Abstract
Immediate diagnosis of human specimen is an essential prerequisites in medical routines. This study aimed to establish a novel cancer diagnostics system based on probe electrospray ionization-mass spectrometry (PESI-MS) combined with statistical data processing. PESI-MS uses a very fine acupuncture needle as a probe for sampling as well as for ionization. To demonstrate the applicability of PESI-MS for cancer diagnosis, we analyzed nine cases of clear cell renal cell carcinoma (ccRCC) by PESI-MS and processed the data by principal components analysis (PCA). Our system successfully delineated the differences in lipid composition between non-cancerous and cancerous regions. In this case, triacylglycerol (TAG) was reproducibly detected in the cancerous tissue of nine different individuals, the result being consistent with well-known profiles of ccRCC. Moreover, this system enabled us to detect the boundaries of cancerous regions based on the expression of TAG. These results strongly suggest that PESI-MS will be applicable to cancer diagnosis, especially when the number of data is augmented.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Noninvasiveness is one of the most important attributes required for diagnostics and therapeutics in clinical medicine. Therefore, nanotechnology prevails in medicine to meet those requirements. We have recently developed probe electrospray ionization (PESI), which uses a very fine needle to collect biological molecules as well as to ionize them [1]. Although mass spectrometry (MS) was introduced to the life sciences about 50 years ago [2], this technique is fundamentally a destructive procedure, for which the specimens have to be homogenized into a solution. In addition, the resultant homogenate is required to go through a multi-step procedure prior to mass spectrometry. Recently, mass spectrometric techniques that do not require sample preparation and vacuum have been developed, such as nanoelectrospray ionization (nanoESI), desorption electrospray ionization (DESI), direct analysis in real time (DART), and rapid evaporative ionization mass spectrometry (REIMS) [3–6]. PESI belongs to this group of techniques, and has several unique strengths when applied to biological samples [7].
PESI is a modified method of electrospray ionization (ESI) [8], and uses a solid needle, which acts as a probe for picking up samples and for ionizing them. PESI-mass spectrometry (PESI-MS) is very useful because it does not require any sample pretreatment and vacuum. Moreover, ion suppression does not occur, even in the presence of high salts [9]. By using a precision actuator system, the stroke of needle probe can be controlled to less than 10 μm from the surface of the sample [10], making it possible to reduce the sample volume to picoliters [11]. Consequently, the invasiveness of PESI-MS is almost negligible. In addition, solid needles are completely free from the clogging that is often cumbersome in nanoESI. By compiling a control program that automatically couples the PESI with MS, we can obtain mass spectra from biological samples in a few seconds.
Our previous studies have demonstrated that PESI-MS can reveal various biomolecules both in plant and animal specimens, such as low molecular weight compounds (e.g., urea, creatinine, and choline), carbohydrates, and lipids [9, 12]. Of particular importance, PESI is very good at detecting the lipid components in human tissues and living animals [10]. Taking into account the fact that cancer calls display changes in lipid composition [13], such as phospholipids [14], sphingolipids [15, 16], and cholesterol sulfates [17], we focused on determining the differences in lipid composition in cancer tissue for diagnoses of human renal cell carcinoma (RCC). We were able to delineate clear differences in spectral patterns with special references to lipids.
2 Materials and Methods
2.1 Preparation of Biological Samples
All examined kidney specimens from RCC patients (n = 9) were retrieved from the pathologic files of Yamanashi University Hospital. Tissue specimens were divided into noncancerous and cancerous regions, and immediately frozen for storage. These tissue specimens were measured by PESI-MS without any pretreatments. To prepare the homogenized samples, tissues were put in equal volumes of 0.9 % NaCl, followed by homogenization with a Dounce homogenizer. Informed consent was obtained from all patients before resection. The sections were examined by two expert pathologists (T.N. and R.K.), whose diagnoses agreed in all cases. All of the procedures concerning human materials and experiments were reviewed and approved by the ethical committee of the University of Yamanashi.
2.2 Histological Staining
Fresh tissues obtained from the surgical resections were routinely processed for paraffin sectioning. Four micrometer-thick sections were cut and stained with hematoxylin and eosin (H&E) [18]. For Sudan IV (Sigma, St. Louis, MO, USA) staining, formalin-fixed tissues were rinsed in water for 24 h. Then, 10 μm-thick frozen sections were cut and collected in distilled water, and subsequently mounted on glass slides with glycerin jelly (Sigma), to which Sudan IV solution was added.
2.3 Quantification of Triacylglycerol
RCC tissues were homogenized in 5 % NP-40 with 20 strokes of a Dounce homogenizer, and boiled for 4 min. The resultant homogenates were centrifuged at 15,000 g for 10 min, and supernatants were assayed by Triglyceride Quantification Kit (BioVision, Mountain View, CA, USA) according to the manufacturer’s instructions. The data were presented as the mean ± standard deviation (SD) of three independent experiments.
2.4 PESI Ion Source
Fine solid acupuncture needles (Seirin, Shizuoka, Japan) were used as sampling probes as well as electrospray emitters throughout the PESI-MS experiments [10, 11]. The needles had a diameter of 140 μm and a sub-μm tip radius. The basic protocol of a previous study was applied [10]. In brief, the needle was moved up and down along the vertical axis in an in-house-made actuator system [9]. The position of the needle was adjusted so that the needle touched the surface of the sample, which was placed on a sample stage.
The sample stage was placed below the sampling orifice, and when the needle was moved down to the lowest position, it touched and stitched into the sample surface at a desirable depth to pick up a small amount of biological fluid. The sample and sample stage were electrically insulated from the ground, and high voltage (HV) from a regulated HV power supply was applied when needle was moved to ionization position. As the needle was moved to the highest position, which was close to the ion-sampling orifice, electrospray was initiated once a sufficient electric field was established. The lowest and highest needle position will henceforth be referred as the sampling and ionization position, respectively. Electrospray commenced when a sufficient electrical field was established. Depending on needle alignment, the applied HV was in the range of 2–2.5 kV.
2.5 Mass Spectrometer
PESI-MS experiments were performed with an orthogonal acceleration time-of-flight mass spectrometer (AccuTOF, JEOL, Tokyo, Japan). The original electrospray ion source was removed, and the interlock was disabled to accommodate the PESI ion source. A linear actuator (Citizen, Chiba, Japan) was used to move the needle up and down along a vertical axis orthogonal to the axis of the ion-sampling orifice. The ions formed in the electrosprayed liquid droplets were sampled through an ion-sampling orifice with a diameter of 0.4 mm into a vacuum chamber. The accumulation time for the ion signal to acquire a single mass spectrum record was 0.4 s, and the measurement was continuously conducted for 5–15 min. Throughout the MS measurements, the needle probe was actuated at a frequency of 3 Hz. Thus, each single record corresponds approximately to the ions detected from a single needle sampling cycle. Mass spectra were measured using the Analog-to-Digital Converter (ADC)/continuous averager ion detection system, and one representative data point from >100 measurements were shown. Statistical analysis was performed by principal component analysis (PCA) (MATLAB ver. R2009a, The MathWorks, Inc. Natick, MA, USA). No background adjustment or smoothing filters were employed. PCA was performed on entire spectra from each tissue sample, and spectral data were used at a bin size of m/z 0.07. We adopted the first three principal components [19].
3 Results
3.1 PESI-MS Revealed Specific Profiles of Renal Cell Carcinoma
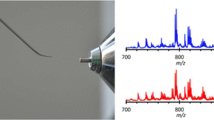
To test whether the PESI-MS is applicable to the diagnosis of malignant tumors, we focused on RCC as it takes on a distinct morphology when viewed as histopathologic specimens [20]. RCC is a unique model for our analyses, as it is known to accumulate triacylglycerol (TAG) in the cytoplasm [21]. Moreover, PESI-MS offers a distinct strength in detecting lipid components in the absence of any pretreatments. Therefore, we intended to identify the spectral pattern of RCC with special references to TAG, whose spectra appear around m/z 900. Tissue samples were freshly isolated from patients and were divided into noncancerous and cancerous regions. To confirm the histopathologic types of tumors, small pieces of tissue were set aside for paraffin sections stained with H&E (Figure 1a and b), which were used as correlative samples to the PESI-MS. As shown in Figure 1b, cancerous region displayed a typical honeycomb appearance, in which the cytoplasm are devoid of eosinophilic materials, showing so-called “clear cell” pattern; these microscopic findings are a hallmark of clear cell RCC (ccRCC). In contrast, noncancerous kidney tissue from the circumferential region of ccRCC exhibited normal arrangements of renal convoluted tubules with occasional renal glomeruli. To differentiate the cancerous region from the noncancerous circumference, specimens were put on a movable sampling stage (Supplementary Figure S1A), and the solid needle probe was positioned to make contact with the tissue to pick up the cellular components on the tip of the needle (Supplementary Figure S1B). Subsequently, HV was applied to the needle, which was synchronized with the needle motion to generate the electrospray (Supplementary Figure S1C). The spectral data were sent to the analysis software (MassCenter, ver. 1.3.10b), and mass spectra were extracted for analysis (Supplementary Figure S1D).
Kidney specimens were subjected to PESI-MS. Paraffin sections of kidney specimens from (a) noncancerous and (b) cancerous regions were stained with H&E. Photomicrograph indicates the typical morphological appearance of ccRCC (b). Bars indicate 100 μm. Insets indicate the low magnification images of specimens. Bars indicate 2.5 mm. Representative patterns of mass spectra from noncancerous (c) and (e) and cancerous (d) and (f) regions. Specimens analyzed by PESI-MS with no sample pretreatment (c) and (d). After direct analysis, specimens were homogenized with a glass homogenizer and analyzed by the same procedure (e) and (f). For molecular species, we expressed them as PC or TAG [X:Y], where X denotes the number of carbon atoms and Y the number of double bonds
As shown in Figure 1c and d, the PESI-MS could successfully generate distinct profiles of the lipid composition of cancerous region, in which TAGs were abundantly detected. Ions originating from phosphatidylcholines (PCs) (e.g., PC[34:1], PC[34:2], PC[36:2]) were shared by both noncancerous and cancerous regions, possibly because they are the principal components of cell membranes. Of note, some TAGs peaks (e.g., TAG[52:2], TAG[54:2], TAG[54:3]) were detected in high abundance only in cancerous region (Figure 1d). To ensure reproducibility of PESI-MS for diagnosis of ccRCC based on their lipid contents, noncancerous and cancerous specimens from eight individual ccRCC cases were analyzed (Supplementary Figure S2). Whilst there were some differences in ion intensity between each case, consistent results were obtained (Supplementary Figures S3 and S4). Moreover, when we homogenized the samples prior to PESI-MS, the spectral profiles were almost consistent with those of intact specimens (Figure 1e and f). These data collectively indicated that PESI-MS could delineate the cellular lipid components despite their intracellular origin. The measurement conditions were set to obtain a sufficient intensity of spectral peaks by a single pickup and ionization process. Mass spectra were essentially identical between the single trials and the averaged data of 1 min (180 measurements), suggesting the robustness and reproducibility of this system.
We next examined the detected TAG derived from the lipid droplet (LD) of the cytoplasm of ccRCC. The LD is an organelle for the storage of neutral lipids and cholesterol ester [22]. As shown in Figure 2a and b, LDs stained with Sudan IV solution, an indicator for neutral lipids, were abundantly expressed in cancerous region, while only a small amount of staining was identified in noncancerous region. Quantitative assay of TAG revealed approximately 2.9-fold deposition of TAG in the region of ccRCC (Figure 2c).
Identification and quantification of TAG. Cryosections of noncancerous (a) and cancerous (b) specimens were stained with Sudan IV solution to detect TAG. Bars indicate 100 μm. (c) The amount of TAG was quantified by an enzyme-based method. The data were expressed as the mean ± SD of three independent experiments. (d) Relative intensity of lipids detected from multiple patients. The data were plotted as a percentage of the highest intensity of peak in lipid spectrum, and were expressed as the mean ± SD of eight independent experiments
3.2 Validation of Diagnosis by Principal Component Analysis
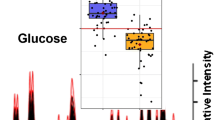
Next, we evaluated the relative abundance of each PC and TAG species from several different specimens (n = 8) (Figure 2d). PCs were detected in noncancerous and cancerous regions, and there were no significant differences in the ion abundance. In the cancerous region, on the contrary, the ion abundances of TAGs were much higher compared with those of noncancerous region (Figure 2d, Table 1, and Supplementary Figure S4). These results indicate that PESI-MS is a useful analytical tool for identification and quantification of lipids composition in pathological specimens.
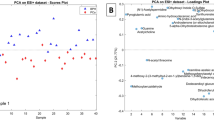
Although individual ions can be used to distinguish cancerous from noncancerous tissues like a tumor maker, we believe that the combinations of multiple ion species may be useful for the diagnosis of malignant tumors, as a single tumor marker is often incompetent for drawing a definitive diagnosis. One of the solutions for achieving our goal is the use of multivariate statistics to reduce the high-dimensional data that is acquired for each sample. In this study, PCA was employed to expand the analyses and identify parameters for diagnosis of ccRCC. PCA is an authentic mathematical method that uses a few composite variables to explain the covariance of multiple variables. Mass spectra of noncancerous and cancerous regions from eight individual ccRCC cases were subjected to PCA using MATLAB software. We took 4,345 eigenvector from m/z 700 to 1,000 as variables, which depicted lipid species. As shown in Figure 3a, mass peaks between 850 and 1,000 indicated higher factor loadings for PC1, showing almost 2-fold elevation compared with those between m/z 700 and 850. This finding was consistent with unique expression pattern of TAG in cancerous region (Figure 2d). While PC1 will be sufficient to discriminate between the noncancerous and cancerous tissues, we made PCA score plot from the first three components (Figure 3c), since the cumulative contribution rate exceeded 90 % at PC3 (Figure 3b).
Principal component analysis (PCA) of lipid spectra. Mass spectra of noncancerous and cancerous regions from eight individual patients were subjected to PCA. (a) Loading plot for PC1 (blue) is overlaid on the mass spectrum of cancerous region (gray). Abscissa represents m/z, whereas right and left ordinate, respectively, denote factor loadings and ion intensity. (b) Table of cumulative contribution rate for PC1 to PC6. (c) Three-dimensional PCA visualization of spectra obtained from noncancerous and cancerous specimens; the axes PC1, PC2, and PC3 correspond to the score of the first three components for each sample, respectively. Dashed plane (arbitrarily selected) indicates the boundary between noncancerous and cancerous regions
Abundant TAGs were reproducibly detected on eight individual ccRCC specimens and, interestingly, quite large differences in the composition of TAG were noted. Presumably, it may due to the differential activity of lipid metabolism among ccRCC cases. Nonetheless, lipid profiles were useful for discriminating cancerous tissue from noncancerous region even when we only employ the first three components of PCA. Therefore, we believe this method with the aid of PCA is very useful in the clinical diagnosis of cancer.
3.3 How Can We Draw a Line Between Noncancerous and Cancerous Tissue?
Very recently, ambient ionization methods have been successfully applied to tissue imaging [23]. PESI-MS also has the potential to be used for tissue imaging [24]. Moreover, we intend to install the PESI-based cancer diagnostics system to the operating room so as to profile biological molecules from human organs during surgery. In imaging MS, the sample surface is continuously scanned while ionizing the sample and recording the resulting mass spectra. Mass spectra are catalogued for each pixel, followed by compilation of data to create a map of molecules over the entire surface of a sample. In this study, to test whether the PESI-MS detects the boundary of cancerous region, line-scanning analysis (a modified method of imaging PESI-MS) was performed on excised kidney tissue composed of noncancerous and cancerous regions (Figure 4). A correlative section of kidney tissue was stained with H&E (Figure 4a) and analyzed by PESI-MS (Figure 4b). Spectra were obtained from 10 consecutive but separate points, as depicted in Figure 4b. The pattern of spectra changed drastically at the boundary between noncancerous and cancerous regions, demonstrating the sensitivity of PESI-based cancer diagnostics system in the diagnosis of malignancies. The ion intensities of PC (m/z 756.61-826.62) were constant in noncancerous and cancerous tissues (Figure 4c), because PC is one of the most common phospholipids in various cell types. In contrast, the ion intensities of TAG (m/z 869.76–925.82) increased specifically in cancerous regions (Figure 4d). As tumor regions per se were well circumscribed with a fibrous capsule, the transition of data at the boundary was very sharp. We assume that these differences may reflect the stage or malignancy of cancer. However, further analysis is essential to verify our hypothesis. On the fibrous capsules surrounding tumor mass, the ion intensity of PC and TAG showed no special tendency in any different points (data not shown). Nevertheless, ion peaks of TAGs are characteristic of cancerous tissue in ccRCC.
Kidney specimens were subjected to line analysis. (a) Paraffin sections of kidney specimens containing non-cancerous and cancerous regions were stained with H&E. (b) Photograph indicates appearance of kidney specimens. Bars indicate 5 mm. Mass spectra were obtained from 10 consecutive points [(b), 1–10]. The relative intensities of lipid peaks of PC at m/z 756.61–826.62 (c) and TAG at m/z 869.76–925.82 (d) are shown
4 Discussion
In this study, we validated the usefulness of a PESI-based cancer diagnostic method [7] to differentiate cancerous tissue, using a set of specific lipid components abundantly expressed in cancerous tissue. These results were further confirmed in renal specimens excised from different clinical cases. PESI uses very fine needles with a tip diameter of less than 1 micrometer, and thus it is a noninvasive technology applicable to living animals [9]. Another advantage of this method is its robustness when using raw materials, as no pretreatment is required prior to measurement. Nevertheless, to introduce this technique to the clinic, we need to consider the following points: (1) reproducibility of spectral data and optimization of measurements; (2) annotation and clustering of spectra for specific histotypes; (3) diagnostic software based on the statistical analyses.
First, in this study, we chose ccRCC as it displays distinct histopathologic features characterized by the transparency of the cytoplasm attributable to accumulated droplets of glycogen, phospholipids, and neutral lipids [25]. Therefore, it is a malignant tumor type suitable to testify the reproducibility of data. While neutral lipids may be used in the diagnosis of ccRCC, this method is far from ideal as it requires time-consuming procedures. In contrast, PESI-MS offers a rapid analytical tool to detect the lipid components without any cumbersome pretreatments. Moreover, detection of TAGs from ccRCC is completed in less than 1 min. Considering the high reproducibility of our data on ccRCC, it is a very robust method in terms of cost, time, and handling. The next issue that must be tackled is to ascertain whether this technique is applicable to more heterogeneous and complex malignant tumor types such as brain tumors. Other groups have succeeded in diagnosing brain tumors by DESI [26, 27]; our method would add something important, such as rapidness and ease in handling.
Second, annotation is a major issue in MS. In standard experiments, the annotation of molecules detected as spectra is one of the most important tasks required in MS. However, when we look at the spectra of different cancers from different origins, we noticed a similarity in the spectral pattern among them, which was not observed in the noncancerous tissue. For example, in cancerous regions of ccRCC, the intensity of ion peaks between m/z 650 and 700 (e.g., 671.6 and 673.6) were higher than in noncancerous regions (Supplementary Figure S3). Although, these peaks have not been annotated (but are presumably a family of phospholipids), the same finding was observed in hepatocellular carcinoma artificially induced in mice (unpublished data). These results indicate that a constellation of lipid peaks between m/z 650 and 700 are regarded as a collective biomarker in certain carcinoma and, therefore, can be utilized in the diagnosis of various cancers. Since PESI-MS also detects glycolipids, carbohydrates, and amino acids [9, 12, 24], it may be possible to discover a novel biomarker composed of several spectral patterns by PESI-MS.
PC contains a phosphocholine head and two fatty acid chains, and TAG contains glycerol backbone and three fatty acid chains. The number of carbon atoms and double bonds determine the subclass of lipids, such as palmitic acid (16:0), oleic acid (18:1), and linoleic acid (18:2). The proportion of lipid composition may reflect the physiological status of cells. For example, a previous study suggested that the lipid composition observed in the urine of breast cancer patients differed from that of healthy patients [28]. In this study, we annotated some representative peaks (Figure 1 and Table 1), and we are now trying to determine the number of carbon atoms and double bonds in all lipid peaks. These numbers may infer cancer-specific lipid profiles, and useful for diagnosis of cancers that do not accumulate TAG. With this in mind, and combined with the concept of collective biomarkers, we will direct our research towards the development of new diagnostic methods for malignant tumors. To achieve this, new systems of data processing will be indispensable.
Third, as ongoing project, we try to develop a new data processing system. In the present study, we analyzed the mass spectra using PCA, by which large number of peaks with high factor loadings were taken as PCs. However, considering the diverse nature of cancer species, which is reflected on the broad distribution of factor loadings over wide range of spectra, PCA has inherent technical limitations in the judgment of cancers. To address this issue, we are now trying to apply the dual penalized logistic regression machine (dPLRM) [29, 30].
5 Conclusion
In this study, we evaluated the usefulness of PESI-MS based cancer diagnosis system. When analyzing the human ccRCC, we could successfully obtain distinct profiles of lipids in cancerous region, which were further ascertained by principal component scores from mass spectrum of eight individual patients. Moreover, PESI-MS could detect the boundary of noncancerous and cancerous regions by line scanning. These two lines of evidence are very useful for establishing a novel cancer diagnostic system based on PESI-MS. Of note, as the profiles of lipid are quite unique to cancerous tissues, we take advantages of its property to construct a database for making diagnosis. For this purpose, we will further accumulate larger number of samples that is essential for successfully establishing a diagnostics system. Furthermore, to demonstrate the versatility of our diagnosis system, we are now trying to diagnose human hepatocellular carcinoma and lymph node metastasis by PESI-MS.
References
Hiraoka, K., Nishidate, K., Mori, K., Asakawa, D., Suzuki, S.: Development of probe electrospray using a solid needle. Rapid Commun. Mass Spectrom. 21, 3139–3144 (2007)
Dalgliesh, C.E., Horning, E.C., Horning, M.G., Knox, K.L., Yarger, K.: A gas-liquid-chromatographic procedure for separating a wide range of metabolites occurring in urine or tissue extracts. Biochem. J. 101, 792–810 (1966)
Wilm, M., Mann, M.: Analytical properties of the nanoelectrospray ion source. Anal. Chem. 68, 1–8 (1996)
Takáts, Z., Wiseman, J.M., Gologan, B., Cooks, R.G.: Mass spectrometry sampling under ambient conditions with desorption electrospray ionization. Science 306, 471–473 (2004)
Cody, R.B., Laramée, J.A., Durst, H.D.: Versatile new ion source for the analysis of materials in open air under ambient conditions. Anal. Chem. 77, 2297–2302 (2005)
Balog, J., Szaniszlo, T., Schaefer, K.C., Denes, J., Lopata, A., Godorhazy, L., Szalay, D., Balogh, L., Sasi-Szabo, L., Toth, M., Takats, Z.: Identification of biological tissues by rapid evaporative ionization mass spectrometry. Anal. Chem. 82, 7343–7350 (2010)
Takeda, S., Yoshimura, K., Hiraoka, K.: Innovations in analytical oncology—status quo of mass spectrometry-based diagnostics for malignant tumor. J. Anal. Oncol. 1, 74–80 (2012)
Fenn, J.B., Mann, M., Meng, C.K., Wong, S.F., Whitehouse, C.M.: Electrospray ionization for mass spectrometry of large biomolecules. Science 246, 64–71 (1989)
Chen, L.C., Nishidate, K., Saito, Y., Mori, K., Asakawa, D., Takeda, S., Kubota, T., Terada, N., Hashimoto, Y., Hori, H., Hiraoka, K.: Application of probe electrospray to direct ambient analysis of biological samples. Rapid Commun. Mass Spectrom 22, 2366–2374 (2008)
Yoshimura, K., Chen, L.C., Yu, Z., Hiraoka, K., Takeda, S.: Real-time analysis of living animals by electrospray ionization mass spectrometry. Anal. Biochem. 417, 195–201 (2011)
Yoshimura, K., Chen, L.C., Asakawa, D., Hiraoka, K., Takeda, S.: Physical properties of the probe electrospray ionization (PESI) needle applied to the biological samples. J. Mass Spectrom. 44, 978–985 (2009)
Yu, Z., Chen, L.C., Suzuki, H., Ariyada, O., Erra-Balsells, R., Nonami, H., Hiraoka, K.: Direct profiling of phytochemicals in tulip tissues and in vivo monitoring of the change of carbohydrate content in tulip bulbs by probe electrospray ionization mass spectrometry. J. Am. Soc. Mass Spectrom. 20, 2304–2311 (2009)
Fernandis, A.Z., Wenk, M.R.: Lipid-based biomarkers for cancer. J. Chromatogr. B. Anal. Technol. Biomed. Life Sci. 877, 2830–2835 (2009)
Riedl, S., Rinner, B., Asslaber, M., Schaider, H., Walzer, S., Novak, A., Lohner, K., Zweytick, D.: In search of a novel target-phosphatidylserine exposed by non-apoptotic tumor cells and metastases of malignancies with poor treatment efficacy. Biochim. Biophys. Acta 1808, 2638–2645 (2011)
Ito, A., Levery, S.B., Saito, S., Satoh, M., Hakomori, S.: A novel ganglioside isolated from renal cell carcinoma. J. Biol. Chem. 276, 16695–16703 (2001)
Biswas, K., Richmond, A., Rayman, P., Biswas, S., Thornton, M., Sa, G., Das, T., Zhang, R., Chahlavi, A., Tannenbaum, C.S., Novick, A., Bukowski, R., Finke, J.H.: M2 expression in renal cell carcinoma: Potential role in tumor-induced T-cell dysfunction. Cancer Res. 66, 6816–6825 (2006)
Kiguchi, K., Iwamori, M., Yamanouchi, S., Ishiwata, I., Saga, M., Amemiya, A.: Coexpression of cholesterol sulfate and cytokeratin as tumor markers in well-differentiated squamous cell carcinoma of the human uterine cervix. Clin. Cancer Res. 4, 2985–2990 (1998)
Takeda, S., Yonekawa, Y., Tanaka, Y., Okada, Y., Nonaka, S., Hirokawa, N.: Left–right asymmetry and kinesin superfamily protein KIF3A: New insights in determination of laterality and mesoderm induction by kif3A–/– mice analysis. J. Cell. Biol. 145, 825–836 (1999)
Zissis, K.D., Brereton, R.G., Dunkerley, S.: Principal components scores and loadings plots for visualization of the electrospray ionization liquid chromatography mass spectra of a mixture of chlorophyll degradation products at different cone voltages. Rapid Commun. Mass Spectrom. 13, 1755–1761 (1999)
Grignon, D.J., Che, M.: Clear cell renal cell carcinoma. Clin. Lab. Med. 25, 305–316 (2005)
Hoffmann, K., Blaudszun, J., Brunken, C., Höpker, W.W., Tauber, R., Steinhart, H.: Lipid class distribution of fatty acids including conjugated linoleic acids in healthy and cancerous parts of human kidneys. Lipids 40, 1057–1062 (2005)
Shibata, M., Yoshimura, K., Furuya, N., Koike, M., Ueno, T., Komatsu, M., Arai, H., Tanaka, K., Kominami, E., Uchiyama, Y.: The MAP1-LC3 conjugation system is involved in lipid droplet formation. Biochem. Biophys. Res. Commun. 382, 419–423 (2009)
Huang, M.Z., Cheng, S.C., Cho, Y.T., Shiea, J.: Ambient ionization mass spectrometry: A tutorial. Anal. Chim. Acta 702, 1–15 (2011)
Chen, L.C., Yoshimura, K., Yu, Z., Iwata, R., Ito, H., Suzuki, H., Mori, K., Ariyada, O., Takeda, S., Kubota, T., Hiraoka, K.: Ambient imaging mass spectrometry by electrospray ionization using solid needle as sampling probe. J. Mass Spectrom. 44, 1469–1477 (2009)
Hajdu, S.I., Savino, A., Hajdu, E.O., Koss, L.G.: Cytologic diagnosis of renal cell carcinoma with the aid of fat stain. Acta. Cytol. 15, 31–33 (1971)
Eberlin, L.S., Dill, A.L., Golby, A.J., Ligon, K.L., Wiseman, J.M., Cooks, R.G., Agar, N.Y.: Discrimination of human astrocytoma subtypes by lipid analysis using desorption electrospray ionization imaging mass spectrometry. Angew. Chem. Int. Ed. Engl. 49, 5953–5956 (2010)
Agar, N.Y., Golby, A.J., Ligon, K.L., Norton, I., Mohan, V., Wiseman, J.M., Tannenbaum, A., Jolesz, F.A.: Development of stereotactic mass spectrometry for brain tumor surgery. Neurosurgery 68, 280–289 (2011)
Min, H.K., Kong, G., Moon, M.H.: Quantitative analysis of urinary phospholipids found in patients with breast cancer by nanoflow liquid chromatography-tandem mass spectrometry: II. Negative ion mode analysis of four phospholipid classes. Anal. Bioanal. Chem. 396, 1273–1280 (2010)
Tanabe, K.: Penalized logistic regression machines: New methods for statistical prediction 1. ISM Cooperative Research Report 143, Estimation and Smoothing Methods in Nonparametric Statistical Models. Institute of Statistical Mathematics, Tokyo: pp. 163–194 (2001a)
Tanabe, K.: Penalized logistic regression machines: New methods for statistical prediction 2. Proceedings of 2001 Workshop on Information Based Induction Science (IBIS2001), Tokyo, Japan, July 2001 pp. 71–76 (2001b)
Acknowledgments
The authors thank A. Iizuka and K. Sawanobori for their assistance with this project. This research was supported by Kato Memorial Bioscience Foundation (to K.Y.), and KAKENHI [23790619; Grant-in-Aid for Young Scientists (B)] (to K.Y.).
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(DOC 2147 kb)
Rights and permissions
About this article
Cite this article
Yoshimura, K., Chen, L.C., Mandal, M.K. et al. Analysis of Renal Cell Carcinoma as a First Step for Developing Mass Spectrometry-Based Diagnostics. J. Am. Soc. Mass Spectrom. 23, 1741–1749 (2012). https://doi.org/10.1007/s13361-012-0447-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13361-012-0447-2