Abstract
A strategy to sequence lysine-containing cyclic peptides by MSn is presented. Doubly protonated cyclic peptides ions are transformed into gold (I) cationized peptide ions via cation switching ion/ion reaction. Gold(I) cationization facilitates the oxidation of neutral lysine residues in the gas phase, weakening the adjacent amide bond. Upon activation, facile cleavage N-terminal to the oxidized lysine residue provides a site-specific ring opening pathway that converts cyclic peptides into acyclic analogs. The ensuing ion contains a cyclic imine as the new N-terminus and an oxazolone, or structural equivalent, as the new C-terminus. Product ions are formed from subsequent fragmentation events of the linearized peptide ion. Such an approach simplifies MS/MS data interpretation as a series of fragment ions with common N- and C-termini are generated. Results are presented for two cyclic peptides, sunflower trypsin inhibitor and the model cyclic peptide, β-Loop. The power of this strategy lies in the ability to generate the oxidized peptide, which is easily identified via the loss of HAuNH3 from [M + Au]+. While some competitive processes are observed, the site of ring opening can be pinpointed to the lysine residue upon MS4 enabling the unambiguous sequencing of cyclic peptides.

Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Cyclic peptides are a class of biomolecules that are more difficult to sequence by mass spectrometry than their linear counterparts. While encountered less often than linear peptides, cyclic peptides represent a body of bioactive natural products and synthetics whose structures must be characterized. Cyclotides, for example, are macrocyclic peptides comprised of a head-to-tail cyclic backbone and three intramolecular disulfide bonds forming a cyclic cysteine knot [1, 2]. This motif instills remarkable thermal, chemical, and enzymatic stability [3, 4] with promising applications in therapeutics and agriculture [5,6,7,8,9,10,11,12,13,14]. Similarly, sunflower trypsin inhibitor (SFTI) analogs, the simplest head-to-tail cyclic peptides containing a single disulfide bond, for example, have been examined as inflammatory bowel disease drug candidates [15], mammalian melanocortin receptor agonists [16], autoantibody scavengers [17], mesotrypsin inhibitors [18], and plasmin inhibitors [19].
Dating back to the early days of cyclic peptide analysis, nuclear magnetic resonance (NMR) techniques have emerged as, perhaps, the primary means of characterization [20,21,22] and remain a popular choice today [23,24,25,26]. However, NMR is not well-suited to sequencing peptides and generally requires multiple milligrams of purified sample. Mass spectrometry-based techniques, on the other hand, are commonly used for peptide sequencing and are attractive for their relatively small sample size and minimal purity requirements. Gross and co-workers first demonstrated the utility of tandem mass spectrometry for cyclic peptide analysis in 1982 [27] and continued to pioneer MS/MS approaches into the early 2000s. Despite its continued use [28,29,30,31], tandem mass spectrometry of cyclic peptides remains challenging, particularly regarding data interpretation. Sequence information of a linear peptide is derived via the predictable dissociation of the peptide ion along the amide backbone. Specifically, b- and y-fragment ions are typically generated upon collisional activation of a linear peptide; these fragment ions can be used to elucidate the primary structure [32]. MSn of cyclic peptides, on the other hand, requires the cleavage of two amide bonds to generate any observable fragment ions. In principle, ring opening can occur at any residue creating linear peptide isomers of identical mass complicating the interpretation of the subsequent product ion spectrum. This spectral complexity is exacerbated with UVPD as a-, b-, c- and x-, y-, z-type ions may be formed [30, 31]. It is widely known, however, that backbone cleavages adjacent to particular amino acids can be preferred under some conditions. For example, the facile opening N-terminal to proline residues upon collisional activation has been exploited to aid in structural elucidation of cyclic peptides by mass spectrometry [33, 34].
The MS/MS sequencing of cyclic peptides can be facilitated through the incorporation of a site-specific ring opening resulting in fragment ions that contain a common N-terminus. For example, several reports of solution-based linearization via enzymatic digestion have appeared [35,36,37]. These methods, however, are not universal as cyclic peptides can show remarkable resistance to enzymatic digestion [3]. Recently, Brodbelt and co-workers have reported an analogous gas-phase strategy to linearize stapled and cyclic peptides by taking advantage of the “ornithine effect” [38, 39]. Conversion of arginine to ornithine is accomplished through solution-phase deguanidination in the presence of hydrazine. Collisional activation of the ornithine containing cyclic peptide resulted in selective fragmentation C-terminal to the ornithine residue, offering a gas-phase approach to site-selective linearization, enhancing cyclic peptide characterization.
In the present work, we use gold (I) cationization to promote site-specific ring opening of cyclic peptides. Helmut Schwarz is a pioneer in the study of gas-phase organometallic chemistry in general [40, 41], and he and his co-workers have described many of the unique characteristics of Au(I) chemistry in the gas phase [42]. The gas-phase organometallic chemistry of gold has been reviewed [43]. We have recently noted that collisional activation of [M + Au]+ precursor ions undergo gas-phase oxidation at neutral lysine residues resulting in a weakened C-N bond N-terminal to lysine [44]. For linear peptides, subsequent activation of the oxidized ion exhibits a facile fragmentation channel competitive with the proline effect. Incorporation of this “weak spot” into cyclic peptide ions via the loss of gold hydride and a molecule of ammonia presents an opportunity to selectively linearize cyclic peptides in the gas phase. Fragment ions containing a common N-terminus upon opening can simplify the MS/MS spectrum and aid in primary structure determination. While competitive ring opening sites can occur, we can further probe the loss of 109 Da to pinpoint the site of linearization at the oxidized lysine residue. This entirely gas-phase approach is demonstrated with sunflower trypsin inhibitor and a model cyclic peptide, β-Loop.
Experimental
Materials and Reagents
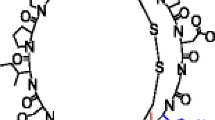
Ammonium bicarbonate, dithiothreitol (DTT), iodoacetamide, and gold (III) chloride were purchased from Sigma-Aldrich (St. Louis, MO, USA). HPLC-grade methanol and Optima LC/MS-grade water were purchased from Fisher Scientific (Fair Lawn, NJ, USA), and acetic acid was purchased from Mallinckrodt (Phillipsburg, NJ, USA). Reduced sunflower trypsin inhibitor (SFTI), cyclo-GRCTKSIPPICFPD, was synthesized by Biomatek (Wilmington, DE, USA). β-Loop, [45] cyclo-GRWQYV(D-Pro)GKFTVQ(D-Pro), was synthesized by the Gellman Lab of the University of Wisconsin Madison.
Sample Preparation
A gold (III) chloride stock solution was diluted to approximately 5 μM with methanol. β-Loop stock solution was prepared at a concentration of 250 μg/mL in an equal mixture of water and methanol. The stock was diluted 20-fold to a concentration of 12.5 μg/mL, approximately 7.5 μM, with 49.5:49.5:1, by volume, water/methanol/acetic acid. The sample was used without further purification.
Approximately 1 mg of SFTI was dissolved in 1 mL of reduction buffer (100 mM ammonium bicarbonate, 7 M urea) and incubated at 55 °C for 45 min with 5 mM DTT. After incubation, the solution was cooled to room temperature and then centrifuged briefly to collect any condensation. Fourteen microliters of a freshly prepared 500 mM iodoacetamide solution was added to the mixture and incubated at room temperature in the dark for 30 min. An additional 5 mM of DTT was added. The mixture was then incubated at room temperature for 15 min in the dark. Ten microliters of the reduced and alkylated SFTI was desalted using a TopTip C-18 desalting column (Glygen, Columbia, MD) as per the manufacturer’s instructions. The final solution concentration was approximately 6.5 μM.
Mass Spectrometry
All data were collected using a QTRAP 4000 hybrid triple quadrupole/linear ion trap mass spectrometer (Sciex, Concord, ON, Canada), previously modified for ion/ion reactions [46]. Reagent anions and analyte cations were introduced into the mass spectrometer via alternately pulsed nano-electrospray ionization (nESI) [47]. Cation switching ion/ion reactions involving gold have been described previously [44, 48, 49]. Briefly, the [AuCl2]− reagent anions were isolated in Q1 and transferred to Q2, followed immediately by injection of the isolated [M + 2H]2+ analyte cations. The ions were mutually stored in Q2 for up to 1000 ms, forming the [M + 2H + AuCl2]+ complex. Beam-type CID of the complex from q2 to Q3 resulted in the loss of two neutral HCl molecules, leaving the gold (I) cationized peptide ions, denoted [M + Au]+. MSn experiments were performed in Q3 where aurated ions were dissociated via resonance excitation at a q value of 0.2. Product ions were mass analyzed via mass-selective axial ejection (MSAE) [50]. Theoretical product ion masses were generated with CycloBranch and all product ions were verified manually [51]. Figure 1 shows the structures of the two cyclic peptides examined in this study.
Results and Discussion
This strategy aims to introduce a weak spot into cyclic peptides by virtue of gas-phase oxidation of neutral lysine residues in gold (I) cationized peptide ions. It has been shown that this oxidation occurs through the consecutive losses of gold hydride and a molecule of ammonia, in either order, generating a cyclic iminium ion and a weaker C-N bond [44]. Consequently, upon collisional activation, cleavage N-terminal to the oxidized lysine residue is observed as a preferred dissociation pathway. For cyclic peptides, it is expected that activation of the oxidized species, [M–H–NH3]+, results in the same site-selective fragmentation as their linear counterparts. This initial cleavage converts the cyclic peptide into an acyclic peptide with an imine at the N-terminus and an oxazolone ion, or structural equivalent (e.g., acylium ion) at the C-terminus. Linearization via the initial cleavage of the peptide results in no change in mass-to-charge. Thus, a second cleavage along the peptide backbone is required to form product ions. The described gas-phase ion/ion strategy is illustrated in Scheme 1.
Sunflower Trypsin Inhibitor
The cation switching ion/ion reaction between doubly protonated reduced and alkylated sunflower trypsin inhibitor and the gold dichloride reagent anion generates the [M + 2H + AuCl2]+ complex ion. Subsequent beam-type CID of the complex generates a prominent aurated peptide ion, [M + Au]+, at m/z 1825.8. As shown in Figure 2a, ion trap CID of the monoisotopically isolated [M + Au]+ ion results in dominant losses of 91 Da, likely arising from the even electron side chain loss of C2H5NOS (91.01 Da) from the carbamidomethyl cysteine residues to form dehydroalanine. Oxidation of lysine is evidenced by the loss of gold hydride and ammonia, designated as HAuNH3 (215.00 Da), thereby yielding the oxidized product indicated as [M–H–NH3]+. The oxidized product ion is present at only 5% relative abundance. Nonetheless, the species was isolated and subjected to additional activation.
CID of the [M–H–NH3]+ species generates the product ion spectrum shown in Figure 2b. The spectrum is comprised of sequence informative fragment ions and small molecule losses. Cyclic peptide fragment ions are labeled according to the nomenclature system of Ngoka and Gross [52]. Briefly, fragment ions are labeled using a four-part descriptor, xnJZ, where x is the type of backbone fragment ion, n is the number of amino acids in the fragment ion, and J/Z designates the site of ring opening. For example, y9TK denotes the y9 fragment ion generated from ring opening between the Thr-Lys bond. We note that a square superscript is associated with fragment ions that are shifted 19 Da lower in mass than their unoxidized counterparts (i.e. [b8–H–NH3]+ versus [b8 + H]+). Product ions formed through the preferential cleavage at lysine are highlighted in red and designated by the “TK” subscript; they represent approximately half of the structurally informative fragments. Among the other fragment ions are the b7IP and \( {\mathrm{b}}_{7\mathrm{DG}}^{\blacksquare } \). These ions are formed from cleavages C-terminal to an aspartic acid and N-terminal to a proline, two selective dissociation channels commonly observed in peptide ion tandem mass spectrometry [53,54,55,56,57].
As mentioned above, oxidized SFTI was present at relatively low abundance due in part to the facile side-chain losses of the carbamidomethyl cysteines. While we could perform the MS3 experiment, in some cases, for reduced and alkylated cyclic peptides, the consecutive losses of 91 Da may divert signal from that associated with the oxidation at the lysine residue. We sought to investigate if oxidation occurs once these competitive fragmentation channels are exhausted. Isolation and CID of the ion generated from the first 91 Da loss from aurated SFTI results in, predominantly, a second loss of 91 Da (Supplemental Fig. S1b), consistent with the fact that there are two alkylated cysteine residues. Subsequent activation of the ion generated by the second loss of 91 Da, resulting in the spectrum shown in Supplemental Fig. S1c, generates the base peak at m/z 1428.8, corresponding to the oxidation of the lysine as indicated by the loss of 215 Da.
The consecutive side-chain losses can be viewed as the gas-phase formation of an SFTI analog in which both cysteine residues are converted to dehydroalanine residues (Supplemental Scheme S1) and are indicated with a prime notation (i.e., C′). Figure 3 shows the spectrum obtained upon collisional activation of the oxidized dehydroalanine containing cyclic peptide ion. While the most abundant peaks correspond to small molecule loss, there is evidence for ring opening at the lysine residue as indicated by the fragment ions labeled in red. A total of seven ions can be assigned to ring opening at Thr-Lys. Additionally, consistent with the formation of dehydroalanine, there are fragment ions related to an initial c/z cleavage adjacent to dehydroalanine [58, 59]. The structures of the acyclic peptides resulting from cleavage at lysine or either dehydroalanine are presented in Supplemental Fig. S2.
MS4 product ion spectrum of [M–H–NH3–91–91]+ formed via collisional activation as shown in Supplemental Fig. S1. M = reduced and alkylated sunflower trypsin inhibitor. Open circles indicate water loss and shaded circles indicate ammonia loss. The lightning bolt indicates the species subjected to CID. Product ions corresponding to opening at lysine are highlighted in red. Fragment ions corresponding to opening at Dha are highlighted in blue and green.
Activation of the oxidized product ion, generated via the loss of HAuNH3 directly from [M + Au]+ or via the loss of HAuNH3 from the [M + Au–91–91]+ ion, proved to be an effective ring opening strategy as demonstrated with sunflower trypsin inhibitor. In both cases, when compared to CID of singly and doubly protonated reduced and alkylated SFTI (Supplemental Fig. S3), an increase in structurally informative product ions was observed with the described Au (I) cationization approach. While this strategy aims to open cyclic peptides at lysine, competitive ring opening sites are observed at proline, aspartic acid, and dehydroalanine. Nonetheless, in both SFTI cases presented, about half of the product ions can be attributed to ring opening at lysine.
β-Loop
Aurated β-Loop was generated via the ion/ion chemistry described above. Oxidation of the lysine residue is observed to be the major process upon collisional activation of gold (I) cationized β-Loop (Figure 4a). The product ion spectrum also shows several aurated fragment ions. To maximize the abundance of the HAuNH3 loss, CID of [M + Au]+ was immediately followed by CID of the gold hydride loss, without isolation. Activation of the [M–H–NH3]+ ion yields an abundant water loss, CO loss, CO2 loss, and y13GK, as well as a number of sequence informative fragments between m/z 1500 and 500.
Figure 4c is an expanded view of Figure 4b over the range of m/z 1500 to m/z 500. Close examination of the product ions in this range reveals the following low-abundance ions originating from the ring opening at the lysine residue: \( {\mathrm{b}}_{5\mathrm{GK}}^{\blacksquare } \), \( {\mathrm{b}}_{7\mathrm{GK}}^{\blacksquare } \), \( {\mathrm{b}}_{8\mathrm{GK}}^{\blacksquare } \), y9GK, \( {\mathrm{b}}_{11\mathrm{GK}}^{\blacksquare } \), and \( {\mathrm{b}}_{12\mathrm{GK}}^{\blacksquare } \). Complicating data interpretation, three of the aforementioned fragment ions are isomeric with other plausible product ions. Specifically, the \( {\mathrm{b}}_{7\mathrm{GK}}^{\blacksquare } \) is isomeric with \( {\mathrm{b}}_{7\mathrm{QP}}^{\blacksquare } \) and \( {\mathrm{b}}_{7\mathrm{PG}}^{\blacksquare } \), \( {\mathrm{b}}_{11\mathrm{GK}}^{\blacksquare } \) is isomeric with \( {\mathrm{a}}_{12\mathrm{PG}}^{\blacksquare } \), and \( {\mathrm{b}}_{12\mathrm{GK}}^{\blacksquare } \) is isomeric with \( {\mathrm{y}}_{12\mathrm{QP}}^{\blacksquare } \). While it is likely that ring opening occurs primarily N-terminal to lysine, the SFTI data and previous reports [33, 34] suggest that cleavage N-terminal to proline could be a competitive dissociation pathway. Therefore, the \( {\mathrm{y}}_{7\mathrm{QP}}^{\blacksquare } \) and \( {\mathrm{y}}_{12\mathrm{QP}}^{\blacksquare } \) fragment ions are plausible products and may therefore contribute to the product ion spectrum.
For unambiguous fragment ion assignment and to pinpoint the site of ring opening at lysine, we further probed the y13GK fragment ion, which is 109 Da lower in mass than the [M–H–NH3]+ precursor ion. As discussed below, this fragment ion is generated by the loss of the oxidized lysine residue from the N-terminus. The utility of fragmenting the ion generated by consecutive losses of 215 Da and 109 Da is first discussed using the linear peptide KGAILPGAILR for illustration (Figure 5). Oxidation of the lysine residue is indicated with the signature loss of 215 Da, HAuNH3 (Figure 5a). Activation of the oxidized [M–H–NH3]+ species is shown in Figure 5b. The base peak arises from the loss of 109 Da, producing the y10 fragment ion, which, in essence, is singly protonated GAILPGAILR. The fragmentation of the y10 fragment ion and fragmentation of singly protonated GAILPGAILR generate identical spectra (compare Figure 5c, d), confirming the loss of 109 Da as the loss of the oxidized lysine residue.
Activation of (a) [M + Au]+, (b) [M–H–NH3]+, and (c) y10 where M = KGAILPGAILR. Activation of (d) [GAILPGAILR + H]+. Open circles indicate water loss and shaded circles indicate ammonia loss. The lightning bolt indicates the species subjected to CID. Lysine residue loss (i.e., 147 Da lower in mass) is represented with a triangle superscript.
Figure 6 demonstrates how the loss of 109 Da can be used to obtain unambiguous sequence information for the β-Loop cyclic peptide. The y13GK fragment ion at m/z 1516.8 represents the acyclic peptide ion [FTVQ(D-Pro)GRWQYV(D-Pro)G + H]+. Isolation and CID of y13GK results in a series of 11 b- and y-fragment ions with unambiguous fragment ion structural assignments. This approach to cyclic peptide analysis successfully sequenced greater than 84% of the β-loop cyclic peptide, missing only fragment ions corresponding to cleavage of the Phe-Thr and Pro-Gly amide linkages. We note here that a loss of 109 Da was observed in the sunflower trypsin inhibitor product ion spectrum (Figure 2b). However, the signal was too low to perform additional stages of interrogation.
Activation of the y13GK fragment ion of Figure 4b. Open circles indicate water loss and shaded circles indicate ammonia loss.
Conclusions
Selective ring opening of two lysine containing cyclic peptides is demonstrated. Ion/ion reactions are used to transform doubly protonated peptides to aurated peptide ions. Oxidation via the loss of HAuNH3 produces a weakened amide bond adjacent to the lysine residue. The unusual redox process that leads to [M–H–NH3]+ from lysine-containing peptides is a characteristic of gold cationization. Collisional activation of the [M–H–NH3]+ species generates numerous fragment ions containing a common cyclic imine N-terminus, indicating a highly facile ring opening pathway. This selectivity simplifies the product ion spectrum as ring opening is localized to a few amide bonds.
Other facile cleavage reactions can compete with the process described above. In the case of sunflower trypsin inhibitor, the competitive pathways include openings N-terminal to proline, N-terminal to dehydroalanine, and C-terminal to aspartic acid as evidenced by the fragment ions annotated DG, IP, IC′, and RC′. Ions generated via opening at dehydroalanine are more prevalent than ions corresponding to opening at aspartic acid and proline. Incorporation of dehydroalanine may present another strategy to selectively open cyclic peptides upon collisional activation. In the case of the β-Loop peptide ion, linearization at lysine is the major process, yet, linearization at proline is also observed. Additionally, several of the proline-related fragment ions are isomeric with product ions opened at lysine. To avoid ambiguities in confident ion assignments arising from possibly isomeric fragments, isolation and activation of the product ion generated by successive losses of 215 Da and 109 Da from the [M–H–NH3]+ ion ensures that the ions arise from loss of an oxidized lysine residue from the N-terminus (viz., the y13GK ion from β-Loop). CID of y13GK, [FTVQ(D-Pro)GRWQYV(D-Pro)G + H]+, cleaves 10 of the 12 amide bonds. This gas-phase strategy for cyclic peptide analysis offers a convenient means of selectively opening cyclic peptides. In favorable cases, when the abundance of the 109 Da loss does not limit the extent to which MS4 can be performed, cyclic peptide linearization can be localized to the lysine residue.
References
Craik, D.J., Daly, N.L., Bond, T., Waine, C.: Plant cyclotides: a unique family of cyclic and knotted proteins that defines the cyclic cystine knot structural motif. J. Mol. Biol. 294, 1327–1336 (1999)
Craik, D.J.: Discovery and applications of the plant cyclotides. Toxicon. 56, 1092–1102 (2010)
Colgrave, M.L., Craik, D.J.: Thermal, chemical, and enzymatic stability of the cyclotide Kalata B1: the importance of the cyclic cystine knot. Biochemistry. 43, 5965–5975 (2004)
Craik, D.J., Conibear, A.C.: The chemistry of cyclotides. J. Org. Chem. 76, 4805–4817 (2011)
Pränting, M., Lööv, C., Burman, R., Göransson, U., Andersson, D.I.: The cyclotide cycloviolacin O2 from Viola odorata has potent bactericidal activity against gram-negative bacteria. J. Antimicrob. Chemother. 65, 1964–1971 (2010)
Gründemann, C., Koehbach, J., Huber, R., Gruber, C.W.: Do Plant Cyclotides Have Potential As Immunosuppressant Peptides? J. Nat. Prod. 75, 167–174 (2012)
Tang, J., Wang, C.K., Pan, X., Yan, H., Zeng, G., Xu, W., He, W., Daly, N.L., Craik, D.J., Tan, N.: Isolation and characterization of cytotoxic cyclotides from Viola tricolor. Peptides. 31, 1434–1440 (2010)
Colgrave, M.L., Huang, Y.-H., Craik, D.J., Kotze, A.C.: Cyclotide Interactions with the Nematode External Surface. Antimicrob. Agents Chemother. 54, 2160 (2010)
Colgrave, M.L., Kotze, A.C., Kopp, S., McCarthy, J.S., Coleman, G.T., Craik, D.J.: Anthelmintic activity of cyclotides: In vitro studies with canine and human hookworms. Acta Tropica. 109, 163–166 (2009)
Daly, N.L., Koltay, A., Gustafson, K.R., Boyd, M.R., Casas-Finet, J.R., Craik, D.J.: Solution structure by NMR of circulin A: a macrocyclic knotted peptide having anti-HIV activity. J. Mol. Biol. 285, 333–345 (1999)
Hallock, Y.F., Sowder, R.C., Pannell, L.K., Hughes, C.B., Johnson, D.G., Gulakowski, R., Cardellina, J.H., Boyd, M.R.: Cycloviolins A−D, Anti-HIV Macrocyclic Peptides from Leonia cymosa1. J. Org. Chem. 65, 124–128 (2000)
Jennings, C., West, J., Waine, C., Craik, D., Anderson, M.: Biosynthesis and insecticidal properties of plant cyclotides: The cyclic knotted proteins from Oldenlandia affinis. Proc. Natl. Acad. Sci. 98, 10614 (2001)
Jennings, C.V., Rosengren, K.J., Daly, N.L., Plan, M., Stevens, J., Scanlon, M.J., Waine, C., Norman, D.G., Anderson, M.A., Craik, D.J.: Isolation, Solution Structure, and Insecticidal Activity of Kalata B2, a Circular Protein with a Twist: Do Möbius Strips Exist in Nature? Biochemistry. 44, 851–860 (2005)
Barbeta, B.L., Marshall, A.T., Gillon, A.D., Craik, D.J., Anderson, M.A.: Plant cyclotides disrupt epithelial cells in the midgut of lepidopteran larvae. Proc. Natl. Acad. Sci. 105, 1221 (2008)
Caceres, C.C., Bansal, P.S., Navarro, S., Wilson, D., Don, L., Giacomin, P., Loukas, A., Daly, N.L.: An engineered cyclic peptide alleviates symptoms of inflammation in a murine model of inflammatory bowel disease. J. Biol. Chem. 292, 10288–10294 (2017)
Durek, T., Cromm, P.M., White, A.M., Schroeder, C.I., Kaas, Q., Weidmann, J., Ahmad Fuaad, A., Cheneval, O., Harvey, P.J., Daly, N.L., Zhou, Y., Dellsén, A., Österlund, T., Larsson, N., Knerr, L., Bauer, U., Kessler, H., Cai, M., Hruby, V.J., Plowright, A.T., Craik, D.J.: Development of novel melanocortin receptor agonists based on the cyclic peptide framework of sunflower trypsin Inhibitor-1. J. Med. Chem. 61, 3674–3684 (2018)
Gunasekera, S., Fernandes-Cerqueira, C., Wennmalm, S., Wähämaa, H., Sommarin, Y., Catrina, A.I., Jakobsson, P.-J., Göransson, U.: Stabilized cyclic peptides as scavengers of autoantibodies: neutralization of anticitrullinated protein/peptide antibodies in rheumatoid arthritis. ACS Chem. Biol. 13, 1525–1535 (2018)
de Veer, S.J., Li, C.Y., Swedberg, J.E., Schroeder, C.I., Craik, D.J.: Engineering potent mesotrypsin inhibitors based on the plant-derived cyclic peptide, sunflower trypsin inhibitor-1. Eur. J. Med. Chem. 155, 695–704 (2018)
Swedberg, J.E., Wu, G., Mahatmanto, T., Durek, T., Caradoc-Davies, T.T., Whisstock, J.C., Law, R.H.P., Craik, D.J.: Highly potent and selective plasmin inhibitors based on the sunflower trypsin Inhibitor-1 scaffold attenuate fibrinolysis in plasma. J. Med. Chem. 62, 552–560 (2019)
Fesik, S.W., Bolis, G., Sham, H.L., Olejniczak, E.T.: Structure refinement of a cyclic peptide from two-dimensional NMR data and molecular modeling. Biochemistry. 26, 1851–1859 (1987)
Coles, M., Sowemimo, V., Scanlon, D., Munro, S.L.A., Craik, D.J.: A conformational study by proton NMR of a cyclic pentapeptide antagonist of endothelin. J. Med. Chem. 36, 2658–2665 (1993)
Mazzeo, M., Isernia, C., Rossi, F., Saviano, M., Pedone, C., Paolillo, L., Benedetti, E., Pavone, V.: Conformational behaviour of a cyclolinopeptide a analogue: two-dimensional NMR study of cyclo(Pro1-Pro-Phe-Phe-Ac6c-IIe-ala-Val8). J. Pept. Sci. 1, 330–340 (1995)
Porter, C., Wilce, J.: NMR analysis of G7-18NATE, a nonphosphorylated cyclic peptide inhibitor of the Grb7 adapter protein. Pept. Sci. 88, 174–181 (2007)
Johnson, M., Liu, M., Struble, E., Hettiarachchi, K.: Characterization of cyclic peptides containing disulfide bonds. J. Pharm. Biomed. Anal. 109, 112–120 (2015)
Northfield, S., Wielens, J., Headey, S., Williams-Noonan, B., Mulcair, M., Scanlon, M., Parker, M., Thompson, P., Chalmers, D.: Cyclic Hexapeptide Mimics of the LEDGF Integrase Recognition Loop in Complex with HIV‐1 Integrase. ChemMedChem. (2018)
Laurencin, M., Simon, M., Fleury, Y., Baudy-Floc'h, M., Bondon, A., Legrand, B.: Selectivity modulation and structure of α/aza-β3 cyclic antimicrobial peptides. Chem. Eur. J. 24, 6191–6201 (2018)
Gross, M.L., McCrery, D., Crow, F., Tomer, K.B., Pope, M.R., Ciuffetti, L.M., Knoche, H.W., Daly, J.M., Dunkle, L.D.: The structure of the toxin from helminthosporium carbonum. Tetrahedron Lett. 23, 5381–5384 (1982)
Tilvi, S., Naik, C.: Tandem mass spectrometry of kahalalides: identification of two new cyclic depsipeptides, kahalalide R and S from Elysia grandifolia. J. Mass Spectrom. 42, 70–80 (2007)
Mohimani, H., Yang, Y.L., Liu, W.T., Hsieh, P.W., Dorrestein, P.C., Pevzner, P.A.: Sequencing cyclic peptides by multistage mass spectrometry. Proteomics. 11, 3642–3650 (2011)
Attard, T.J., Carter, M.D., Fang, M., Johnson, R.C., Reid, G.E.: Structural Characterization and Absolute Quantification of Microcystin Peptides Using Collision-Induced and Ultraviolet Photo-Dissociation Tandem Mass Spectrometry. J. Am. Soc. Mass. Spectrom. 1–14 (2018)
Parsley, N.C., Kirkpatrick, C.L., Crittenden, C.M., Rad, J.G., Hoskin, D.W., Brodbelt, J.S., Hicks, L.M.: PepSAVI-MS reveals anticancer and antifungal cycloviolacins in Viola odorata. Phytochemistry. 152, 61–70 (2018)
Paizs, B., Suhai, S.: Fragmentation pathways of protonated peptides. Mass Spectrom. Rev. 24, 508–548 (2005)
Tomer, K.B., Crow, F.W., Gross, M.L., Kopple, K.D.: Fast-atom bombardment combined with tandem mass spectrometry for the determination of cyclic peptides. Anal. Chem. 56, 880–886 (1984)
Hitzeroth, G., Vater, J., Franke, P., Gebhardt, K., Fiedler, H.-P.: Whole cell matrix-assisted laser desorption/ionization time-of-flight mass spectrometry and in situ structure analysis of streptocidins, a family of tyrocidine-like cyclic peptides. Rapid Commun. Mass Spectrom. 19, 2935–2942 (2005)
Poth, A.G., Colgrave, M.L., Philip, R., Kerenga, B., Daly, N.L., Anderson, M.A., Craik, D.J.: Discovery of cyclotides in the Fabaceae plant family provides new insights into the cyclization, evolution, and distribution of circular proteins. ACS Chem. Biol. 6, 345–355 (2011)
Chan, L.Y., He, W., Tan, N., Zeng, G., Craik, D.J., Daly, N.L.: A new family of cystine knot peptides from the seeds of Momordica cochinchinensis. Peptides. 39, 29–35 (2013)
Narayani, M., Chadha, A., Srivastava, S.: Cyclotides from the indian medicinal plant Viola odorata (Banafsha): identification and characterization. J. Nat. Prod. 80, 1972–1980 (2017)
Crittenden, C.M., Parker, W.R., Jenner, Z.B., Bruns, K.A., Akin, L.D., McGee, W.M., Ciccimaro, E., Brodbelt, J.S.: Exploitation of the ornithine effect enhances characterization of stapled and cyclic peptides. J. Am. Soc. Mass Spectrom. 27, 856–863 (2016)
McGee, W.M., McLuckey, S.A.: The ornithine effect in peptide cation dissociation. J. Mass Spectrom. 48, 856–861 (2013)
Eller, K., Schwarz, H.: Organometallic in the gas phase. Chem. Rev. 91, 1121–1177 (1991)
Schwarz, H.: Relativistic effects in gas-phase ion chemistry: an experimentalist’s view. Angew. Chem. Int. Ed. 42, 4442–4454 (2003)
Schröder, D., Schwarz, H., Hrušák, J., Pyykkö, P.: Cationic gold (I) complexes of xenon and of ligands containing the donor atoms oxygen, nitrogen, phosphorous, and sulfur. Inorg. Chem. 37, 624–632 (1998)
O’Hair, R.A.J., “Mass spectrometry of organogold compounds” in The Chemistry of Organogold Compounds, Rappoport, Z.; Liebman, J.F.; Marek, I. (eds.) John Wiley & Sons, Ltd: Chichester, UK, Chapter 4, 2014, 57–105
Foreman, D.J., Betancourt, S.K., Pilo, A.L., McLuckey, S.A.: Novel peptide ion chemistry associated with gold (I) cationization: preferential cleavage at lysine residues. Int. J. Mass Spectrom. 427, 114–122 (2018)
Espinosa, J.F., Gellman, S.H.: A designed β-hairpin containing a natural hydrophobic cluster. Angew. Chem. Int. Ed. 39, 2330–2333 (2000)
Xia, Y., Wu, J., McLuckey, S.A., Londry, F.A., Hager, J.W.: Mutual storage mode ion/ion reactions in a hybrid linear ion trap. J. Am. Soc. Mass Spectrom. 16, 71–81 (2005)
Liang, X., Xia, Y., McLuckey, S.A.: Alternately pulsed nanoelectrospray ionization/atmospheric pressure chemical ionization for ion/ion reactions in an electrodynamic ion trap. Anal. Chem. 78, 3208–3212 (2006)
Gunawardena, H.P., O'Hair, R.A.J., McLuckey, S.A.: Selective disulfide Bond cleavage in gold(I) cationized polypeptide ions formed via gas-phase ion/ion cation switching. J. Proteome Res. 5, 2087–2092 (2006)
Mentinova, M., McLuckey, S.A.: Cleavage of multiple disulfide bonds in insulin via gold cationization and collision-induced dissociation. Int. J. Mass Spectrom. 308, 133–136 (2011)
Londry, F.A., Hager, J.W.: Mass selective axial ion ejection from a linear quadrupole ion trap. J. Am. Soc. Mass Spectrom. 14, 1130–1147 (2003)
Novák, J., Lemr, K., Schug, K.A., Havlíček, V.: CycloBranch: de novo sequencing of nonribosomal peptides from accurate product ion mass spectra. J. Am. Soc. Mass Spectrom. 26, 1780–1786 (2015)
Ngoka, L.C., Gross, M.L.: A nomenclature system for labeling cyclic peptide fragments. J. Am. Soc. Mass Spectrom. 10, 360–363 (1999)
Yu, W., Vath, J.E., Huberty, M.C., Martin, S.A.: Identification of the facile gas-phase cleavage of the Asp-Pro and Asp-Xxx peptide bonds in matrix-assisted laser desorption time-of-flight mass spectrometry. Anal. Chem. 65, 3015–3023 (1993)
Sullivan, A.G., Brancia, F.L., Tyldesley, R., Bateman, R., Sidhu, K., Hubbard, S.J., Oliver, S.G., Gaskell, S.J.: The exploitation of selective cleavage of singly protonated peptide ions adjacent to aspartic acid residues using a quadrupole orthogonal time-of-flight mass spectrometer equipped with a matrix-assisted laser desorption/ionization source. Int. J. Mass Spectrom. 210-211, 665–676 (2001)
Bleiholder, C., Suhai, S., Harrison, A.G., Paizs, B.: Towards Understanding the Tandem Mass Spectra of Protonated Oligopeptides. 2: The Proline Effect in Collision-Induced Dissociation of Protonated Ala-Ala-Xxx-Pro-Ala (Xxx = Ala, Ser, Leu, Val, Phe, and Trp). J. Am. Soc. Mass. Spectrom. 22, 1032–1039 (2011)
Schwartz, B.L., Bursey, M.M.: Some proline substituent effects in the tandem mass spectrum of protonated pentaalanine. Biol. Mass Spectrom. 21, 92–96 (1992)
Vaisar, T., Urban, J.: Probing proline effect in CID of protonated peptides. J. Mass Spectrom. 31, 1185–1187 (1996)
Pilo, A.L., Peng, Z., McLuckey, S.A.: The dehydroalanine effect in the fragmentation of ions derived from polypeptides. J. Mass Spectrom. 51, 857–866 (2016)
Peng, Z., Bu, J., McLuckey, S.A.: The generation of dehydroalanine residues in protonated polypeptides: ion/ion reactions for introducing selective cleavages. J. Am. Soc. Mass Spectrom. 28, 1765–1774 (2017)
Acknowledgements
This work was supported by the National Institutes of Health (NIH) under Grant GM R37-45372. Vanessa M. Kung of the Gellman Lab at the University of Wisconsin is acknowledged for synthesis of the β-Loop cyclic peptide and Samuel H. Gellman is acknowledged for providing the peptide to our laboratory.
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic Supplementary Material
ESM 1
(DOCX 1385 kb)
Rights and permissions
About this article
Cite this article
Foreman, D.J., Lawler, J.T., Niedrauer, M.L. et al. Gold(I) Cationization Promotes Ring Opening in Lysine-Containing Cyclic Peptides. J. Am. Soc. Mass Spectrom. 30, 1914–1922 (2019). https://doi.org/10.1007/s13361-019-02247-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13361-019-02247-x