Abstract
Ion mobility spectrometry-mass spectrometry (IMS-MS) has demonstrated the ability to characterize structures of weakly-bound peptide assemblies. However, these assemblies can potentially dissociate during the IMS-MS measurement if they undergo energetic ion-neutral collisions. Here, we investigate the ability of tandem-trapped ion mobility spectrometry-mass spectrometry (TIMS-TIMS-MS) to retain weakly-bound peptide assemblies. We assess ion heating and dissociaton in the tandem-TIMS instrument using bradykinin and its assemblies as reference systems. Our data indicate that non-covalent bradykinin assemblies are largely preserved in TIMS-TIMS under carefully selected operating conditions. Importantly, we observe quadruply-charged bradykinin tetramers, which attests to the “softness” of our instrument.

Graphical Abstract
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Macromolecular assemblies play crucial roles in many biological processes [1]. Protein and peptide assemblies in particular are often intermediates in disease pathogeneses [2,3,4], for example, during formation of amyloid fibrils [5] or viral capsids [6]. Ion mobility spectrometry-mass spectrometry (IMS-MS) has demonstrated great utility to investigate these assemblies by separating assembly states by differences in their mass-to-charge ratios and conformations of the same assembly state by differences in their ion mobilities K.
Moreover, the measured ion mobility K yields the momentum transfer cross section of the ion [7, 8] according to
where z is the charge of the analyte ion; T and N0 are the temperature and reduced number density of the buffer gas; kB is the Boltzmann constant; μ is the reduced mass of the ion-neutral pair; and K0 is the reduced ion mobility, which is trivially related to K [7]. These cross sections Ω are useful to propose potential structures for the assembly by computational approaches [9, 10].
Because IMS-MS detects ions in the gas phase, the structure of protein systems detected by IMS will generally differ from their biologically active solution structures. While it is now established that “soft” ion mobility spectrometry measurements do not typically detect equilibrium gas phase structures of protein systems [11,12,13,14,15,16,17,18], what is less clear is how closely the detected ions resemble the native state present in solution. This is a general question that applies to all biological species detected by IMS-MS, but relatively little is secured about how structures of non-covalent assemblies change after exposure to the gas phase environment. Obviously, the most drastic change would be if assemblies dissociated into their subunits during the course of the IMS-MS measurement.
Intuitively, the stronger the intermolecular interaction between the assembly units, the less prone the assembly is to dissociate during the IMS-MS measurement. Electrostatic interactions between moieties with permanent multiple moments, such as charges (monopoles) or dipole moments, are among the strongest intermolecular interactions [19]. Induction interactions, attractive forces between moieties with permanent multipole moments and moieties that are polarizable, can also lead to strong non-covalent interactions [19]. It thus does not come as a surprise that the general approach of using IMS-MS as a tool to study structures of non-covalent assemblies can be traced back to early work on ion-neutral clusters [20] or salts nanocrystals [21]. Salts are held together by electrostatic interactions between oppositely charged monopoles whereas binding in ion neutral-complexes arises mainly from electrostatic and induction forces between the charged ion and permanent and/or induced multipole moments on the neutral partner.
With the advent of “soft” electrospray [22] and matrix-assisted laser desorption ionization [23] methods, peptides and proteins could be lifted gently into the gas phase and were found to strongly bind to species with permanent multipole moments such as metal cations [24] or water [25]. Simultaneously, several groups explored the ability to characterize stoichiometry and/or the quaternary structure of protein complexes by mass spectrometry [26,27,28]. While many of these protein complexes are among the most strongly bound non-covalent assemblies, even multiply-charged assemblies of the small peptide bradykinin were found to be sufficiently stable in the gas phase so that their cross sections could be measured by ion mobility spectrometry [29]. The significance of these cross section measurements for peptide and protein assembly systems arises from the fact that the relationship between the cross section and the number of assembly units revealed the overall topology of the assembly.
Moreover, cross sections measured by “soft” IMS-MS instruments for assemblies of the intrinsically-disordered protein α-synuclein implicated in Parkinson’s disease were shown to strongly depend on the solution pH [30]. Such solution-dependency of assembly structures detected by IMS indicated that the assemblies did not have sufficient time and energy to evolve into their equilibrium gas phase structure within the time-scale of the measurement. Similar observations were made for other peptides including islet amylin precursor protein [9, 31] or amyloid-β [5]. Hence, the cross sections recorded for these systems could be used to propose the mechanisms by which these assemblies form in the solution. Moreover, IMS-MS succeeded in characterizing structures of presumed toxic intermediate assemblies of amyloid-β [5] and the conformational transitions of assemblies during formation of amyloid-like fibrils of peptides [32] and amino acids [33], thus providing insight into how Alzheimer drug candidates modulate amyloid assemblies [34].
The above considerations underscore that cross sections measured by “soft” ion mobility spectrometry methods [11] are useful for characterizing structures and formation mechanism of peptide assemblies in the solution phase. However, one potential problem of these applications of IMS-MS is that the assembly ions could dissociate if they become vibrationally activated by energetic ion-neutral collisions during the IMS-MS measurement. Such “ion heating” processes are well-known to occur in MS [35, 36] and, if not operated under low-field conditions, in IMS instrumentation [37,38,39]. What is less characterized, however, is how structures of non-covalent assemblies change due to ion heating in post-IMS instrument components. If assemblies remain intact throughout the ion mobility experiment but dissociate afterwards, then the measured ion mobility K0 corresponds to the intact assembly while the mass m and charge z correspond to the dissociated assembly units. Hence, if assemblies dissociate after ion mobility separation, then cross sections calculated via Eq. (1) will be incorrect and spurious ions appear as observed on traveling-wave [40] or drift tube systems [41, 42].
We recently developed a tandem-trapped IMS (tandem-TIMS) instrument [43] by coupling two trapped IMS (TIMS) [44,45,46,47] devices. Tandem-TIMS has great potential for structural studies of biological assemblies because it enables mobility-selective gating and activation of ions followed by mobility analysis. However, tandem-TIMS is embedded in a Qq-TOF mass spectrometer where activating ion-neutral collisions can occur. Here, ions traverse several ion funnels with intermediate pressure regimes, a hexapole ion guide, a quadrupole, and even a collision cell after separation in tandem-TIMS. While we showed that TIMS [37] and tandem-TIMS [43] devices can preserve native-like structures of the monomeric protein ubiquitin if operated under sufficiently “soft” conditions, it remains unclear to what extent non-covalent assemblies of peptides are retained.
Here, we investigate under which operating conditions, and to what extent non-covalent assemblies of peptides can be characterized with TIMS-Qq-TOF and tandem-TIMS-Qq-TOF instruments. To this end, we characterize ion heating effects in post-tandem-TIMS instrument components using the nonapeptide bradykinin. Bradykinin is a well-suited reference system for this purpose: activation parameters have been reported for fragmentation pathways of several monomer charge states [48, 49], with water-loss from the triply-charged monomer being particularly active. Hence, bradykinin is a very sensitive probe for ion heating. Further, oligomeric species observed on the “gentle” drift tubes in the Clemmer [29] and Bowers [50] groups are strongly consistent with each other, suggesting that bradykinin is suitable as a reference system to also probe retention of non-covalent peptide assemblies. While we focus our discussion on our newly-developed tandem-TIMS system, post-TIMS ion optics are identical to the single-TIMS instruments and the results are thus comparable. Our analysis indicates that peptide assemblies can be retained with a tandem-TIMS-Qq-TOF instrument if the tandem-TIMS device as well as all post-tandem-TIMS components are optimized to minimize the energy of ion-neutral collisions. In particular, our data recorded under carefully optimized operating conditions unambiguously confirm the presence of long-postulated quadruply-charged bradykinin tetramers [29]. Nevertheless, we emphasize that energy barriers of assembly dissociation pathways depend on the specific system investigated. Therefore, when investigating less characterized assembly species, we recommend that the analyst asserts that assemblies remain intact throughout the entire measurement process using the approach discussed here.
Experimental
Sample Preparation
High concentration electrospray tune mix (Agilent Technologies, Santa Clara, CA, composed of perfluorated phosphazenes) was used as purchased. Bradykinin was purchased from Sigma (St. Louis, MO). A 10 μM solution of bradykinin in 50:50 water:methanol (LC/MS grade, Fisher Scientific, Waltham, MA and Acros Organics, Pittsburgh, PA) was prepared as described [50]. This solution was then used for TIMS-MS and tandem-TIMS-MS analysis.
Tandem-TIMS Operation
Measurements were performed on TIMS-Qq-TOF and tandem-TIMS-Qq-TOF instruments described elsewhere [37, 43]. Schematics of the instruments are given in Figure 1 and Figure S5 (Supporting Information). Post-TIMS ion optics are identical among the two instruments and the results are comparable. For completeness, Section S2 (Supporting Information) shows data for bradykinin obtained on the TIMS-Qq-TOF instrument. Samples were directly infused into the electrospray ionization (ESI) source operating in positive ion mode by a 250-μL gas tight syringe (Hamilton, Franklin, MA) at a flow rate of 180 μL/h. The capillary voltage was set to 3.5 kV with an end plate offset of 500 V. Nitrogen buffer gas was produced from a nitrogen generator (Peak Scientific, NM32-LA-MS-230V). The temperature of the nitrogen desolvation gas was set to 304 K and monitored at the entrance of the resistive glass capillary by means of a thermocouple element and a thermal imaging gun. Ions produced from electrospray enter the tandem-TIMS analyzer through a resistive glass capillary and are deflected into the entrance funnel of the first TIMS device (TIMS-1, see Figure 1). Ions traverse an entrance funnel, TIMS analyzer tunnel, and leave TIMS-1 through an exit funnel. Subsequently, they traverse two apertures and enter the second TIMS analyzer (TIMS-2). As reported, TIMS-2 is dimensionally identical to TIMS-1, i.e., ions traverse an entrance funnel, the TIMS-2 mobility analyzer, and leave TIMS-2 through a second exit funnel. In the experiments used here, the electric potential on the entrance of the analyzer tunnel in TIMS-2 was ramped over a range of 70 V at a rate of 0.23 V/ms at a TOF pulse length of 0.316 ms. The amplitude of the rf-potentials were set to 225 Vpp (TIMS-1) and 250 Vpp (TIMS-2), respectively. The bias between the hexapole and quadrupole was set to 5 V, and the bias between the quadrupole and collision cell was 1 V. Other TIMS-TIMS settings were taken from previous studies [37, 43] and are known to preserve native-like structures of ubiquitin (Section S1.1, Supporting Information). In particular, all measurements were carried out in “forward flow” mode, i.e., with the exit pressure of TIMS-1 larger than the entrance pressure of TIMS-2 and with zero electric potential differences in the interface between TIMS-1 and TIMS-2. Ions are thus passively transported through the interface region as they are only dragged from TIMS-1 into TIMS-2 by the flowing gas.
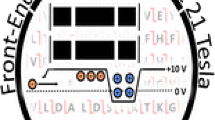
Tandem-TIMS-Qq-TOF instrument. Ions are generated by ESI and traverse a transfer capillary before entering TIMS-TIMS. Once ions are eluted from TIMS-TIMS, they traverse several ion funnels, a hexapole ion guide, a quadrupole, and a collision cell before reaching the TOF mass analyzer for detection (Ion lenses used to focus ions prior entering the quadrupole and TOF devices are shaded.) Energetic ion-neutral collisions in post-TIMS components can take place primarily in the TIMS–hexapole interface, where the pressure changes gradually from intermediate pressures (~ 0.5–2 mbar) to ~ 10−4 mbar, and in the collision cell (shaded in red). If intact dimers are produced from ESI and remain intact throughout the ion mobility measurement then the measured mobility corresponds to the mobility of the dimer, K0,dimer. If these dimers dissociate into monomers after the ion mobility measurement (for example in the shaded regions), then ions will be detected with the mass and charge of the monomers but with the mobility of the dimer. Consequently, cross sections calculated via Eq. (1) will suggest presence of spurious ions
Post-TIMS Operation
We varied the potential differences ΔΦ1 (between the TIMS-2 exit funnel and funnel F2; 0–5 V), ΔΦ2 (across funnel F2; 2–8 V), and ΔΦ3 (between the exit of funnel F2 and the hexapole ion guide; 1–11 V) to determine effects on transmitting and preserving assemblies (see Table 1 and Table S1 in the Supporting Information). The potential difference between the hexapole and the analytical quadrupole was set to 5 V. The potential difference between the quadrupole and collision cell was kept at 1 V.
Ion Mobility Calibration
Ion mobilities and cross sections were calibrated as described [44, 51, 52] using perfluorated phosphazenes contained in Agilent ESI tuning mix (m/z 322, 622, 922, 1522, 2122). Reduced ion mobilities (1.369, 1.016, 0.841, 0.642, 0.530 cm2/Vs for m/z 322, 622, 922, 1522, 2122, respectively) were taken from a previous report [52] based on the cross sections reported by Stow et al. [53]. Notice that each spectrum was calibrated immediately after acquisition under identical operating settings.
Results and Discussion
General Remarks on Ion Heating in Post-TIMS Ion Optics
If intact assemblies elute from the tandem-TIMS device, they traverse several instrument components before they arrive at the TOF mass analyzer (Figure 1). As the assembly ions traverse these instrument components, they could gain internal energy (“ion heating”) and dissociate. We expect ion heating to occur primarily in the TIMS-hexapole interface (see inset Figure 1) and in the collision cell due to the pressures and applied electric fields. In the TIMS-hexapole interface, the pressure drops from the TIMS-2 exit funnel (~ 0.5–2 mbar) to funnel F2 (~ 0.05 mbar) and the hexapole (~ 5·10−4 mbar). The mean-free-path between the buffer gas particles thus increases from about 100 μm (0.5 mbar) to ~ 1 mm (0.05 mbar) and ~ 10 cm (5·10−4 mbar). Additionally, electric potentials are applied over short distances (~ 2–20 mm) which means that even small voltage differences correspond to high electric fields. Consequently, ions may be accelerated to high velocities in this instrument region and dissociate due to energetic collisions with gas particles.
Once an assembly ion exits tandem-TIMS, then the mobility of the ion corresponds to the intact assembly regardless of whether or not the assembly remains intact until it reaches the detector. Thus, if assemblies dissociate in any of the post-TIMS components, then the detected (dissociated) ions have a different mass and/or charge than the precursor assembly. Thus, cross sections calculated via Eq. (1) will be incorrect, because the mobility corresponds to the intact assembly but the mass and charge correspond to the detected (dissociated) ions. Consequently, the ions will be detected as spurious ions as discussed [40,41,42] and lead to errors in the assembly formation mechanism.
Ion Heating in the TIMS-Hexapole Interface
A reference system well-suited to characterize ion heating is bradykinin, a nonapeptide with amino acid sequence RPPGFSPFR. Bradykinin monomer ions produced from electrospray ionization are typically charge states 1+ to 3+ (m/z 1061, 531, 354) [29, 48,49,50, 54]. Charge state 3+ in particular is highly sensitive to collisional-activation because of a water-loss pathway with a low activation barrier [48, 49]. Hence, we can probe the extent of collisional-activation in post-TIMS components by assessing the activity of these low-energy fragmentation pathways. Additionally, multiply-charged bradykinin assemblies were reported by IMS-MS [29, 50]. Most dominant were doubly-charged dimers (n/z 2/2) and triply-charged trimers (n/z 3/3) for m/z 1061, but ion mobility suggests presence of even quadruply-charged bradykinin tetramers [29]. Moreover, triply-charged bradykinin dimers (n/z 2/3; m/z 707) were observed by IMS-MS [29, 50]. Because these reports show that the abundances of these assemblies do not depend strongly on the ion mobility instrumentation or on the ion sources used, these species appear well suited to assess the ability of IMS-MS instrumentation to retain intact peptide assemblies.
We record bradykinin mass spectra for different voltages ΔΦ1, ΔΦ2, and ΔΦ3 (see Table S1 and Figure S1, Supporting Information). The doubly-charged monomer m/z 531 dominates the mass spectrum when collisions in the TIMS-hexapole interface are most energetic (Figure 2a). But the spectrum essentially lacks m/z 707 (triply charged dimer, n/z 2/3) and m/z 354 (triply charged monomer, n/z 1/3) observed by others [29, 48, 50].
Dissociation of bradykinin species for different activation voltages in the TIMS-hexapole interface. (a) Mass spectrum recorded under energetic collisions in the TIMS-hexapole interface. Abundance of m/z 354 (triply-charged monomer) is negligible whereas m/z 348 is present. (b) Ion mobility spectrum of m/z 348 is identical to m/z 354 recorded under low-voltage settings. This suggests that m/z 348 is produced by water-loss from m/z 354 in post-TIMS instrument components. (c) m/z 348 increases, while m/z 354 decreases, in abundance as the collision energy in the TIMS-hexapole interface is increased. This provides further evidence that m/z 348 is produced by water-loss from m/z 354 in post-TIMS instrument components. (d) The mass spectrum recorded under low-energy collisions shows bradykinin species m/z 354, 531, 707, 1061, albeit only minor abundance of m/z 707 (triply-charged dimer)
The presence of m/z 348, which is a triply charged ion that corresponds to [m/z 354-H2O], indicates that m/z 354 is collisionally-energized and dissociates by loss of water. Notice that the arrival time distribution of m/z 348 is identical to m/z 354 obtained at low voltages (Figure 2b). This indicates that m/z 354 loses water to form m/z 348 after ion mobility separation. Further, m/z 348 increases, while m/z 354 decreases, in abundance as we increase the collision energy in the TIMS-hexapole interface (Figure 2c). This indicates that the loss of water from m/z 354 occurs in the TIMS-hexapole interface. We draw two conclusions from our observations. First, energetic collisions in the TIMS-hexapole interface sufficiently energize m/z 354 to activate its water-loss pathway. Second, low-energy collisions reduce ion heating to the extent that even this highly reactive water-loss pathway is rendered inactive. Notice that the mass spectrum recorded under low-energy collisions in the TIMS-hexapole interface (Figure 2d) reveals all bradykinin species discussed by Bowers and Clemmer (though m/z 531 predominates) [29, 50]. What is particularly important is that these operating conditions are gentle enough to essentially deactivate the low-energy water-loss pathway from the triply-charged monomer.
We now investigate if bradykinin assemblies are preserved in the tandem-TIMS analyzer and post-TIMS components. Figure 3 shows tandem-TIMS spectra recorded for m/z 1061 under high- and low-energy collisions in the TIMS-hexapole interface (see Figure S2, Supporting Information, for more details). The spectrum recorded for the most energetic collisions shows several singly-charged features (ta ≈ 211 ms, 240 ms, 250–263 ms, 267 ms with reduced ion mobilities K0 ≈ 0.65, 0.83, 0.95–1.04, and 1.15 cm2/Vs, respectively). This spectrum would thus imply that bradykinin exists in distinct monomer conformations with cross sections ranging from 177 Å2 (ta ≈ 267 ms) to 318 Å2 (ta ≈ 211 ms). We note two caveats in these data. First, Bowers [50] and Clemmer [29] unambiguously report doubly-charged dimers (n/z 2/2) and triply-charged trimers (n/z 3/3) for m/z 1061, and Clemmer’s data even indicate presence of a quadruply charged tetramer [29]. By contrast, Figure 3a exclusively shows singly-charged monomers. Second, a cross section of 177 Å2 (nitrogen buffer gas) is unrealistically small for bradykinin, which contains nine residues including arginines and prolines with bulky side chains. As a comparison, even tetraglycine with only four, tightly folded glycine residues has a cross section of 153 Å2 in nitrogen [55]. Moreover, as we reduce the collision energy in the TIMS-hexapole interface, the peaks in the m/z 1061 spectrum retain their ion mobilities but their charge states change (Figure 3b).
TIMS spectra recorded for m/z 1061 under high- and low-energy collisions in the TIMS-hexapole interface. (a) Exclusively singly-charged monomers are observed for m/z 1061 under energetic collisions in the TIMS-hexapole interface. (b) Under low-energy collisions, the charge states change and reveal dimers, trimers, and tetramers. A spurious ion visible as a highly compact monomer with a reduced ion mobility K0 ≈ 1.15 cm2/Vs is still observed. (Note that small variations in the arrival times of the observed species are caused by small pressure fluctuations but that the ion mobilities of the species do not change)
Figure 4 compares the isotopic patterns for the various features present in the “soft” m/z 1061 spectrum (Figure 3b) to the isotopic patterns calculated for bradykinin dimers (n/z 2/2), trimers (n/z 3/3), and tetramers (n/z 4/4). The peak positions and relative abundances of the experimental mass spectra closely agree with the simulated spectra (Figure 4b–d). Note that the adjacent n/z 3/3 trimer and 4/4 tetramer peaks are not baseline-separated in the TIMS device (Figure 4a), which likely causes the slight deviation between the experimental and simulated isotopic patterns for n/z 4/4 (Figure 4d). Overall, however, the experimental and simulated isotopic patterns agree closely. Thus, Figure 4 reveals presence of a doubly-charged dimer n/z 2/2 (ta ~ 240 ms, 491 Å2), a triply charged trimer n/z 3/3 (ta ~ 253 ms, 643 Å2) and a quadruply-charged tetramer n/z 4/4 (ta ~ 260, 782 Å2) in addition to a singly-charged monomer (ta ~ 211 ms, 318 Å2) and the spurious peak at ta ~ 267 ms. This “soft” TIMS spectrum now qualitatively agrees with those reported by Bowers [50] and Clemmer [29], except for the spurious ion.
Isotopic patterns for various regions in m/z 1061 ATD recorded under low-energy collisions (setting “B”). (a) ATD for m/z 1061 without m/z-selection under low-energy setting “B.” Peak positions used for extraction of isotopic pattern analysis for the m/z 1061 ATD are indicated. The apparent monomer feature visible at ta ~ 267 is a spurious ion coming from fragmentation of the 2BK/+3 dimer at m/z 707. For details, see Table S2 and Figure S2. (b) Extracted isotopic pattern for the peak at ta ~ 240 ms agrees strongly with the calculated isotopic pattern for a double-charged bradykinin dimer. (c) Extracted isotopic pattern for the peak at ta ~ 253 ms agrees strongly with the calculated isotopic pattern for a triply-charged bradykinin trimer. (d) Extracted isotopic pattern for the peak at ta ~ 260 ms agrees strongly with the calculated isotopic pattern for a quadruply-charged bradykinin tetramer. (The sharp peak ~ 211 ms corresponds to the bradykinin monomer)
We rationalize our observations as follows. Bradykinin assemblies are formed by ESI; they remain intact during ion mobility separation in tandem-TIMS; and they then enter the TIMS-hexapole interface. If the assemblies undergo energetic collisions with buffer gas particles in this region, they gain sufficient vibrational energy to dissociate into singly-charged monomer ions (Figure 3a). These are then detected by the mass analyzer at the arrival time given by the mobility of the intact precursor assembly. Nonetheless, low-energy collisions in the TIMS-hexapole interface sufficiently reduce heating and prevent dissociation of the assemblies (Figure 3b). Note also that we are the first to unambiguously confirm the long-postulated [29] presence of a bradykinin tetramer by an isotopic distribution (Figure 4), which underscores the ability to retain assemblies in our tandem-TIMS-Qq-TOF instrument. The data further show that presence of the spurious monomer ion is largely independent of the collision-energy in the TIMS-hexapole interface.
Ion Heating in the Collision Cell
The preceding discussion showed that we observed a spurious, highly compact spurious monomer for m/z 1061 (1.15 cm2/Vs, Figure 3b). Presence of this spurious ion was largely independent of the collision energy in the TIMS-hexapole interface, and persistent even under conditions that retained the quadruply charged tetramer n/z 4/4. The spurious ion vanishes, however, when we select m/z 1061 in the quadrupole (Figure 5e). This observation shows that the spurious ion does not exist prior to the quadrupole. Our data also show that this ion vanishes when we turn off the collision gas, and thus prevent energetic ion-neutral collisions, in the collision cell (Figure S3, Supporting Information). These observations suggest that the spurious ion is produced by dissociation of a different m/z in the collision cell, instead of in the TIMS-hexapole interface or the pusher region of the time-of-flight mass analyzer.
TIMS spectra for various bradykinin species under low-energy collisions in the TIMS-hexapole interface. (a) Arrival time distributions observed for m/z 1061 and m/z 531 produced by fragmentation of m/z 707 in the collision cell. (b–e) Arrival time distributions measured for bradykinin species m/z 354, m/z 531, m/z 707, m/z 1061 without quadrupole selection (red trace) are compared to arrival time distributions recorded after m/z selection in the quadrupole (black trace). Differences between the traces reveal ions that are produced in the collision cell. The comparison reveals that, under the conditions used, the only spurious ion created by fragmentation of a bradykinin species in post-TIMS instrument components is the artifact ion apparent as a highly compact monomer ion in the m/z 1061 spectrum (e, asterisk)
To identify the precursor of the spurious ion, we select m/z 707, m/z 531, and m/z 354 in the quadrupole (Figure S4, Supporting Information) and probe if these ions dissociate in the collision cell. Fragment ions are not observed for m/z 531. Water-loss is the only fragmentation pathway active for m/z 354, the activity of which is however negligible under the conditions used here. By contrast, m/z 707 dissociates into m/z 1061 and m/z 531. Further, the arrival time (ta ~ 268 ms) of the m/z 1061 fragment ion generated from m/z 707 corresponds well to the spurious monomer ion in the m/z 1061 spectrum (ta ~ 267 ms, see Figure 5a and e). The data thus indicate that the spurious species in the m/z 1061 spectrum is produced from fragmentation of m/z 707 (triply charged dimer) in the collision cell.
Nevertheless, quadrupole selection experiments of all bradykinin species (m/z 354, 531, 707, 1061, Figure 5b–e) show that the only spurious peak produced in the collision cell is the discussed spurious monomer in the m/z 1061 spectrum. Moreover, previously reported drift tube cross sections for bradykinin m/z 531 (348 Å2 and 344 Å2) [55, 56] agree well with ours (349 Å2 and 344 Å2, Figure 5; see also Table S2). Note that cross sections for m/z 354 are consistent with those reported by Park [51], and the spectra agree overall with those reported by Russell & Clemmer [54] and Bowers [50]. Furthermore, if desired, the collision energy in the collision cell can be further reduced by replacing the nitrogen collision gas with helium gas. These observations are not consistent with the idea that the bradykinin species observed in all these instruments arise from particularities during the electrospray process and/or the instrument components.
We are intrigued by the finding that our ion mobility spectra for the different bradykinin assemblies are highly consistent with those reported by Clemmer [29] and Bowers [50]. Intuitively, one might expect that the bradykinin assemblies may not be present in solution and that they are formed during the electrospray process. Hence, one might expect that the degree of oligomer formation depends on a number of experimental conditions including the electrospray ionization source (such as droplet size or electrospray voltage) or sample concentration. However, the experimental data (Figure 5) show that the bradykinin assembly species do not strongly depend on the experimental conditions of the IMS-MS measurements. Bowers and Clemmer both observe the same assembly species as we do with the main difference being that the relative abundances differ. Cross sections, even the relative abundances of multimeric to monomeric species in the m/z 1061 ion mobility spectrum, are overall consistent among the various instruments. Indeed, our tandem-TIMS-Qq-TOF instrument, which differs from the Bowers’ and Clemmer instruments in a number of aspects, even confirms the presence of a quadruply charged tetramer that was previously proposed by Clemmer [29]. These observations are not consistent with the idea that the bradykinin species observed in all these instruments arise from particularities of the electrospray process and/or the instrument operating components. In this regard, we note that Woods and Schultz previously discussed that non-covalent assemblies of small peptides may have functional biological relevance [42, 57].
We draw three conclusions from our measurements: (1) Bradykinin assemblies are preserved during ion mobility separation in a tandem-TIMS device when operated under “soft” conditions; (2) Bradykinin assemblies dissociate after elution from tandem-TIMS if they undergo energetic ion-neutral collisions; and (3) Minimizing the ion-neutral collision energy in post-TIMS components largely prevents fragmentation of bradykinin assemblies.
Summary and Conclusions
Our objective here was to determine how non-covalent peptide assemblies can be retained in a tandem-TIMS-Qq-TOF instrument. Basic considerations suggest that assemblies eluted from tandem-TIMS can undergo energetic collisions and dissociate in the TIMS-hexapole interface and in the collision cell. We demonstrated that assemblies of the nonapeptide bradykinin are largely retained after minimizing the ion-neutral collision energy in the TIMS-hexapole region and the collision cell. We unambiguously confirmed a bradykinin tetramer, which attests to the ability of retaining non-covalent assemblies throughout the tandem-TIMS-Qq-TOF instrument when carefully operated. We conclude that tandem-TIMS-Qq-TOF instruments are useful to study non-covalent peptide assemblies if both the tandem-TIMS device and post-TIMS ion optics are carefully optimized to minimize the energy of ion-neutral collisions. Nevertheless, we emphasize that energy barriers of assembly dissociation pathways depend on the specific system investigated. Therefore, when investigating less characterized assembly species, we recommend that the analyst asserts that assemblies remain intact throughout the entire measurement process using the approach discussed here.
References
Alberts, B.: The cell as a collection of protein machines: preparing the next generation of molecular biologists. Cell. 92, 291–294 (1998)
Koenen, R.R., von Hundelshausen, P., Nesmelova, I.V., Zernecke, A., Liehn, E.A., Sarabi, A., Kramp, B.K., Piccinini, A.M., Paludan, S.R., Kowalska, M.A., Kungl, A.J., Hackeng, T.M., Mayo, K.H., Weber, C.: Disrupting functional interactions between platelet chemokines inhibits atherosclerosis in hyperlipidemic mice. Nat. Med. 15, 97–103 (2009)
Kirkitadze, M.D., Bitan, G., Teplow, D.B.: Paradigm shifts in Alzheimer’s disease and other neurodegenerative disorders: the emerging role of oligomeric assemblies. J. Neurosci. Res. 69, 567–577 (2002)
Ross, C.A., Poirier, M.A.: Protein aggregation and neurodegenerative disease. Nat. Med. 10, S10–S17 (2004)
Bernstein, S.L., Dupuis, N.F., Lazo, N.D., Wyttenbach, T., Condron, M.M., Bitan, G., Teplow, D.B., Shea, J.-E., Ruotolo, B.T., Robinson, C.V., Bowers, M.T.: Amyloid-β protein oligomerization and the importance of tetramers and dodecamers in the aetiology of Alzheimer’s disease. Nat. Chem. 1, 326–331 (2009)
Uetrecht, C., Barbu, I.M., Shoemaker, G.K., van Duijn, E., Heck, A.J.R.: Interrogating viral capsid assembly with ion mobility–mass spectrometry. Nat. Chem. 3, 126–132 (2011)
Mason, E.A., McDaniel, E.W.: Transport properties of ions in gases. Wiley-VCH, Weinheim (1988)
Revercomb, H.E., Mason, E.A.: Theory of plasma chromatography/gaseous electrophoresis. Review. Anal. Chem. 47, 970–983 (1975)
Dupuis, N.F., Wu, C., Shea, J.-E., Bowers, M.T.: The amyloid formation mechanism in human IAPP: dimers have β-strand monomer−monomer interfaces. J. Am. Chem. Soc. 133, 7240–7243 (2011)
Ruotolo, B.T., Benesch, J.L.P., Sandercock, A.M., Hyung, S.-J., Robinson, C.V.: Ion mobility–mass spectrometry analysis of large protein complexes. Nat. Protoc. 3, 1139–1152 (2008)
Bleiholder, C., Bowers, M.T.: The solution assembly of biological molecules using ion mobility methods: from amino acids to amyloid β-protein. Annu. Rev. Anal. Chem. 10, 365–386 (2017)
Myung, S., Badman, E.R., Lee, Y.J., Clemmer, D.E.: Structural transitions of electrosprayed ubiquitin ions stored in an ion trap over ∼ 10 ms to 30 s. J. Phys. Chem. A. 106, 9976–9982 (2002)
Allen, S.J., Eaton, R.M., Bush, M.F.: Structural dynamics of native-like ions in the gas phase: results from tandem ion mobility of cytochrome c. Anal. Chem. 89, 7527–7534 (2017)
Wyttenbach, T., Bowers, M.T.: Structural stability from solution to the gas phase: native solution structure of ubiquitin survives analysis in a solvent-free ion mobility–mass spectrometry environment. J. Phys. Chem. B. 115, 12266–12275 (2011)
El-Baba, T.J., Woodall, D.W., Raab, S.A., Fuller, D.R., Laganowsky, A., Russell, D.H., Clemmer, D.E.: Melting proteins: evidence for multiple stable structures upon thermal denaturation of native ubiquitin from ion mobility spectrometry-mass spectrometry measurements. J. Am. Chem. Soc. 139, 6306–6309 (2017)
Laszlo, K.J., Munger, E.B., Bush, M.F.: Folding of protein ions in the gas phase after cation-to-anion proton-transfer reactions. J. Am. Chem. Soc. 138, 9581–9588 (2016)
Jhingree, J.R., Beveridge, R., Dickinson, E.R., Williams, J.P., Brown, J.M., Bellina, B., Barran, P.E.: Electron transfer with no dissociation ion mobility–mass spectrometry (ETnoD IM-MS). The effect of charge reduction on protein conformation. Int. J. Mass Spectrom. 413, 43–51 (2017)
Koeniger, S.L., Merenbloom, S.I., Sevugarajan, S., Clemmer, D.E.: Transfer of structural elements from compact to extended states in unsolvated ubiquitin. J. Am. Chem. Soc. 128, 11713–11719 (2006)
Jeziorski, B., Moszynski, R., Szalewicz, K.: Perturbation theory approach to intermolecular potential energy surfaces of van der Waals complexes. Chem. Rev. 94, 1887–1930 (1994)
Kemper, P.R., Bushnell, J., Van Koppen, P., Bowers, M.T.: Binding energies of cobalt(1+)-hydrogen-methane-ethane (Co+.cntdot.(H2/CH4/C2H6)1,2,3) clusters. J. Phys. Chem. 97, 1810–1817 (1993)
Hudgins, R.R., Dugourd, P., Tenenbaum, J.M., Jarrold, M.F.: Structural transitions in sodium chloride nanocrystals. Phys. Rev. Lett. 78, 4213–4216 (1997)
Fenn, J.B., Mann, M., Meng, C.K., Wong, S.F.: Electrospray ionization for mass spectrometry of large biomolecules. Science. 246, 64–71 (1989)
Karas, M., Bachmann, D., Hillenkamp, F.: Influence of the wavelength in high-irradiance ultraviolet laser desorption mass spectrometry of organic molecules. Anal. Chem. 57, 2935–2939 (1985)
Schnier, P.D., Price, W.D., Strittmatter, E.F., Williams, E.R.: Dissociation energetics and mechanisms of leucine enkephalin (M+H)+ and (2M+X)+ ions (X=H, Li, Na, K, and Rb) measured by blackbody infrared radiative dissociation. J. Am. Soc. Mass Spectrom. 8, 771–780 (1997)
Wyttenbach, T., Bowers, M.T.: Hydration of biomolecules. Chem. Phys. Lett. 480, 1–16 (2009)
Loo, J.A.: Studying noncovalent protein complexes by electrospray ionization mass spectrometry. Mass Spectrom. Rev. 16, 1–23 (1997)
Rostom, A.A., Robinson, C.V.: Detection of the intact GroEL chaperonin assembly by mass spectrometry. J. Am. Chem. Soc. 121, 4718–4719 (1999)
van den Heuvel, R.H., Heck, A.J.: Native protein mass spectrometry: from intact oligomers to functional machineries. Curr. Opin. Chem. Biol. 8, 519–526 (2004)
Counterman, A.E., Valentine, S.J., Srebalus, C.A., Henderson, S.C., Hoaglund, C.S., Clemmer, D.E.: High-order structure and dissociation of gaseous peptide aggregates that are hidden in mass spectra. J. Am. Soc. Mass Spectrom. 9, 743–759 (1998)
Bernstein, S.L., Liu, D., Wyttenbach, T., Bowers, M.T., Lee, J.C., Gray, H.B., Winkler, J.R.: Alpha-Synuclein: stable compact and extended monomeric structures and pH dependence of dimer formation. J. Am. Soc. Mass Spectrom. 15, 1435–1443 (2004)
Dupuis, N.F., Wu, C., Shea, J.-E., Bowers, M.T.: Human islet amyloid polypeptide monomers form ordered β-hairpins: a possible direct amyloidogenic precursor. J. Am. Chem. Soc. 131, 18283–18292 (2009)
Bleiholder, C., Dupuis, N.F., Wyttenbach, T., Bowers, M.T.: Ion mobility–mass spectrometry reveals a conformational conversion from random assembly to β-sheet in amyloid fibril formation. Nat. Chem. 3, 172–177 (2011)
Do, T.D., de Almeida, N.E.C., LaPointe, N.E., Chamas, A., Feinstein, S.C., Bowers, M.T.: Amino acid metaclusters: implications of growth trends on peptide self-assembly and structure. Anal. Chem. 88, 868–876 (2016)
Bleiholder, C., Do, T.D., Wu, C., Economou, N.J., Bernstein, S.S., Buratto, S.K., Shea, J.-E., Bowers, M.T.: Ion mobility spectrometry reveals the mechanism of amyloid formation of Aβ(25–35) and its modulation by inhibitors at the molecular level: epigallocatechin Gallate and Scyllo-inositol. J. Am. Chem. Soc. 135, 16926–16937 (2013)
Konijnenberg, A., Butterer, A., Sobott, F.: Native ion mobility-mass spectrometry and related methods in structural biology. Biochim. Biophys. Acta BBA - Protein Proteomics. 1834, 1239–1256 (2013)
McLuckey, S.A., Goeringer, D.E.: Slow heating methods in tandem mass spectrometry. J. Mass Spectrom. 32, 461–474 (1997)
Liu, F.C., Kirk, S.R., Bleiholder, C.: On the structural denaturation of biological analytes in trapped ion mobility spectrometry – mass spectrometry. Analyst. 141, 3722–3730 (2016)
Morsa, D., Gabelica, V., De Pauw, E.: Fragmentation and isomerization due to field heating in traveling wave ion mobility spectrometry. J. Am. Soc. Mass Spectrom. 25, 1384–1393 (2014)
Voronina, L., Masson, A., Kamrath, M., Schubert, F., Clemmer, D., Baldauf, C., Rizzo, T.: Conformations of prolyl–peptide bonds in the bradykinin 1–5 fragment in solution and in the gas phase. J. Am. Chem. Soc. 138, 9224–9233 (2016)
Gillig, K.J., Chen, C.-H.: Critical examination of gas-phase protein conformation/multimer ion formation by electrospray ion mobility-mass spectrometry. Anal. Chem. 85, 2177–2182 (2013)
Arndt, J.R., Kondalaji, S.G., Maurer, M.M., Parker, A., Legleiter, J., Valentine, S.J.: Huntingtin N-terminal monomeric and multimeric structures destabilized by covalent modification of heteroatomic residues. Biochemistry. 54, 4285–4296 (2015)
Woods, A.S., Koomen, J.M., Ruotolo, B.T., Gillig, K.J., Russel, D.H., Fuhrer, K., Gonin, M., Egan, T.F., Schultz, J.A.: A study of peptide—peptide interactions using MALDI ion mobility o-TOF and ESI mass spectrometry. J. Am. Soc. Mass Spectrom. 13, 166–169 (2002)
Liu, F.C., Ridgeway, M.E., Park, M.A., Bleiholder, C.: Tandem trapped ion mobility spectrometry. Analyst. 143, 2249–2258 (2018)
Hernandez, D.R., DeBord, J.D., Ridgeway, M.E., Kaplan, D.A., Park, M.A., Fernandez-Lima, F.: Ion dynamics in a trapped ion mobility spectrometer. Analyst. 139, 1913–1921 (2014)
Michelmann, K., Silveira, J.A., Ridgeway, M.E., Park, M.A.: Fundamentals of trapped ion mobility spectrometry. J. Am. Soc. Mass Spectrom. 26, 14–24 (2015)
Silveira, J.A., Michelmann, K., Ridgeway, M.E., Park, M.A.: Fundamentals of trapped ion mobility spectrometry part II: fluid dynamics. J. Am. Soc. Mass Spectrom. 27, 585–595 (2016)
Bleiholder, C.: Towards measuring ion mobilities in non-stationary gases and non-uniform and dynamic electric fields (I). Transport equation. Int. J. Mass Spectrom. 399–400, 1–9 (2016)
Butcher, D.J., Asano, K.G., Goeringer, D.E., McLuckey, S.A.: Thermal dissociation of gaseous bradykinin ions. J. Phys. Chem. A. 103, 8664–8671 (1999)
Schnier, P.D., Price, W.D., Jockusch, R.A., Williams, E.R.: Blackbody infrared radiative dissociation of bradykinin and its analogues: energetics, dynamics, and evidence for salt-bridge structures in the gas phase. J. Am. Chem. Soc. 118, 7178–7189 (1996)
Kemper, P.R., Dupuis, N.F., Bowers, M.T.: A new, higher resolution, ion mobility mass spectrometer. Int. J. Mass Spectrom. 287, 46–57 (2009)
Silveira, J.A., Ridgeway, M.E., Park, M.A.: High resolution trapped ion mobility spectrometery of peptides. Anal. Chem. 86, 5624–5627 (2014)
Chai, M., Young, M.N., Liu, F.C., Bleiholder, C.: A transferable, sample-independent calibration procedure for trapped ion mobility spectrometry (TIMS). Anal. Chem. 90, 9040–9047 (2018)
Stow, S.M., Causon, T.J., Zheng, X., Kurulugama, R.T., Mairinger, T., May, J.C., Rennie, E.E., Baker, E.S., Smith, R.D., McLean, J.A., Hann, S., Fjeldsted, J.C.: An interlaboratory evaluation of drift tube ion mobility–mass spectrometry collision cross section measurements. Anal. Chem. 89, 9048–9055 (2017)
Pierson, N.A., Chen, L., Valentine, S.J., Russell, D.H., Clemmer, D.E.: Number of solution states of bradykinin from ion mobility and mass spectrometry measurements. J. Am. Chem. Soc. 133, 13810–13813 (2011)
Bleiholder, C., Johnson, N.R., Contreras, S., Wyttenbach, T., Bowers, M.T.: Molecular structures and ion mobility cross sections: analysis of the effects of He and N 2 buffer gas. Anal. Chem. 87, 7196–7203 (2015)
Bush, M.F., Hall, Z., Giles, K., Hoyes, J., Robinson, C.V., Ruotolo, B.T.: Collision cross sections of proteins and their complexes: a calibration framework and database for gas-phase structural biology. Anal. Chem. 82, 9557–9565 (2010)
Woods, A.S., Jackson, S.N., Lewis, E.K., Egan, T., Muller, L., Tabet, J.-C., Schultz, J.A.: MALDI/post ionization-ion mobility mass spectrometry of noncovalent complexes of dopamine receptors’ epitopes. J. Proteome Res. 12, 1668–1677 (2013)
Acknowledgements
The authors thank Mark E. Ridgeway and Melvin A. Park for thoughtful discussions. Support from the Florida State University is gratefully acknowledged (C.B.). This material is based upon work supported by the National Science Foundation under CHE–1654608 (C.B.).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare that they have no conflicts of interest.
Electronic Supplementary Material
ESM 1
(PDF 978 kb)
Rights and permissions
About this article
Cite this article
Kirk, S.R., Liu, F.C., Cropley, T.C. et al. On the Preservation of Non-covalent Peptide Assemblies in a Tandem-Trapped Ion Mobility Spectrometer-Mass Spectrometer (TIMS-TIMS-MS). J. Am. Soc. Mass Spectrom. 30, 1204–1212 (2019). https://doi.org/10.1007/s13361-019-02200-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13361-019-02200-y