remarkable Consistency in Yolk Lipid Profiles
The coexistence of C18:1 (9) and C18:1 (11), found in all C18:1-containing lipids, was the most prevalent pair of C=C location isomers in zebrafish embryos. We monitored eight C18:1-containing lipids, namely LPC 18:1, PC 18:0_18:1, PC 18:1_18:2, PC 18:1_18:3, PC 18:1_20:5, PC 18:1_20:4, PC 18:1_22:6, and PC 18:1_22:5, in the yolk of single embryos by measuring the relative amount of C18:1 Δ9 and Δ11 isomers. Our data revealed that isomer compositions of these lipids were remarkably stable over the entire developmental process, except for LPC 18:1 (Figure 4e). In addition, for PCs that contained C18:1 and another polyunsaturated fatty acyl (e.g., C20:5 or C22:6), the relative amounts of C18:1 (9) and C18:1 (11) were roughly equal. By contrast, C18:1 (9) was significantly elevated in PCs containing a saturated fatty acyl (e.g. C16:0 or C18:0), suggesting a selectivity towards the incorporation of fatty acyl C=C location isomer in PC biosynthesis, which was affected by the other fatty acyl attached to the PC backbone.
Lastly, to evaluate inter-day reproducibility, we compared the fatty acyl and C=C location isomer compositions of two sets of yolk samples collected on different days, both at 24 hpf stage. As shown in Figure S7, day-to-day variations were essentially negligible, strongly suggesting that the PC isomer compositions were tightly regulated metabolically. In this study, small standard deviations (SDs, typically < 3%) were consistently found in the determination of isomer compositions. It is the high accuracy and analytical reproducibility for lipid isomer analysis that has enabled the monitoring of subtle lipid alterations with high precision.
Conclusions
In summary, we have achieved for the first time a deep lipidomic monitoring with C=C specificity for a developing zebrafish embryo from zygote to early larval stage. Relative amounts of C=C location isomers of unsaturated PCs were extremely consistent in the yolk as the nutrient source. Although the embryo body, as the destination of lipid transport, shared the same set of unsaturated lipids with yolk throughout embryogenesis, alterations in the compositions of PC fatty acyl and C=C location isomers were found in the blastomeres and maintained a homeostasis in early-stage embryogenesis (prior to 4hpf), which strongly suggested that the lipid remodeling process took place even at the C=C location level to maintain a specific composition of lipid structural moieties essential for membrane construction, cell division, and differentiation. Additionally, the divergent lipid isomer profiles of embryo head and tail from 24 hpf to 5 dpf were the results of both specific developmental needs and gradual depletion of yolk lipids. Altogether, our study opens a new avenue for detailed structural analysis and monitoring of lipids in the dynamic context of vertebrate development. More importantly, as lipid functions are ultimately determined by the lipid structure, this study should promote the functional studies of lipids in biological systems
References
van Meer, G., Voelker, D.R., Feigenson, G.W.: Membrane lipids: where they are and how they behave. Nat. Rev. Mol. Cell Biol. 9, 112–124 (2008)
Sturmey, R.G., Reis, A., Leese, H.J., McEvoy, T.G.: Role of fatty acids in energy provision during oocyte maturation and early embryo development. Reprod. Domest. Anim. 44(Suppl 3), 50–58 (2009)
Elouarrat, D., van der Velden, Y.U., Jones, D.R., Moolenaar, W.H., Divecha, N., Haramis, A.P.: Role of phosphatidylinositol 5-phosphate 4-kinase alpha in zebrafish development. Int. J. Biochem. Cell Biol. 45, 1293–1301 (2013)
Bieberich, E.: Ceramide in stem cell differentiation and embryo development: novel functions of a topological cell-signaling lipid and the concept of ceramide compartments. J. Lipids. 2011, 610306 (2011)
Ferreira, C.R., Saraiva, S.A., Catharino, R.R., Garcia, J.S., Gozzo, F.C., Sanvido, G.B., Santos, L.F., Lo Turco, E.G., Pontes, J.H., Basso, A.C., Bertolla, R.P., Sartori, R., Guardieiro, M.M., Perecin, F., Meirelles, F.V., Sangalli, J.R., Eberlin, M.N.: Single embryo and oocyte lipid fingerprinting by mass spectrometry. J. Lipid Res. 51, 1218–1227 (2010)
Gonzalez-Serrano, A.F., Pirro, V., Ferreira, C.R., Oliveri, P., Eberlin, L.S., Heinzmann, J., Lucas-Hahn, A., Niemann, H., Cooks, R.G.: Desorption electrospray ionization mass spectrometry reveals lipid metabolism of individual oocytes and embryos. PLoS One. 8, e74981 (2013)
Ferreira, C.R., Jarmusch, A.K., Pirro, V., Alfaro, C.M., Gonzalez-Serrano, A.F., Niemann, H., Wheeler, M.B., Rabel, R.A., Hallett, J.E., Houser, R., Kaufman, A., Cooks, R.G.: Ambient ionisation mass spectrometry for lipid profiling and structural analysis of mammalian oocytes, preimplantation embryos and stem cells. Reprod. Fertil. Dev. 27, 621–637 (2015)
Tian, H., Fletcher, J.S., Thuret, R., Henderson, A., Papalopulu, N., Vickerman, J.C., Lockyer, N.P.: Spatiotemporal lipid profiling during early embryo development of Xenopus laevis using dynamic ToF-SIMS imaging. J. Lipid Res. 55, 1970–1980 (2014)
Rustam, Y.H., Reid, G.E.: Analytical challenges and recent advances in mass spectrometry based lipidomics. Anal. Chem. 90, 374–397 (2018)
Porta Siegel, T., Ekroos, K., Ellis, S.R.: Reshaping lipid biochemistry by pushing barriers in structural lipidomics. Angew. Chem. Int. Ed. 58, 6492–6501 (2019)
Williams, P.E., Klein, D.R., Greer, S.M., Brodbelt, J.S.: Pinpointing double bond and sn-positions in glycerophospholipids via hybrid 193 nm ultraviolet photodissociation (UVPD) mass spectrometry. J. Am. Chem. Soc. 139, 15681–15690 (2017)
Paine, M.R.L., Poad, B.L.J., Eijkel, G.B., Marshall, D.L., Blanksby, S.J., Heeren, R.M.A., Ellis, S.R.: Mass spectrometry imaging with isomeric resolution enabled by ozone-induced dissociation. Angew. Chem. Int. Ed. 57, 10530–10534 (2018)
Deng, J., Yang, Y., Liu, Y., Fang, L., Lin, L., Luan, T.: Coupling Paterno-Buchi reaction with surface-coated probe nanoelectrospray ionization mass spectrometry for in vivo and microscale profiling of lipid C=C location isomers in complex biological tissues. Anal. Chem. 91, 4592–4599 (2019)
Howe, K., Clark, M.D., Torroja, C.F., Torrance, J., Berthelot, C., Muffato, M., Collins, J.E., Humphray, S., McLaren, K., Matthews, L., McLaren, S., Sealy, I., Caccamo, M., Churcher, C., Scott, C., Barrett, J.C., Koch, R., Rauch, G.J., White, S., Chow, W., Kilian, B., Quintais, L.T., Guerra-Assuncao, J.A., Zhou, Y., Gu, Y., Yen, J., Vogel, J.H., Eyre, T., Redmond, S., Banerjee, R., Chi, J., Fu, B., Langley, E., Maguire, S.F., Laird, G.K., Lloyd, D., Kenyon, E., Donaldson, S., Sehra, H., Almeida-King, J., Loveland, J., Trevanion, S., Jones, M., Quail, M., Willey, D., Hunt, A., Burton, J., Sims, S., McLay, K., Plumb, B., Davis, J., Clee, C., Oliver, K., Clark, R., Riddle, C., Elliot, D., Threadgold, G., Harden, G., Ware, D., Begum, S., Mortimore, B., Kerry, G., Heath, P., Phillimore, B., Tracey, A., Corby, N., Dunn, M., Johnson, C., Wood, J., Clark, S., Pelan, S., Griffiths, G., Smith, M., Glithero, R., Howden, P., Barker, N., Lloyd, C., Stevens, C., Harley, J., Holt, K., Panagiotidis, G., Lovell, J., Beasley, H., Henderson, C., Gordon, D., Auger, K., Wright, D., Collins, J., Raisen, C., Dyer, L., Leung, K., Robertson, L., Ambridge, K., Leongamornlert, D., McGuire, S., Gilderthorp, R., Griffiths, C., Manthravadi, D., Nichol, S., Barker, G., Whitehead, S., Kay, M., Brown, J., Murnane, C., Gray, E., Humphries, M., Sycamore, N., Barker, D., Saunders, D., Wallis, J., Babbage, A., Hammond, S., Mashreghi-Mohammadi, M., Barr, L., Martin, S., Wray, P., Ellington, A., Matthews, N., Ellwood, M., Woodmansey, R., Clark, G., Cooper, J., Tromans, A., Grafham, D., Skuce, C., Pandian, R., Andrews, R., Harrison, E., Kimberley, A., Garnett, J., Fosker, N., Hall, R., Garner, P., Kelly, D., Bird, C., Palmer, S., Gehring, I., Berger, A., Dooley, C.M., Ersan-Urun, Z., Eser, C., Geiger, H., Geisler, M., Karotki, L., Kirn, A., Konantz, J., Konantz, M., Oberlander, M., Rudolph-Geiger, S., Teucke, M., Lanz, C., Raddatz, G., Osoegawa, K., Zhu, B., Rapp, A., Widaa, S., Langford, C., Yang, F., Schuster, S.C., Carter, N.P., Harrow, J., Ning, Z., Herrero, J., Searle, S.M., Enright, A., Geisler, R., Plasterk, R.H., Lee, C., Westerfield, M., de Jong, P.J., Zon, L.I., Postlethwait, J.H., Nusslein-Volhard, C., Hubbard, T.J., Roest Crollius, H., Rogers, J., Stemple, D.L.: The zebrafish reference genome sequence and its relationship to the human genome. Nature. 496, 498–503 (2013)
MacRae, C.A., Peterson, R.T.: Zebrafish as tools for drug discovery. Nat. Rev. Drug Discov. 14, 721–731 (2015)
Roscioli, T., Kamsteeg, E.J., Buysse, K., Maystadt, I., van Reeuwijk, J., van den Elzen, C., van Beusekom, E., Riemersma, M., Pfundt, R., Vissers, L.E., Schraders, M., Altunoglu, U., Buckley, M.F., Brunner, H.G., Grisart, B., Zhou, H., Veltman, J.A., Gilissen, C., Mancini, G.M., Delree, P., Willemsen, M.A., Ramadza, D.P., Chitayat, D., Bennett, C., Sheridan, E., Peeters, E.A., Tan-Sindhunata, G.M.: de Die-Smulders, C.E., Devriendt, K., Kayserili, H., El-Hashash, O.A., Stemple, D.L., Lefeber, D.J., Lin, Y.Y., van Bokhoven, H.: Mutations in ISPD cause Walker-Warburg syndrome and defective glycosylation of alpha-dystroglycan. Nat. Genet. 44, 581–585 (2012)
Golzio, C., Willer, J., Talkowski, M.E., Oh, E.C., Taniguchi, Y., Jacquemont, S., Reymond, A., Sun, M., Sawa, A., Gusella, J.F., Kamiya, A., Beckmann, J.S., Katsanis, N.: KCTD13 is a major driver of mirrored neuroanatomical phenotypes of the 16p11.2 copy number variant. Nature. 485, 363–367 (2012)
Tobin, D.M., Roca, F.J., Oh, S.F., McFarland, R., Vickery, T.W., Ray, J.P., Ko, D.C., Zou, Y., Bang, N.D., Chau, T.T., Vary, J.C., Hawn, T.R., Dunstan, S.J., Farrar, J.J., Thwaites, G.E., King, M.C., Serhan, C.N., Ramakrishnan, L.: Host genotype-specific therapies can optimize the inflammatory response to mycobacterial infections. Cell. 148, 434–446 (2012)
Bakkers, J.: Zebrafish as a model to study cardiac development and human cardiac disease. Cardiovasc. Res. 91, 279–288 (2011)
Gut, P., Reischauer, S., Stainier, D.Y.R., Arnaout, R.: Little fish, big data: zebrafish as a model for cardiovascular and metabolic disease. Physiol. Rev. 97, 889–938 (2017)
Farrell, J.A., Wang, Y., Riesenfeld, S.J., Shekhar, K., Regev, A., Schier, A.F.: Single-cell reconstruction of developmental trajectories during zebrafish embryogenesis. Science. 360, 6392, eaar3131 (2018)
Wagner, D.E., Weinreb, C., Collins, Z.M., Briggs, J.A., Megason, S.G., Klein, A.M.: Single-cell mapping of gene expression landscapes and lineage in the zebrafish embryo. Science. 360, 981–987 (2018)
Lucitt, M.B., Price, T.S., Pizarro, A., Wu, W., Yocum, A.K., Seiler, C., Pack, M.A., Blair, I.A., Fitzgerald, G.A., Grosser, T.: Analysis of the zebrafish proteome during embryonic development. Mol. Cell. Proteomics. 7, 981–994 (2008)
Holtta-Vuori, M., Salo, V.T., Nyberg, L., Brackmann, C., Enejder, A., Panula, P., Ikonen, E.: Zebrafish: gaining popularity in lipid research. Biochem. J. 429, 235–242 (2010)
Anderson, J.L., Carten, J.D., Farber, S.A.: Zebrafish lipid metabolism: from mediating early patterning to the metabolism of dietary fat and cholesterol. Methods Cell Biol. 101, 111–141 (2011)
Miyares, R.L., de Rezende, V.B., Farber, S.A.: Zebrafish yolk lipid processing: a tractable tool for the study of vertebrate lipid transport and metabolism. Dis. Model. Mech. 7, 915–927 (2014)
Zeituni, E.M., Farber, S.A.: Studying lipid metabolism and transport during zebrafish development. Methods Mol. Biol. 1451, 237–255 (2016)
Dutta, A., Sinha, D.K.: Zebrafish lipid droplets regulate embryonic ATP homeostasis to power early development. Open Biol. 7, (2017)
Yao, Y., Sun, S., Wang, J., Fei, F., Dong, Z., Ke, A.W., He, R., Wang, L., Zhang, L., Ji, M.B., Li, Q., Yu, M., Shi, G.M., Fan, J., Gong, Z., Wang, X.: Canonical Wnt signaling remodels lipid metabolism in zebrafish hepatocytes following Ras oncogenic insult. Cancer Res. 78, 5548–5560 (2018)
Fraher, D., Sanigorski, A., Mellett, N.A., Meikle, P.J., Sinclair, A.J., Gibert, Y.: Zebrafish embryonic lipidomic analysis reveals that the yolk cell is metabolically active in processing lipid. Cell Rep. 14, 1317–1329 (2016)
Duenas, M.E., Essner, J.J., Lee, Y.J.: 3D MALDI mass spectrometry imaging of a single cell: spatial mapping of lipids in the embryonic development of zebrafish. Sci. Rep. 7, 14946 (2017)
Pirro, V., Guffey, S.C., Sepulveda, M.S., Mahapatra, C.T., Ferreira, C.R., Jarmusch, A.K., Cooks, R.G.: Lipid dynamics in zebrafish embryonic development observed by DESI-MS imaging and nanoelectrospray-MS. Mol. BioSyst. 12, 2069–2079 (2016)
Babin, P.J., Thisse, C., Durliat, M., Andre, M., Akimenko, M.-A., Thisse, B.: Both apolipoprotein E and A-I genes are present in a nonmammalian vertebrate and are highly expressed during embryonic development. Proc. Natl. Acad. Sci. U. S. A. 94, 8622–8627 (1997)
Otis, J.P., Zeituni, E.M., Thierer, J.H., Anderson, J.L., Brown, A.C., Boehm, E.D., Cerchione, D.M., Ceasrine, A.M., Avraham-Davidi, I., Tempelhof, H., Yaniv, K., Farber, S.A.: Zebrafish as a model for apolipoprotein biology: comprehensive expression analysis and a role for ApoA-IV in regulating food intake. Dis. Model. Mech. 8, 295–309 (2015)
Büchi, G., Inman, C.G., Lipinsky, E.S.: Light-catalyzed organic reactions. I. The reaction of carbonyl compounds with 2-methyl-2-butene in the presence of ultraviolet light. J. Am. Chem. Soc. 76, 4327–4331 (1954)
Ma, X., Xia, Y.: Pinpointing double bonds in lipids by Paterno-Buchi reactions and mass spectrometry. Angew. Chem. Int. Ed. 53, 2592–2596 (2014)
Ma, X., Chong, L., Tian, R., Shi, R., Hu, T.Y., Ouyang, Z., Xia, Y.: Identification and quantitation of lipid C=C location isomers: a shotgun lipidomics approach enabled by photochemical reaction. Proc. Natl. Acad. Sci. U. S. A. 113, 2573–2578 (2016)
Bednarik, A., Bolsker, S., Soltwisch, J., Dreisewerd, K.: An on-tissue Paterno-Buchi reaction for localization of carbon-carbon double bonds in phospholipids and glycolipids by matrix-assisted laser-desorption-ionization mass-spectrometry imaging. Angew. Chem. Int. Ed. 57, 12092–12096 (2018)
Zhang, W., Zhang, D., Chen, Q., Wu, J., Ouyang, Z., Xia, Y.: Online photochemical derivatization enables comprehensive mass spectrometric analysis of unsaturated phospholipid isomers. Nat. Commun. 10, 79 (2019)
Xu, T., Pi, Z., Song, F., Liu, S., Liu, Z.: Benzophenone used as the photochemical reagent for pinpointing C=C locations in unsaturated lipids through shotgun and liquid chromatography-mass spectrometry approaches. Anal. Chim. Acta. 1028, 32–44 (2018)
Jeck, V., Korf, A., Vosse, C., Hayen, H.: Localization of double-bond positions in lipids by tandem mass spectrometry succeeding high-performance liquid chromatography with post-column derivatization. Rapid Commun. Mass Spectrom. 33, 86–94 (2019)
Esch, P., Heiles, S.: Charging and charge switching of unsaturated lipids and apolar compounds using Paterno-Buchi reactions. J. Am. Soc. Mass Spectrom. 29, 1971–1980 (2018)
Waldchen, F., Becher, S., Esch, P., Kompauer, M., Heiles, S.: Selective phosphatidylcholine double bond fragmentation and localisation using Paterno-Buchi reactions and ultraviolet photodissociation. Analyst. 142, 4744–4755 (2017)
Korf, A., Jeck, V., Schmid, R., Helmer, P.O., Hayen, H.: Lipid species annotation at double bond position level with custom databases by extension of the MZmine 2 open-source software package. Anal. Chem. 91, 5098–5105 (2019)
Wäldchen, F., Spengler, B., Heiles, S.: Reactive matrix-assisted laser desorption/ionization mass spectrometry imaging using an intrinsically photoreactive Paternò–Büchi matrix for double-bond localization in isomeric phospholipids. J. Am. Chem. Soc. 141, 11816–11820 (2019)
Kimmel, C.B., Ballard, W.W., Kimmel, S.R., Ullmann, B., Schilling, T.F.: Stages of embryonic development of the zebrafish. Dev. Dyn. 203, 253–310 (1995)
Bligh, E.G., Dyer, W.J.: A rapid method of total lipid extraction and purification. Can. J. Biochem. Physiol. 37, 911–917 (1959)
Ren, J., Franklin, E.T., Xia, Y.: Uncovering structural diversity of unsaturated fatty acyls in cholesteryl esters via photochemical reaction and tandem mass spectrometry. J. Am. Soc. Mass Spectrom. 28, 1432–1441 (2017)
Marshall, D.L., Criscuolo, A., Young, R.S.E., Poad, B.L.J., Zeller, M., Reid, G.E., Mitchell, T.W., Blanksby, S.J.: Mapping unsaturation in human plasma lipids by data-independent ozone-induced dissociation. J. Am. Soc. Mass Spectrom. 30 (9), 1621–1630 (2019)
Guijas, C., Meana, C., Astudillo, A.M., Balboa, M.A., Balsinde, J.: Foamy monocytes are enriched in cis-7-hexadecenoic fatty acid (16:1n-9), a possible biomarker for early detection of cardiovascular disease. Cell Chem. Biol. 23, 689–699 (2016)
Ma, W., Wu, J.H., Wang, Q., Lemaitre, R.N., Mukamal, K.J., Djoussé, L., King, I.B., Song, X., Biggs, M.L., Delaney, J.A., Kizer, J.R., Siscovick, D.S., Mozaffarian, D.: Prospective association of fatty acids in the de novo lipogenesis pathway with risk of type 2 diabetes: the Cardiovascular Health Study. Amer. J. Clin. Nutri. 101, 153–163 (2014)
Dai, L., Gonçalves, C.M.V., Lin, Z., Huang, J., Lu, H., Yi, L., Liang, Y., Wang, D., An, D.: Exploring metabolic syndrome serum free fatty acid profiles based on GC–SIM–MS combined with random forests and canonical correlation analysis. Talanta. 135, 108–114 (2015)
Houten, S.M., Wanders, R.J.: A general introduction to the biochemistry of mitochondrial fatty acid beta-oxidation. J. Inherit. Metab. Dis. 33, 469–477 (2010)
Acknowledgements
We acknowledge Simin Cheng for her help on the LC-PB-MS/MS platform. This research was financially supported by the National Natural Science Foundation of China (Grants 21705091, 21621003, and 21727813) and the Ministry of Science and Technology of the People’s Republic of China (Grant 2016YFF0100301).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Fish maintenance and embryo raising followed the ethical guidelines of the Tsinghua University Animal Care and Use Committee.
Conflict of Interest
The authors declare that they have no conflicts of interest.
Electronic supplementary material
ESM 1
(DOCX 10.1 MB)
Rights and permissions
About this article
Cite this article
Zhao, X., Chen, J., Zhang, W. et al. Lipid Alterations during Zebrafish Embryogenesis Revealed by Dynamic Mass Spectrometry Profiling with C=C Specificity. J. Am. Soc. Mass Spectrom. 30, 2646–2654 (2019). https://doi.org/10.1007/s13361-019-02334-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13361-019-02334-z