Abstract
microRNAs are non-coding molecules, approximately 22 nucleotides in length, that regulate various cellular processes. A growing body of evidence has suggested that their dysregulated expression is involved in the pathogenesis of diverse diseases, including diabetes mellitus type 2 (DM2). Early onset of this chronic and complex metabolic disorder is frequently undiagnosed, leading to the development of severe diabetic complications. Notably, DM2 prevalence is rising globally and an increasing number of articles demonstrate that DM2 susceptibility, development, and progression differ between males and females. Therefore, this paper discusses the role of microRNAs as a source of novel diagnostic biomarkers for DM2 and aims to underline the importance of sex disparity in biomarkers research. Taking into account an urgent need for the development of sex-specific diagnostic strategies in DM2, recent results have shown that circulating miRNAs are promising candidates for sex-biased biomarkers.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
International Diabetes Federation (IDF) projects that the worldwide prevalence of diabetes among adults (20–79 years old) will systematically rise to finally reach about 700 million people by 2045 (Saeedi et al. 2019). Notably, many cases of diabetes are undiagnosed until the appearance of complications (Association 2003), which may lead to severe disorders development such as cardiovascular diseases, renal failure, retinopathy, neuropathy, and even lower extremity amputations (Susan van et al. 2010; Schmidt 2018). This data shows that diabetes continues to be one of the most crucial public health issues (Saeedi et al. 2019). Diabetes mellitus type 2 (T2DM or DM2) represents the majority of all diabetes cases (Chatterjee et al. 2017). This chronic metabolic disorder is characterized by hyperglycemia along with insulin resistance and usually relative insulin deficiency (Association 2014). Currently, an increasing number of studies indicate that susceptibility and pathogenesis of DM2 are sex-biased (Kautzky-Willer et al. 2016). In general, the interplay of biological, environmental, social, or even lifestyle factors contributes to observed sex differences in DM2 (Kautzky-Willer et al. 2016). Moreover, the results of the clinical research from different populations confirm the significance of sex disparity in DM2. For example, comparative analysis in the European population showed that men develop DM2 at an earlier age and at a lower body mass index than women (Logue et al. 2011). A population-based study from Italy indicated a greater impact of diabetes on cardiovascular mortality in women (Ballotari et al. 2015). Recently, Li et al. 2021 demonstrated that the most common panel of anthropometric and biochemical parameters better predicts DM2 in women than in men from the Hainan population. In addition, utilizing conventional biomarkers such as glucose homeostasis-related parameters, lipid classes, and glycated hemoglobin displays some limitations such as not full specificity for DM2 and presence in subjects already suffering from metabolic disorders (Urdea et al. 2009; Bhatia et al. 2015). Given all the abovementioned reports, there is an urgent need to find novel diagnostic biomarkers, especially those indicating the early onset of DM2 and taking into account sex differences.
To date, many studies have proved that molecules classified as non-coding RNAs (ncRNAs) may serve as a valuable source of prognostic, diagnostic, and even predictive biomarkers for many human diseases (Borga et al. 2019). Non-coding RNAs play important role in a variety of fundamental biological processes such as development, metabolism, cell proliferation, and apoptosis; therefore, their aberrant expression is associated with the development of various diseases (Wang et al. 2019). Interestingly, the results of the Encyclopedia of DNA Elements project reported that many coding and non-coding transcripts are processed into steady-stable precursors of small non-coding RNA (Dunham et al. 2012). Notably, microRNAs (miRNA) are a group of small non-coding RNA, approximately 22 nucleotides in length (Bartel 2018), which exert an important regulatory role in cells usually by inhibiting the expression of genes at the post-transcriptional level (Bartel 2004). A growing body of literature shows that many miRNAs are dysregulated in patients with DM2 and may play role in pathological mechanisms underlying the development of diabetes complications (He et al. 2017; Shi et al. 2020). Therefore, in this article, we discuss the potential role of miRNA as a source of biomarkers for DM2. In addition, with the growing emphasis on sex as a relevant biological variable (Mannon et al. 2020), we also underscore the importance of sex differences in biomarkers research.
miRNA biogenesis and association with DM2
According to the miRBase database (v22), the human genome encodes more than 2500 miRNAs (Kozomara et al. 2019). Their genes are located in the protein-coding or non-coding part of the genome and could be transcribed by polymerase II (Lee et al. 2004) or III (Borchert et al. 2006). Maturation of miRNA requires stepwise processing (Lee et al. 2002) and takes place via a canonical (Bartel 2004) or non-canonical pathway (Miyoshi et al. 2010; Abdelfattah et al. 2014). However, the synthesis of the majority of miRNAs is via the canonical pathway (Fig. 1), in which firstly polymerase II transcribes a long primary miRNA (pri-miRNA) in the nucleus (Kim et al. 2009; O’Brien et al. 2018). This transcript has at least one hairpin structure and could be caped and polyadenylated (Cai et al. 2004). Furthermore, the microprocessor complex, consisting of RNA binding protein DiGeorge Syndrome Critical Region 8 (DGCR8) and an RNase III enzyme, Drosha, generates from pri-miRNAs the hairpin-like precursor miRNAs (pre-miRNAs), ~ 70 nt in length (Lee et al. 2003; O’Brien et al. 2018). The next step requires translocation of pre-miRNA to the cytoplasm by an exportin 5 (XPO5)/RanGTP complex (Lund et al. 2004), where the RNase III endonuclease, Dicer process pre-miRNA into miRNA duplex, ~ 22 nt in length (Han et al. 2004). One strand of this duplex miRNA usually serves as mature miRNA (Han et al. 2004) and is incorporated into an RNA-induced silencing complex, RISC (Bartel 2004). Based on miRNA sequence, RISC binds into target mRNA transcript, usually within 3’ untranslated region (3’ UTR), leading to repression of translation or transcript degradation (Kim et al. 2009). More detailed information about miRNA biogenesis, function, and mechanism of action could be found in (Bartel 2004, 2018; Krol et al. 2010; O’Brien et al. 2018).
Biogenesis of miRNA via canonical pathway and polymorphism of miRNAs and their associated genes. Polymerase II (Pol II) transcribes a long primary miRNA (pri-miRNA) in the nucleus. Then, the microprocessor complex, consisting of RNA binding protein DiGeorge Syndrome Critical Region 8 (DGCR8) and Drosha, generates hairpin-like precursor miRNAs (pre-miRNAs) from pri-miRNAs. pre-miRNA is translocated into the cytoplasm by an exportin 5 (XPO5)/RanGTP complex. In the next step, Dicer process pre-miRNA into miRNA duplex. One strand of this duplex miRNA is incorporated into RISC. After that, RISC can bind into a 3’ untranslated region (3’ UTR) of target mRNA, leading to repression of translation or transcript degradation. Polymorphism of genes that are engaged in miRNA expression and maturation may lead to an increased risk of diabetes mellitus type 2 development
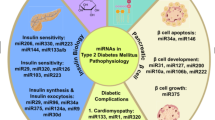
Interestingly, miRNAs are predicted to regulate the majority of human genes (Friedman et al. 2009). Moreover, the regulatory network of miRNAs and their targets is very complex, as one miRNA could regulate multiple genes and one gene could be controlled by many miRNAs (Peter 2010). Current findings demonstrate that miRNAs exert a role in the pathogenesis of diabetes mellitus type 2 (reviewed in Yaribeygi et al. 2018; Deng and Guo 2019). For example, given that insulin resistance is the main reason for type 2 diabetes, miRNAs can modulate proteins engaged in insulin signaling, thus affecting insulin resistance and glucose homeostasis (Deng and Guo 2019). In addition, they play role in pancreatic β-cell development and survival (Filios and Shalev 2015) and even regulate lipid metabolism (Agbu and Carthew 2021). Studies conducted in various populations have shown that polymorphism in miRNA genes is associated with DM2 and diabetes complications (Elfaki et al. 2019). Moreover, Zhu et al. (2021) have reported that in sex-stratified analysis, single-nucleotide polymorphism (SNP) in miRNA genes, let-7a-1 rs13293512, and miR-27a rs895819, was associated with increased risk of DM2 in the male subjects from the Chinese population. Overall, polymorphism in miRNA genes (localized within a sequence of mature miRNA, pre-miRNA, pri-mRNA, and regulatory regions of miRNA) may lead to changes in mature miRNAs structure and their expression level resulting in dysregulated expression of miRNA’s target genes (Gong et al. 2014). The impact of genetic variants on elevated susceptibility to DM2 and its vascular complications is also observed in genes encoding proteins engaged in miRNA biogenesis which could potentially lead to changes in the miRNA processing (Wen et al. 2019). Wen et al. (2019) have found that rs13078 SNP in gene encoding DICER1 is associated with decreased susceptibility of DM2, but rs14035 SNP in gene encoding RAN resulted in a higher risk of developing macrovascular complications in the Southern Chinese Population. Given that miRNA realize their function via binding to their target mRNAs, polymorphism in 3’ UTR of mRNAs may affect miRNA binding sites leading to dysregulation of protein expression (Gong et al. 2014). For example, Zhang et al. (2013a) have proved that rs3811463 polymorphism in the let-7 target region of the Lin28 gene is associated with increased susceptibility to DM2 in females from the Chinese Han Population. In general, regardless of cause and involved mechanism, aberrant expression and function of miRNA are linked with DM2 development.
Circulating miRNA as potential biomarkers for DM2
Apart from miRNAs that exert regulatory function inside cells, scientists have discovered extracellular, circulating miRNAs (Chim et al. 2008). It appears that miRNAs could be secreted by cells in extracellular vesicles or are transported bound with different proteins such as Argonaut proteins and high-density lipoproteins (HDL) (Nik Mohamed Kamal and Shahidan 2020). Moreover, their expression is detected in various physiological fluids and may change under different pathophysiological conditions (Weber et al. 2010). In addition, high stability, resilience to endogenous ribonucleases, along with the possibility of detection by available methods, makes circulating miRNA suitable candidates as biomarkers (Chen et al. 2008) for various diseases including cancers (George and Mittal 2010; Yu et al. 2011; Lan et al. 2015), diabetes (Guay and Regazzi 2013; Mirra et al. 2015), cardiovascular diseases (Xu et al. 2012; Kondkar and Abu-Amero 2015; Felekkis and Papaneophytou 2020), and neurodegenerative disorders (Grasso et al. 2014). A recent review and bioinformatics analysis have shown that expression of circulating miRNA miR-30a-5p, miR-30d-5p, and miR-30c-5p is associated with glucose metabolism, inflammation, platelet reactivity, and endothelial dysfunction in diabetes type 2 and underscores their utility as biomarkers for detection and progression of disease (Pordzik et al. 2019). Interestingly, among genes regulated in DM2, PRKAR1A (protein kinase cAMP-dependent type I regulatory subunit alpha), may be one of the most significant targets of miRNAs involved in glucose metabolism, insulin signaling, and blood coagulation. In the cells, protein kinase A is engaged in proliferation and cell growth (Pordzik et al. 2019). These findings are in line with the results of previous studies indicating that miRNA could be used as potential biomarkers for DM2 (Zhu and Leung 2015). As a result of the meta-analysis of DM2 miRNA expression profiling studies, Zhu and Leung (2015) identified eight circulating miRNAs (miR-103, miR-107, miR-132, miR-144, miR-142-3p, miR-29a, miR-34a, and miR-375) that may serve as promising biomarkers. These miRNAs regulate processes such as insulin secretion and signaling, or adipogenesis (Zhu and Leung 2015).
Considering the importance of early detection of DM2, continuous efforts are being made to find miRNAs that enable distinguishing the pre-diabetes state from the early onset of disease or identify individuals with a high risk of DM2 development (Bhatia et al. 2015) (Table 1). For example, Yan et al. (2020) built a multi-parameter diagnostic model to discriminate the impaired glucose regulation from DM2. Specifically, the expression levels of miR-148b, miR-223, miR-130a, and miR-19a in the serum may potentially be used for distinguishing patients with DM2 from that suffering from impaired glucose regulation (Yan et al. 2020). Zhang et al. (2013b) have shown that miR-126 had significantly reduced expression levels in DM2 susceptible individuals and diagnosed DM2 patients in comparison to controls which may make miR-126 useful to identify individuals with a higher risk of DM2. In turn, Jiménez-Lucena et al. (2018) in a prospective study have demonstrated that coupling the expression values of 9 miRNAs with a conventional biomarker, glycated hemoglobin (HbA1c), enables prediction of the DM2 development with high accuracy.
During DM2, diabetes-induced vascular dysfunction may lead to macrovascular complications manifesting with cardiovascular diseases, as well as microvascular complications that are associated with the development of retinopathy, neuropathy, and nephropathy (Beckman and Creager 2016). Examples of circulating miRNAs that may serve as biomarkers for various diabetes vascular complications are presented in Table 2. Prevention of diabetes complications requires the selection of appropriate medication which, in an ideal situation, would be tailored to the patient. It occurs that circulating miRNAs could also be used to predict the effects of DM2 therapy (Table 3). After 1 year of treatment with glucagon-like peptide 1 receptor agonists (GLP1-RA), patients with initially higher expression levels of eight circulating miRNAs had a better response to therapy than those with lower levels (Formichi et al. 2021). In addition to the abovementioned examples, the recent results showed that circulating miRNA may be utilized to select a personalized diet model to prevent the development of DM2. In the case of patients with cardiovascular diseases, initial high miR-150 along with low miR-29a, miR-28-3p, and miR-126 in subjects on a Mediterranean diet, and low initial level of miR-145 in subjects on low-fat high-complex carbohydrate diet, was associated with a higher risk of development DM2 (Jimenez-Lucena et al. 2021). Therefore, the profile of circulating miRNAs may be also used to select a group of patients for a particular treatment in DM2 (Formichi et al. 2021) and nutritional therapy to lower the risk of disease development (Jimenez-Lucena et al. 2021).
Sex difference in circulating miRNA in DM2
Scientists have discovered the sex-biased expression of microRNA in many pathological conditions, including metabolic diseases (Sharma and Eghbali 2014). For example, Wang et al. (2013) demonstrated that elevated expression level of circulating let-7 g and miR-221 was associated with metabolic syndrome. This disorder is characterized by clinical features such as abdominal adiposity, insulin resistance, hypertension, dyslipidemia, inflammation, and a prothrombotic state, and is linked with an increased risk of DM2 development. Further analysis revealed a sex-specific expression pattern of both investigated miRNAs. Their level was elevated in the serum of women with metabolic syndrome in comparison to healthy females, while this pattern was not reported in men (Wang et al. 2013).
Surprisingly, not every study takes into account sex differences in the circulating miRNA expression of DM2 patients, even though sex hormones, as well as X-linked genes, may potentially regulate their expression (Corral-Fernández et al. 2013; Sharma and Eghbali 2014). Interestingly, results obtained by Prabu et al. (2015) showed that there are associations between the sex of patients with pre-diabetic or DM2 state and circular miRNA expression. Moreover, the aberrant expression of some circulating miRNA may be found only in one sex. In the beginning, they noticed that in Asian Indian female and male patients analyzed together, miR-142-3p were not differentially expressed between pre-diabetes or DM2, and controls. In turn, when the analysis of miRNA expression was sex stratified, an increased level of the abovementioned miRNA was shown in the samples of pre-diabetic women versus healthy samples. In addition, circulating miR-423-5p and miR-128 altered expression was confirmed in comparison of pre-diabetes with control only in men or women, respectively. Comparing DM2 with the control samples has shown that an increased level of miR-374a-5p was found only in females (Prabu et al. 2015). More recent results of the research also emphasized the sex differences in circulating miRNA expression level but in the population of Israeli Arab and Jewish patients. Although the studied group was genetically heterogenous, Meerson et al. (2019) have found that expression levels of miR-146a-5p, miR-16–2-3p, miR-126-5p, miR-30d, and miR-423 may serve as biomarkers to distinguish early from complicated DM2. Further sex-stratified analysis showed their better diagnostic accuracy for men than women (Meerson et al. 2019). Last but not least research demonstrates that the sex-specific expression level of miRNA could be analyzed together with other markers to indicate diabetes complications. Heart failure with preserved ejection fraction (HFpEF) is an example of diabetes microvascular complications that have more prevalence in women and are frequently undiagnosed at the early stage of development (Florijn et al. 2020). Interestingly, the elevated expression level of a protein marker of microvascular injury angiopoietin-2 (Ang-2) only in diabetic females with HFpEF along with lower expression levels of miR-224 and miR-452 in diabetic males with HFpEF (Florijn et al. 2020) shows their potential to serve as sex-biased biomarkers. In summary, all the mentioned results demonstrate the need for the development of sex-specific diagnostic strategies in diabetes mellitus type 2, as well as show the possibility of using circulating miRNAs as sex-biased biomarkers. In addition, recent studies have shown the role of microRNAs, identified as candidates for sex-dependent biomarkers, in different types of cells involved in DM2 pathogenesis (Table 4). For example, they are engaged in cellular processes such as apoptosis, proliferation, oxidative stress, and regulation of lactate transport (Chen et al. 2021a; Blum et al. 2019; Luo et al. 2020; Xiao et al. 2019; Zhao et al. 2021).
However, there is also an interesting biomarkers searching strategy that focuses on those miRNAs which are expressed in both sexes. Considering that the previous studies suggested greater prevalence and severity of diabetic retinopathy (DR) in males with early diagnosed DM2 (Ozawa et al. 2015), Greco et al. (2020) included only male subjects in the initial screening of potential circulating miRNA biomarkers. Subsequently, they validated the aberrant expression of selected miRNAs in the samples obtained from both male and female subjects that led to findings that in both sexes the most up-regulated miRNA is miR-1281 in comparison to controls. Further analysis confirmed that circulating miR-1281 may serve as a potential biomarker for early diagnosis of DR in males and females (Greco et al. 2020).
A potential mechanism underlying sex difference in circular miRNA expression
To date, mechanisms underlying different expressions of miRNA between females and males are not fully recognized. Based on the literature review, Florijn et al. proposed two mechanisms that may be involved in the sex-biased expression of circulating miRNA: (1) incomplete X-chromosome inactivation leading to biallelic expression of miRNA, and in addition, (2) regulation of miRNA transcription and processing by estrogens (Florijn et al. 2018; 2020). However, these processes may only partially explain the differential expressions of miRNA between sexes. The epigenetic mechanisms also contribute to the regulation of miRNA expression (Bianchi et al. 2017). For example, Sun et al. (2014) suggested that hypomethylation of miR-375 gene promoter may be associated with its overexpression in the plasma of DM2 patients. Another study has shown that in pancreatic islets, methylation of 160 sites annotated to miRNA on chromosome X and only 3 autosomal miRNA genes, was sex dependent. Further analysis indicated that miR-660 and miR-532, encoded on chromosome X, displayed lower methylation levels and higher expression levels in females pancreatic islets (Hall et al. 2014); therefore, the different methylation patterns of miRNA genes in females and males may contribute to the observed sex-dependent level of circulating miRNAs. Interestingly, Ciccacci et al. (2020) suggested that rs11888095 SNP in gene encoding miR-128a may contribute to its higher expression in peripheral blood mononuclear cells from patients with diabetic polyneuropathy. In addition, the recent results from miRNA gene and miRNA target gene polymorphic variants analyses showed that even a single-nucleotide polymorphism could have different impacts between sexes (Zhang et al. 2013a; Zhu et al. 2021). Notably, the effects of polymorphic variants of miRNA genes may be modulated by interactions with polymorphic variants of other miRNAs or by the influence of environmental factors such as smoking or dyslipidemia (Zhu et al. 2021). Considering, the abovementioned results, additional studies are needed to elucidate the potential role of genetic variants and epigenetic mechanisms in sex-dependent aberrant expression of circulating miRNA in DM2.
Conclusion and future perspective
Circulating miRNAs are promising candidates as biomarkers for screening and diagnosis of early onset, progression, and complications of DM2 or even for the selection of individually tailored medications and therapy. However, some studies have reported discrepancies in the analyzed expression level of circulating miRNA and their utility as biomarkers in different populations (Bhatia et al. 2015; Yu et al. 2020). The establishment and validation of protocols for detection and assessment of the clinical value of investigated circulating miRNA will enable overcoming these problems. Moreover, recent findings indicate that sex may affect the results obtained in biomarkers research for DM2. The approach in which males and females are analyzed as one group may lead to overlooking some important biomarkers or finding biomarkers more precisely predicting diseases in one sex. A better understanding of the mechanism underlying sex difference in DM2 susceptibility, development, and progression would enable the selection of the most accurate circulating miRNA biomarkers and transfer those findings into clinical practice.
Data availability
N/A.
Code availability
N/A.
References
Abdelfattah AM, Park C, Choi MY (2014) Update on Non-Canonical microRNAs. Biomol Concepts 5:275–287. https://doi.org/10.1515/bmc-2014-0012
Agbu P, Carthew RW (2021) MicroRNA-mediated regulation of glucose and lipid metabolism. Nat Rev Mol Cell Biol 22:425–438. https://doi.org/10.1038/s41580-021-00354-w
Al-Hayali MA, Sozer V, Durmus S et al (2019) Clinical value of circulating microribonucleic acids miR-1 and miR-21 in evaluating the diagnosis of acute heart failure in asymptomatic type 2 diabetic patients. Biomolecules 9:E193. https://doi.org/10.3390/biom9050193
Al-Kafaji G, Al-Mahroos G, Abdulla Al-Muhtaresh H et al (2017) Circulating endothelium-enriched microRNA-126 as a potential biomarker for coronary artery disease in type 2 diabetes mellitus patients. Biomarkers 22:268–278. https://doi.org/10.1080/1354750X.2016.1204004
Al-Muhtaresh HA, Salem AH, Al-Kafaji G (2019) Upregulation of circulating cardiomyocyte-enriched miR-1 and miR-133 associate with the risk of coronary artery disease in type 2 diabetes patients and serve as potential biomarkers. J Cardiovasc Transl Res 12:347–357. https://doi.org/10.1007/s12265-018-9857-2
Amr KS, Abdelmawgoud H, Ali ZY et al (2018) Potential value of circulating microRNA-126 and microRNA-210 as biomarkers for type 2 diabetes with coronary artery disease. Br J Biomed Sci 75:82–87. https://doi.org/10.1080/09674845.2017.1402404
An Y, Zhang C, Xu F et al (2020) Increased urinary miR-196a level predicts the progression of renal injury in patients with diabetic nephropathy. Nephrol Dial Transplant 35:1009–1016. https://doi.org/10.1093/ndt/gfy326
Association AD (2003) Standards of medical care for patients with diabetes mellitus. Diabetes Care 26:s33–s50. https://doi.org/10.2337/diacare.26.2007.S33
Association AD (2014) Diagnosis and classification of diabetes mellitus. Diabetes Care 37:S81–S90. https://doi.org/10.2337/dc14-S081
Ballotari P, Ranieri SC, Luberto F et al (2015) Sex differences in cardiovascular mortality in diabetics and nondiabetic subjects: a population-based study (Italy). Int J Endocrinol 2015:e914057. https://doi.org/10.1155/2015/914057
Bartel DP (2018) Metazoan MicroRNAs. Cell 173:20–51. https://doi.org/10.1016/j.cell.2018.03.006
Bartel DP (2004) MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 116:281–297. https://doi.org/10.1016/S0092-8674(04)00045-5
Beckman JA, Creager MA (2016) Vascular complications of diabetes. Circ Res 118:1771–1785. https://doi.org/10.1161/CIRCRESAHA.115.306884
Bhatia P, Raina S, Chugh J, Sharma S (2015) miRNAs: early prognostic biomarkers for type 2 diabetes mellitus? Biomark Med 9:1025–1040. https://doi.org/10.2217/bmm.15.69
Bianchi M, Renzini A, Adamo S, Moresi V (2017) Coordinated actions of MicroRNAs with other epigenetic factors regulate skeletal muscle development and adaptation. Int J Mol Sci 18:840. https://doi.org/10.3390/ijms18040840
Blum A, Meerson A, Rohana H et al (2019) MicroRNA-423 may regulate diabetic vasculopathy. Clin Exp Med 19:469–477. https://doi.org/10.1007/s10238-019-00573
Borchert GM, Lanier W, Davidson BL (2006) RNA polymerase III transcribes human microRNAs. Nat Struct Mol Biol 13:1097–1101. https://doi.org/10.1038/nsmb1167
Borga C, Meeran SM, Fassan M (2019) Non-coding RNAs, a real Next-Gen Class of Biomarkers? Noncoding RNA Res 4:80–81. https://doi.org/10.1016/j.ncrna.2019.10.001
Cai X, Hagedorn CH, Cullen BR (2004) Human microRNAs are processed from capped, polyadenylated transcripts that can also function as mRNAs. RNA 10:1957–1966. https://doi.org/10.1261/rna.7135204
Catanzaro G, Besharat ZM, Chiacchiarini M et al (2018) Circulating MicroRNAs in elderly type 2 diabetic patients. Int J Endocrinol 2018:e6872635. https://doi.org/10.1155/2018/6872635
Chatterjee S, Khunti K, Davies MJ (2017) Type 2 diabetes. Lancet 389:2239–2251. https://doi.org/10.1016/S0140-6736(17)30058-2
Chen X, Ba Y, Ma L et al (2008) Characterization of microRNAs in serum: a novel class of biomarkers for diagnosis of cancer and other diseases. Cell Res 18:997–1006. https://doi.org/10.1038/cr.2008.282
Chen H, Guo Y, Cheng X (2021a) Long non-cording RNA XIST promoted cell proliferation and suppressed apoptosis by miR-423-5p/HMGA2 axis in diabetic nephropathy. Mol Cell Biochem 476:4517–4528. https://doi.org/10.1007/s11010-021-04250-x
Chen Q, He Y, Wang X et al (2021b) LncRNA PTGS2 regulates islet β-cell function through the miR-146a-5p/RBP4 axis and its diagnostic value in type 2 diabetes mellitus. Am J Transl Res 13:11316–11328
Chim SSC, Shing TKF, Hung ECW et al (2008) Detection and characterization of placental microRNAs in maternal plasma. Clin Chem 54:482–490. https://doi.org/10.1373/clinchem.2007.097972
Ciccacci C, Latini A, Colantuono A et al (2020) Expression study of candidate miRNAs and evaluation of their potential use as biomarkers of diabetic neuropathy. Epigenomics 12:575–585. https://doi.org/10.2217/epi-2019-0242
Corral-Fernández NE, Salgado-Bustamante M, Martínez-Leija ME et al (2013) Dysregulated miR-155 expression in peripheral blood mononuclear cells from patients with type 2 diabetes. Exp Clin Endocrinol Diabetes 121:347–353. https://doi.org/10.1055/s-0033-1341516
Deng J, Guo F (2019) MicroRNAs and type 2 diabetes. ExRNA 1:36. https://doi.org/10.1186/s41544-019-0038-5
Dunham I, Kundaje A, Aldred SF et al (2012) An integrated encyclopedia of DNA elements in the human genome. Nature 489:57–74. https://doi.org/10.1038/nature11247
Elfaki I, Mir R, Mir MM et al (2019) Potential impact of MicroRNA gene polymorphisms in the pathogenesis of diabetes and atherosclerotic cardiovascular disease. J Pers Med 9:51. https://doi.org/10.3390/jpm9040051
Felekkis K, Papaneophytou C (2020) Challenges in using circulating Micro-RNAs as biomarkers for cardiovascular diseases. Int J Mol Sci 21:561. https://doi.org/10.3390/ijms21020561
Filios SR, Shalev A (2015) β-cell MicroRNAs: small but powerful. Diabetes 64:3631–3644. https://doi.org/10.2337/db15-0831
Florijn BW, Bijkerk R, van der Veer EP, van Zonneveld AJ (2018) Gender and cardiovascular disease: are sex-biased microRNA networks a driving force behind heart failure with preserved ejection fraction in women? Cardiovasc Res 114:210–225. https://doi.org/10.1093/cvr/cvx223
Florijn BW, Valstar GB, Duijs JMGJ et al (2020) Sex-specific microRNAs in women with diabetes and left ventricular diastolic dysfunction or HFpEF associate with microvascular injury. Sci Rep 10:13945. https://doi.org/10.1038/s41598-020-70848-8
Formichi C, Fignani D, Nigi L et al (2021) Circulating microRNAs signature for predicting response to GLP1-RA therapy in type 2 diabetic patients: a pilot study. Int J Mol Sci 22:9454. https://doi.org/10.3390/ijms22179454
Friedman RC, Farh KK-H, Burge CB, Bartel DP (2009) Most mammalian mRNAs are conserved targets of microRNAs. Genome Res 19:92–105. https://doi.org/10.1101/gr.082701.108
George GP, Mittal RD (2010) MicroRNAs: potential biomarkers in cancer. Indian J Clin Biochem 25:4–14. https://doi.org/10.1007/s12291-010-0008-z
Gong W, Xiao D, Ming G et al (2014) Type 2 diabetes mellitus-related genetic polymorphisms in microRNAs and microRNA target sites. J Diabetes 6:279–289. https://doi.org/10.1111/1753-0407.12143
Grasso M, Piscopo P, Confaloni A, Denti MA (2014) Circulating miRNAs as biomarkers for neurodegenerative disorders. Molecules 19:6891–6910. https://doi.org/10.3390/molecules19056891
Greco M, Chiefari E, Accattato F et al (2020) MicroRNA-1281 as a novel circulating biomarker in patients with diabetic retinopathy. Front Endocrinol (lausanne) 11:528. https://doi.org/10.3389/fendo.2020.00528
Guay C, Regazzi R (2013) Circulating microRNAs as novel biomarkers for diabetes mellitus. Nat Rev Endocrinol 9:513–521. https://doi.org/10.1038/nrendo.2013.86
Hall E, Volkov P, Dayeh T et al (2014) Sex differences in the genome-wide DNA methylation pattern and impact on gene expression, microRNA levels and insulin secretion in human pancreatic islets. Genome Biol 15:522. https://doi.org/10.1186/s13059-014-0522-z
Han J, Lee Y, Yeom K-H et al (2004) The Drosha-DGCR8 complex in primary microRNA processing. Genes Dev 18:3016–3027. https://doi.org/10.1101/gad.1262504
He Y, Ding Y, Liang B et al (2017) A systematic study of dysregulated MicroRNA in type 2 diabetes mellitus. Int J Mol Sci 18:456. https://doi.org/10.3390/ijms18030456
Jiang Q, Lyu X-M, Yuan Y, Wang L (2017) Plasma miR-21 expression: an indicator for the severity of type 2 diabetes with diabetic retinopathy. Biosci Rep 37:BSR20160589. https://doi.org/10.1042/BSR20160589
Jimenez-Lucena R, Alcala-Diaz JF, Roncero-Ramos I et al (2021) MiRNAs profile as biomarkers of nutritional therapy for the prevention of type 2 diabetes mellitus: From the CORDIOPREV study. Clin Nutr 40:1028–1038. https://doi.org/10.1016/j.clnu.2020.06.035
Jiménez-Lucena R, Rangel-Zúñiga OA, Alcalá-Díaz JF et al (2018) Circulating miRNAs as predictive biomarkers of type 2 diabetes mellitus development in coronary heart disease patients from the CORDIOPREV Study. Mol Ther Nucleic Acids 12:146–157. https://doi.org/10.1016/j.omtn.2018.05.002
Kautzky-Willer A, Harreiter J, Pacini G (2016) Sex and gender differences in risk, pathophysiology and complications of type 2 diabetes mellitus. Endocr Rev 37:278–316. https://doi.org/10.1210/er.2015-1137
Kim VN, Han J, Siomi MC (2009) Biogenesis of small RNAs in animals. Nat Rev Mol Cell Biol 10:126–139. https://doi.org/10.1038/nrm2632
Kondkar AA, Abu-Amero KK (2015) Utility of circulating MicroRNAs as clinical biomarkers for cardiovascular diseases. Biomed Res Int 2015:e821823. https://doi.org/10.1155/2015/821823
Kozomara A, Birgaoanu M, Griffiths-Jones S (2019) miRBase: from microRNA sequences to function. Nucleic Acids Res 47:D155–D162. https://doi.org/10.1093/nar/gky1141
Krol J, Loedige I, Filipowicz W (2010) The widespread regulation of microRNA biogenesis, function and decay. Nat Rev Genet 11:597–610. https://doi.org/10.1038/nrg2843
La Sala L, Mrakic-Sposta S, Tagliabue E et al (2019) Circulating microRNA-21 is an early predictor of ROS-mediated damage in subjects with high risk of developing diabetes and in drug-naïve T2D. Cardiovasc Diabetol 18:18. https://doi.org/10.1186/s12933-019-0824-2
Lan H, Lu H, Wang X, Jin H (2015) MicroRNAs as potential biomarkers in cancer: opportunities and challenges. Biomed Res Int 2015:e125094. https://doi.org/10.1155/2015/125094
Lee Y, Ahn C, Han J et al (2003) The nuclear RNase III Drosha initiates microRNA processing. Nature 425:415–419. https://doi.org/10.1038/nature01957
Lee Y, Jeon K, Lee J-T et al (2002) MicroRNA maturation: stepwise processing and subcellular localization. EMBO J 21:4663–4670. https://doi.org/10.1093/emboj/cdf476
Lee Y, Kim M, Han J et al (2004) MicroRNA genes are transcribed by RNA polymerase II. EMBO J 23:4051–4060. https://doi.org/10.1038/sj.emboj.7600385
Li T, Quan H, Zhang H et al (2021) Type 2 diabetes is more predictable in women than men by multiple anthropometric and biochemical measures. Sci Rep 11:6062. https://doi.org/10.1038/s41598-021-85581-z
Logue J, Walker J, Colhoun H et al (2011) Do men develop type 2 diabetes at lower body mass indices than women? Diabetologia 54:3003–3006. https://doi.org/10.1007/s00125-011-2313-3
Lund E, Güttinger S, Calado A et al (2004) Nuclear export of MicroRNA precursors. Science 303:95–98. https://doi.org/10.1126/science.1090599
Luo M, Li R, Deng X et al (2015) Platelet-derived miR-103b as a novel biomarker for the early diagnosis of type 2 diabetes. Acta Diabetol 52:943–949. https://doi.org/10.1007/s00592-015-0733-0
Luo E, Wang D, Yan G et al (2020) The NF-κB/miR-425-5p/MCT4 axis: a novel insight into diabetes-induced endothelial dysfunction. Mol Cell Endocrinol 500:110641. https://doi.org/10.1016/j.mce.2019.110641
Mannon EC, Ray SC, Ryan MJ, Sullivan JC (2020) Does sex matter?: an update on the implementation of sex as a biological variable in research. Am J Physiol Ren Physiol 318:F329–F331. https://doi.org/10.1152/ajprenal.00575.2019
Meerson A, Najjar A, Saad E et al (2019) Sex differences in plasma MicroRNA biomarkers of early and complicated diabetes mellitus in Israeli Arab and Jewish patients. Noncoding RNA 5:32. https://doi.org/10.3390/ncrna5020032
Mirra P, Raciti GA, Nigro C et al (2015) Circulating miRNAs as intercellular messengers, potential biomarkers and therapeutic targets for Type 2 diabetes. Epigenomics 7:653–667. https://doi.org/10.2217/epi.15.18
Miyoshi K, Miyoshi T, Siomi H (2010) Many ways to generate microRNA-like small RNAs: non-canonical pathways for microRNA production. Mol Genet Genomics 284:95–103. https://doi.org/10.1007/s00438-010-0556-1
Morrison KR, Solly EL, Shemesh T et al (2021) Elevated HDL-bound miR-181c-5p level is associated with diabetic vascular complications in Australian Aboriginal people. Diabetologia 64:1402–1411. https://doi.org/10.1007/s00125-021-05414-6
Nik Mohamed Kamal NNSB, Shahidan WNS (2020) Non-exosomal and exosomal circulatory MicroRNAs: which are more valid as biomarkers? Front Pharmacol 10:1500. https://doi.org/10.3389/fphar.2019.01500
O’Brien J, Hayder H, Zayed Y, Peng C (2018) Overview of MicroRNA biogenesis, mechanisms of actions, and circulation. Front Endocrinol (lausanne) 9:402. https://doi.org/10.3389/fendo.2018.00402
Ozawa GY, Bearse MA, Adams AJ (2015) Male-female differences in diabetic retinopathy? Curr Eye Res 40:234–246. https://doi.org/10.3109/02713683.2014.958500
Pescador N, Pérez-Barba M, Ibarra JM et al (2013) Serum circulating microRNA profiling for identification of potential type 2 diabetes and obesity biomarkers. PLoS ONE 8:e77251. https://doi.org/10.1371/journal.pone.0077251
Peter ME (2010) Targeting of mRNAs by multiple miRNAs: the next step. Oncogene 29:2161–2164. https://doi.org/10.1038/onc.2010.59
Pordzik J, Jakubik D, Jarosz-Popek J et al (2019) Significance of circulating microRNAs in diabetes mellitus type 2 and platelet reactivity: bioinformatic analysis and review. Cardiovasc Diabetol 18:113. https://doi.org/10.1186/s12933-019-0918-x
Prabu P, Rome S, Sathishkumar C et al (2015) Circulating MiRNAs of “Asian Indian Phenotype” identified in subjects with impaired glucose tolerance and patients with type 2 diabetes. PLoS ONE 10:e0128372. https://doi.org/10.1371/journal.pone.0128372
Saeedi P, Petersohn I, Salpea P et al (2019) Global and regional diabetes prevalence estimates for 2019 and projections for 2030 and 2045: results from the International Diabetes Federation Diabetes Atlas, 9th edition. Diabetes Res Clin Pract 157:107843. https://doi.org/10.1016/j.diabres.2019.107843
Santovito D, Toto L, De Nardis V et al (2021) Plasma microRNA signature associated with retinopathy in patients with type 2 diabetes. Sci Rep 11:4136. https://doi.org/10.1038/s41598-021-83047-w
Schmidt AM (2018) Highlighting diabetes mellitus: the epidemic continues. Arterioscler Thromb Vasc Biol 38:e1–e8. https://doi.org/10.1161/ATVBAHA.117.310221
Sharma S, Eghbali M (2014) Influence of sex differences on microRNA gene regulation in disease. Biol Sex Differ 5:3. https://doi.org/10.1186/2042-6410-5-3
Shi R, Chen Y, Liao Y et al (2020) Research status of differentially expressed noncoding RNAs in type 2 diabetes patients. Biomed Res Int 2020:e3816056. https://doi.org/10.1155/2020/3816056
Sun K, Chang X, Yin L et al (2014) Expression and DNA methylation status of microRNA-375 in patients with type 2 diabetes mellitus. Mol Med Rep 9:967–972. https://doi.org/10.3892/mmr.2013.1872
van Susan D, Beulens JWJ, van der Schouw YT et al (2010) The global burden of diabetes and its complications: an emerging pandemic. Eur J Cardiovasc Prev Rehabil 17:s3–s8. https://doi.org/10.1097/01.hjr.0000368191.86614.5a
Urdea M, Kolberg J, Wilber J et al (2009) Validation of a multimarker model for assessing risk of type 2 diabetes from a five-year prospective study of 6784 Danish People (Inter99). J Diabetes Sci Technol 3:748–755
Wang J, Zhu S, Meng N et al (2019) ncRNA-encoded peptides or proteins and cancer. Mol Ther 27:1718–1725. https://doi.org/10.1016/j.ymthe.2019.09.001
Wang Y-T, Tsai P-C, Liao Y-C et al (2013) Circulating microRNAs have a sex-specific association with metabolic syndrome. J Biomed Sci 20:72. https://doi.org/10.1186/1423-0127-20-72
Weber JA, Baxter DH, Zhang S et al (2010) The MicroRNA spectrum in 12 body fluids. Clin Chem 56:1733–1741. https://doi.org/10.1373/clinchem.2010.147405
Wen Z, Zou X, Xie X et al (2019) Association of polymorphisms in miRNA processing genes with type 2 diabetes mellitus and its vascular complications in a Southern Chinese population. Front Endocrinol 10:461. https://doi.org/10.3389/fendo.2019.00461
Xiao Q, Zhao Y, Xu J et al (2019) NFE2/miR-423-5p/TFF1 axis regulates high glucose-induced apoptosis in retinal pigment epithelial cells. BMC Mol and Cell Biol 20:39. https://doi.org/10.1186/s12860-019-0223-2
Xu J, Zhao J, Evan G et al (2012) Circulating microRNAs: novel biomarkers for cardiovascular diseases. J Mol Med 90:865–875. https://doi.org/10.1007/s00109-011-0840-5
Yan L-N, Zhang X, Xu F et al (2020) Four-microRNA signature for detection of type 2 diabetes. World J Clin Cases 8:1923–1931. https://doi.org/10.12998/wjcc.v8.i10.1923
Yan S, Wang T, Huang S et al (2016) Differential expression of microRNAs in plasma of patients with prediabetes and newly diagnosed type 2 diabetes. Acta Diabetol 53:693–702. https://doi.org/10.1007/s00592-016-0837-1
Yaribeygi H, Katsiki N, Behnam B et al (2018) MicroRNAs and type 2 diabetes mellitus: molecular mechanisms and the effect of antidiabetic drug treatment. Metabolism 87:48–55. https://doi.org/10.1016/j.metabol.2018.07.001
Yu D-C, Li Q-G, Ding X-W, Ding Y-T (2011) Circulating MicroRNAs: potential biomarkers for cancer. Int J Mol Sci 12:2055–2063. https://doi.org/10.3390/ijms12032055
Yu J, Su W, Zhang X et al (2020) MicroRNAs in type 2 diabetes mellitus: important for the pathogenesis but uncertain as biomarkers. J Diabetes 12:697–700. https://doi.org/10.1111/1753-0407.12772
Yuan Y, Peng W, Liu Y, Xu Z (2019) Circulating miR-130 and its target PPAR-γ may be potential biomarkers in patients of coronary artery disease with type 2 diabetes mellitus. Mol Genet Genomic Med 7:e909. https://doi.org/10.1002/mgg3.909
Zang J, Maxwell AP, Simpson DA, McKay GJ (2019) Differential expression of urinary exosomal MicroRNAs miR-21-5p and miR-30b-5p in individuals with diabetic kidney disease. Sci Rep 9:10900. https://doi.org/10.1038/s41598-019-47504-x
Zhang J, Zhang L, Fan R et al (2013a) The polymorphism in the let-7 targeted region of the Lin28 gene is associated with increased risk of type 2 diabetes mellitus. Mol Cell Endocrinol 375:53–57. https://doi.org/10.1016/j.mce.2013.04.022
Zhang T, Lv C, Li L et al (2013b) Plasma miR-126 is a potential biomarker for early prediction of type 2 diabetes mellitus in susceptible individuals. Biomed Res Int 2013:761617. https://doi.org/10.1155/2013/761617
Zhao N, Luo Q, Lin R et al (2021) MiR-142-3p ameliorates high glucose-induced renal tubular epithelial cell injury by targeting BOD1. Clin Exp Nephrol 25:1182–1192. https://doi.org/10.1007/s10157-021-02102-y
Zhu H, Leung SW (2015) Identification of microRNA biomarkers in type 2 diabetes: a meta-analysis of controlled profiling studies. Diabetologia 58:900–911. https://doi.org/10.1007/s00125-015-3510-2
Zhu Z, Zhang Y, Bai R et al (2021) Association of genetic polymorphisms in MicroRNAs with type 2 diabetes mellitus in a Chinese population. Front Endocrinol 11:1139. https://doi.org/10.3389/fendo.2020.587561
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Ethics approval
N/A.
Consent to participate
N/A.
Consent for publication
N/A.
Competing interests
The authors declare no competing interests.
Additional information
Communicated by Michal Witt
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Kraczkowska, W., Stachowiak, L., Pławski, A. et al. Circulating miRNA as potential biomarkers for diabetes mellitus type 2: should we focus on searching for sex differences?. J Appl Genetics 63, 293–303 (2022). https://doi.org/10.1007/s13353-021-00678-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13353-021-00678-5