Abstract
Participants of the Second International Workshop (WS) on human chorionic gonadotropin (hCG) of the International Society of Oncology and Biomarkers Tissue Differentiation 7 (ISOBM TD-7) have characterized in detail a panel of 69 antibodies (Abs) directed against hCG and hCG-related variants that were submitted by eight companies and research groups. Specificities of the Abs were determined using the First WHO International Reference Reagents for six hCG variants, i.e., hCG, hCGn, hCGβ, hCGβn, hCGβcf, and hCGα, which are calibrated in SI units, and hLH. Molecular epitope localizations were assigned to the ISOBM-mAbs by comparing ISOBM-Ab specificity, sandwich compatibility, and mutual inhibition profiles, to those of 17 reference monoclonal (m)Abs of known molecular epitope specificities. It appeared that 48 Abs recognized hCGβ-, 8 hCGα-, and 13 αβ-heterodimer-specific epitopes. Twenty-seven mAbs were of pan hCG specificity, two thereof with no (<0.1 %; epitope β1), 12 with low (<1.0 %; epitopes β2/4), and 13 with high (>>1 %; epitopes β3/5) hLH cross-reactivity. The majority of hCGβ epitopes recognized were located in two major antigenic domains, one on the peptide chain of the tips of β-sheet loops 1 and 3 (epitopes β2–6; 27 mAbs) and the second around the cystine knot (e.g., epitopes β1, β7, and β10; 9 mAbs). Four mAbs recognized epitopes on hCGβcf-only (e.g., epitopes β11 and β13) and six mAbs epitopes on the remote hCGβ-carboxyl-terminal peptide (epitopes β8 and β9 corresponding to amino acids 135–144 and 111–116, respectively). For routine diagnostic measurements, methods are used that either detect hCG-only, hCGβ-only, or hCG together with hCGβ or hCG together with hCGβ and hCGβcf. Sandwich assays that measure hCG plus hCGβ and eventually hCGβcf should recognize the protein backbone of the analytes preferably on an equimolar basis, should not cross-react with hLH and not be susceptible to blunting of signal by nonmeasured variants like hCGβcf. Such assays can be constructed using pairs of mAbs directed against the cystine knot-associated epitope β1 (Asp10, Asp60, and Gln89) in combination with epitopes β2 or β4 located at the top of β-sheet loops 1 + 3 of hCGβ involving aa hCGβ20-25 + 68-77. In summary, the results of the First and Second ISOBM TD-7 WSs on hCG provide the basis for harmonization of specificities and epitopes of mAbs to be used in multifunctional and selective diagnostic hCG methods for different clinical purposes.
Similar content being viewed by others
Objective
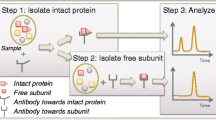
Improving between method comparability for measurement of the heterogeneous glycoprotein hCG requires harmonization of epitopes of the Abs used and broad consensus about assay specificity. To address this, the Second International Society of Oncology and Biomarkers Tissue Differentiation 7 (ISOBM TD-7) Workshop (WS) by a three-step algorithm characterized and epitope typed 69 Abs directed against hCG and variants submitted by diagnostic companies and research groups. The results of this WS in combination with those of the First WS enable recommendations to be made regarding epitope combinations to be used for the design of immunoassays for hCG and its variants [1].
Introduction
Physiology, protein structure, and posttranslational protein backbone variants of hCG
The glycoprotein hormone hCG is essential for maintaining pregnancy. Physiologically, it is produced and secreted by the placental trophoblast and pathophysiologically by trophoblastic cancers and by germ cell tumors of the testis and ovary [2].
hCG is a protein heterodimer consisting of hCGα noncovalently linked to the hCGβ subunit. As all glycoprotein hormone (GPH) subunits, hCGα and hCGβ share structural homology with members of the cystine knot growth factor superfamily that includes nerve growth factor, platelet-derived growth factor and transforming growth factor beta [3]. The common structural cystine knot motif consists of two disulfide bridges that link adjacent antiparallel strands of the single peptide chain to form a ring that is axially permeated by a third disulfide bond. This central cystine knot determines the three-dimensional structure of hCGα and hCGβ. On one side of the knot, there are two neighboring hairpin-like peptide loops 1 and 3, which, in hCGβ, are stabilized by a disulfide bond between Cys 23 and Cys72. The single larger loop 2 is located on the opposite side of the knot [3].
The subunits are noncovalently linked in antiparallel, i.e., a head-to-toe fashion, such that loops 1 + 3 of one subunit are adjacent to loop 2 of the other subunit [3]. Loops 1 and 3 of either subunit and the hCGβ cystine knot, respectively are the most important antigenic regions [1].
The hCGβ genes have developed from an ancestral LHβ gene by gene duplications and mutations [4]. The hCGβ protein is 145 amino acids (aa) in length and encoded by 4 genes and 2 alleles (CGβ6/7, CGβ3/9, CGβ5, and CGβ8), while hLHβ is encoded by a single gene, CGβ4, on chromosome 19q13.3. Thus, hCGβ and hLHβ are highly similar in protein sequence (>85 %) and are immunologically closely related. Furthermore, LH and hCG activate the same receptor. The major structural difference between hCGβ and hLHβ is a carboxyl-terminal peptide extension of hCGβ (hCGβCTP) encompassing aa 113–145. hCGβCTP evolved through a read-through event due to a mutational loss of the stop codon at the genomic level and the incorporation of a hitherto untranslated gene sequence into the coding region [5]. Antibodies recognizing epitopes on hCGβCTP are used in a number of highly specific hCG assays [6]. A single gene on human chromosome 12q21.1-23 encodes the α-subunit, which is 92 aa in length and common to all four human GPHs [7].
hCG is heterogeneous with respect to protein backbone structure and carbohydrate content and is best considered as a complex family of hCG variants occurring in body fluids and tissues. The unambiguous nomenclature for the most important hCG forms of the protein backbone developed by the International Federation of Clinical Chemistry (IFCC) Working Group for Standardization of hCG Determinations is used here (Table 1 and Fig. 1) [1, 8, 9].
Schematic representation of human chorionic gonadotropin β (hCGβ) protein backbone variants and molecular epitope localizations on assembled and free hCGβ (amino acids, aa hCGβ1-145), hCGβ core fragment (hCGβcf, aa hCGβ6-40 + β55-92), and the carboxyl-terminal peptide (hCGβCTP, aa hCGβ109/113-145). Modified according to [1] with permission (INN). Antigenic determinants are diagrammatically represented on the linear aa sequence. Non-assembled hCGβ carries nine epitopes (β1–β9), seven are present also on the hCGαβ-heterodimer (β1–β5, β8, β9), and all, except those on the hCGβCTP (β8, β9), are located within the amino acid sequences (aa) of hCGβcf. Four additional specific epitopes are present on hCGβcf only (β10–β13) but not on intact hCGβ and hCG. All epitopes that are located on core hCGβ (aa 1–112) are conformationally dependent and determined by the tertiary protein structure. Important residues contributing to these epitopes at the primary sequence level were identified by selective mutational analyses: Pro24, Val25, Arg68, Gly71, and Gly75 contribute to epitope β3, aa Lys20, Glu21, Gln22, Gly75, and Asn77 to free subunit epitope β6 and Arg68 to structurally overlapping epitopes β2, β3, β4, and β5 [22, 42]. hCG-specific epitope β1 is built up by cystine-knot associated Arg10 and Arg60 and to a minor extent Gln89 as it does in epitope β7. Asp61 plays a role in free subunit epitopes β6 and β7 [43]. Major antigenic regions of hCGβCTP are rather linear in nature and determined by the primary structure: aa hCGβ133-144 comprising epitope β8, that is substructured into β8,1 to β8,3 and aa hCGβ113-116 corresponding to epitope β9 [21, 24, 29, 50, 71]. Numbers represent positions of amino acid residues in the peptide chain. The metabolic product hCGβcf consists of two peptide fragments that are linked via five disulfide bonds (depicted by S–S), and its N-linked carbohydrate antennae are truncated. Open circles N-linked glycans, filled circles O-linked glycans
Glycosylation isoforms
hCG subunit folding, assembly, intracellular trafficking, secretion, receptor activation, and half-life in serum is dependent on glycosylation [10]. Both hCG subunits are glycosylated: hCGα contains two N-glycosylation sites at Asn52 and Asn78 that are either mono-, bi-, or triantennary or are sometimes missing. Most N-linked carbohydrate antennae at Asn13 and Asn30 of hCGβ are of the bi-antennary type, but malignancy-associated hCG increasingly carries triantennary carbohydrates at Asn30 and fucosylation at Asn13 (Fig. 2). Four putative O-glycosylation sites are located at Ser121 (core-2), Ser127 (core-1), Ser132 (core-1), and Ser138 (core-1) on the hCGβCTP. The pregnancy associated core-1 glycans on Ser127 and Ser132 are frequently replaced by core-2 glycans in hCG synthesized in early pregnancy and by tumors [11].
Glycosylation variants of hCGβ (according to [11]). In pregnancy-derived hCGβ, the N-linked carbohydrates are of the biantennary type. O-Glycosylation of hCGβ at Ser 121 always contains a biantennary core-2 and at Ser 138 a core-1 structure with one or two sialic acids. Malignancy-derived hCG and very early pregnancy hCG as compared to middle-to-late pregnancy hCGβ is characterized by increased content of triantennary complex-type N-linked carbohydrates attached to hCGβ Asn 30 and fucosylated carbohydrates attached to Asn 13. “Hyperglycosylated” hCGβ contains an increased proportion of triantennary N-linked carbohydrates (Asn 30); core-2 type O-glycans at Ser 127, Ser 132, and Ser 138; and fucosylated Asn 13-linked glycan. Some glycosylation sites were not glycosylated in some variants (Ser 138, Ser 121, and Asn 13). Immunoassays for hCG-h based on mAb B152 recognize the encircled glycan at Ser 132 and surrounding peptide structure. The major differences in carbohydrate antennae composition between early and mid-to-late pregnancy- and malignancy-derived hCG are depicted in red. Filled square GlcNAc, filled diamond Fuc, empty square GalNAc, empty circle Man, filled circle Gal, empty diamond NeuAc
Due to variability in branching of carbohydrate antennae and terminal sialylation (8–15 sialic acids), numerous isoforms exist [11, 12]. The relative proportions of more extensively glycosylated and terminally sialylated glycosylation variants change with advancing pregnancy, tumor progression, and between different tumors [13–15]. Consequently, acidic variants (av) of hCG (avhCG) produced by testicular cancer [16], by other tumors, and in early pregnancy [13] have very low isoelectric points (pIs) and higher MWs [15, 17, 18]. The expression avhCG, describing hCG with complex extensively terminally sialylated carbohydrate antennae, was later replaced by the term “hyperglycosylated” hCG (hCG-h) [19]. Presently, hCG-h is defined as hCG isoforms carrying a biantennary core-2 O-glycan on Ser132 that is detected by immunoassays using a mAb-designated B152 (Fig. 2) [20].
hCG epitopes
Elucidation of the three-dimensional structure of hCG [3] provided the basis for assignment of immunologically and biologically important domains to the molecular surface of hCG and hCG-related molecules. Several strategies were pursued to resolve epitope distribution and arrangement as well as identification of immunodominant regions. Epitope localization and sharing of epitopes among hCG, hCG-variants, subunits, and related hormones like LH and subunits were determined using molecular chimeras, hCG metabolites, homologous and heterologous glycoprotein hormones and subunits, chemically modified hormones, proteolytic hormone fragments and synthetic peptides, including peptide scanning, and most importantly by site-specific mutagenesis of hCGβ. It is important to mention that in three independent laboratories with different sets of mAbs and analytical techniques similar epitopes and antigenic domains were defined (for reviews, see [1, 21].
In previous studies, 26 epitopes on hCG and hCG-related molecules were defined (for reviews, see [1, 20, 22]). Sixteen epitopes are located on the intact holo hormone hCG (epitopes β1–β5, β8, and α9; α1–α5; and c1–c4). Seven of these are present on both free and assembled hCGβ (β1–β5, β8, and α9; Fig. 1 and Table 2).
Antibodies against epitopes β1–β5 recognize a wide range of hCG and hCGβ variants (pan hCG-mAbs), including hCG + hCGn + hCGβ + hCGβn + hCGβcf. In a first step, epitopes recognized by these mAbs can be discerned by their cross-reactivity with hLH or hLHβ: Abs directed against (1) epitope β1 have no (<0.1 %), (2) epitopes β2 and β4 <1%, and (3) epitopes β3 and β5 >>1 % hLH cross-reactivity. Epitopes β1–β5 are distributed among two antigenic domains: (1) the cystine knot (epitope β1) and (2) hCGβ loops 1 + 3 comprising the neighbouring epitopes β2–β6. Epitopes β8 andβ9 are located on the hCGβCTP and by definition are specific for hCG and hCGβ [1, 23, 24].
A number of epitopes are of restricted variant specificity. Abs against such epitopes are useful for variant-selective immunoassays designed to measure hCG, hCG + hCGn, hCGβ, hCGβ + hCGβcf, hCGβcf, or hCGα, respectively, in the presence of excess of other hCG protein backbone variants and GPHs.
Epitopes β6 (hCGβ loops 1 + 3 related) and β7 (cystine knot related) are shared by free hCGβ and hCGβcf but not by holo-hCG. Epitope β14, which is related to the core region of hCGβ (amino acids 1–112) and maybe cystine knot-related, was defined in the First ISOBM WS. It is specific for free hCGβ and not shared by hCGβcf [1]. Four hCGβcf-specific epitopes β10–β13 (of which β10 and β12 probably are hCGβ cystine knot-related, PB unpublished data) are not present on hCGβ, hCGβn, hCG, hCGn, or on hLH/hLHβ/hLHβcf. Two epitopes, α6 (hCGα loop 2 related) and α7 (hCGα carboxyl-terminal related), are specific for nonassembled hCGα.
Some additional Abs recognize epitopes defined only broadly at the molecular level, e.g., additional c- or β-mAbs. Within antigenic domains there seem to be epitopes that remain to be defined more precisely [1].
Algorithm to define hCG epitopes of ISOBM mAbs (INN)
For the Second ISOBM TD-7 WS on hCG, a previously reported epitope mapping algorithm [22], which was also used in similar form in the First ISOBM TD-7 hCG WS [1] has been further refined to reliably characterize the specificities and epitopes of the 69 ISOBM-Abs. This three-step algorithm involves:
-
1.
Determination of intraspecies Ab specificities with hCG, hCG-related variants [six First International Reference Reagents (IRR) preparations], hLH and synthetic hCGβCTP peptides to enable grouping of the mAbs according to their main specificities (α-, β-, and c-mAbs), and tentative assignment of epitopes by comparing specificity profiles to those of reference mAbs with known epitope recognition
-
2.
Confirmation of epitope recognition and spatial arrangement of epitopes using sandwich assays with mutual antigen recognition or inhibition by pairs of mAbs to provide information about epitope disparity or identity/vicinity
-
3.
Cross-referencing of the ISOBM-Abs’ reaction profiles in specificity and sandwich assays to those of reference mAbs recognizing previously defined epitopes.
This approach frequently enabled definitive assignment of epitopes at the molecular level. In rare cases where no exact molecular localization could be determined due to the lack of appropriate reference mAbs, additional circumstantial evidence, e.g., mutual steric inhibition in simultaneous antigen recognition with mAbs of known epitope localization or recognition of breakdown products like hCGβcf or inter-species cross-reactivity, was used to elucidate the epitope’s antigenic domain [1].
Materials and methods
ISOBM-Abs: codes and descriptions
Sixty-nine ISOBM-TD-7 Abs were submitted by eight participants to Dr. Kjell Nustad at the Central Laboratory, Norwegian Radium Hospital (NRH), Oslo, Norway (see Table 7 in Appendix). The Abs were assigned code numbers ISOBM-382 to 450. The panel contained (1) 42 Abs to be tested for molecular epitope recognition, (2) 10 mAbs that were previously specificity- and epitope-typed in the First ISOBM TD-7 WS on hCG as blinded internal controls: ISOBM-403 is identical to reference mAb -435 and corresponds to ISOBM-265 and ISOBM-274 in the First WS, ISOBM-411 is identical to -275 (First WS), ISOBM-415 to -281; ISOBM-416 to -273; ISOBM-417 to -276; ISOBM-418 to -280; ISOBM-419 to -271; ISOBM-420 to -264 and -277; ISOBM-422 to -272, and ISOBM-424 to -279 [1] (Appendix); and (3) 17 reference mAbs (ISOBM-434–450) of known specificity and epitope recognition provided by Dr. Peter Berger from the Institute for Biomedical Aging Research, Innsbruck (INN), Austria. The hybridomas producing mouse reference mAbs (INN-mAbs) against hCG, hCGβ, hCGβcf, and hCGβCTP were established as previously described [23, 25–31] and specificity, affinity, and epitope analyses by a panel of immunochemical techniques (for reviews, see [1, 22]).
The 17 reference mAbs were directed against 15 epitopes on hCG and hCG-related molecules (Table 2) (for reviews, see [1, 22]). Ten epitopes were located on intact hCG (epitopes β1–β5 and β8; c1–c4), and six of these shared by hCGβ (β1–β5, and β8). Two Abs recognized epitopes on hCGβ plus hCGβn plus hCGβcf (β6 and β7). Three reference mAbs against epitopes β10, β11, and β13 recognized exclusively hCGβcf. MAb FB12 recognizing hCGβCTP epitope β9 [32], ISOBM-278 (epitope (β8 type 1, β8,1) ISOBM-277 (epitope β8 type 2, β8,2) and ISOBM-267 (epitope β14) [1] were additionally used as control reagents. No reference or control mAbs for epitopes α1–α7, β12, and β8/3) were applied in the specificity and epitope typing experiments.
The Abs were checked for purity by sodium dodecyl sulfate polyacrylamide gel electrophoresis (SDS-PAGE), the protein content determined by measuring the absorbance at 280 nm (1 mg/mL = 1.43), aliquoted and 1 mg of each sent to the laboratories of the workshop participants performing the experimental work: Dr. Phil Hemken, Diagnostic Research and Development, Abbott Diagnostics (ABB); Dr. Elisabeth Paus, Radiumhospitalet, Oslo University Hospital, (NRH); Dr. Ulf-Håkan Stenman (UHS), Helsinki University Central Hospital; and Dr. Wilson Stewart, Ninewells Hospital and Medical School, Dundee (NHD).
First international reference reagents for hCG and hCG variants
The new international standards for hCG, nicked hCG (hCGn), hCGα, hCGβ, hCGβn, and hCGβcf were purified and characterized by the IFCC Working Group for Standardization of hCG Determinations [9] and adopted by the WHO as the First IRR for hCG and related variants [33]. The material is intended for use in the calibration of immunoassays in substance concentrations, i.e., moles per liter [6]. One milligram each of the six First IRRs for hCG and related molecules were kindly supplied by the NIBSC (Dr. Catharine Sturgeon, CS) to each of the participants and used to characterize the 69 ISOBM-Abs (Table 3).
For iodination, FRET and BIAcore® specificity and affinity determinations the carrier-free frozen concentrates (FC) of the six First IRRs were used: hCG (FC 99/688), hCGn (FC 99/642), hCGβ (FC 99/650), hCGβn (FC 99/692), hCGβcf (FC 99/708), and hCGα (FC 99/720).
Other hormones and peptides
Human LH (hLH-I-1) AFP4345B for iodination was obtained from National Hormone & Peptide Program, USA. The peptide hCGβCTP135-145, PGPSDTPILPQ, was ordered from AltaBioscience, UK. The peptide hCGβCTP109-145, TCDDPRFQDSSSSKAPPPSLPSPSRLPGPSDTPILPQ, was provided by Dr. Jean-Michel Bidart.
Biochemical characterization of the mAbs (ABB, NRH)
The mAbs were biochemically characterized by gel permeation chromatography–high performance chromatography (GPC-HPLC; ABB; Online Resource 1), SDS-PAGE under reducing (ABB; Online Resource 2) and non-reducing conditions (NRH; Online Resource 3), Ab isotyping (ABB, NRH; Online Resource 4), isoelectric focusing (IEF; ABB; Online Resource 5), and finally mass spectrometry (MS; ABB; Online Resource 6), which was utilized for further characterization of Ab samples where double heavy or double light chain bands were observed using SDS-PAGE testing.
Determination of Ab specificity, affinity, and epitope localization (ABB, NRH)
The main specificity profiles of mAbs were determined (1) by direct binding RIA (DB-RIA) with 125I-labeled hormones and hormone fragments with excess Ab (Online Resources 7, 8) and (2) with competitive ligand analysis (CLA), a RIA format, wherein the binding between 125I-hCG and serial diluted Abs is competed with fixed concentrations of the six First IRRs of hCG and hCG-related molecules and hLH (75/552), respectively (Online Resource 9). Cross-reactivity of the ISOBM-Abs with hLH was determined by titration RIA (NRH) by comparing titers of 125I-labeled hCG versus 125I-labeled LH (Online Resource 10). Epitope recognition on the hCGβCTP by ISOBM-Abs was evaluated by competitive RIA with synthetic peptides (NRH; Online Resource 11). Ab affinities were determined by Forster Resonance Energy Transfer (FRET) (ABB; Online Resource 12) and by BIAcore® (NHD; Online Resource 13). For elucidation of the spatial arrangement of epitopes, Ab compatibility in antigen recognition was evaluated by sandwich RIA (NRH; Online Resource 14).
Results
Biochemical characterization of the mAbs (ABB; NRH)
Gel permeation chromatography–high performance chromatography (ABB)
The homogeneity of the mAbs determined by GPC ranged from 57 to 99 % with varying degrees of aggregation and low MW contaminants (Fig. 3a; Online Resource 15). All samples exhibited low MW peaks possibly caused by buffer components (azide, citrate, DTT, etc.).
Biochemical characterization of ISOBM-435 by GPC-HPLC (a), reduced SDS-PAGE (b), and IEF (c). a Percent purity by GPC-HPLC analysis is 96 % with 7 % aggregation and a small amount (2 %) of low molecular weight (LMW) contaminants. The LMW material could be sodium azide and residual albumin, but this was not confirmed. b Reduced SDS-PAGE analysis shows a combined heavy and light chain purity of 96 %. The first strip has been enhanced to increase the image contrast to highlight a faint band (<1 %) that has a similar molecular weight as the albumin standard (66 kDa). The second strip utilized the auto-scale feature available with Quantity One software (Bio-Rad). A 1 % band is noted at approximately 76 kDa, and a 3 % band is also seen at approximately 47 kDa. c IEF reveals a tight pI range of 5.9–6.0.
Sodium dodecyl sulfate polyacrylamide gel electrophoresis (NRH, ABB)
SDS-PAGE analysis was performed under nonreducing (Online Resource 16a; NRH) and reducing conditions (Online Resource 16b; ABB). The purity of the mAbs determined as the proportion of heavy and light chains relative to all protein bands ranged from 84 to 100 % (Online Resource 16b). An example for slight albumin impurity is shown in Fig. 3b. The appearance of double chains could be due to glycosylation differences [34, 35], amino acid residue issues [34, 35], or the presence of more than one Ab in the sample (Online Resource 16b).
Isoelectric focusing (ABB)
The isoelectric point (pI) of the Abs ranged from 5.0 to 7.7 (Fig. 3c; Online Resource 17). Those for ISOBM-397, ISOBM-418, and ISOBM-431 could not be determined. The smearing or absence of bands was probably due to low solubility of these Abs at the low ion strength in IEF.
Isotyping (ABB)
Isotyping was performed on samples displaying double heavy or light chains in SDS-PAGE analysis (Table 4).
Mass spectrometry analysis (ABB)
Apart from the following exceptions most ISOBM-Abs gave expected results. ISOBM-382 had double light chains, and more than one group of heavy chains present following deglycosylation, suggesting presence of two mAbs. ISOBM-385 had two groups of heavy chains due to glycosylation and ISOBM-388 two nonglycosylated light chains due to amino acid residue differences. ISOBM-400 and ISOBM-406 had more than one group of light chains due to differences in glycosylation and ISOBM-426 complex heavy chains and ISOBM-450 complex light chains (Online Resource 18).
Ab affinities, specificities, and epitope localizations
Ab specificities (NRH)
Based on the results of DB-RIAs with 125I-labeled hCG, hCG-variants, and hLH tracers, the 69 ISOBM-Abs were categorized according to their main specificities (α-, β-, and c-mAbs) (Fig. 4): Antibodies either recognized (a) assembled and/or free hCGα (hCGα-mAbs, n = 8; α epitopes were not determined) or (b) assembled and/or free hCGβ or hCGβ metabolites such as hCGβcf (hCGβ-mAbs, n = 48; epitopes β1–β13), or (c) exclusively the intact ± nicked hCGαβ heterodimer, but not the free subunits or metabolic variants thereof (c-mAbs, n = 13; epitopes c1–c4).
Specificity profiles of the ISOMB-Abs of the Second TD-7 WS recognizing hCG and hCGβ variants (a), hCG-only and hCGα, respectively (b) were determined by binding of iodinated tracers to excess of Ab (DB-RIA) (NRH). ISOBM-mAbs were classified according to their main specificities and their epitopes recognized on the basis of cross-reactivity patterns with hCG, hCG-variants, and hLH: (1) β-mAbs corresponding to epitopes β1–β13, (2) c-mAbs recognizing epitopes c1–c4 on holo-hCG only, and (3) α-mAbs. a MAbs directed against epitopes β1–β5 are pan-hCG reagents recognizing hCG and hCGβvariants but differ in their cross-reactivity with hLH: β1 mAbs are highly specific for hCG and show no hLH cross-reactivity (<0.1 %), β2 and β4 show very low hLH reactivity (<1 %), whereas β3 and β5 strongly cross-react (>>1 %). Epitopes β6 and β7 are specific for uncombined hCGβ, hCGβn, and hCGβcf. MAbs against epitope β8 at the very carboxyl-terminal end of hCGβCTP do not cross-react with hCGβcf and hLH but recognize all other hCG variants except for those lacking the CTP. These mAbs constantly show a low bindable fraction of the tracers as only approximately 50 % of the tracers can be bound specifically. This is in contrast to the β1–β5 mAbs. ISOBM-418 seems to be directed against epitope β9 as already typed previously in the First WS (ISOBM-280, [1]. Epitopes β10–β13 are specific for hCGβcf as no other hCG variants or hLH are recognized by the respective mAbs. b c-mAbs directed against epitopes determined by the quaternary structure of hCG either do not (c1 and c2) or do recognize hCGn (c3 and c4) [56]. The apparent hCGn cross-reactivity of c1 and c2 mAbs is due to a cross-contamination of this preparation with non-nicked hCG (approximately 20 %) [1]. The presence of non-nicked hCG and recognition by the ISOBM-mAbs of the two-nicked forms in hCGn were investigated in detail by LC-MS/MS (see accompanying publication by H. Lund). Epitope c3 (ISOBM-446 = INN-hCG-45, reference mAb) is highly specific for hCG + hCGn. ISOBM-mAb 433 that has the same specificity pattern might be directed against a fifth sterically independent c-epitope as shown by sandwich assay. The exact molecular localization of epitope c4 on hCG is not known, but it is remote from the other c-epitopes. In the First ISOBM TD-7 WS, ISOBM-424 has been characterized (ISOBM-279) and classified as c4 specific [1]. The α-mAbs have not been investigated in detail as to their epitope recognition. As they readily recognized iodinated tracers (in contrast to α3- and α5-mAbs), they should be directed against the epitope cluster α1/α2/α4 with the exception of ISOBM-404 that is free hCGα-specific and therefore presumably recognizing the subunit assembly region of hCGα (aa hCGα33-42). Minor apparent cross-reactivity with hCGn is owed to a cross-contamination of hCGα in that preparation. hLH cross-reactivity of ISOBM-404 might be due to dissociation of highly purified hLH that is observed during testing (PB, personal observation). DB-RIA with 125I-tracers: results are expressed as maximum specific binding in percent of the “bindable fraction” of added tracer (NRH) [26]. Italics RIA titration experiments (NRH): Results are expressed as percent hLH cross-reactivities compared to hCG; asterisk 125I-tracers; gray background significant cross-reactions. Superscripted a Apparent cross-reactivities with hCGn of ISOBM-447–438 are caused by an approximate 20 % cross-contamination of intact hCG (see accompanying publication by H. Lund) and superscripted b of ISOBM-383–404 due to a suchlike with hCGα that is contained in hCGβn; superscripted c ISOBM-404: apparent cross-reactivity is probably caused by slight dissociation of α-subunit in hLH. Section symbol Competitive RIA with hCGβ109-145 vs. hCGβ*, percent cross-reactivity. Double section symbol Competitive RIA with hCGβ135-145 vs. hCGβ*, percent cross-reactivity
Comparing specificity profiles of ISOBM-Abs to those of reference mAbs permitted preliminary epitope assignment. To discern mAbs against epitopes β1–β5, hLH cross-reactivity was determined by titration RIAs with 125I-hLH (Fig. 4). Recognition of hCGβCTP was investigated by competitive RIA using synthetic peptides derived from hCGβCTP (Fig. 4a; Online Resource 19). No mAbs against epitope β14 (hCGβ specific) were identified in this ISOBM panel (Fig. 4a).
Ab affinities and specificities as determined by FRET (ABB)
The ISOBM-Abs were grouped according to their specificity profiles based on affinity for hCG, hCGβ, hCGβcf, and hLH (Fig. 5) determined by FRET. Affinities for the major hCG variants and hLH, reported as dissociation constants (K d), ranged from subpicomolar values (0.3 pmol/L for hCGβ of ISOBM-429) to ≥50 nmol/L. The latter value indicated that binding was very weak or not detectable.
Affinity and specificity of the ISOBM-Abs as determined by FRET (ABB). Preliminary assignment of epitopes was done by comparing the specificity profiles of the ISOBM-Abs to those of reference mAbs. Specificities based on affinity of mAbs against hCGα could not be determined with hCG, hCGβ, and hCGβcf
Only nine Abs (ISOBM-387, ISOBM-399, ISOBM-401, ISOBM-414, ISOBM-416, ISOBM-427, ISOBM-428, ISOBM-429, and ISOBM-444) expressed high affinities (K d, ≤50 pmol/L) for any of the four antigens tested (hCG, hCGβ, hCGβcf, and hLH). It is striking that six of these mAbs recognize the major antigenic domain on the tips of hCGβ loops 1 + 3. An exception was the hCGβcf-specific mAb ISOBM-444 (reference mAb INN-hCG-106), the epitope of which (β11) does not overlap with epitopes β2–β5 on hCGβ loops 1 + 3 nor with the cystine knot-associated epitopes β1 and β7. Thus, this epitope is remote from either cluster. This is an interesting mAb for highly sensitive and specific measurement of hCGβcf in particular in combination with β2-mAbs [9, 36].
Another interesting observation is that all three sheep mAbs, ISOBM-428, ISOBM-429 and ISOBM-430, were in the high affinity group. ISOBM-430 was not tested by the FRET technology but by titration RIA. All four sheep Abs (three mAbs and polyclonal ISOBM-431) were directed against hCGβ loops 1 + 3 epitope β5 that is shared by hLH and, therefore, in principle, do not seem suitable for hCG measurement. Nevertheless, ISOBM-429 seems to have tolerably low cross-reactivity with hLH (Figs. 4a and 5 and Online Resource 20), but its suitability for use in hCG + hCGβ variant measurement might still be hampered by preferential recognition of hCGβ.
Most Abs, including all directed against the cystine knot-associated epitopes β1 and β7, the hCGβCTP epitope β8 types 1 and 2, showed moderate affinities (50 pmol/L–5 nmol/L) against their primary target hCG variant. Low affinities could be observed for three reasons: (1) the primary antigenic target hCG variant of the mAb in question was not among the antigens tested, as is the case of mAbs against uncombined hCGα (ISOBM-404), or (2) genuine low affinity to the primary target antigens, e.g., ISOBM-418/280 (hCGβCTP epitope β9, aa hCGβ113-116) and ISOBM-443 and 448 against hCGβcf, and (3) FRET labeling affected binding of Abs (ISOBM-445 to hCGβcf; ISOBM-399, ISOBM-436, ISOBM-412, and ISOBM-421 to hLH) (Fig. 5). This is also the case with 125I-labeling of epitopes α3 and α5 [37].
Ab affinities and specificities as determined by BIAcore® (NHD)
The specificity patterns of the ISOBM-Abs were determined based on their affinity for hCG, hCGβ, and hCGβcf in BIAcore®. The affinities (dissociation constants; K d) ranged from picomolar values (<10 pM for hCG of ISOBM-399) to >10 nM. An affinity of <100 nM was observed for 43 of the Abs for either a single or a combination of the three antigens tested (hCG, hCGβ, and hCGβcf). Five of these Abs, ISOBM-427 (epitope β2), ISOBM-399, ISOBM-423, ISOBM-441 (all three epitope β3), and ISOBM-428 (epitope β5) had affinities of <10pM for the antigen. It is striking that all of these recognize the tops of hCGβ loops 1 + 3; thus, all epitopes were located within the same antigenic domain. Assignment of epitopes to the ISOBM-Abs was achieved by comparing their specificity profiles to those of reference mAbs (Online Resource 20).
The affinity of many of the ISOBM-Abs appeared to be higher than determined by FRET analysis. This could perhaps be a consequence of having the antigen in a bound form on the BIAcore® chip rather than in a fluid state. The affinity for this immobilized form of antigen may result in an overestimate of affinity.
Ab specificities as determined by CLA (NHD)
In the CLA approach, ISOBM-Abs were titrated against 125I-hCG and in parallel competed with a fixed amount (0.5 pmol/mL) of hCG, hCG-variants, and hLH, respectively. A shift of the Ab dilution curves to a lower titre indicated cross-reactivity of this competitor with the Ab. Based on the CLA results, 34 ISOBM-Abs, either recognized (a) assembled and/or free hCGβ metabolites such as hCGβcf (hCGβ-mAbs, n = 24; epitopes β1–β9) or (b) exclusively hCG ± hCGβ, but not the free subunits (c-mAbs, n = 10; epitopes c1–c4). Epitope assignment was achieved by comparing profiles of the reference mAbs with ISOBM-Abs (Online Resource 20).
No CLA analysis was possible for 35 ISOBM-Abs, which had very low or no signal, which suggested that the Ab did not recognize 125I-hCG or could not be competed with the amounts utilized.
Epitope classification by sandwich assays (NRH)
IRMA-like sandwich assays were performed to confirm preliminary Ab epitope classifications by specificity assays and to determine epitope localization by comparison with reference mAbs. Characteristic reaction patterns were observed when solid-phase bound ISOBM-mAbs were tested for their ability to sandwich hCG or hCGβ with the panel of reference mAbs directed against epitopes β1–β9 (Fig. 6a) and c1–c4 (Fig. 6b). Patterns observed agreed with previously determined epitope locations for the reference mAbs [28, 38].
Classification and spatial relationship of ISOBM-mAb epitopes. Two-site IRMA-like sandwich assay experiments with a chessboard-like matrix of antibody pairs tested for their ability to simultaneously bind hCGβ (99/650) for hCGβ-mAbs (a) and hCG (99/688) for holo hCG-mAbs (b) (NRH). Reference Abs for epitopes β1–β9 and c1–c4 served as 125I-labeled detection reagents, respectively. Reaction profiles of the solid-phase ISOBMii mAbs with the detection reference mAbs were cross-matched to that of solid-phase reference mAbs the molecular epitope specificity of which had previously been defined [1]. Similar reaction profiles were interpreted as epitope identity or neighborhood of mAbs. a The compatibility patterns of pairs of mAbs do not only reveal epitope affiliation of single mAbs but also disclose hCGβ epitope arrangement in larger antigenic domains consisting of one or more epitopes. Abs directed against epitopes located within the same antigenic domain are generally mutually exclusive in hCGβ recognition, whereas those the epitopes of which are located in different domains are compatible. Three major antigenic domains were identified on hCGβ: (1) the domain on the tips of hCGβ loops 1 + 3 encompassing epitopes β2–β6 (2) the cystine knot associated domain including hCG specific epitope β1, hCGβ + hCGβcf specific epitope β7, and a structurally related hCGβ-only specific epitope epitope β14 located on core hCGβ1-112 and characterized by a single mAb, and (3) hCGβCTP epitopes β8 and β9 remote from the other domains. MAbs against all hCGβ loops 1 + 3 associated epitopes β2–β6 are compatible with the hCG-specific cystine knot-associated epitope β1 and vice versa. Within antigenic domains not all epitopes can be discerned by distinct reaction profiles. As an example, although β1 and β7 show identical patterns in sandwich assays and are not compatible with each other, they are definitely recognizing different but adjacent epitopes as β1-mAbs are pan-hCGβ-mAbs recognizing a broad spectrum of hCG-variants and in contrast β7-mAbs are highly selective for hCGβ + hCGβcf and would not recognize, e.g., hCG (see, e.g., DB-RIA, Fig. 4). A second example are mAbs against epitopes β4 (ISOBM-419 and ISOBM-445) and β5 (ISOBM-428, ISOBM-429, ISOBM-430, ISOBM-431, and ISOBM-442) having an identical compatibility profile, i.e., nicely work with mAbs against epitopes β1 and β7–9 but not with β2–β6. These epitopes can be discerned by their variant recognition profiles whereby β4 mAbs are specific for hCG (≤1 % cross-reactivity with hLH) and β5 mAbs strongly cross-react with hLH (>>1 %) in titration and competitive RIA (Fig. 4). β3-mAbs, although showing a similar reaction pattern as other mAbs directed to hCGβ loops 1 + 3 associated epitopes (β2, β4, β5, and β6), seems to be remote from the free subunit specific epitope β6 and not compatible with hCGβCTP113-116 located epitope β9 at the beginning of hCGβCTP. Such spatial vicinity between the hCGβCTP and hCGβ loop 3 has already been postulated previously [72]. As expected, the epitope of β8-mAbs located at the very carboxyl-terminal end of hCGβ (aa hCGβ141-144; [24]) is compatible with all other epitopes. In the first ISOBM TD-7 WS, a new epitope β14 was observed represented by a single mAb (ISOBM-267) that exclusively recognized core hCGβ [1] and that now appeared compatible with all hCGβ locate epitopes except for epitope β1, thus seems to be remote from any other core hCGβ epitope. ISOBM-406 according to its sandwich pattern (no compatibility with cystine knot epitopes β1 and β7) seems to be cystine knot associated. b c-mAbs show variant reaction patterns among themselves. The heterodimeric epitopes c1–c3 are located in the same antigenic domain thus are not compatible with each other. c4 is clearly remote from that domain as as it is compatible with c1 to c3-mAbs. ISOBM-433 recognizes a previously structurally not defined epitope that is highly hCG specific as is ISOBM-446 (epitope c3) (Fig. 4). ISOBM-389 a highly hCG specific c-mAb that according to BIAcore® analyses rapidly dissociates (K d = 13E−03), ISOBM-397 and ISOBM-418 (very low affinity in FRET analyses) did not perform well as capture mAbs in this type of assay and were negative throughout (not shown). Reactions classified as positive (mean + 2 standard deviations) are depicted as closed squares. Noncompatible mAb pairs are shown as white squares
Compatibility of Ab pairs in sandwich assays indicated that their epitopes were spatially distinct, e.g., epitopes β1 versus β2–β6 and vice versa (Fig. 6a). Identical or highly similar compatibility patterns of Abs to that of reference or other mAbs indicated recognition of identical or of neighboring epitopes within the same antigenic domain: e.g., the cystine knot-related epitopes β1 and β7, or epitopes β2, β4, and β5 on hCGβ loops 1 + 3. Epitopes within a particular antigenic domain can be easily discerned by cross-reactivity patterns with hCG variants and LH from various species [1]. Thus, although β1- and β7-mAbs show identical compatibility patterns in sandwich assays and are not compatible with each other (Fig. 6a), they recognize different but spatially adjacent cystine knot-related epitopes reflected by differing variant recognition patterns: β1-mAbs recognize a broad spectrum of hCG-variants whereas β7-mAbs are highly selective for hCGβ, hCGβn ± hCGβcf and do not recognize hCG (Fig. 4).
Antigenic domains and epitope maps of hCG and hCGβ (INN)
Results of the three approaches for epitope typing are summarized in Table 5. In Fig. 7, the ISOBM-mAbs are assigned to the three-dimensional epitope maps of hCGβ (a) and hCG (b), which were established with the reference mAbs previously [1]. The Abs grouped according to epitope recognition are listed in Table 8 in Appendix.
Epitope maps of hCG, hCGβ, and variants (INN) (modified according to [1], with permission) were previously constructed based on the epitopes recognized by the reference mAbs. The identification of reference mAb epitopes was performed by direct binding, competitive and sandwich RIA and ELISA with hormones of various species, hormones subunits, metabolic breakdown products, and synthetic peptides (for reviews, see [1, 22]). Furthermore, on the basis of molecular modeling of crystallographic data of hCG and subsequent mutational analyses to assign epitopes to particular amino acids, epitopes of reference mAbs and, by comparison, epitopes of ISOBM-mAbs could be superimposed on the molecular model of hCGβ. a Assignment of ISOBM-mAbs to epitopes on the molecular model of hCGβ/hCGβn/hCGβcf/hCGβCTP. Reaction profiles of the ISOBMii mAbs in specificity and sandwich assays were compared to that of reference mAbs. It appeared that the most immunogenic region of hCGβ is determined by the peptide sequences that correspond to hCGβcf. In particular, the tips of beta-sheet loops 1 + 3 corresponding to hCGβ20-25 + 68–77 comprise the major antigenic domain (epitopes β2–β6) that is recognized by high affinity mAbs. The only hCG-specific epitope on core hCGβ is β1 located around the center of the molecule corresponding to part of the cystine knot (aa hCGβ10,60,89). Adjacent to epitope β1, the hCGβ/hCGβcf-specific epitope β7 is also located in this region (aa hCGβ61,89) [43]. Thus, pairs of antibodies against these two epitopes are not compatible in sandwich type assays (Fig. 6a) [24]. hCGβCTP epitopes β9 and β8 are located at either end of the hCGβCTP, whereby β9 might be close to epitope β3 (Fig. 6a) [72]. b Epitope map of hCG. ISOBM-mAbs were assigned to epitopes on a ribbon representation of the molecular model of hCG [3]. hCGα and epitopes thereon are depicted in blue, hCGβ and its epitopes in green. Conformationally (c) dependent epitopes determined by the quaternary structure of hCG are shown in red. Note the major antigenic clusters of epitopes on the top of beta sheet loops 1 and 3 of hCGα (α1/α2/α4 and α3/α5) and of hCGβ (β2– β5), the central cystine knot-based epitope cluster encompassing highly hCG-specific β1 and c-epitopes (c3), the latter having a share on loop 3 of hCGβ, that in turn are confluent with the α1/α2/α4 epitope cluster. The hCGβCTP epitopes are located on both of its ends at aa hCGβ113-116 (epitope β9) and aa hCGβ133-144 (epitope β8)
In the ISOBM panel, 48 out of 69 were β-Abs. The major antigenic domain on hCGβ located on the tips of the neighboring β-sheet loops 1 and +3 encompassing aa hCGβ20-25 and 68–77 (epitopes β2–β6) was recognized by 27 of the β-Abs. Of these, 12 Abs recognize epitopes β2 or β4 (β2/4), 13 epitopes β3 or β5 (β3/5), and 2 epitope β6. Epitopes β2–β5 are pan hCG specific, i.e., present on hCG, hCGn, hCGβ hCGβn, and hCGβcf (Fig. 4a), whereas β6 is present only on hCGβ, hCGβn, and hCGβcf.
The cystine knot-associated antigenic domain comprises a number of epitopes that are recognized by 10 of the 48 β-mAbs (including ISOBM-397 results of which are ambiguous). ISOBM-267 that is hCGβ specific and its epitope cystine knot-related (epitope β14) was used as a control mAb: The pan-hCG epitope, β1 (hCGβ Arg10, Arg60, and Gln89), which is not shared by hLH, was recognized by two ISOBM-mAbs (ISOBM-403 and reference mAb ISOBM-435). It is spatially close to epitope β7 (hCGβ Asp61 and Gln89) against which six mAbs (including ISOBM-397) were directed. MAbs classified as β7 recognize either hCGβ + hCGβn + hCGβcf or mainly hCGβ + hCGβn (ISOBM-407). The cystine knot-related epitope β10 recognized by reference mAb ISOBM-448 is hCGβcf specific. One mAb (ISOBM-406) reacted with a not specified cystine knot epitope (Fig. 6a). This mAb is of restricted pan-hCG specificity and does not recognize hCGβcf (Fig. 4a).
In addition to the above-mentioned mAb ISOBM-448 (cystine knot related epitope β10), 4 of the 48 β-Abs recognize epitopes located on hCGβcf only (epitope β11, ISOBM-384 and ISOBM-444; epitope β13, ISOBM-443; and one non-coded hCGβcf epitope, ISOBM-393).
Six of the 48 β-Abs are directed against the hCGβCTP. The linear antigenic region (aa hCGβ137–144; epitope β8) at the very end of the hCGβCTP is recognized by four mAbs; type 2). Three of these (ISOBM-395, ISOBM-413, and ISOBM-420) are mAbs against epitope β8,2 recognizing glycosylated hCGβ much better than the nonglycosylated synthetic peptide. Thus, epitope β8,2 might be influenced by glycans on Ser132 and/or Ser138 [1, 39]. One mAb (ISOBM-450; epitope β8,1) recognizes both antigens to the same extent. Two mAbs, 394 and 418, may be directed against epitope β9. One β-mAb (ISOBM-392) could not be classified but it does not seem to be located on hCGβCTP (Figs. 4a and 7a).
Thirteen of the 69 mAbs reacted with c-epitopes: c1 (n = 2), c2 (n = 6), c3 (n = 1), c4 (n = 2), c (n = 1; ISOBM-433; new noncoded c-epitope). One c-mAb could not be classified (ISOBM-389).
Eight out of 69 mAbs are directed against hCGα. Six recognize assembled and one, ISOBM-404, which has been prepared by immunization with hCGα (Stenman et al., unpublished data), recognizes only free hCGα. The exact molecular localization of the hCGα mAbs was not elucidated (Figs. 4b and 7b).
Discussion
Topography of hCG epitopes
Epitopes and antibodies
By definition, epitopes are molecular structures dependent on the existence of complementary Abs. Not the entire surface of a glycoprotein like hCG is antigenic. Against certain molecular areas no Abs exist as they are immunologically inert, e.g., due to insufficient T cell help, or sterically not accessible due to protein folding or shielding by glycans. In contrast, other areas representing structurally inherent epitopes, which are characterized by high solvent accessibility and high protrusion indices, are often sites of Ab recognition [40]. hCGβ cystine knot-associated residues Arg10 and Gln89 (epitopes β1 and β7), hCGα loop 1 residues Pro16, Phe17, and Phe 18 (epitopes α1, α2,and α4) and the antigenic domain on hCGβloops 1 + 3 comprising aa 20–25 + 68–75 (epitopes β2–β6) all bulge away from the molecule forming prominent surfaces that are the major antigenic domains of hCG [3, 22, 37, 41–43]. There is a good chance that irrespective of the immunized species these molecular structures will be recognized as epitopes [38]. For example, the immunodominant antigenic domain on top of hCGβ beta-sheet loops 1 and 3 is recognized by Abs derived from mice and sheep as shown in the present study and interestingly by Abs from humans and rabbits (PB, unpublished observations). Moreover, hLH cross-reactive mAb B206 directed against an epitope within this cluster, presumably epitope β3/5, inhibited 40–90 % of the binding of human antisera to hCG [44].
The definition of epitopes by Abs and recognition of the multitudes of possible amino acid combinations within an inherently antigenic structure/domain is dependent on and restricted by the combinatorial repertoire of the VDJ and VJ immunoglobulin heavy and light chains gene segments, respectively, and the cellular capacity to mature the paratope of a given Ab to optimally fit the antigenic surface. This repertoire of Ab specificity varies with individual immune responses, haplotypes, and species. Not every amino acid combination within an antigenic domain will therefore be recognized by Abs of any individual or species. Thus, the repertoire of Ab specificities and corresponding epitopes within an antigenic domain is very large but still somewhat restricted as shown by the present and previous studies. For example, the antigenic domain on hCGβ loops 1 + 3 is recognized by large panels of Abs that differ slightly in hCG variant recognition, hLH cross-reactivity, affinity, etc. This has been shown to be due to variability in amino acid recognition within the antigenic domain [1].
It is striking that this antigenic region, aa hCGβ20–25 + 68–75 on the tips of loops 1 + 3, comprises 16 amino acids, a number that reasonably well corresponds to the surface covered by a single complementary paratope of an Ab whereby two to three amino acids that vary from Ab to Ab provide most of the binding energy and fine specificity [45]. Consequently, dozens of ISOBM-mAbs and Abs of other panels directed against hCGβ loops 1 + 3 epitopes β2–β5 do not behave uniformly in their recognition of the approximately 15 potential contact amino acids composing discontinuous epitopes, even though they cover more or less the same surface with their paratope [43]. Thus, all differences in affinity, specificity, and hLH cross-reactivity of numerous antibodies directed against this major antigenic region seem to have their basis in variability of preferential recognition of a few amino acids, providing binding energy within very similar or even identical sets of amino acids covered by the Abs’ paratopes.
The surface area of an epitope that is covered by a cylinder-like antigen binding site of an Ab is approximately 700 Å2 in size [38, 46], whereby the radius of the antibody binding domain is 8–10 Å and the radius of the epitope covering area is 15 Å irrespective of Ab specificity [45]. X-ray crystallography studies revealed that core hCG, i.e., hCG without hCGβCTP, has a length of 75 Å and a width of 30–35 Å [3, 47] corresponding to a surface area of approximately 8,200 Å2. As some regions on assembled hCGβ, such as the stems of β-sheet loops 1 + 3, are not recognized by any anti-hCG-mAbs [1, 18, 48], the total epitope-covered area on core hCG could be in the range of 5,000 Å2 theoretically accommodating simultaneous binding of up to seven Abs to spatially independent epitopes. The minimal spatial requirement for sterical compatibility of two mAbs is that the respective epitopes are approximately 20–30 Å apart. In fact preliminary experiments showed that at least five radiolabeled mAbs against epitopes β1 + β3 + α2 + α3 + c4 were able to bind to core hCG simultaneously [38].
Glycosylation and epitopes
With two exceptions, glycosylation has little effect on hCG’s immunological make-up, although the glycans, which are hydrophilic in nature and thus surface exposed, represent approximately 30–35 % of its total molecular mass. The exceptions are glycans at the very end of hCGβCTP and in the stem region of hCGβ loop 1. The 14 epitopes on core hCG, which is lacking hCGβCTP, are dependent on the protein backbone. Neither desialylation, deglycosylation [48], partial natural deglycosylation as in the case of the metabolic product hCGβcf [49], nor intense glycosylation as shown with highly acidic pI variants of pregnancy- and tumor-derived hCG have essential effects on Ab recognition by the reference mAbs [17, 18]. In addition, the number and the relative spatial location of epitopes do not differ between the isoforms [1, 18, 48].
The peptidic stem region of assembled hCGβ loop 1, which accommodates the two large N-linked glycans at hCGβAsn13 and Asn30 that are spatially near the hCGα glycan at Asn52 [3], is not recognized by any mAb in the panels of anti-hCG-mAbs of the previous and the present study. Thus, the immune response seems to be attenuated by the N-linked glycans in this region of hCGβ loop 1 [1, 18, 48].
A mAb (B152) that was not included in this study recognizes hCG with a core-2 O-glycan at Ser 132 and surrounding peptide structures [50, 51]. Its epitope, which we termed β8,3, is spatially related to epitope β8,2 that also seems to be influenced by the glycans on Ser 132 and/or Ser 138 [1, 29].
Some hCG assays have been claimed to underestimate hCG-h [52]. However, these results have been obtained with an hCG-h preparation that also was completely nicked (C5) [39]. Thus, the results most probably reflected failure to recognize hCGn rather than hyperglycosylated hCG.
Epitopes on assembled and/or free hCGβ (β1–β9, β14) and hCGβcf only (β10–β13)
The immunodominant structure of hCG and hCGβ-related molecules is the molecular region corresponding to hCGβcf, which has lost its N-terminus, the long loop 2, most of its N-linked carbohydrate antennae, and the hCGβCTP with all O-linked glycans but has retained its protein backbone configuration [53]. Thus, numerous mAbs against epitopes β1–β7 recognize hCGβ, hCGβn, and hCGβcf. However, one mAb (ISOBM-407) did not react with hCGβcf.
The epitopes on assembled and/or free hCGβ (β1–β9, β14) are located in three molecular regions: (1) hCGβ cystine knot, (2) tips of hCGβ loops 1 + 3, and (3) hCGβCTP.
The cystine knot-associated antigenic domain includes epitope β1 involving aa hCGβArg10 + Arg60 and possibly Gln89 that sterically are in close proximity to each other [42, 43]. hCGβArg10 and Gln89 are unique to hCG and not shared by hLH. This presumably explains why epitope β1 is highly specific for hCG and its variants and therefore is not present on hLH or hLHβ [26]. Due to its superior specificity, it is highly valuable for hCG/hCGβ-variant measurement by immunoassay with no interference by hLH or hLHβ [1].
The assumed location of epitope β7 on hCGβ, hCGβn, and hCGβcf is based both on mutational analyses and vicinity analysis by sandwich assays: It is associated with the cystine knot, present on hCGβcf, and Asp61 and Gln89 have a role in this epitope. Thus, in sandwich type assays, β7-mAbs are not compatible with β1-mAbs (Fig. 6a) [1, 22, 24].
MAbs against the cystine knot epitope β7 recognize hCGβcf in addition to hCGβ. ISOBM-407 is an exception to this, although other parameters match with epitope β7, it shows an exceptionally low cross-reactivity with hCGβcf (Fig. 4) and thus seems to be suitable for measurement of hCGβ in urine in the presence of high levels of hCGβcf. The assignment of hCGβ specific epitope β14 to the cystine knot antigenic domain is based on circumstantial evidence as mAb ISOBM-267 defined in the First ISOBM TD-7 WS to recognize epitope β14 is not compatible with hCGβcystine knot-related epitope β1 but with all other hCGβ-related epitopes (Fig. 6a). Two hCGβcf epitopes β10 and β12 are also cystine knot-associated (PB, unpublished data). An additional cystine knot-related epitope is represented by mAb ISOBM-406.
Antibodies directed against the major hCGβ antigenic domain on loops 1 and 3 are of significantly higher affinity compared to those against other antigenic regions of hCGβ[1, 21, 54]. MAbs against epitopes β2–β5 recognize a wide spectrum of hCG and hCGβ-related variants (hCG, hCGn, hCGβ, hCGβn, and hCGβcf) [1, 17, 18]. MAbs against epitopes β3 and β5 additionally react well with hLH and hLHβ, whereas epitopes β2 and β4 are specific for hCG and hCGβ variants (<1 % hLH and hLHβ cross-reactivity) and thus highly suitable for specific measurement of hCG and hCGβ variants (Fig. 4) [1, 26].
In summary, β-epitopes located on the protein core hCGβ1-112 are discontinuous in nature, determined by the tertiary protein structure, present on hCGβcf, and arranged in antigenic domains associated with the cystine knot and on the tips of loops 1 + 3. MAbs directed against these epitopes are of adequate affinity and suitable for immunoassay applications.
hCGβ-related epitopes not determined by hCGβcf or core hCGβ1–112 are located in two major regions on the hCGβCTP (aa hCGβ113–145). The immunodominant linear antigenic region at the very end of the hCGβCTP consists of aa hCGβ133–144 and encompasses epitope β8 that is composed of epitope variants β8,1, β8,2, and β8,3) [29, 55]. It partially seems to be influenced by glycans on Ser132 and/or Ser138 (epitopes β8,2 and β8,3) [29] [1, 50]. One mAb in this WS (epitope β8,1; ISOBM-450) and four mAbs in the First WS recognized nonglycosylated synthetic peptides and glycosylated hCGβ equally [1]. Epitope β9 at aa hCGβ113–116 [21, 24] was recognized by two mAbs (Fig. 7a) whereby ISOBM-394 was of high and ISOBM-418 of very low affinity (Fig. 5).
When immunizing with the glycoprotein hCG, the vast majority of antibodies will be generated against composite epitopes on hCGα or the core region of hCGβ (aa 1–112) but only rarely against linear peptide sequences of low structural order like the hCGβCTP. MAbs against hCGβCTP are generally of fairly low affinity. Nevertheless, they are used in diagnostic sandwich-type immunoassays as they do not cross-react with hLH (Fig. 4).
hCGα epitopes (α1–α7)
In the panel of ISOBM-mAbs, 8 of 69 recognize hCGα epitopes. One of these mAbs, ISOBM-404, seems to be specific for free hCGα, and it is speculated that it might recognize the sequence hCGα33-42 on the single loop 2. As no reference hCGα-mAbs (Table 2) were included, a detailed assignment of epitopes was not possible.
Epitopes on the hCG αβ-heterodimer (c1–c4)
At least four epitopes (c1–c4) are present only on hCG ± hCGn but not on either free subunit or hCGβcf [21, 26, 28]. Detailed analysis of hCG and hCGn recognition by the ISOBM-mAbs was performed by liquid chromatography mass spectrometry (LC-MS/MS) (see accompanying publication by H. Lund). Epitopes c1 (reference mAb INN-hCG-10) and c2 (reference mAbs INN-hCG-40 and INN-hCG-53) are (1) dependent on intact hCG and thus sensitive to nicking of assembled hCGβ loop 2, (2) not compatible in sandwich-type assays with the cystine knot-related hCGβ epitope β1 (aa hCGβArg10 + Arg60 and possibly Gln89) [38], and (3) incompatible with mAbs recognizing epitope cluster α1, α2, and α4 [38] on loop 1 in the region of aa hCGα13-22. Amino acids hCGβ44-48 in loop 2 and hCGα loop 1 have been shown by X-ray crystallography to be in close proximity as the subunits are assembled in a head-to-toe fashion [3]. It is striking that in sandwich assays c1-mAbs show identical reactivity patterns as α1- and α2-mAbs reflecting sterical epitope relatedness [28, 38].
MAbs against epitope c3 are sterically related to epitope c2, highly specific versus hLH as well as non-combined intact and modified subunits (<1 % cross-reactivity), not influenced by nicking of assembled hCGβ loop 2, and thus recognize hCGn and hCG equally [1, 56](Fig. 4). They are therefore highly suitable for simultaneous measurement hCG and hCGn (Table 6).
The exact molecular localization of epitope c4 has not been resolved yet. It is present on hCGn and hLH, remote from and thus sterically compatible with all other c-epitopes and to a minor extent determined by hCGβ as shown by low cross-reactivity [1, 26, 38]. A variant of the c4-epitope represented by ISOBM-424 (=ISOBM-279, First ISOBM TD-7 WS) that is not shared with hLH (cross-reactivity <0.1 %) seems to exist. A presumably fifth highly specific c-epitope has been observed in sandwich assays wherein mAb ISOBM-433 is compatible with mAbs to c1–c4 (Fig. 6b). Its molecular localization is unknown. ISOBM-389 is a c-mAb that could not be epitope typed but, according to its specificity profile analyzed by LC-MS/MS, might be a c2 mAb (see accompanying publication by H. Lund).
Method-specific recognition of hCG and hCG variants
Sandwich-type assays measuring hCG alone or in combination with free hCGβ and metabolites are used for detection of pregnancy, pregnancy-related disorders, trophoblastic disease, and various other female and male tumors [2]. Detailed knowledge of the epitopes recognized by the Abs used facilitates development of assays providing better comparability of the results between methods. It has been suggested that assays that are multifunctional with respect to clinical use should (1) recognize in an equimolar fashion hCG and hCGβ protein backbone and glycosylation variants, (2) not cross-react with hLH or derivatives, and (3) not be prone to signal blunting by non-measured variants, e.g., caused by excess hCGβcf leading to false low results [36]. This is a problem when hCG in urine is measured with sandwich assays utilizing a mAb against core hCGβ1-112 in combination with an anti-hCGβCTP mAb [57].
While assays measuring hCG and all hCGβ-related variants are useful as first line methods, for diagnosis of pregnancy and cancer, it is often advantageous to specifically measure only selected variants [2]. Thus, specific hCGβ assays are used for first trimester Down’s syndrome screening and also for diagnosis of testicular [58, 59] and nontrophoblastic cancers, 20–50 % of which produce only hCGβ but not hCG [60–62]. However, the concentrations are mostly low, and the assays used need to be highly sensitive. Assays for hCGβ that are intended for first trimester screening of Down’s syndrome need to be insensitive to interferences by an approximately 100-fold excess of hCG and tuned to measure fairly high concentrations. They are therefore of limited utility for the diagnosis of nontrophoblastic cancers.
Elevated plasma concentrations of hCGβ are reflected by high levels of hCGβcf in urine [63], and specific assay of this form has been used for diagnosis of nontrophoblastic cancer [62, 64, 65] and for the characterization of the First IRR for hCGβcf [9]. However, commercial assays are not available presently.
Candidate epitopes for measurement of hCG and hCGβ
Assays specifically recognizing hCG, hCGβ, and related variants can be constructed using a combination of two pan hCGβ mAbs with identical specificity profiles [66], i.e., with one partner directed against epitopes β2 or β4 (the hCGβloops 1 and 3 domain) combined with a mAb-recognizing epitope β1 (the hCGβ cystine knot domain; Table 6).
In the two ISOBM TD-7 WSs, 50 of 96 Abs were shown to recognize hCG + hCGβ and 23 of these did not recognize hLH. Theoretically, any of the five mAbs directed against epitope β1 around the cystine knot could be combined with any of the 18 mAbs against epitopes β2 or β4 on loops 1 + 3 for construction of multifunctional assays. Epitopes β1 and β2/4 are shared by all important hCG and hCGβ protein backbone variants and glycosylation isoforms including hCG-h and hCGβ-h [17, 18]. MAbs against these two discrete epitopes are highly specific for hCG with <0.1 and <1 % cross-reaction for hLH for epitopes β1 and β2/4, respectively. No other epitope combination provided assays with equally wide and identical recognition of hCG and hCGβ variants and high specificity versus hLH.
While Abs recognizing these epitopes provide desirable specificity, variable affinity for hCG variants (Fig. 5) may cause nonequimolar recognition of hCG and hCG variants in different methods [6]. Although assay specificity can be predicted on the basis of mAb specificity profiles and epitope recognition [66], ultimate performance can only be evaluated with the final assay. An additional source of method variability in hCG measurement that cannot be fully predicted is that of Ab synergy, which may vary between different Ab pairs [67].
Alternative epitopes for measurement of hCG and/or hCGβ and variants
Few manufacturers provide information about the epitope specificities of Abs used in their assays, but due to variable recognition of the First IRR preparations for hCG and variants, it is obvious that different epitope combinations are used in the major commercial assays [6]. In addition to the epitope combination β1–β2/4, other combinations are possible for the construction of assays for hCG and variants, e.g., epitopes β8,1 and β2, β1–α5, α4–β2, etc., but none of them will fulfill all three above-mentioned criteria. However, the frequently used β8 and β2 combination does not pose problems as long as serum specimen are measured that do not contain hCGβcf, truncated hCG or truncated hCGβ, or clipped hCGβCTP.
For selective measurement of hCG or free hCGβ or hCGβcf certain epitope combinations can be suggested: for hCG (no recognition of hLH or noncombined subunits), a mAb against epitope c2 or c3 can be combined with one against β2/4 (Table 6). Alternatively, β1–α3 combinations [66] or β2/4 combined with a tracer mAb against an α epitope are possible [68]. These designs eliminate cross-reactions with free subunits but are sensitive to interferences by free subunits and hCGβcf. For measurement of free hCGβ, a mAb to epitope β7 or β14, and for hCGβcf, a mAb to epitope β11 can be combined with one to epitope β2/4. For hCGα, combinations of mAbs against epitopes α6 and α5 are recommended [69] (Table 6).
A unique mAb coded B152 is used for the measurement of hCG-h that carries a core-2 glycan on Ser132 located on hCGβCTP [20]. However, the clinical utility of assays using this mAb remains to be established [70].
Future perspectives: harmonization of hCG and/or hCGβ and variant measurement
Considerable reduction in between-method and between-laboratory variability in results can be achieved by a number of measures: (1) the establishment and usage of a clear nomenclature of hCG and its variants [1, 8]; (2) endorsement of that nomenclature to define what hCG-assay measure [1, 8, 60]; (3) characterization of diagnostic assays with the new six First IRRs calibrated in SI units that were adopted by WHO for immunoassay standardization [6]; (4) standardization of methods with the highly pure new WHO Fifth IS for hCG encoded 07/36,4 which is identical to the First IRR for hCG 99/688; (5) harmonization of mAb epitopes used in diagnostic methods for hCG, hCGβ, and their variants; and (6) the establishment of reference methods for the various forms of hCG [8], which will be supported by the detailed knowledge on Ab epitope recognition reported in the present study.
References
Berger P, Sturgeon C, Bidart JM, Paus E, Gerth R, Niang M, et al. The isobm td-7 workshop on hCG and related molecules. Towards user-oriented standardization of pregnancy and tumor diagnosis:assignment of epitopes to the three-dimensional structure of diagnostically and commercially relevant monoclonal antibodies directed against human chorionic gonadotropin and derivatives. Tumor Biol. 2002;23:1–38.
Stenman UH, Alfthan H, Hotakainen K. Human chorionic gonadotropin in cancer. Clin Biochem. 2004;37:549–61.
Lapthorn AJ, Harris DC, Littlejohn A, Lustbader JW, Canfield RE, Machin KJ, et al. Crystal structure of human chorionic gonadotropin. Nature. 1994;369:455–61.
Talmadge K, Boorstein WR, Fiddes JC. The human genome contains seven genes for the beta-subunit of chorionic gonadotropin but only one gene for the beta-subunit of luteinizing hormone. DNA. 1983;2:281–9.
Fiddes JC, Goodman HM. The cdna for the beta-subunit of human chorionic gonadotropin suggests evolution of a gene by readthrough into the 3′-untranslated region. Nature. 1980;286:684–7.
Sturgeon CM, Berger P, Bidart JM, Birken S, Burns C, Norman RJ, et al. Differences in recognition of the 1st who international reference reagents for hCG-related isoforms by diagnostic immunoassays for human chorionic gonadotropin. Clin Chem. 2009;55:1484–91.
Fiddes JC, Goodman HM. Isolation, cloning and sequence analysis of the cDNA for the alpha-subunit of human chorionic gonadotropin. Nature. 1979;281:351–6.
Stenman UH, Bidart JM, Birken S, Mann K, Nisula B, O’Connor J. Standardization of protein immunoprocedures. Choriogonadotropin (cg). Scand J Clin Lab Invest Suppl. 1993;216:42–78.
Birken S, Berger P, Bidart JM, Weber M, Bristow A, Norman R, et al. Preparation and characterization of new who reference reagents for human chorionic gonadotropin and metabolites. Clin Chem. 2003;49:144–54.
Merz WE, Krause JM, Roig J, Singh V, Berger P. Nonassembled human chorionic gonadotropin subunits and {alpha}{alpha}-homodimers use fast-track processing in the secretory pathway in contrast to {alpha}{beta}-heterodimers. Endocrinology. 2007;148:5831–41.
Valmu L, Alfthan H, Hotakainen K, Birken S, Stenman UH. Site-specific glycan analysis of human chorionic gonadotropin beta-subunit from malignancies and pregnancy by liquid chromatography–electrospray mass spectrometry. Glycobiology. 2006;16:1207–18.
Toll H, Berger P, Hofmann A, Hildebrandt A, Oberacher H, Lenhof HP, et al. Glycosylation patterns of human chorionic gonadotropin revealed by liquid chromatography-mass spectrometry and bioinformatics. Electrophoresis. 2006;27:2734–46.
Wide L, Lee JY, Rasmussen C. A change in the isoforms of human chorionic gonadotropin occurs around the 13th week of gestation. J Clin Endocrinol Metab. 1994;78:1419–23.
Lopata A, Oliva K, Stanton PG, Robertson DM. Analysis of chorionic gonadotrophin secreted by cultured human blastocysts. Mol Hum Reprod. 1997;3:517–21.
Kovalevskaya G, Birken S, Kakuma T, Ozaki N, Sauer M, Lindheim S, et al. Differential expression of human chorionic gonadotropin (hCG) glycosylation isoforms in failing and continuing pregnancies: preliminary characterization of the hyperglycosylated hCG epitope. J Endocrinol. 2002;172:497–506.
Mann K, Schneider N, Hoermann R. Thyrotropic activity of acidic isoelectric variants of human chorionic gonadotropin from trophoblastic tumors. Endocrinology. 1986;118:1558–66.
Berger P, Schwarz S, Spottl G, Wick G, Mann K. Variants of human chorionic gonadotropin from pregnant women and tumor patients recognized by monoclonal antibodies. J Clin Endocrinol Metab. 1993;77:347–51.
Lottersberger C, Hoermann R, Mann K, Schwarz S, Berger P. Tumor- and pregnancy-derived isoforms of human chorionic gonadotropin: biological and diagnostic relevance. Horm Res. 2003;59:125–34.
Elliott MM, Kardana A, Lustbader JW, Cole LA. Carbohydrate and peptide structure of the alpha- and beta-subunits of human chorionic gonadotropin from normal and aberrant pregnancy and choriocarcinoma. Endocrine. 1997;7:15–32.
Birken S. Specific measurement of o-linked core 2 sugar-containing isoforms of hyperglycosylated human chorionic gonadotropin by antibody b152. Tumour Biol. 2005;26:131–41.
Bidart JM, Birken S, Berger P, Krichevsky A. Immunochemical mapping of hCG and hCG-related molecules. Scand J Clin Lab Invest Suppl. 1993;216:118–36.
Berger P, Bidart JM, Delves PS, Dirnhofer S, Hoermann R, Isaacs N, et al. Immunochemical mapping of gonadotropins. Mol Cell Endocrinol. 1996;125:33–43.
Bidart JM, Troalen F, Bohuon CJ, Hennen G, Bellet DH. Immunochemical mapping of a specific domain on human choriogonadotropin using anti-protein and anti-peptide monoclonal antibodies. J Biol Chem. 1987;262:15483–9.
Dirnhofer S, Madersbacher S, Bidart JM, Ten Kortenaar PB, Spottl G, Mann K, et al. The molecular basis for epitopes on the free beta-subunit of human chorionic gonadotrophin (hCG), its carboxyl-terminal peptide and the hCG beta-core fragment. J Endocrinol. 1994;141:153–62.
Kofler R, Kalchschmid E, Berger P, Wick G. Production and characterization of monoclonal antibodies against bovine luteinizing hormone. Immunobiology. 1981;160:196–207.
Kofler R, Berger P, Wick G. Monoclonal antibodies against human chorionic gonadotropin (hCG): I. Production, specificity, and intramolecular binding sites. Am J Reprod Immunol. 1982;2:212–6.
Stahli C, Miggiano V, Stocker J, Staehelin T, Haring P, Takacs B. Distinction of epitopes by monoclonal antibodies. Methods Enzymol. 1983;92:242–53.
Berger P. molecular morphology of placental and pituitary hormones: epitope mapping with monoclonal antibodies. Wien Klin Wochenschr. 1985;97:573–81.
Bidart JM, Bellet DH, Alberici GF, Van Besien F, Bohuon C. The immune response to a synthetic peptide analogous to the 109–145 beta hCG carboxyl-terminus is directed against two major and two minor regions. Mol Immunol. 1987;24:339–45.
Berger P, Panmoung W, Khaschabi D, Mayregger B, Wick G. Antigenic features of human follicle stimulating hormone delineated by monoclonal antibodies and construction of an immunoradiomometric assay. Endocrinology. 1988;123:2351–9.
Berger P, Klieber R, Panmoung W, Madersbacher S, Wolf H, Wick G. Monoclonal antibodies against the free subunits of human chorionic gonadotrophin. J Endocrinol. 1990;125:301–9.
Bidart JM, Ozturk M, Bellet DH, Jolivet M, Gras-Masse H, Troalen F, et al. Identification of epitopes associated with hCG and the beta hCG carboxyl terminus by monoclonal antibodies produced against a synthetic peptide. J Immunol. 1985;134:457–64.
Bristow A, Berger P, Bidart JM, Birken S, Norman R, Stenman UH, et al. Establishment, value assignment, and characterization of new who reference reagents for six molecular forms of human chorionic gonadotropin. Clin Chem. 2005;51:177–82.
Tsai PK, Bruner MW, Irwin JI, Ip CC, Oliver CN, Nelson RW, et al. Origin of the isoelectric heterogeneity of monoclonal immunoglobulin h1b4. Pharm Res. 1993;10:1580–6.
Perkins M, Theiler R, Lunte S, Jeschke M. Determination of the origin of charge heterogeneity in a murine monoclonal antibody. Pharm Res. 2000;17:1110–7.
Madersbacher S, Berger P. Antibodies and immunoassays. Methods. 2000;21:41–50.
Dirnhofer S, Lechner O, Madersbacher S, Klieber R, de Leeuw R, Wick G, et al. Free alpha subunit of human chorionic gonadotrophin: Molecular basis of immunologically and biologically active domains. J Endocrinol. 1994;140:145–54.
Schwarz S, Berger P, Wick G. The antigenic surface of human chorionic gonadotropin as mapped by murine monoclonal antibodies. Endocrinology. 1986;118:189–97.
Birken S, Krichevsky A, O’Connor J, Schlatterer J, Cole L, Kardana A, et al. Development and characterization of antibodies to a nicked and hyperglycosylated form of hCG from a choriocarcinoma patient: Generation of antibodies that differentiate between pregnancy hCG and choriocarcinoma hCG. Endocrine. 1999;10:137–44.
Novotny J, Handschumacher M, Haber E, Bruccoleri RE, Carlson WB, Fanning DW, et al. Antigenic determinants in proteins coincide with surface regions accessible to large probes (antibody domains). Proc Natl Acad Sci USA. 1986;83:226–30.
Charlesworth MC, Bergert ER, Morris JC, McCormick DJ, Ryan RJ. The antigenic structure of the human glycoprotein hormone alpha-subunit: Iii Solution- and solid-phase mapping using synthetic peptides. Endocrinology. 1991;128:2907–15.
Jackson AM, Klonisch T, Lapthorn AJ, Berger P, Isaacs NW, Delves PJ, et al. Identification and selective destruction of shared epitopes in human chorionic gonadotropin beta subunit. J Reprod Immunol. 1996;31:21–36.
Charrel-Dennis M, Jackson AM, Lund T, Lapthorn AJ, Berger P, Roitt IM, et al. The major hormone-specific discontinuous epitopes on human chorionic gonadotrophin. J Mol Endocrinol. 2004;32:571–81.
Deshmukh US, Pal R, Talwar GP, Gupta SK. Antibody response against epitopes on hCG mapped by monoclonal antibodies in women immunized with an anti-hCG vaccine and its implications for bioneutralization. J Reprod Immunol. 1993;25:103–17.
Novotny J, Bruccoleri R, Newell J, Murphy D, Haber E, Karplus M. Molecular anatomy of the antibody binding site. J Biol Chem. 1983;258:14433–7.
Atassi MZ, Smith JA. A proposal for the nomenclature of antigenic sites in peptides and proteins. Immunochemistry. 1978;15:609–10.
Wu H, Lustbader JW, Liu Y, Canfield RE, Hendrickson WA. Structure of human chorionic gonadotropin at 2.6 a resolution from mad analysis of the selenomethionyl protein. Structure. 1994;2:545–58.
Schwarz S, Krude H, Klieber R, Dirnhofer S, Lottersberger C, Merz WE, et al. Number and topography of epitopes of human chorionic gonadotropin (hCG) are shared by desialylated and deglycosylated hCG. Mol Cell Endocrinol. 1991;80:33–40.
Birken S, Kovalevskaya G, O’Connor J. Metabolism of hCG and hLH to multiple urinary forms. Mol Cell Endocrinol. 1996;125:121–31.
Kovalevskaya G, Birken S, Kakuma T, O’Connor JF. Early pregnancy human chorionic gonadotropin (hCG) isoforms measured by an immunometric assay for choriocarcinoma-like hCG. J Endocrinol. 1999;161:99–106.
Birken S, Yershova O, Myers RV, Bernard MP, Moyle W. Analysis of human choriogonadotropin core 2 o-glycan isoforms. Mol Cell Endocrinol. 2003;204:21–30.
Cole LA, Sutton JM, Higgins TN, Cembrowski GS. Between-method variation in human chorionic gonadotropin test results. Clin Chem. 2004;50:874–82.
Birken S, Armstrong EG, Kolks MA, Cole LA, Agosto GM, Krichevsky A, et al. Structure of the human chorionic gonadotropin beta-subunit fragment from pregnancy urine. Endocrinology. 1988;123:572–83.
Berger P, Sturgeon C. Human chorionic gonadotropin isoforms and their epitopes: diagnostic utility in pregnancy and cancer. Expert OpinMedDiagn. 2008;2:1347–64.
Dirnhofer S, Klieber R, De Leeuw R, Bidart JM, Merz WE, Wick G, et al. Functional and immunological relevance of the COOH-terminal extension of human chorionic gonadotropin beta: implications for the who birth control vaccine. FASEB J. 1993;7:1381–5.
Hoermann R, Berger P, Spoettl G, Gillesberger F, Kardana A, Cole LA, et al. Immunological recognition and clinical significance of nicked human chorionic gonadotropin in testicular cancer. Clin Chem. 1994;40:2306–12.
Grenache DG, Greene DN, Dighe AS, Fantz CR, Hoefner D, McCudden C, et al. Falsely decreased human chorionic gonadotropin (hCG) results due to increased concentrations of the free beta subunit and the beta core fragment in quantitative hCG assays. Clin Chem. 2010;56:1839–44.
Madersbacher S, Berger P, Mann K, Kuzmists R, Wick G. Diagnostic value of free subunits of serum chorionic gonadotropin in testicular cancer. Lancet. 1990;336:630–1.
Lempiainen A, Stenman UH, Blomqvist C, Hotakainen K. Free beta-subunit of human chorionic gonadotropin in serum is a diagnostically sensitive marker of seminomatous testicular cancer. Clin Chem. 2008;54:1840–3.
Stenman UH, Tiitinen A, Alfthan H, Valmu L. The classification, functions and clinical use of different isoforms of hCG. Hum Reprod Update. 2006;12:769–84.
Marcillac I, Troalen F, Bidart JM, Ghillani P, Ribrag V, Escudier B, et al. Free human chorionic gonadotropin beta subunit in gonadal and nongonadal neoplasms. Cancer Res. 1992;52:3901–7.
Alfthan H, Haglund C, Roberts P, Stenman UH. Elevation of free beta subunit of human choriogonadotropin and core beta fragment of human choriogonadotropin in the serum and urine of patients with malignant pancreatic and biliary disease. Cancer Res. 1992;52:4628–33.
Alfthan H, Stenman UH. Pregnancy serum contains the beta-core fragment of human choriogonadotropin. J Clin Endocrinol Metab. 1990;70:783–7.
Norman RJ, Buck RH, Aktar B, Mayet N, Moodley J. Detection of a small molecular species of human chorionic gonadotropin in the urine of patients with carcinoma of the cervix and cervical intraepithelial neoplasia: comparison with other assays for human chorionic gonadotropin and its fragments. Gynecol Oncol. 1990;37:254–9.
O’Connor JF, Schlatterer JP, Birken S, Krichevsky A, Armstrong EG, McMahon D, et al. Development of highly sensitive immunoassays to measure human chorionic gonadotropin, its beta-subunit, and beta core fragment in the urine: application to malignancies. Cancer Res. 1988;48:1361–6.
Schwarz S, Berger P, Wick G. Epitope-selective, monoclonal-antibody-based immunoradiometric assays of predictable specificity for differential measurement of choriogonadotropin and its subunits. Clin Chem. 1985;31:1322–8.
Klonisch T, Panayotou G, Edwards P, Jackson AM, Berger P, Delves PJ, et al. Enhancement in antigen binding by a combination of synergy and antibody capture. Immunology. 1996;89:165–71.
Stenman UH, Alfthan H, Ranta T, Vartiainen E, Jalkanen J, Seppala M. Serum levels of human chorionic gonadotropin in nonpregnant women and men are modulated by gonadotropin-releasing hormone and sex steroids. J Clin Endocrinol Metab. 1987;64:730–6.
Madersbacher S, Klieber R, Mann K, Marth C, Tabarelli M, Wick G, et al. Free alpha-subunit, free beta-subunit of human chorionic gonadotropin (hCG), and intact hCG in sera of healthy individuals and testicular cancer patients. Clin Chem. 1992;38:370–6.
Lempiainen A, Hotakainen K, Blomqvist C, Alfthan H, Stenman UH. Hyperglycosylated human chorionic gonadotropin in serum of testicular cancer patients. Clin Chem. 2012;58:1123–9.
Klonisch T, Delves PJ, Berger P, Panayotou G, Lapthorn AJ, Isaacs NW, et al. Relative location of epitopes involved in synergistic antibody binding using human chorionic gonadotropin as a model. Eur J Immunol. 1996;26:1897–905.
Charrel-Dennis M, Terrazzini N, McBride JD, Kaye P, Martensen PM, Justesen J, et al. The human chorionic gonadotropin-beta arginine68 to glutamic acid substitution fixes the conformation of the c-terminal peptide. Mol Endocrinol. 2005;19:1803–11.
Acknowledgments
We would like to thank the International Society of Oncology and BioMarkers for organizing the Tissue Differentiation (TD)-7 Workshop on hCG and Related Molecules, the International Federation of Clinical Chemistry (IFCC) Working Group for Standardization of hCG, the National Institute for Biological Standards and Control (NIBSC) (South Mimms, UK), and the National Hormone & Peptide Program (NHPP), USA, for the supply of hormones and hormone variants. We would like to thank the following at Abbott Laboratories for their valued contributions: B.L. Dowell, S.Y. Tetin, and J.R. Fishpaugh. The NHS National Services Scotland, National Services Division, supported the work of Wilson Stewart. The Austrian Science Fund (National Research Network, S9307B05) in part supported the work of Peter Berger.
Author information
Authors and Affiliations
Consortia
Corresponding author
Additional information
The International Society of Oncology and Biomarkers Tissue Differentiation (TD)-7 Workshop on hCG and Related Molecules.
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(PDF 20194 kb)
Appendix
Appendix
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution License which permits any use, distribution, and reproduction in any medium, provided the original author(s) and the source are credited.
About this article
Cite this article
Berger, P., Paus, E., Hemken, P.M. et al. Candidate epitopes for measurement of hCG and related molecules: the second ISOBM TD-7 workshop. Tumor Biol. 34, 4033–4057 (2013). https://doi.org/10.1007/s13277-013-0994-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13277-013-0994-6