Abstract
Introduction
The α-glucosidase inhibitor acarbose is an efficacious medicine for the treatment and prevention of type 2 diabetes mellitus (T2DM). However, the response of gut microbiota to acarbose is important, as the microbiota may have a critical role in the development of metabolic diseases, and acarbose is metabolized exclusively within the gastrointestinal tract. We explored the changes in the proportion and diversity of gut microbiota before and after treatment with acarbose in patients with prediabetes.
Methods
We designed a randomized, double-blind, controlled crossover trial in which 52 Chinese patients with prediabetes by an oral glucose tolerance test (OGTT) with a BMI of 18–35 kg/m2 were randomly allocated to treatment with acarbose or placebo. Gut microbiota characterizations were determined with 16S rDNA-based high-throughput sequencing.
Results
Of the 52 participants who entered the study, 40 (76.9%) completed the protocol. On the basis of the operational taxonomic unit (OTU) profiles, a total of 107 OTUs were significantly altered after acarbose treatment, with 76 (71%) assigned to the order of Clostridiales. Ruminococcaceae (15 OTUs) and Lachnospiraceae (22 OTUs) decreased in response to acarbose, and 48 OTUs increased by 12.8-fold, including Lactobacillaceae (8 of 9 belonging to Lactobacillus), Ruminococcaceae (6 of 11 belonging to Faecalibacterium), and Veillonellaceae (8 of 15 belonging to Dialister). At genera level, five flourished after treatment with acarbose, including Lactobacillus and Dialister, while Butyricicoccus, Phascolarctobacterium, and Ruminococcus were inhibited.
Conclusion
This study suggests that the benefits of acarbose for T2DM may correlate with the selective modulation of the gut microbiota.
Trial Registration
Chinese Clinical Trial Register number, ChiCTR-TTRCC-13004112.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Population-based studies have clearly shown that subjects with prediabetes will more likely develop diabetes and have increased risk of cardiovascular disease (CVD). Thus, one of the major reasons to recommend therapeutic intervention for individuals with diabetes or prediabetes is the potential to reduce the long-term increased risk of CVD. Over the last 20 years, a series of clinical trials have shown that interventions differ in their effects on CVD outcomes even though they each successfully slowed the progression to diabetes [1,2,3]. As an α-glucosidase inhibitor, acarbose delays the enzymatic breakdown of carbohydrates in the small intestine and, thus, reduces postprandial hyperglycemia. Many randomized trials have shown that treatment with acarbose was associated with reduction in the risk of progression to diabetes and reduction in the incidence of newly diagnosed CVD events [3,4,5]. In a randomized controlled trial that compared acarbose with metformin as initial treatment for type 2 diabetes mellitus (T2DM), both treatments significantly increased glucagon-like peptide-1 (GLP-1) concentration and decreased glucagon from 24 weeks onwards [6]. However, the reason why acarbose confers additional benefits besides its antidiabetic effect remains unknown.
The composition and activity of the gut microbiota codevelop with the host from birth and is subject to a complex interplay that depends on the host genome, nutrition, and lifestyle. Accumulating evidence indicates that the gut microbiota is involved in host metabolism by increasing energy extraction, immune system modulation, and altering lipid metabolism, which play an intricate role in metabolic disorders, such as obesity and diabetes [7, 8]. One recent metagenome-wide association study showed that patients with T2DM were characterized by a decrease in the abundance of some universal butyrate-producing bacteria and an increase in various opportunistic pathogens [7]. We even observed changes in the proportion and diversity of gut microbiota in the early prediabetes period [8]. Furthermore, gut microflora may also influence drug bioavailability and toxicological incidence via microbial-dependent drug metabolism. In high-fat diet (HFD)-fed mice given the prebiotic oligofructose, Cani et al. [9] showed that Bifidobacterium spp. significantly and positively correlated with improved glucose tolerance, insulin secretion, decreased endotoxemia, and proinflammatory cytokine level [9]. Similarly, Shin et al. [10] found that HFD-fed mice showed a higher abundance of the mucin-degrading bacterium Akkermansia after treatment with metformin. As acarbose is metabolized exclusively within the gastrointestinal tract, principally by intestinal bacteria and digestive enzymes, we hypothesize that modulation of gut microbiota may contribute to one of the mechanisms mediating its antidiabetic effect and confer additional benefits. To test this hypothesis, we designed a randomized, double-blind, controlled crossover trial in which the clinical effects and the changes of gut microbiota before and after treatment with acarbose and placebo were compared.
Methods
Subjects
In this study, we screened men and women aged between 30 and 60 years who had a body mass index (BMI) of between 18 and 35 kg/m2 by a baseline 75-g oral glucose tolerance test (OGTT) in Beijing, China and enrolled 52 participants (21 male and 31 female) with prediabetes according to WHO criteria 1998 [11]. Patients were excluded if they had one of the following: renal and/or hepatic diseases, gastrointestinal tract diseases, psychiatric disorders, neoplasia, or any other disease that could interfere with the outcomes of the study. In addition, none of the participants had taken a pharmacological dose of antibiotics, hormones, anti-obesity agents, lipid-lowering drugs, hypoglycemic agents, or probiotics at least 1 month before initiation of the study. The study complied with the Declaration of Helsinki and was approved by the ethics committee of Peking University People’s Hospital, and informed consent was obtained from each participant. Trial Registration: Chinese Clinical Trial Register number, ChiCTR-TTRCC-13004112.
Design
The study was a randomized, double-blind, controlled crossover trial in which the clinical effects and changes of gut microbiota in response to acarbose and placebo were compared. It consisted of a screening visit, 4 weeks of treatment with acarbose or placebo, a 4-week wash period in which all participants received placebo, and 4 weeks of treatment with the alternative medication (Fig. 1). Eligible participants were randomly allocated to one of the two treatment sequences: acarbose/placebo (sequence 1) or placebo/acarbose (sequence 2). Allocation was performed by a person who was independent of the research team using a computer-generated randomization list. The participants were instructed to take the study medication with the first bite of each meal. To keep the known gastrointestinal side effects of acarbose (e.g., flatulence, diarrhea, or abdominal cramps) to a minimum, the acarbose was administered at 50 mg once daily in the morning on days 1–3, 50 mg twice daily on days 4–7, and thereafter 50 mg three times daily. The study medications were provided by Bayer HealthCare Company Ltd.
Study design. The study consisted of a screening visit, 4 weeks of treatment with acarbose or placebo (stage 1), a 4-week wash period in which all participants received placebo, and then 4 weeks of treatment with the alternative medication (stage 2). Eligible participants (n = 52) were randomly allocated to one of the two treatment sequences
Procedures and Laboratory Analyses
At the start of the study 75-g OGTT was undertaken for each eligible participant with venous blood samples obtained in the normal manner. At each follow-up visit, every participant was asked to have a standard meal with the same dose of medication as the previous visits by measuring plasma glucose levels at 0 and 2 h after meal. Plasma glucose levels were measured by the glucose oxidase method, and serum insulin levels were determined by electrochemiluminescence immunoassays (Elecsys 2010 system, Roche Diagnostics Ltd, Basel, Switzerland). Glycated hemoglobin (HbA1c) was measured at the baseline and after each stage by high-performance liquid chromatography (Ultra2 HbA1c Detector, PRIMUS Corporation, Atlanta, GA, USA; normal range 4–6%). Total serum cholesterol (TC), low-density lipoprotein cholesterol (LDL-C), high-density lipoprotein cholesterol (HDL-C), triglycerides (TG), C-reactive protein (CRP), and liver and renal function indicators were measured using an automatic biochemical analyzer. For each participant, two fecal samples were collected using sterile cups after defecation at each visit. One sample was used for routine testing to exclude bloody stools, pus, or parasitic infections. Another fecal sample was brought to the laboratory within 2 h and stored at −80 °C for microbiota analysis.
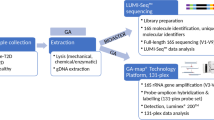
DNA Extraction and Pyrosequencing
DNA extraction and PCR amplification of each fecal sample were performed as described previously [12]. Briefly, the fecal material was diluted in PBS and homogenized with an easyMIX®Lab Blender (AES Chemunex, Bruz Cedex, France). Then all cells were treated with lysozyme at 37 °C for 15 min. Subsequently, the lysates were mixed with binding buffer and the gDNA was purified using resin columns followed the protocol of the commercial kit iNtRON, Kyung-gi, Korea. The universal bacterial forward primer 341F (5′-CCTACGGGNGGCWGCAG-3′) and the reverse primer 926R (5′-CCGTCAATTCMTTTRAGT-3′) were used to amplify the V3-V5 region of the 16S ribosomal RNA (rRNA) gene with the following parameters: initial denaturation at 94 °C for 3 min, followed by 30 cycles of 94 °C for 45 s, 56 °C for 1 min, and 72 °C for 1 min with a final extension at 72 °C for 10 min. PCR products were selected by excising the DNA band of the correct size and purification using the Wizard® SV Gel and PCR Clean-Up System (Promega, St. Louis, MO, USA). A 10-base barcode was added to the 5′ end of the 341F primer to differentiate the PCR amplicons into different samples. Emulsion PCR was conducted according to Roche’s protocols. Sequencing was performed on a 454 GS FLX Titanium pyrosequencer (454 Life Sciences, Branford, CT, USA) at BGI-Shenzhen. All sequences derived from this project were deposited in the European Nucleotide Archive (http://www.ebi.ac.uk/ena/) under accession number ERP009531.
Bioinformatics Analysis
Reads processing and operational taxonomic unit (OTU) analyses were performed as described in our previous study [8]. Briefly, we only kept clean reads longer than 200 bp for OTU analysis after removing tags and primers and reads with ambiguous bases. The sequencing noise was corrected with the PyroNoise algorithm in Mothur (version 1.21) with default parameters as previously described [13]. Preclustering was conducted with CD-HIT to bin all identical reads, and all reads sequenced for the 3′ end of 16S rDNA were reversed [14]. All representatives derived from preclustering were selected and clustered into OTUs using QIIME (version 1.5.0) with default parameters, using the Greengenes reference set as in Ref. [15]. All OTUs were assigned to taxon on the basis of the representative sequence with the RDP classifier (version 2.3) at a confidence of 0.8. A taxonomic profile based on the annotation of OTU was generated using PERL script as previously described [16]. To evaluate if the gut microflora profile became similar to that of healthy people, we compared the taxonomic profile of subjects treated with acarbose with that of healthy and T2DM individuals from our previous study [8].
Statistical Analysis
Clinical data were analyzed on an intention-to-treat basis. Data were expressed as mean ± standard deviation (SD) or median (minimum, maximum). The treatment effect of acarbose versus placebo in metabolic outcomes (FPG, 2 h plasma glucose [2HPG], FINS, 2HINS, TG, CH, HDL-c, LDL-c, CRP) were evaluated by linear mixed models for imbalanced crossover design studies (PROC MIXED, version 9.1.3; SAS Institute Inc, Cary, NC, USA) and were corrected for age and sex. The arm of the trial (treatment) and the intervention period were explicitly modeled as fixed factors, as was the sequence that addressed whether there were carry-over effects. Subjects were treated as random. Statistical significance was set at 0.05, and all statistical analyses were performed using SAS statistical software (version 9.1.3; SAS Institute Inc, Cary, NC, USA).
Kruskal–Wallis rank sum test and Wilcoxon rank sum test were adopted to compare the relative abundances of gut microbiota between the groups. Alpha diversity, including Chao 1, abundance-based coverage estimator (ACE), Simpson, Shannon, and npShannon indices, were calculated with Mothur. The correlation between microbial profiles and the clinical data were calculated on the basis of Spearman’s method, and the P value were adjusted with Bonferroni correction. Principal component analysis (PCA) was used to analyze the beta diversity between the groups with R package ade4 v1.6.2 and gmodels v2.15.4.
Results
Of the 52 participants who entered the study, 40 (76.9%) completed the protocol. The most common reason for discontinuation was adverse gastrointestinal tract events, such as flatulence and increased frequency of bowel movements. No serious adverse events related to acarbose were reported. The clinical characteristics of the study groups at baseline are shown in Table 1. Mean FPG concentration at baseline was 5.92 mmol/L with a mean 2HPG of 10.53 mmol/L, and the mean HbA1c level was 6.29% (45.26 mmol/mol). Table 2 shows the evaluation of the crossover trial using a linear mixed model, which indicated that there were no significant differences in the FPG and 2HPG levels between the acarbose and placebo groups. However, acarbose decreased 2HINS and TG levels (Table 2). When the arm of the trial and the intervention period were explicitly modeled as fixed factors, no carry-over effect was found.
Overall Characteristics of Gut Microbiota in Patients with Prediabetes
A total of 1,544,664 clean pyrosequencing reads (>200 bp, average of 9092 sequences per sample) were obtained after discarding sequences with no primer or barcode sequences, and pyrosequencing noise removal based on the flow information. All clean tags were clustered into 6841 OTUs at the 97% global similarity level, and an average of 435 OTUs was estimated for each sample. One third of all OTUs (2608) occurred in no more than one sample, but the total abundance of these OTUs contributed to only 0.33% of all reads. Together with the alpha diversity (including the Shannon, Chao 1, and ACE1 indices), most of the diversity was captured (Table 3). A total of 141 genera were detected using an RDP classifier with a bootstrap cutoff of 80%, comprising 78.50% of the total tags. The two most abundant genera were Bacteroides (belonging to Bacteroidetes) and Faecalibacterium (belong to Firmicutes), with a median abundance of 19.48 and 8.97%, respectively (Fig. 2). The most abundant phyla included Firmicutes (68.53% of all reads), Bacteroidetes (27.85% of all reads), Proteobacteria (1.98% of all reads), and Actinobacteria (0.98% of all reads).
Most abundant genera identified in all 156 biosamples. All samples were included to exhibit the distribution of microbes in the population we selected. For each genus, the median abundance among all 156 biosamples was computed and presented as the log abundance. Only top 20 genera with high median abundance were plotted
Structural Changes of Gut Microbiota after Treatment with Acarbose
PCA was conducted to compare the composition of the two groups at the baseline of stage 1, and no differences could be observed regarding both the OTUs and genera (Fig. S1). This was validated by comparing the alpha diversity with a Wilcoxon rank sum test, including the Chao 1, ACE, Shannon, npShannon, and Simpson indices (Table 3). All the alpha diversity indices, except for the Simpson index, did not significantly change after the treatment with placebo (Table 3).
Genera Differently Enriched Before and After Treatment with Acarbose
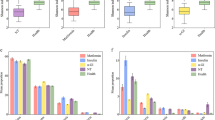
To estimate the effect of acarbose on microbiota structure accurately, the genus and OTU abundance profiles were calculated for subjects who finished the whole trial by PCA analysis after normalization (Fig. 3). A total of 108 genera were identified, and nearly 40% (43 genera) were at relatively higher abundance after treatment with acarbose, accounting for 58.6% of the total abundance on average. Of the genera that flourished with acarbose, higher abundance was observed for five genera, including Lactobacillus and Dialister (Fig. 4 and Table 4). In contrast, significantly lower abundance after acarbose treatment was observed in six genera, including Butyricicoccus, Phascolarctobacterium, and Ruminococcus (P < 0.05; Fig. 4 and Table 4).
Differential OTU Enrichment with Acarbose
A total of 107 OTUs were significantly altered by acarbose, and more than 90% of these OTUs had an abundance of greater than 1.12 × 10−5. Almost all of these OTUs could be assigned to a specific order by an RDP classifier with a confidence of greater than 0.8, except two OTUs that were assigned a specific phylum or domain. Among the OTUs that changed in response to acarbose, 76 (71.03%) could be assigned to the order Clostridiales (phylum Firmicutes). In addition, 59 OTUs were significantly inhibited after intake of acarbose (P ≤ 0.05), accounting for 11.53 and 3.18% of the total abundance of gut microflora before and after administration of acarbose, respectively. The OTUs that decreased in response to acarbose mainly were Ruminococcaceae (15 OTUs) and Lachnospiraceae (22 OTUs). In contrast, 48 OTUs increased in response to acarbose by an average of 12.8-fold, and included Lactobacillaceae (9 OTUs), Ruminococcaceae (11 OTUs), and Veillonellaceae (15 OTUs). It is interesting to note that more than half of the significantly different OTUs of these clusters belong to a single genus. For example, a total of nine OTUs belonging to Lactobacillaceae were determined to be enriched after the administration of acarbose, while eight could be assigned to Lactobacillus accurately by RDP classifier.
Correlation Between Microbiota Composition and Clinical Indicators
To determine if there are some associations between acarbose-mediated changes in gut microbiota composition and that of the physiological index, the correlation between physiological indices and OTU and genera were calculated by the Spearman method, separately (Fig. S2). Thirteen OTUs correlated with clinical parameters, including HbA1c, PBG, 2HINS, HDL, and CH (Table S1). Of the 13 OTUs identified, nine belonged to the order of Clostridiales. OTU4558 (Subdoligranulum), OTU5143 (Erysipelotrichaceae), and OTU3989 (Phascolarctobacterium) were correlated with at least one of the lipid indices. OTU4558 (Subdoligranulum) and OTU1563 (Dialister) were negatively related with HbAlc. Specifically, the abundance of OTU4558 was significantly upregulated by acarbose, increasing from 1.20 × 10−3 to 3.99 × 10−3 (P = 0.03). We also observed a significant negative correlation between 2HINS and OTU7869 (Roseburia).
Nine genera, including Coprobacillus, Bifidobacterium, and Lactococcus, were correlated with metabolic indices (Fig. S3, Table S1). Four cholesterol-related indices, TG, HDL, LDL, and CH, were affected by intestinal microbiota, and a strong correlation between Coprobacillus and HDL was observed in the present study (Spearman correlation coefficient of −0.537, P = 8.26 × 10−5). Furthermore, the average abundance of this genus was slightly decreased from 3.31 × 10−2 to 2.32 × 10−2 after acarbose intake. In addition, TG was negatively correlated with Lactococcus (Spearman correlation coefficient of −0.465, and P = 8.5 × 10−4). As for the glucose-related indices, HbAlc and FINS were correlated with the microbiota profiles: the correlation between FINS and Bifidobacterium was −0.375 (P = 9.5 × 10−3), and both Asaccharobacter and Escherichia/Shigella were correlated with HbAlc (Spearman correlation coefficients of −0.410 and 0.387, respectively).
Differences in Microbiota Composition Between Prediabetes Patients Treated with Acarbose and Healthy Subjects
As in our previous work [8], we also recognized that the proportion and diversity of gut microbiota are different according to glucose intolerance status. Comparison of the microbial composition in prediabetes treated with acarbose and in subjects with normal glucose tolerance (NGT, 44 subjects) and in those with newly diagnosed T2DM (13 subjects) was performed (Table S2). After normalization, only the genera/OTUs with an occurrence rate more than 10% were used (Fig. S4). Acarbose altered the gut microbiota profile at both the genus and OTU levels in prediabetes. However, the profile induced by acarbose was different from that observed in healthy subjects as well as that observed in untreated T2DM patients.
Discussion
As a widely used α-glucosidase inhibitor, acarbose is effective in lowering blood glucose level in patients with T2DM by delaying and reducing the digestion of complex carbohydrates through competitive inhibition of pancreatic α-amylase and a variety of α-glucosides. These carbohydrates will be fermented by microbiota, which will change the composition of the intestinal microbiota significantly as was shown in the present study. Indeed, many of the OTUs that greatly increased in response to acarbose belong to short chain fatty acid (SCFA)-producing taxa, such as Faecalibacterium, Prevotella, and Lactobacillus.
Compared with baseline levels, the abundance of Lactobacillus increased significantly with acarbose, which may be due to its high tolerance to the low pH environment that is maintained by the enhanced fermentation of gut microbiota [17]. Previous studies have described the beneficial effects of Lactobacillus to human health, including improving T2DM, antidiabetic and anti-inflammatory effects, and reducing body weight [18,19,20]. In addition, strains of Lactobacillus isolated from the human gut also have potential α- and β-glucosidase inhibitory activities and can reduce blood glucose responses in vivo [21].
In addition to Lactobacillus, we observed that one OTU belonging to Dialister, which was in higher abundance following acarbose, was negatively correlated with HbA1c. Thus, some species of Dialister may play a role in the regulation of glucose metabolism. This genus is often isolated from clinical samples, including those derived from the oral cavity, periodontitis, brain abscesses, bacteremia of vaginal origin, and human feces [22,23,24,25]. Although this genus is non-fermentative, and all clinical strains are asaccharolytic and non-reactive in conventional biochemical tests, some strains, including D. invisus, D. propionicifaciens, and D. succinatiphilus, could be stimulated by succinate [25,26,27]. It was also noted that some species of Megasphaera, which also thrived following acarbose treatment, could produce a larger amount of succinic acid by glucose fermentation [28]. In addition, a higher amount of complex carbohydrate-utilizing genes present in the human gut isolates that contained Megasphaera, including glycosyl hydrolase and amylomaltase [29], suggest that it may play a key role in the utilization of carbohydrates that cannot be directly degraded and utilized by the human digestive system, including the non-digested carbohydrates in acarbose-treated individuals. Megasphaera could potentially transform carbohydrates into SCFAs, including butyrate, formate, acetate, valerate, and caproate, which is beneficial for Lactobacillus growth. Taken together, acarbose modulates intestinal microbes.
The gut microbiome exerts major effects on metabolic balance in obesity, metabolic syndrome, liver steatosis, and the host’s immune system [30,31,32,33]. We have previously shown that FINS concentration was negatively correlated with the diversity of gut microbiota, suggesting that insulin resistance may affect the composition of gut microbiota. In addition, compositional changes of human gut microbiota in response to weight status have been shown [30, 32]. One of the possible mechanisms of the effect of acarbose on body weight might be related to improved microbiota structure. Moreover, given that bacterial DNA can be identified in more than half of the atherosclerotic plaques analyzed [34, 35], the gut microbiota is now recognized as a causal factor in atherosclerosis. Clinical studies have shown that acarbose significantly improved glycemic control and lowered known CVD risk factors, including TG levels, BMI, insulin levels, and systolic blood pressure [3, 4]. In the present study, plasma glucose concentrations were not significantly different between acarbose and placebo groups, which may be due to the lower dose of the drug and the short time at which acarbose was administered. However, three OTUs were correlated with at least one of the lipid indices, and seven OTUs were associated with glucose metabolism indicators, including, PBG and 2HINS. In view of the potential harmful effects of some antidiabetic agents, we speculate that the structural modulation of gut microbiota during administration of acarbose may partly contribute to the additional benefits regarding CVD risk.
SCFAs fermented by the intestinal bacteria from dietary fiber play an important role in energy metabolism, maintaining intestinal integrity, prevention of endotoxemia, and ability of intestinal gluconeogenesis to produce glucose itself [31, 36,37,38]. Interestingly, oral administration of butyrate was found to improve insulin sensitivity and increase energy expenditure by enhancing mitochondrial function in mice [39]. Many previous studies have reported that acarbose can modulate the fermentation and SCFA productions derived by gut microflora [40, 41]. We also noted that a study focused on the effect of acarbose on fecal nutrients found that pH decreased and total SCFA, butyrate, and acetate output were markedly increased [42]. Another study based on double-blind experiments found that serum butyrate significantly increased after a 4-month administration of acarbose in subjects with impaired glucose tolerance [43].
Acarbose had a much stronger effect on a few clades, especially Clostridiales. In addition, compared with the baseline group, all the significantly increased OTUs belonged to Lactobacillaceae, suggesting that the taxa belonging to Lactobacillaceae have the same response to acarbose. Interestingly, the reduced average abundance of the six genera (2.52%) was almost perfectly compensated by an increase in five other genera (2.06%) in response to acarbose, which may imply a complicated relationship that corresponds with changes in nutrition. We previously reported that dysbiosis in gut microbiota occurs early before the prediabetes stage [8]. Thus, interventions that can revert the gut microbiota of patients with prediabetes to a normal state will be ideal. However, the results of the present study showed that acarbose did not return the dysbiosis observed in prediabetic patients to a normal state, suggesting that the regulatory effect of acarbose on the composition of intestinal microbiota was selective and competitive. Nevertheless, it is possible that a longer treatment with acarbose may have different outcomes.
In addition, several associations between antidiabetic drugs and altered gut microbiota composition have been reported in recent years. Lee et al. [44] and Forslund et al. [45] found that the gut microbiota and their metablic pathways were influenced by metformin treatment. Su et al. [46] reported that acarbose treatment can increase the content of gut Bifidobacterium longum and Enterococcus faecalis in Chinese T2DM patients. Notably, the changes of gut microbiota will be more complex when combined antidiabetic treatments were used, especially in patients with long diabetes course or complicated by other chronic disease. So, in this study, monotherapy in prediabetes gave us the chance to investigate the pure effect of acarbose on gut microbiota.
Discussion of the findings from this study would be incomplete without recognition of the related limitations. First, although we use a controlled crossover design to increase the test power, the sample size was small. Second, the participants who were lost to follow-up were not included in the data analysis and may be a possible source of bias. Third, this study failed to observe a significant reduction in plasma glucose levels with acarbose, which may be due to the short observation time, the lower dose of the drug, and the placebo effect. Fourth, in Asia, rice forms a major component of the diet, and the dietary contribution of carbohydrates is high. In view of the perceived difference in efficacy of acarbose in reducing glucose level between white and Asian populations, it is unclear whether similar results will be obtained in other groups. Thus, further studies with larger sample sizes and in other ethnic populations are needed.
Conclusion
This is the first randomized, double-blind, controlled study to explore the effects of acarbose on the intestinal microbiota. In this study, we demonstrated that the diversity and composition of the gut microbiota changed significantly during acarbose treatment in prediabetes individuals. Furthermore, the changes of microbiota showed strong correlations with the levels of various metabolic indicators. However, the microbiota profile induced by acarbose treatment was different from that in healthy subjects. These findings support that the antidiabetic and additional effects of acarbose may be mediated by its selective modulation of the microbiota.
References
Li G, Zhang P, Wang J, et al. Cardiovascular mortality, all-cause mortality, and diabetes incidence after lifestyle intervention for people with impaired glucose tolerance in the Da Qing diabetes prevention study: a 23-year follow-up study. Lancet Diabetes Endocrinol. 2014;2:474–80.
Knowler WC, Barrett-Connor E, Fowler SE, et al. Reduction in the incidence of type 2 diabetes with lifestyle intervention or metformin. N Engl J Med. 2002;346:393–403.
Chiasson JL, Josse RG, Gomis R, et al. Acarbose for prevention of type 2 diabetes mellitus: the STOP-NIDDM randomised trial. Lancet. 2002;359:2072–7.
Chiasson JL, Josse RG, Gomis R, et al. Acarbose treatment and the risk of cardiovascular disease and hypertension in patients with impaired glucose tolerance: the STOP-NIDDM trial. JAMA. 2003;290:486–94.
Hanefeld M, Cagatay M, Petrowitsch T, Neuser D, Petzinna D, Rupp M. Acarbose reduces the risk for myocardial infarction in type 2 diabetic patients: meta-analysis of seven long-term studies. Eur Heart J. 2004;25:10–6.
Yang W, Liu J, Shan Z, et al. Acarbose compared with metformin as initial therapy in patients with newly diagnosed type 2 diabetes: an open-label, non-inferiority randomized trial. Lancet Diabetes Endocrinol. 2014;2:46–55.
Qin J, Li Y, Cai Z, et al. A metagenome-wide association study of gut microbiota in type 2 diabetes. Nature. 2012;490:55–60.
Zhang X, Shen D, Fang Z, et al. Human gut microbiota changes reveal the progression of glucose intolerance. PLoS One. 2013;8:e71108.
Cani PD, Bibiloni R, Knauf C, et al. Changes in gut microbiota control metabolic endotoxemia-induced inflammation in high-fat diet-induced obesity and diabetes in mice. Diabetes. 2008;57:1470–81.
Shin NR, Lee JC, Lee HY, et al. An increase in the Akkermansia spp. population induced by metformin treatment improves glucose homeostasis in diet-induced obese mice. Gut. 2014;63:727–35.
Alberti KG, Zimmet PZ. Definition, diagnosis and classification of diabetes mellitus and its complications. Part 1: diagnosis and classification of diabetes mellitus provisional report of a WHO consultation. Diabet Med. 1988;15:539–53.
Chun J, Kim KY, Lee JH, Choi Y. The analysis of oral microbial communities of wild-type and toll-like receptor 2-deficient mice using a 454 GS FLX Titanium pyrosequencer. BMC Microbiol. 2010;10:10.
Quince C, Lanzen A, Davenport RJ, Turnbaugh PJ. Removing noise from pyrosequenced amplicons. BMC Bioinform. 2011;12:38.
Li W, Godzik A. Cd-hit: a fast program for clustering and comparing large sets of protein or nucleotide sequences. Bioinformatics. 2006;22:1658–9.
Caporaso JG, Kuczynski J, Stombaugh J, et al. QIIME allows analysis of high-throughput community sequencing data. Nat Methods. 2010;7:335–6.
Wang Q, Garrity GM, Tiedje JM, Cole JR. Naive Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl Environ Microbiol. 2007;73:5261–7.
Holt PR, Atillasoy E, Lindenbaum J, et al. Effects of acarbose on fecal nutrients, colonic pH, and short-chain fatty acids and rectal proliferative indices. Metabolism. 1996;45:1179–87.
Yadav H, Jain S, Sinha PR. Antidiabetic effect of probiotic dahi containing Lactobacillus acidophilus and Lactobacillus casei in high fructose fed rats. Nutrition. 2007;23:62–8.
Naito E, Yoshida Y, Makino K, et al. Beneficial effect of oral administration of Lactobacillus casei strain Shirota on insulin resistance in diet-induced obesity mice. J Appl Microbiol. 2011;110:650–7.
Kang JH, Yun SI, Park HO. Effects of Lactobacillus gasseri BNR17 on body weight and adipose tissue mass in diet-induced overweight rats. J Microbiol. 2010;48:712–4.
Panwar H, Calderwood D, Grant IR, Grover S, Green BD. Lactobacillus strains isolated from infant faeces possess potent inhibitory activity against intestinal alpha- and beta-glucosidases suggesting anti-diabetic potential. Eur J Nutr. 2014;53:1465–74.
Contreras A, Doan N, Chen C, Rusitanonta T, Flynn MJ, Slots J. Importance of Dialister pneumosintes in human periodontitis. Oral Microbiol Immunol. 2000;15:269–72.
Rousée JM, Bermond D, Piémont Y, et al. Dialister pneumosintes associated with human brain abscesses. J Clin Microbiol. 2002;40:3871–3.
Pierre Lepargneur J, Dubreuil L, Levy J. Isolation of Dialister pneumosintes isolated from a bacteremia of vaginal origin. Anaerobe. 2006;12:274–5.
Morotomi M, Nagai F, Sakon H, Tanaka R. Dialister succinatiphilus sp. nov. and Barnesiella intestinihominis sp. nov., isolated from human faeces. Int J Syst Evol Microbiol. 2008;58:2716–20.
Jumas-Bilak E, Jean-Pierre H, Carlier JP, et al. Dialister micraerophilus sp. nov. and Dialister propionicifaciens sp. nov., isolated from human clinical samples. Int J Syst Evol Microbiol. 2005;55:2471–8.
Joossens M, Huys G, Cnockaert M, et al. Dysbiosis of the faecal microbiota in patients with Crohn’s disease and their unaffected relatives. Gut. 2011;60:631–7.
Lanjekar VB, Marathe NP, Ramana VV, Shouche YS, Ranade DR. Megasphaera indica sp. nov., an obligate anaerobic bacteria isolated from human faeces. Int J Syst Evol Microbiol. 2014;64:2250–6.
Shetty SA, Marathe NP, Lanjekar V, Ranade D, Shouche YS. Comparative genome analysis of Megasphaera sp. reveals niche specialization and its potential role in the human gut. PLoS One. 2013;8:e79353.
Turnbaugh PJ, Ley RE, Mahowald MA, Magrini V, Mardis ER, Gordon JI. An obesity-associated gut microbiome with increased capacity for energy harvest. Nature. 2006;444:1027–31.
Cani PD, Amar J, Iglesias MA, et al. Metabolic endotoxemia initiates obesity and insulin resistance. Diabetes. 2007;56:1761–72.
Furet JP, Kong LC, Tap J, et al. Differential adaptation of human gut microbiota to bariatric surgery-induced weight loss: links with metabolic and low-grade inflammation markers. Diabetes. 2010;59:3049–57.
Hooper LV, Littman DR, Macpherson AJ. Interactions between the microbiota and the immune system. Science. 2012;336:1268–73.
Ott SJ, El Mokhtari NE, Musfeldt M, et al. Detection of diverse bacterial signatures in atherosclerotic lesions of patients with coronary heart disease. Circulation. 2006;113:929–37.
Koren O, Spor A, Felin J, et al. Human oral, gut, and plaque microbiota in patients with atherosclerosis. Proc Natl Acad Sci USA. 2011;108:4592–8.
Tremaroli V, Bäckhed F. Functional interactions between the gut microbiota and host metabolism. Nature. 2012;489:242–9.
Delaere F, Duchampt A, Mounien L, et al. The role of sodium-coupled glucose co-transporter 3 in the satiety effect of portal glucose sensing. Mol Metab. 2012;2:47–53.
De Vadder F, Kovatcheva-Datchary P, Goncalves D, et al. Microbiota-generated metabolites promote metabolic benefits via gut-brain neural circuits. Cell. 2014;156:84–96.
Gao Z, Yin J, Zhang J, et al. Butyrate improves insulin sensitivity and increases energy expenditure in mice. Diabetes. 2009;58:1509–17.
Hiele M, Ghoos Y, Rutgeerts P, Vantrappen G. Effects of acarbose on starch hydrolysis. Study in healthy subjects, ileostomy patients, and in vitro. Dig Dis Sci. 1992;37:1057–64.
Wolever TM, Radmard R, Chiasson JL, et al. One-year acarbose treatment raises fasting serum acetate in diabetic patients. Diabet Med. 1995;12:164–72.
Weaver GA, Tangel CT, Krause JA, et al. Acarbose enhances human colonic butyrate production. J Nutr. 1997;127:717–23.
Wolever TM, Chiasson JL. Acarbose raises serum butyrate in human subjects with impaired glucose tolerance. Br J Nutr. 2000;84:57–61.
Lee H, Ko G. Effect of metformin on metabolic improvement and gut microbiota. Appl Environ Microbiol. 2014;80:5935–43.
Forslund K, Hildebrand F, Nielsen T, et al. Disentangling type 2 diabetes and metformin treatment signatures in the human gut microbiota. Nature. 2015;528:262–6.
Su B, Liu H, Li J, et al. Acarbose treatment affects the serum levels of inflammatory cytokines and the gut content of bifidobacteria in Chinese patients with type 2 diabetes mellitus. J Diabetes. 2015;7:729–39.
Acknowledgements
This work was supported through a grant from the National High Technology Research and Development Program (863 Program, 2012AA02A509). National Key Basic Research Program of China (973 Program, 2011CB504000), and Beijing Science and Technology Committee Funding (D131100005313008). We thank all the participants for agreeing to join this study. We are grateful to the staff at PKU Diabetes Centre for their assistance with subject recruitment, conducting the clinical protocol, and laboratory testing. We are indebted to the technicians of BGI-Shenzhen for their practical work in this study.
All authors meet the International Committee of Medical Journal Editors (ICMJE) criteria for authorship for this manuscript, take responsibility for the integrity of the work as a whole, and have given final approval for the version to be published.
Author Contributions
Conceived and designed the study: X. Zhang, Y. Chen, and L. Ji.
Analyses and interpretation of the data: X. Zhang, Z. Fang, C. Zhang, H. Xia, and Z. Jie.
Wrote the paper: X. Zhang, Z. Fang, and L. Ji.
Critically revised the manuscript for important intellectual content: All authors.
Disclosures
Xiuying Zhang, Zhiwei Fang, Chunfang Zhang, Huihua Xia, Zhuye Jie, Xueyao Han, Yingli Chen, and Linong Ji have nothing to disclose.
Compliance with Ethics Guidelines
The study was approved by the local institutional review board. All procedures followed were in accordance with the ethical standards of the responsible committee on human experimentation (institutional and national) and with the Helsinki Declaration of 1975, as revised in 2008. Informed consent was obtained from all patients for being included in the study.
Open Access
This article is distributed under the terms of the Creative Commons Attribution-NonCommercial 4.0 International License (http://creativecommons.org/licenses/by-nc/4.0/), which permits any noncommercial use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
Author information
Authors and Affiliations
Corresponding author
Additional information
X. Zhang and Z. Fang contributed equally to this work.
Enhanced content To view enhanced content for this article go to http://www.medengine.com/Redeem/7D77F0600D879758.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (https://creativecommons.org/licenses/by/4.0), which permits use, duplication, adaptation, distribution, and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Zhang, X., Fang, Z., Zhang, C. et al. Effects of Acarbose on the Gut Microbiota of Prediabetic Patients: A Randomized, Double-blind, Controlled Crossover Trial. Diabetes Ther 8, 293–307 (2017). https://doi.org/10.1007/s13300-017-0226-y
Received:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13300-017-0226-y