Abstract
Background
Staphylococcus aureus is a major human pathogen, that can lead to various community- and hospital-acquired infections. RinA is a transcription activator of S. aureus phage φ 11 involved in phage packaging and virulence gene transfer. However, little is known about the molecular mechanism of RinA in the regulation of virulence.
Objective
We aimed to explore a novel contribution of RinA in the regulation of virulence and provide a new drug target in the treatment of S. aureus infections.
Methods
The specific functions of RinA in S. aureus were analyzed by the methods of growth curve, real-time quantitative PCR (RT-qPCR), subcellular localization, electrophoretic mobility shift assay (EMSA), infection model of Galleria mellonella larvae and the mouse subcutaneous abscess model.
Results
In this study, we demonstrated that RinA is a protein evenly distributed in the cytoplasm of S. aureus, and its deletion could cause the growth defects. RT-qPCR and EMSA determined that rinA could negatively regulate the expression of sarA by directly binding to its promoter, and vice versa. The Galleria mellonella larvae infection and mouse subcutaneous abscess models revealed that the rinA mutant strain exhibited obvious virulence defects. When sarA is knocked out, the virulence of S.aureus had no significantly changes whether rinA is knocked out or not.
Conclusion
Our fndings demonstrated that phage transcription activator RinA regulates S. aureus virulence by governing sarA expression.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
S. aureus is a major human pathogen that causes multiple diseases such as skin infections, bacteremia, pneumonia, endocarditis, and toxic shock (Lee et al. 2018). Furthermore, owing to their adaptability to different environmental conditions, hospital- and community-acquired infections are surging (Dayan et al. 2016). The main treatment for S. aureus relies heavily on the use of antibiotics. However, the emergence of drug-resistant strain brought a more serious burden for the treatment of S. aureus infections (Lakhundi and Zhang 2018). S. aureus can produce an arsenal of virulence factors, such as adhesion proteins, superantigens, pore-forming toxins, ADP-ribosylating toxins and proteases. Moreover, multiple complex virulence regulatory systems including the quorum-sensing system, two-component system and the SarA protein family, have evolved. These virulence factors and virulence regulatory systems together contribute to the high pathogenicity of S. aureus (Bronner et al. 2004; Jenul and Horswill 2019).
The SarA protein family plays a crucial role in regulating the virulence of S. aureus (Cheung et al. 2008). The staphylococcal accessory regulator locus (sarA) encodes a DNA-binding protein that regulates the expression of more than 100 genes (Sterba et al. 2003). SarA can indirectly regulate the expression of virulence genes, including α-hemolysin-, protein A-, and fibronectin-binding proteins in an agr-dependent manner, furthermore, it can directly bind to the promoter regions of virulence genes to up-regulate or down-regulate their expressions (Chien Y et al. 1999; Pan and Simon 1998; Yueh-tyng Chien and Cheung 1998). Additionally, the mutation of sarA could decrease biofilm formation and prevent the accumulation of extracellular toxins (Zielinska et al. 2012).
The RinA family contains two proteins, RinA and RinB, RinA is basic DNA-binding protein that directly activates gene expression, while RinB is acidic and does not appear to regulate gene expression, however, RinB may play a minor role in activation such as providing stability to RinA (Ferrer et al. 2011; Ye and Lee 1993). The late operons of many phages control the morphogenetic and lysis genes of S. aureus, previous studies have shown that the RinA family of phage-encoded proteins are activators required for the transcription of late operon in a crowd of temperate staphylococcal phages (Ferrer et al. 2011; Quiles-Puchalt et al. 2013). To control the expression of the late operon, the RinA protein binds to a DNA repeat region situated upstream of the terminase small subunit gene (terS) (Quiles-Puchalt et al. 2014). Moreover, the deletion of rinA abolishes the formation of functional phage particles and significantly reduces the transfer of phage and virulence factors encoded by the pathogenicity islands (Ferrer et al. 2011). The control of phage-mediated transfer of virulence factor is a conserved mechanism that regulates horizontal gene transfer (HGT), which significantly impacts the evolution of bacterial pathogens between microorganisms (Olszak et al. 2017). Phages are one of the types of S.aureus Mobile genetic elements (MGEs) which can impact expression of virulence determinants by either positive or negative lysogenic conversion. (Malachowa and DeLeo 2010). In horizontal gene transfer between bacteria, phages play an essential role by acting as gene transfer vectors and carrying virulence factors, these two functions can be observed in S. aureus phages (McCarthy et al. 2012). Although RinA acts upon the transfer of virulence factors, its specific impact on the S. aureus pathogenicity is unknown. Moreover, whether the regulation of RinA in S. aureus virulence depends on sarA expression also needs to be uncovered.
Herein, it was demonstrated that RinA could negatively regulate the expression of sarA by directly binding to its promoter region, and vice versa. Additionally, the contribution of RinA to the pathogenicity of S. aureus was further confirmed by using the infective G. mellonella larvae and the mouse subcutaneous abscess model. In summary, the results revealed that RinA positively regulates the virulence of S. aureus in a SarA dependent manner. These insights on the underlying molecular mechanisms can guide the development of novel anti-infective strategies.
Materials and methods
Bacterial strains, plasmids, and growth conditions
The bacterial strains and plasmids used in this study are listed in Table 1. Escherichia coli was grown (220 rpm) in a lysogeny broth (LB) medium or on lysogeny broth agar (LA), while S. aureus was grown (220 rpm) in a tryptic soy broth (TSB) medium or on tryptic soy agar (TSA) at 37℃. The appropriate antibiotics were used for plasmid selection and maintenance. They were added to the bacterial cultures at the following concentrations: 150 μg/mL ampicillin sodium salt or 50 μg/mL kanamycin sulfate for E. coli or 15 μg/mL chloromycetin for S. aureus.
Construction of S. aureus mutant strains
Plasmid pBTs were utilized to construct the rinA mutant strain. The upstream and downstream regions of rinA from the genomic DNA of S. aureus NCTC8325 were amplified with primers rinA-LB-F/R and rinA-RB-F/R. Next, the products were connected by overlap extension PCR. The resulting fragments were digested with KpnI and SalI and then cloned into pBTs. Afterward, the resulting plasmid pBTsΔrinA was electroporated into S. aureus strain RN4220 for modification and then transformed into S. aureus strain NCTC8325. An allelic replacement mutant was selected using a previously described method (Bae and Schneewind 2006) and was further confirmed by PCR and sequencing. The sarA mutant strains and sarA rinA double mutant strains were constructed in the same way. The sarA mutant and rinA sarA double mutant strains were constructed using a similar strategy by introducing the plasmid pBTsΔsarA into the wild-type (WT) and rinA mutant strains, respectively. All primers used in this study are listed in Table 2.
Construction of the complementation strain
To construct the rinA chromosomal complementation, the DNA fragment of the full-length rinA ORF and the flank upstream and downstream regions were amplified from the genomic DNA of S. aureus NCTC8325 with primers C-rinA-LB-F/R and C-rinA-RB-F/R. Next, the resulting fragments were digested with restriction enzymes and then cloned into pBTs. The resulting plasmid pBTs-LB-rinA-RB was first electroporated into S. aureus strain RN4220 for modification, and then transformed into the rinA mutant strain. The allelic replacement complementation strain was selected using the same method described above and was further confifirmed by PCR and sequencing.
Total RNA isolation, cDNA generation, and real-time quantitative reverse transcription-PCR
Overnight cultures of S. aureus were diluted to an OD600 of 0.05 in TSB, grown to the indicated cell density and collected. Afterward, the collected cells were processed with RNAiso plus (TaKaRa) and lysed with 0.1 mm-diameter-silica beads in the FastPrep-24 system (MP Biomedicals) to extract the total RNA. The PrimeScript 1st Strand cDNA synthesis kit (TaKaRa) was employed to synthesize cDNA. Then, RT-qPCR was performed with SYBR Ex Taq premix (TaKaRa) using the StepOne real-time PCR system (Applied Biosystems) to quantify the relative gene expression. pta was used as the internal reference gene to normalize the gene expression (Wang and Sun 2021).
Growth curves
Overnight cultures of S. aureus were diluted to an OD600 of 0.02 in TSB, added into a 96-well plate, and grown at 37℃ with shaking (220 rpm). The optical density was measured at 600 nm every hour using a microplate reader (Elx800; Bio-Tek). When the bacteria reached the stationary phase (about 12 h), the measurement was stopped.
Subcellular localization of RinA
To detect the cellular localization of RinA, the DNA fragment of the full-length rinA ORF was amplified using PCR, and RinA was fused with eGFP at its C-terminal. The fusion gene was controlled by S. aureus S10 promoter. The RinA::eGFP fluorescent expression strain was confirmed by PCR and sequencing. An inverted confocal laser scanning microscope (FV1000, Olympus) was utilized for observation. An argon ion laser (Ex = 488 nm, Em = 515–530 nm) was used to observe fluorescent signals of eGFP. Finally, the confocal images were captured using the FV10-ASW 4.2 Viewer software (Olympus).
Expression and purification of RinA
The 6-His-tagged RinA was expressed and purified using standard procedures. The DNA fragment of the full-length rinA ORF was amplified from the genomic DNA of S. aureus NCTC8325 using primers PrinA-F/PrinA-R, cloned into the expression vector pET28a to obtain the plasmid pET28a-rinA, and transformed into E. coli BL21 (DE3). The transformant was grown in LB medium at 37 °C to an OD600 of 0.6–0.8 and induced with 0.5 mM isopropyl-β-D-1-thiogalactopyr-anoside (IPTG) at 16 °C for 12 h. Next, the cells were harvested, resuspended in a protein buffer (50 mM Tris–HCl, 300 mM NaCl, pH 8.0) and lysed by sonication on ice. An Ni–NTA agarose solution (Novagen) was used to purify the protein with His tag at the C-terminus. The bound protein was eluted with an elution buffer (50 mM Tris–HCl, 300 mM NaCl, 300 mM imidazole, pH 8.0). The imidazole in the eluate was removed using a protein buffer, and the purified proteins were stored at − 80 °C until use. SDS-PAGE analyzed the purity of the protein and the protein concentration was determined using the bicinchoninic acid (BCA) assay with bovine serum albumin as the standard.
Electrophoretic mobility shift assay
The FAM-labeled DNA fragment containing the promoter region of the target gene was amplified from the genomic DNA of S. aureus strain NCTC8325. The unlabeled fragment of the promoter was added as specific competitors, while that of the pta ORF was added as non-specific competitors. The FAM-labeled probe, different concentrations of proteins, and the protein buffer (50 mM Tris–HCl, 300 mM NaCl, pH 8.0) were mixed in 10 μL and incubated for 30 min at room temperature. After incubation, the mixtures were electrophoresed in a 5% native polyacrylamide gel in 1 × Tris–borate-EDTA (TBE) buffer. Finally, the band shifts were observed under the multifunctional laser imager.
Infection model of G. mellonella larvae
Overnight cultures of S. aureus were diluted to an OD600 of 0.05 in TSB, then the late-logarithmic phase (OD600 = 6) bacterial solution was collected and diluted with PBS. Next, 20 μL was injected into the middle of the second gastropod (107 CFU/mL). Larvae injected with 20 μL PBS were used as the control group. The larvae were incubated at 37℃ in the dark and observed at 24 h intervals for a week. The larvae were considered dead when they repeatedly failed to respond to physical stimuli. The primary outcome for the infection model of G. mellonella larvae was the rapidity and extent of mortality of the larvae assessed by the log-rank test.
Mouse model of subcutaneous abscess
Outbred, immunocompetent female BALB/c mice between 5 and 6 weeks old were purchased from Gem Pharmatech Technology Company and raised to 6–8 weeks old in a specific sterile laboratory. The hair on both sides of each mouse's back was shaved off with a razor. The overnight cultures of S. aureus were diluted to an OD600 of 0.05 in TSB. The late-logarithmic phase (OD600 = 6) bacterial solution was then collected, washed twice, and diluted in sterile PBS. Afterward, the mice were anesthetized with 1% sodium pentobarbital and inoculated using subcutaneous injections in the bilateral back flanks with 5 × 107 S. aureus cells in 50 μL PBS. The areas of abscess formation and skin lesions were monitored at 24 h intervals for 7 days and the skin lesions’ sizes were calculated using the maximal length × width. Each skin lesion (7 days after infection) from the mouse was excised and ground with a grinding rod in 1 mL PBS in iron gauze until it was homogenized. The number of CFU recovered from each lesion was then counted by serial dilution and plated onto LA plates. Next, the skin lesions were placed in 4% PFA Fix Solution for histopathological analyses. Paraffin embedding and hematoxylin and eosin (H&E) staining were performed by the Wuhan Servicebio Technology.
Ethics statement
The use and care of mice in the present study strictly followed the guidelines adopted by the Ministry of Health of the People's Republic of China in June 2004. The protocol was approved by the Institutional Animal Care and Use Committee of the University of Science and Technology of China (USTCACUC182301015).
Statistics
Statistical analyses were performed with Student’s t test or Mantel-Cox test. All error bars depict the standard errors of the means. Differences with a p value of 0.05 or less were considered significant. Significance was defined as *p < 0.05; **p < 0.01; ***p < 0.001; ****p < 0.0001. All experiments were performed in triplicate. For insect survival analysis, Kaplan–Meier survival curve was generated and analyzed for statistical significance with GraphPad Prism 7.0. Statistical details for each experiment can be found in the Figure Legends.
Results
Construction of rinA mutant
To explore the functions of RinA, the rinA disruption mutant was constructed by knocking out rinA from the genomic DNA of S. aureus NCTC8325 and the genome complementation strain was constructed by complementing rinA into the rinA disruption mutant (Figs. 1A, B). Moreover, the transcription levels of rinA during the different growth phases in the wild-type (WT) were detected by RT-qPCR. The results demonstrated that the transcription level of rinA was highest in the late-logarithmic phase and lower in the early-logarithmic and mid-logarithmic phase (Fig. 1C).
Construction of rinA mutant. A Diagram of rinA knockout strategy. B Evaluation of PCR products by agarose gel electrophoresis, from left to right were: 1 kb DNA Maker, wild-type, rinA mutant strain, rinA complementation strain. C The transcription level of rinA in different growth phases (OD600 = 0.5, 2, 6), the internal reference gene is pta
Disruption of rinA renders growth defect of S. aureus
To determine the effects of RinA on the growth of S. aureus, the growth curve of the wild-type, rinA mutant and rinA complementation strains were measured. The results revealed that after 3 h, the growth rate of the rinA mutant strain was significantly weaker than the wild-type and rinA complementation strains (Fig. 2A). Since, rsgA, ccpA, relA, and srrA have been reported to affect the growth of S. aureus (Corrigan et al. 2016; Gentry et al. 2000; Pragman et al. 2007; Seidl et al. 2006), the transcription levels of these genes in the wild-type and rinA mutant strains were investigated. The results revealed that the transcription level of ccpA in the rinA mutant was significantly reduced compared to the wild-type (Fig. 2B). Taken together, the data suggested that RinA may be involved in the positive regulation of S. aureus growth in a ccpA dependent manner.
RinA deficiency slows down the growth rate of S. aureus. A The growth curve of the wild-type, rinA mutant and rinA complementation strains. The results were obtained from three independent experiments performed. Data is presented as the mean ± SD. Two-tailed Student’s t-test was used for the comparison of statistical significance. The wild-type strain was used as the reference. *p < 0.05; **p < 0.01; ***p < 0.001. B The transcription levels of genes related to growth in the wild-type and rinA mutant were evaluated through RT-qPCR. The means and standard deviations were then calculated. Data is presented as the mean ± SD. Two-tailed Student’s t test is used for the comparison of statistical significance. The wild-type strain was used as the reference. *p < 0.05
Localization of RinA in S. aureus
The secondary structures of the RinA sequences were predicted using the PSIPRED online software (Buchan and Jones 2019) and the amino acid sequence analysis suggested that the secondary structures were mainly in the form of helixes (Fig. 3A). It has been reported that the localization of a protein in cells is closely related to its function. Therefore, the change of the subcellular location of a protein will affect its biological functions (Pishchany et al. 2009). To observe the localization of RinA, a RinA::eGFP fluorescent expression strain was constructed. Then, the growth curve was measured and the result showed that RinA::eGFP strain exhibited a similar phenotype to wild-type S. aureus, suggesting that fused with eGFP doesn’t impaire the function of RinA (Fig. 3B). The laser confocal microscope revealed that RinA was evenly distributed in the cytoplasm of S. aureus (Fig. 3C).
Localization of RinA in S. aureus. A PSIPRED online software predicted the predicted secondary structures of RinA. B The growth curve of the wild-type and RinA::eGFP fluorescent expression strain. The results were obtained from three independent experiments performed. Data is presented as the mean ± SD. Two-tailed Student’s t test was used for the comparison of statistical significance. The wild-type strain was used as the reference. *p < 0.05; **p < 0.01. C RinA::eGFP localized to the cytoplasm. The scale bar is 2 µm
RinA negatively regulates the expression of sarA by binding directly to the promoter and vice versa.
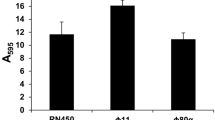
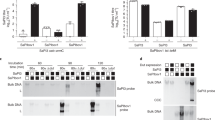
To investigate whether RinA was involved in regulating the virulence of S. aureus, the transcription levels of the virulence factors in the wild-type and rinA mutant strains were detected by RT-qPCR. The results determined that only the transcription level of sarA was significantly increased in the rinA mutant strain among all the genes detected. The transcription levels of other virulence factors such as RNAIII, saeR, sigB, agrA, agrC and hla had no significant changes (Fig. 4A). To that end, EMSA assay was utilized to further explore the regulatory mechanism. RinA was purified in vitro to obtain higher purity proteins (Fig. 4B). EMSA showed that RinA could directly bind to the sarA promoter to retard the migration of DNA probe, and as the protein concentration gradually increased, the phenomenon became more apparent. This binding could be disrupted using approximately 100-fold unlabeled sarA promoter fragment, while a 100-fold excess of unlabeled coding sequence fragment of pta did not have the same effect (Fig. 4C). To verify whether the regulatory function of RinA is affected by SarA, a sarA mutant strain was constructed, and the transcription level of rinA in sarA mutant was detected by RT-qPCR. The results revealed that the transcription level of rinA in the sarA mutant strain increased significantly compared to the wild-type (Fig. 4D). An EMSA assay was performed to establish whether SarA could regulate RinA by binding directly to the promoter. The SarA was purified in vitro to obtain higher purity proteins (Fig. 4E) and EMSA assay determined that sarA could directly bind to the rinA promoter to retard the migration of DNA probe (Fig. 4F). Taken together, these results indicated that rinA negatively regulates the expression of sarA by binding directly to the promoter and vice versa.
RinA negatively regulates the expression of sarA by binding directly to the promoter and vice versa. A The transcription levels of virulence genes in the wild-type and rinA mutant strains. The means and standard deviations were then calculated. Data is presented as the mean ± SD. Two-tailed Student’s t test is used for the comparison of statistical significance. The wild-type strain was used as the reference. **p < 0.01. B SDS-PAGE analysed of the purity RinA protein. C EMSA assay of the purified RinA with the FAM-labelled DNA fragments of sarA. Increasing concentrations of purified RinA and 1 fmol the FAM-labelled sarA probe were used in the reactions. The unlabelled probes were added as a specific competitor, and the unlabelled fragment of pta ORF region was added as a nonspecific competitor. D The transcription level of rinA in the late-logarithmic phase in the wild-type and sarA mutant strains. The means and standard deviations were then calculated. Data is presented as the mean ± SD. Two-tailed Student’s t test is used for the comparison of statistical significance. The wild-type strain was used as the reference. **p < 0.01. E SDS-PAGE analysed of the purity SarA protein. F EMSA assay of the purified SarA with the FAM-labelled DNA fragments of rinA. Increasing concentrations of purified SarA and 1 fmol the FAM-labelled rinA probe were used in the reactions. The unlabelled probes were added as a specific competitor, and the unlabelled fragment of pta ORF region was added as a nonspecific competitor
Knockout of rinA reduces the pathogenicity of S. aureus
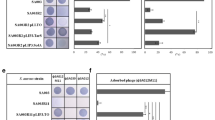
We respectively selected the G. mellonella larvae infection and the mouse subcutaneous abscess models to verify the effects of RinA on the pathogenicity of S. aureus. The wild-type, rinA mutant and rinA complementation strains were inserted into the larvae, and the survival of the G. mellonella larvae was recorded for 7 consecutive days after the injection. The results demonstrated that compared with the rinA mutant strain-injected larvae, the fatality rate of G. mellonella larvae injected with the wild-type and complementation strains were significantly higher (Fig. 5). Next, the mouse subcutaneous abscess model was employed to test the pathogenicity, and the results revealed that compared with the wild-type and complementation strains, the subcutaneous abscess area of mice inoculated with the rinA mutant strain was significantly reduced (Figs. 6A, B). Then, we examined bacterial colonization of the skin lesions, and the analysis found that the number of bacterial colonization in the abscessed skins of mice inoculated with the rinA mutant strain was significantly lower than the wild-type (Fig. 6C). Furthermore, the histological examination of the wild-type and complementation strains exhibited more severe tissue destruction and inflammatory responses in the abscesses (Fig. 6D). Overall, these findings demonstrate that RinA plays a vital role in S. aureus pathogenicity.
Knockout of rinA reduces the pathogenicity of G.mellonella larvae. 1 × 107 CFU of the wild-type, rinA mutant and complementation strains were injected into G. mellonella larvae, PBS were used as the control group, and the survival of the G. mellonella larvae was recorded for 7 consecutive days after the injection. Data is presented as the mean ± standard deviations. Log-rank test is used for the comparison of statistical significance. The wild-type strain was used as the reference. ****p < 0.0001
Knockout of rinA reduces the pathogenicity of S. aureus in the mouse subcutaneous abscess model. The mice were treated with 50 µL of PBS containing 5 × 107 CFU of the wild-type, rinA mutant, and complementation strains, PBS were used as the control group. The treatment was administered in both flanks of the back by subcutaneous injection (n = 6–8). The abscess areas were measured daily using a caliper. A The photographic images of representative abscesses in mice 7 days after infection. B Abscess formation and skin lesion areas were monitored at 24 h intervals for 7 days. The sizes of the skin lesions were calculated by the maximal length × width. The error bars indicate the standard errors of the means, obtained from three biological replicates. Data is presented as the mean ± standard deviation. Two-tailed Student’s t test is used for the comparison of statistical significance. ***p < 0.001. C The number of CFU recovered at 7 days after infection was determined. ***p < 0.001. D Representative images of histological analysis (H&E staining) for samples from (A). The scale bar is 50 µm
RinA regulates S.aureus virulence by governing sarA expression.
We selected the G. mellonella larvae infection to explain whether the mechanism of RinA regulates S. aureus virulence is related to SarA. The sarA mutant and sarA rinA double mutant strains were inserted into the larvae, and the survival of the G. mellonella larvae was recorded for 7 consecutive days after the injection. The results demonstrated that no significant difference was observed in the fatality rate of the sarA mutant and sarA rinA double mutant strains (Fig. 7), that means When sarA is knocked out, the virulence of S.aureus had no significantly changes whether rinA is knocked out or not. These results indicated that RinA regulates S.aureus virulence by governing sarA expression.
RinA regulates S.aureus virulence by governing sarA expression. 1 × 107 CFU of the sarA mutant and sarA rinA double mutant strains were injected into G. mellonella larvae, PBS were used as the control group, and the survival of the G. mellonella larvae was recorded for 7 consecutive days after the injection. Data is presented as the mean ± standard deviations. Log-rank test is used for the comparison of statistical significance. The wild-type strain was used as the reference. NS, not significant (p > 0.05)
Discussion
S. aureus is an opportunistic pathogen, whose colonization in humans can cause skin infections, pneumonia, bacteremia and other diseases (Horino and Hori 2020). Both antibiotic resistance and an absence of a working vaccine complicate the treatment of S. aureus infections (Cheung et al. 2021). Unsurprisingly, S. aureus infections correlate with considerable morbidity and mortality (Rasigade et al. 2014). The reason why S. aureus can cause serious infections is closely related to its production of virulence factors and its virulence regulatory factors. The studies on the global virulence regulators, including the Agr system, SarA family, and Sae two-component system, have shown that the virulence regulation network of S. aureus is highly complex (Bronner et al. 2004; Jenul and Horswill 2019).
Bacteria can acquire virulence genes through horizontal gene transfer, thus facilitating the evolution of virulence in pathogens (Dell'Annunziata et al. 2021; Juhas 2015). RinA is an important member of the RinA family that controls the transfer of virulence genes in Gram-positive bacteria. Deleting rinA resulted in the decrease of phage titers and significantly reduced the transfer of SAPI-encoded virulence factors, which play essential roles in bacterial evolution and pathogenesis (Ferrer et al. 2011). Together, these studies have provided a thorough understanding of RinA in the transfer of virulence genes. However, the exhaustive regulatory mechanisms mediated by RinA in virulence control are still unclear. Herein, a novel contribution of RinA in the regulation of virulence in S. aureus NCTC8325 was investigated.
The subcellular protein localization showed that RinA is evenly distributed in the cytoplasm of S. aureus. Protein subcellular localization is directly related to the disease, and proteins targeting similar subcellular localization tend to participate in mutual protein–protein interactions (PPIs) (Park et al. 2011). PPIs are vital for biological processes and are attractive pathological targets for drug discovery (Rosell and Fernandez-Recio 2018). When proteins are involved in a common biological pathway or process with disease-associated proteins, it is very plausible that they are themselves disease-associated (Barabasi et al. 2011). Hence, to understand the specific impact of RinA on the pathogenic mechanism of S.aureus, the subcellular localization of RinA is crucial.
Bacterial virulence is highly dynamic and context-dependent (Diard and Hardt 2017). The first metabolic dimension associated with virulence was growth. The standard virulence evolution theory assumes that virulence factors are maintained because of increasing bacterial growth within hosts and that growth influences key traits important for in vivo infection (Brown et al. 2012; Kim et al. 2021). Our results determined that the disruption of rinA causes growth defects in S. aureus. This growth defect may also be one of the reasons for the decline of pathogenicity in animal models.
Our study also illustrated how rinA negatively regulates the expression of sarA by binding directly to the promoter and vice versa. Moreover, SarA can indirectly regulate the virulence of S. aureus in an agr-dependent manner to increase the transcription levels of RNAII and RNAIII by binding to the P2 and P3 promoter regions of the agr locus (Cheung et al. 1997). In our experiments, the transcription levels of agr and RNAIII in wild-type and rinA mutant strains were not significantly different, suggesting that RinA might regulate the expression of virulence genes in an agr-independent manner in S. aureus NCTC8325. SarA can directly regulate the virulence of S. aureus by binding to the promoter regions of virulence factors, up-regulating virulence genes such as hla, fnbA and tsst, or down-regulating virulence genes such as sspA, aur and can (Cheung and Zhang 2002; Oscarsson et al. 2006). Since the pathogenicity of S. aureus was significantly reduced when rinA was mutated, it was speculated that the virulence regulation of S. aureus by RinA was achieved by modulating the transcription of sarA to affect the expression of the virulence genes. Interestingly, it was uncovered that RinA and SarA could regulate each other. However, this conjecture needs to be verified by further studies.
This study confirmed the critical role of RinA in S. aureus. It was established that the transcription level of rinA significantly increased in the late-logarithmic phase and that RinA was evenly distributed in the cytoplasm of S. aureus. Furthermore, the knockout of rinA significantly reduced the growth of S. aureus. RT-qPCR and EMSA revealed that rinA could negatively regulate the expression of sarA by directly binding to its promoter region and vice versa. The contribution of RinA to the pathogenicity of S. aureus was confirmed by using the infection model of G. mellonella larvae and the mouse model of subcutaneous abscess. These findings provide new insights into the regulatory mechanisms of virulence gene expression and the pathogenesis of S. aureus.
Data availability
All relevant data are within the manuscript and further information and requests for resources and reagents should be directed to and will be fulfilled by the corresponding author, Yujie Li (lyj2020@ustc.edu.cn).
References
Bae T, Schneewind O (2006) Allelic replacement in Staphylococcus aureus with inducible counter-selection. Plasmid 55:58–63. https://doi.org/10.1016/j.plasmid.2005.05.005
Barabasi AL, Gulbahce N, Loscalzo J (2011) Network medicine: a network-based approach to human disease. Nat Rev Genet 12:56–68. https://doi.org/10.1038/nrg2918
Bronner S, Monteil H, Prevost G (2004) Regulation of virulence determinants in Staphylococcus aureus: complexity and applications. FEMS Microbiol Rev 28:183–200. https://doi.org/10.1016/j.femsre.2003.09.003
Brown SP, Cornforth DM, Mideo N (2012) Evolution of virulence in opportunistic pathogens: generalism, plasticity, and control. Trends Microbiol 20:336–342. https://doi.org/10.1016/j.tim.2012.04.005
Buchan DWA, Jones DT (2019) The PSIPRED Protein Analysis Workbench: 20 years on. Nucleic Acids Res 47:W402–W407. https://doi.org/10.1093/nar/gkz297
Cheung AL, Zhang G (2002) Global regulation of virulence determinants in Staphylococcus aureus by the SarA protein family. Front Biosci 7:1825–1842. https://doi.org/10.2741/A882
Cheung AL, Bayer MG, Heinrichs JH (1997) sar Genetic determinants necessary for transcription of RNAII and RNAIII in the agr locus of Staphylococcus aureus. J Bacteriol 179:3963–3971. https://doi.org/10.1128/jb.179.12.3963-3971.1997
Cheung AL, Nishina KA, Trotonda MP, Tamber S (2008) The SarA protein family of Staphylococcus aureus. Int J Biochem Cell Biol 40:355–361. https://doi.org/10.1016/j.biocel.2007.10.032
Cheung GYC, Bae JS, Otto M (2021) Pathogenicity and virulence of Staphylococcus aureus. Virulence 12:547–569. https://doi.org/10.1080/21505594.2021.1878688
Chien Y-T, ACMaAL, Cheung AL, (1998) SarA level is a determinant of agr activation in Staphylococcus aureus. Mol Microbiol 30:991–1001. https://doi.org/10.1046/j.1365-2958.1998.01126.x
Chien Y, Manna AC, Projan SJ, AL. C, (1999) SarA, a global regulator of virulence determinants in Staphylococcus aureus, binds to a conserved motif essential for sar-dependent gene regulation. J Biol Chem 274:37169–37176. https://doi.org/10.1074/jbc.274.52.37169
Corrigan RM, Bellows LE, Wood A, Grundling A (2016) ppGpp negatively impacts ribosome assembly affecting growth and antimicrobial tolerance in Gram-positive bacteria. Proc Natl Acad Sci U S A 113:E1710-1719. https://doi.org/10.1073/pnas.1522179113
Dayan GH, Mohamed N, Scully IL, Cooper D, Begier E, Eiden J, Jansen KU, Gurtman A, Anderson AS (2016) Staphylococcus aureus: the current state of disease, pathophysiology and strategies for prevention. Expert Rev Vaccines 15:1373–1392. https://doi.org/10.1080/14760584.2016.1179583
Dell’Annunziata F, Folliero V, Giugliano R, De Filippis A, Santarcangelo C, Izzo V, Daglia M, Galdiero M, Arciola CR, Franci G (2021) Gene transfer potential of outer membrane vesicles of gram-negative bacteria. Int J Mol Sci 22:5985. https://doi.org/10.3390/ijms22115985
Diard M, Hardt WD (2017) Evolution of bacterial virulence. FEMS Microbiol Rev 41:679–697. https://doi.org/10.1093/femsre/fux023
Ferrer MD, Quiles-Puchalt N, Harwich MD, Tormo-Mas MA, Campoy S, Barbe J, Lasa I, Novick RP, Christie GE, Penades JR (2011) RinA controls phage-mediated packaging and transfer of virulence genes in Gram-positive bacteria. Nucleic Acids Res 39:5866–5878. https://doi.org/10.1093/nar/gkr158
Gentry D, Li T, Rosenberg M, McDevitt D (2000) The rel gene is essential for in vitro growth of Staphylococcus aureus. J Bacteriol 182:4995–4997. https://doi.org/10.1128/JB.182.17.4995-4997.2000
Horino T, Hori S (2020) Metastatic infection during Staphylococcus aureus bacteremia. J Infect Chemother 26:162–169. https://doi.org/10.1016/j.jiac.2019.10.003
Hu J, Zhang X, Liu X, Chen C, Sun B (2015) Mechanism of reduced vancomycin susceptibility conferred by walK mutation in community-acquired methicillin-resistant Staphylococcus aureus strain MW2. Antimicrob Agents Chemother 59:1352–1355. https://doi.org/10.1128/AAC.04290-14
Jenul C, Horswill AR (2019) Regulation of Staphylococcus aureus Virulence. Microbiol Spectr 6:10. https://doi.org/10.1128/microbiolspec.GPP3-0031-2018
Juhas M (2015) Horizontal gene transfer in human pathogens. Crit Rev Microbiol 41:101–108. https://doi.org/10.3109/1040841X.2013.804031
Kim GL, Hooven TA, Norambuena J, Li B, Boyd JM, Yang JH, Parker D (2021) Growth and stress tolerance comprise independent metabolic strategies critical for Staphylococcus aureus Infection. mbio. https://doi.org/10.1128/mBio.00814-21
Lakhundi S, Zhang K (2018) Methicillin-Resistant Staphylococcus aureus: molecular characterization, evolution, and epidemiology. Clin Microbiol Rev 31:e00020-e18. https://doi.org/10.1128/CMR.00020-18
Lee AS, de Lencastre H, Garau J, Kluytmans J, Malhotra-Kumar S, Peschel A, Harbarth S (2018) Methicillin-resistant Staphylococcus aureus. Nat Rev Dis Primers 4:18033. https://doi.org/10.1038/nrdp.2018.33
Malachowa N, DeLeo FR (2010) Mobile genetic elements of Staphylococcus aureus. Cell Mol Life Sci 67:3057–3071. https://doi.org/10.1007/s00018-010-0389-4
McCarthy AJ, Witney AA, Lindsay JA (2012) Staphylococcus aureus temperate bacteriophage: carriage and horizontal gene transfer is lineage associated. Front Cell Infect Microbiol 2:6. https://doi.org/10.3389/fcimb.2012.00006
Olszak T, Latka A, Roszniowski B, Valvano MA, Drulis-Kawa Z (2017) Phage life cycles behind bacterial biodiversity. Curr Med Chem 24:3987–4001. https://doi.org/10.2174/0929867324666170413100136
Oscarsson J, Tegmark-Wisell K, Arvidson S (2006) Coordinated and differential control of aureolysin (aur) and serine protease (sspA) transcription in Staphylococcus aureus by sarA, rot and agr (RNAIII). Int J Med Microbiol 296:365–380. https://doi.org/10.1016/j.ijmm.2006.02.019
Pan FC, Simon JF (1998) Role of SarA in virulence determinant production and environmental signal transduction in Staphylococcus aureus. J Bacteriol 180:6232–6241. https://doi.org/10.1128/JB.180.23.6232-6241.1998
Park S, Yang JS, Shin YE, Park J, Jang SK, Kim S (2011) Protein localization as a principal feature of the etiology and comorbidity of genetic diseases. Mol Syst Biol 7:494. https://doi.org/10.1038/msb.2011.29
Pishchany G, Dickey SE, Skaar EP (2009) Subcellular localization of the Staphylococcus aureus heme iron transport components IsdA and IsdB. Infect Immun 77:2624–2634. https://doi.org/10.1128/IAI.01531-08
Pragman AA, Herron-Olson L, Case LC, Vetter SM, Henke EE, Kapur V, Schlievert PM (2007) Sequence analysis of the Staphylococcus aureus srrAB loci reveals that truncation of srrA affects growth and virulence factor expression. J Bacteriol 189:7515–7519. https://doi.org/10.1128/JB.00547-07
Quiles-Puchalt N, Tormo-Mas MA, Campoy S, Toledo-Arana A, Monedero V, Lasa I, Novick RP, Christie GE, Penades JR (2013) A super-family of transcriptional activators regulates bacteriophage packaging and lysis in Gram-positive bacteria. Nucleic Acids Res 41:7260–7275. https://doi.org/10.1093/nar/gkt508
Quiles-Puchalt N, Martinez-Rubio R, Ram G, Lasa I, Penades JR (2014) Unravelling bacteriophage varphi11 requirements for packaging and transfer of mobile genetic elements in Staphylococcus aureus. Mol Microbiol 91:423–437. https://doi.org/10.1111/mmi.12445
Rasigade JP, Dumitrescu O, Lina G (2014) New epidemiology of Staphylococcus aureus infections. Clin Microbiol Infect 20:587–588. https://doi.org/10.1111/1469-0691.12718
Rosell M, Fernandez-Recio J (2018) Hot-spot analysis for drug discovery targeting protein-protein interactions. Expert Opin Drug Discov 13:327–338. https://doi.org/10.1080/17460441.2018.1430763
Seidl K, Stucki M, Ruegg M, Goerke C, Wolz C, Harris L, Berger-Bachi B, Bischoff M (2006) Staphylococcus aureus CcpA affects virulence determinant production and antibiotic resistance. Antimicrob Agents Chemother 50:1183–1194. https://doi.org/10.1128/AAC.50.4.1183-1194.2006
Sterba KM, Mackintosh SG, Blevins JS, Hurlburt BK, Smeltzer MS (2003) Characterization of Staphylococcus aureus SarA binding sites. J Bacteriol 185:4410–4417. https://doi.org/10.1128/JB.185.15.4410-4417.2003
Wang W, Sun B (2021) VraCP regulates cell wall metabolism and antibiotic resistance in vancomycin-intermediate Staphylococcus aureus strain Mu50. J Antimicrob Chemother 76:1712–1723. https://doi.org/10.1093/jac/dkab113
Ye ZH, Lee CY (1993) Cloning, sequencing, and genetic characterization of regulatory genes, rinA and rinB, required for the activation of staphylococcal phage phi 11 int expression. J Bacteriol 175:1095–1102. https://doi.org/10.1128/jb.175.4.1095-1102.1993
Zielinska AK, Beenken KE, Mrak LN, Spencer HJ, Post GR, Skinner RA, Tackett AJ, Horswill AR, Smeltzer MS (2012) sarA-mediated repression of protease production plays a key role in the pathogenesis of Staphylococcus aureus USA300 isolates. Mol Microbiol 86:1183–1196. https://doi.org/10.1111/mmi.12048
Acknowledgements
This work was supported by the National Key Research and Development Program of China (2021YFC2300300), the Fundamental Research Funds for the Central Universities (YD9100002013) and the National Natural Science Foundation of China (31870126).
Author information
Authors and Affiliations
Contributions
MJ and YL performed most of the experiments and prepared the manuscript. YL analyzed, interpreted the data, and modified the manuscript. YL, SX and TP participated in the animal experiments. BS and YL supervised the project and obtained funding. All authors discussed the data and read the manuscript.
Corresponding author
Ethics declarations
Conflict of Interest
The authors have no financial conflicts of interest to declare.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Jiang, M., Li, Y., Sun, B. et al. Phage transcription activator RinA regulates Staphylococcus aureus virulence by governing sarA expression. Genes Genom 45, 191–202 (2023). https://doi.org/10.1007/s13258-022-01352-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13258-022-01352-8