ABSTRACT
The recent human infection with avian influenza virus revealed that H9N2 influenza virus is the gene donor for H7N9 and H10N8 viruses infecting humans. The crucial role of H9N2 viruses at the animal-human interface might be due to the wide host range, adaptation in both poultry and mammalian, and extensive gene reassortment. As the most prevalent subtype of influenza viruses in chickens in China, H9N2 also causes a great economic loss for the poultry industry, even under the long-term vaccination programs. The history, epidemiology, biological characteristics, and molecular determinants of H9N2 influenza virus are reviewed in this paper. The contribution of H9N2 genes, especially RNP genes, to the infection of humans needs to be investigated in the future.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
INTRODUCTION
H9N2 influenza virus has become endemic in different types of terrestrial poultry in multiple countries on the Eurasian continent, resulting in great economic losses due to reduced egg production or high mortality associated with co-infection with other pathogens (Lee and Song, 2013; Shanmuganatham et al., 2013; Sun et al., 2010b; Zhao et al., 2013). Epidemiological and genetic studies revealed that the hemagglutinin (HA) gene of the H9N2 influenza viruses could be divided into Eurasian avian and American avian lineages. The Eurasian avian lineage involved three distinct lineages, including A/chicken/Beijing/1/94-like (BJ/94-like), A/quail/Hong Kong/G1/97-like (G1-like), and A/duck/Hong Kong/Y439/97 (Y439-like). BJ/94-like and G1-like viruses have been prevalent mainly in China since the mid-1990s (Sun et al., 2010b).
HISTORY AND EPIDEMIOLOGY OF H9N2 VIRUS IN AVIAN SPECIES IN CHINA
In China, which is regarded as an epicenter of avian influenza viruses, the H9N2 virus has been detected in multiple avian species, including chicken, duck, quail, pheasant, partridge, pigeon, silky chicken, chukar, and egret (Wang et al., 2012a; Xu et al., 2007).
The first outbreak of the H9N2 influenza virus in China occurred in Guangdong province of Southern China during November 1992 to May 1994; the outbreak affected, amongst others, seventeen chicken farms and two rare bird farms. These H9N2 viruses killed broilers with mortality of 10%–40%, and reduced the laying rates by 14%–75% (Chen et al., 1994; Zhang et al., 1994); however, they only caused mild flu-like symptoms in specified pathogen free (SPF) chickens in the laboratory (Chen et al., 1994). After this outbreak, the H9N2 infection sporadically occurred in chickens, ducks, and goose and spread to Northern China (Chen and Chen, 1999). The H9N2 influenza virus spread to most provinces of China within two months in 1998, and is now the most prevalent subtype of influenza viruses in chickens in China (Pan et al., 2012).
H9N2 infections occur throughout the whole year, with lower morbidity in the summer. The isolation rate of H9N2 influenza virus in apparently healthy chickens, ducks, and other minor poultry species (such as quail, partridges, chukar, pheasant, and guinea fowl) in live poultry markers between 2000 and 2005 were 2.5%, 0.18%, and 4.7%, respectively (Xu et al., 2007). The isolation rate of H9N2 virus in the poultry in live poultry markets in Shanghai province of China was 2.6% between 2008 and 2010 (Wang et al., 2014).
CLINICAL SIGNS IN CHICKENS OF H9N2 VIRUS
The H9N2 influenza virus can be transmitted by air droplet, dust, feed, or water (Liu, 2012). On its own, the H9N2 influenza did not induce obvious clinical signs or deaths in chickens (Sun et al., 2010b). Chickens usually seemed to be healthy; few showed depression and ruffled feathers. Increased oral mucus was observed in the infected chickens (Sun et al., 2010b). At autopsy, local pulmonary consolidation and petechiae in throat, trachea, and/or intestine has been observed. The virus replicated itself efficiently in tracheas, while the inoculated chickens rarely shed detectable viruses in the cloacal swabs (Li et al., 2005).
H9N2 influenza virus makes chickens more susceptible to secondary infections, especially Escherichia coli infections with a mortality rate of at least 10%. In addition, the trachea or bronchi are easily embolized by mucus when the ventilation is poor, leading to severe respiratory disease and death. Chickens are immunized with H9N2 vaccines in China. Therefore, outbreak of H9N2 infections is common between vaccination. Commercial broilers are most commonly infected with H9N2 virus because most of them were very young and kept under poor conditions. About 60% of the poorly ventilated intensive broiler farms are infected by the H9N2 influenza virus. The morbidity of H9N2 infected laying hens is about 10%. If the laying hens were infected by H9N2 influenza during the laying period, the laying rate reduced to about 20%, and the birds were characterized by severe follicular congestion, and secretions are observed in fallopian tubes and uterus (Zhang, 2013).
VACCINATION APPLICATION OF H9N2 VIRUS IN CHINA
To prevent H9N2 infection in chickens, China implemented long-term vaccination programs in chicken farms as early as 1998 (Li et al., 2005; Zhang et al., 2008). Most of the H9N2 vaccines are inactivated vaccines. At least over twenty different commercial vaccines are used in China, with the vaccines are frequently updated. However, H9N2 avian influenza viruses continues to persist in chicken populations, even in vaccinated flocks (Zhang et al., 2008).
ANTIGENIC DRIFT OF H9N2 VIRUS IN CHINA
H9N2 viruses isolated from chickens in China underwent antigenic drift to evolve into distinct antigenic groups (Sun et al., 2012). This antigenic drift might have led to immunization failure and may explain the current prevalence of the H9N2 influenza virus in China.
The identification of antigenic sites of H9 is important for monitoring antigenic variants and developing effective vaccines. Although the crystal structure of H9 has been determined (Ha et al., 2001), detailed H9 antigenic epitopes have not been elucidated. The identification of amino acids in H9 antigenic sites revealed different distribution of antigenic areas among other subtypes. Unlike the subtypes analyzed so far, H9 hemagglutinin does not contain an antigenic site corresponding to site A in H3 hemagglutinin (Kaverin et al., 2004). Multiple amino acid positions in HA protein related to the antigenicity of H9N2 viruses were identified, most of which located in the distal head of the HA trimer (see Fig. 1) (Kaverin et al., 2004; Okamatsu et al., 2008; Wan et al., 2014).
INTERSPECIES TRANSMISSION OF H9N2 VIRUS IN CHINA
Occasional cases of H9N2 infecting humans have been reported in Southern China and Hong Kong (see Table 1), but there has been no evidence for human-to-human transmission (Uyeki et al., 2002). Patients presented with a mild and typical human flu-like illness that can easily be overlooked (Butt et al., 2005; Gou et al., 2000). In fact, serologic surveillance revealed that the number of humans infected by H9N2 virus were much higher than that of the confirmed cases. Poultry workers are considered to be at high risk of infection with avian influenza due to their frequent exposure to chickens. About 2.3%–4.6% of poultry workers had antibodies against H9 (Huang et al., 2013; Yu et al., 2013).
H9N2 influenza viruses in poultry also transmitted to pigs, generating variants with novel antigenic and genetic characteristics (Cong et al., 2007; Peiris et al., 2001). Recently, an avian origin H9N2 influenza virus was isolated from dogs in Southern China, and the positive rates of serums from dogs in Southern China were 20.21% in 2010, 28.98% in 2011, and 44.85% in 2012; suggesting the circulation of H9N2 virus among dogs (Sun et al., 2013a). The enlarged host range of the H9N2 influenza virus underscored the potential public threat.
GENETIC AND BIOLOGICAL EVOLUTION OF THE H9N2 VIRUS IN CHINA
Phylogenetic analysis indicated that the HA and neuroaminidase (NA) genes of most H9N2 viruses in China belong to BJ/94-like and G1-like lineages. The hosts of BJ/94-like viruses included chickens, ducks, other minor poultry species, swine, and humans, while G1-like viruses were only isolated from minor poultry and humans in Southern China. Compared with the surface genes (HA and NA), the lineages of six internal genes show a greater diversity, indicating frequent reassortment of these genes. At least 35 genotypes, with some genotypes consisting of triple or quadruple reassortants, were found in poultry in China (Sun et al., 2010b; Xu et al., 2007).
The number of H9N2 viruses, possessing HA proteins with an alanine to serine substitution at the P5 cleavage site (position 316, by H9 numbering) and a three-amino-acid deletion in the NA stalk (positions 61–63) in China was increasing (Sun et al., 2010b; Xu et al., 2007) (see Table 3). The HA-A316S mutation increased the HA cleavage efficiency, and the short stalk NA improved the NA enzyme activity and the virus release from erythrocytes. Each mutation or a combination of these mutations increased the virulence of H9N2 virus in chickens and mice (Sun et al., 2013b). In China, increasingly H9N2 influenza viruses with HA-Q226L substitution were found. HA-226L allowed H9N2 viruses to preferentially infect nonciliated cells expressing mainly SA-ɑ-2,6-Gal receptors and to grow more efficiently in human airway epithelial cultures maintained at the air-liquid interface (Wan and Perez, 2007). Additionally, the substitutions in M2 protein associated with drug-resistance were observed in Chinese H9N2 viruses since 1998 (Sun et al., 2010b).
It is noteworthy that H9N2 viruses present are with increasing adaption to both chickens and mammalians. Our chicken experiment showed that the titers of the H9N2 viruses in the tracheas of the inoculated and contact birds isolated after 2003 were hundred to thousand times higher than those in the earlier isolates (Sun et al., 2010b). According to an experiment performed by Li et al., H9N2 viruses isolated before 2000 could not be recovered from any of the organs of inoculated mice, and the mice stayed healthy and kept gaining weight. By contrast, those isolated between 2000 and 2002 induced disease signs and weight loss (up to 20% weight loss) (Li et al., 2005). Some H9N2 viruses isolated between 2007 and 2009 were reported to be highly lethal to mice and able to spread systemically, comparable with the highly pathogenic avian influenza virus (HPAI) (Bi et al., 2010; Deng et al., 2010). However, few reports similar to those of Bi et al. (2010) and Deng et al. (2010) have been published. The possibility of contamination of the H9N2 viruses isolated in the above-mentioned two studies with HPAI or bacteria cannot be excluded.
VIRULENCE, TRANSMISSIBILITY, AND THE MOLECULAR MECHANISM OF THE H9N2 VIRUS IN ANIMAL MODELS
H9N2 influenza virus replicated efficiently in several animal models, including chickens, quails, BALB/c mice (Li et al., 2005), guinea pigs (Sun et al., 2010a), and ferrets (Ku et al., 2014) (see Table 2)—these animals are natural host species of the H9N2 virus. Quail were, without showing any signs of disease, more susceptible to the G1-lineage of the H9N2 virus than turkeys; while turkeys were associated with worse clinical conditions after being infected with G1-like H9N2 viruses (Bonfante et al., 2013). Some isolates could transmit among guinea pigs and ferrets via direct contact, but not by respiratory droplets (Lv et al., 2012; Wan et al., 2008) (see Table 2). Rhesus macaque, a non-human primate model, infected by the H9N2 virus, presented with biphasic fever and viral pneumonia, and the virus replicated in both their upper and lower respiratory tracts (Zhang et al., 2013a). The dogs and cats were also susceptible to avian H9N2 virus via the upper respiratory tract, and the virus could transmit between cats but not between dogs (Zhang et al., 2013b).
The H9N2 virus with PB2-D253N and PB2-Q591K had greater polymerase activity, induced more tumor necrosis factor alpha (TNF-ɑ) in human macrophages, replicated better in differentiated normal human bronchial epithelial (NHBE) cells, and were more pathogenic in mice (Mok et al., 2011). Serial passages of H9N2 influenza virus in the animal models suggested that threat of H9N2 virus for public health might be increasing. An H9N2 virus could replicate and transmit in quail, but replicated poorly and did not transmit in chickens (Hossain et al., 2008). After 23 serial passages in quail, followed by 10 serial passages in chickens; this virus became very efficient at replicating and transmitting in quail and chickens. Moreover, the passaged viruses, unlike the parent virus, were able to readily infect mice and showed faster replication kinetics in tissue culture. These results are in agreement with the notion that adaptation of H9 viruses to land-based birds can lead to strains with expanded host range. A mouse-adapted H9N2 virus, generated by serial lung-to-lung passage, gained improved growth characteristics in mammalian cells, extended tissue tropism in mice, and was lethal for mice (Wu et al., 2009). We found that the combination of M147L and E627K on PB2 protein was responsible for the enhanced viral replication ability and increased virulence in mice of a mouse-adapted H9N2 virus (Wang et al., 2012b) (see Table 3).
Because of the wide host range of H9N2, the possibility of reassortment between H9N2 and other influenza viruses in the future cannot be excluded. High genetic compatibility was found between H9N2 and H1N1/09 influenza viruses (Sun et al., 2011), and reassortants carrying the HA gene alone or in combination with the NA gene of an H9N2 virus in the H1N1/09 genetic background were respiratory droplet-transmissible between ferrets (Kimble et al., 2014; Kimble et al., 2011). Sorrell et al. found that following adaptation in the ferret, a reassortant virus carrying the surface proteins of an avian H9N2 into a human H3N2 backbone could gain respiratory transmissibility, and the adaptive changes in HA1-T189A and HA2-G192R were demonstrated to be necessary for respiratory droplet transmission (Sorrell et al., 2009) (see Table 3).
H9N2 AS GENE DONOR TO NOVEL INFLUENZA VIRUSES
H9N2 influenza virus has been recognized to reassort with multiple other subtypes, including H6N1, H6N2, and H5N1 viruses (Shi et al., 2014; Sun et al., 2010b) (see Fig. 2). The G1-like H9N2 viruses were the likely donors of the six internal genes of the H5N1 viruses causing the bird flu outbreak which transmitted to humans in Hong Kong in 1997 (Guan et al., 1999), and the BJ/94-like H9N2 viruses provided the six internal genes to H7N9 and H10N8 viruses which emerged in humans in China since 2013 (Cui et al., 2014; Liu et al., 2014). Moreover, H7N9 influenza viruses continued to reassort with circulating H9N2 viruses, resulting in multiple genotypes of H7N9 viruses (Wu et al., 2013).
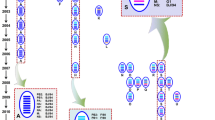
Phylogenetic tree of the PB2 gene of representative influenza viruses which are genetically related to H9N2 viruses in China. The unrooted phylogenetic tree was generated with MEGA 5.0, using the distance based neighbor-joining method. The reliability of the tree was assessed by bootstrap analysis with 1000 replications. The analysis was based on nucleotides 10–2241. H9N2 viruses are shown in blue
THE FUTURE
The reassortant viruses derived from the H9N2 virus posed a great threat to public health. Compared with early isolates, recent H9N2 influenza viruses are much easier to infect mammals. The contribution of H9N2 genes, especially ribonucleoprotein (RNP) genes, to the infection in human needs to be determined. Additionally, to prevent and control avian influenza, veterinary biosecurity on farms need to be increased, rather than vaccine application alone, and live poultry markets should be shut down.
References
Bi J, Deng G, Dong J, Kong F, Li X, Xu Q, Zhang M, Zhao L, Qiao J (2010) Phylogenetic and molecular characterization of H9N2 influenza isolates from chickens in Northern China from 2007–2009. PLoS One 5:e13063
Bonfante F, Patrono LV, Aiello R, Beato MS, Terregino C, Capua I (2013) Susceptibility and intra-species transmission of the H9N2 G1 prototype lineage virus in Japanese quail and turkeys. Vet Microbiol 165:177–183
Butt KM, Smith GJ, Chen H, Zhang LJ, Leung YH, Xu KM, Lim W, Webster RG, Yuen KY, Peiris JS et al (2005) Human infection with an avian H9N2 influenza A virus in Hong Kong in 2003. J Clin Microbiol 43:5760–5767
Chen BL, Zhang Z, Chen WB (1994) The study of avian influenza: I. The isolation and preliminary serological identification of avian influenza virus in chicken. Chin J Vet Med 20:3–5
Chen F, Chen XC (1999) The genetic analysis of Avian influenza A/chicken/Beijing/1/96 (H9N2) strain. Chin J Vet Med 21:130–133
Cong YL, Pu J, Liu QF, Wang S, Zhang GZ, Zhang XL, Fan WX, Brown EG, Liu JH (2007) Antigenic and genetic characterization of H9N2 swine influenza viruses in China. J Gen Virol 88:2035–2041
Cui L, Liu D, Shi W, Pan J, Qi X, Li X, Guo X, Zhou M, Li W, Li J et al (2014) Dynamic reassortments and genetic heterogeneity of the human-infecting influenza A (H7N9) virus. Nat Commun 5:3142
Deng G, Bi J, Kong F, Li X, Xu Q, Dong J, Zhang M, Zhao L, Luan Z, Lv N et al (2010) Acute respiratory distress syndrome induced by H9N2 virus in mice. Archiv Virol 155:187–195
Guan Y, Shortridge KF, Krauss S, Webster RG (1999) Molecular characterization of H9N2 influenza viruses: were they the donors of the “internal” genes of H5N1 viruses in Hong Kong? Proc Natl Acad Sci USA 96:9363–9367
Guo Y, Li J, Cheng X (1999) [Discovery of men infected by avian influenza A (H9N2) virus]. Zhonghua shi yan he lin chuang bing du xue za zhi = Zhonghua shiyan he linchuang bingduxue zazhi =. Chin J Exp Clin Virol 13:105–108
Guo Y, Xie J, Wang M (2000) [A strain of influenza A H9N2 virus repeatedly isolated from human population in China]. Zhonghua shi yan he lin chuang bing du xue za zhi = Zhonghua shiyan he linchuang bingduxue zazhi =. Chin J Exp Clin Virol 14:209–212
Guo Y, Xie J, Wu K, Dong J, Wang M, Zhang Y, Guo J, Chen J, Chen Z, Li Z (2002) [Characterization of genome of A/Guangzhou/333/99(H9N2) virus]. Zhonghua shi yan he lin chuang bing du xue za zhi = Zhonghua shiyan he linchuang bingduxue zazhi =. Chin J Exp Clin Virol 16:142–145
Ha Y, Stevens DJ, Skehel JJ, Wiley DC (2001) X-ray structures of H5 avian and H9 swine influenza virus hemagglutinins bound to avian and human receptor analogs. Proc Natl Acad Sci USA 98:11181–11186
Hossain MJ, Hickman D, Perez DR (2008) Evidence of expanded host range and mammalian-associated genetic changes in a duck H9N2 influenza virus following adaptation in quail and chickens. PLoS One 3:e3170
Huang R, Wang AR, Liu ZH, Liang W, Li XX, Tang YJ, Miao ZM, Chai TJ (2013) Seroprevalence of avian influenza H9N2 among poultry workers in Shandong Province, China. Eur J Clin Microbiol Infect Dis 32:1347–1351
Kaverin NV, Rudneva IA, Ilyushina NA, Lipatov AS, Krauss S, Webster RG (2004) Structural differences among hemagglutinins of influenza A virus subtypes are reflected in their antigenic architecture: analysis of H9 escape mutants. J Virol 78:240–249
Kimble JB, Sorrell E, Shao H, Martin PL, Perez DR (2011) Compatibility of H9N2 avian influenza surface genes and 2009 pandemic H1N1 internal genes for transmission in the ferret model. Proc Natl Acad Sci U S A 108:12084–12088
Kimble JB, Angel M, Wan H, Sutton TC, Finch C, Perez DR (2014) Alternative reassortment events leading to transmissible H9N1 influenza viruses in the ferret model. J Virol 88:66–71
Ku KB, Park EH, Yum J, Kim HM, Kang YM, Kim JC, Kim JA, Kim HS, Seo SH (2014) Transmissibility of novel H7N9 and H9N2 avian influenza viruses between chickens and ferrets. Virology 450–451:316–323
Lee DH, Song CS (2013) H9N2 avian influenza virus in Korea: evolution and vaccination. Clin Exp Vaccine Res 2:26–33
Li C, Yu K, Tian G, Yu D, Liu L, Jing B, Ping J, Chen H (2005) Evolution of H9N2 influenza viruses from domestic poultry in Mainland China. Virology 340:70–83
Liu KY (2012) Epidemicity and control of H9N2 subtype avian influenza virus. Anim Husb Feed Sci 33:116
Liu D, Shi W, Gao GF (2014) Poultry carrying H9N2 act as incubators for novel human avian influenza viruses. Lancet 383:869
Lv J, Wei B, Yang Y, Yao M, Cai Y, Gao Y, Xia X, Zhao X, Liu Z, Li X et al (2012) Experimental transmission in guinea pigs of H9N2 avian influenza viruses from indoor air of chicken houses. Virus Res 170:102–108
Mok CK, Yen HL, Yu MY, Yuen KM, Sia SF, Chan MC, Qin G, Tu WW, Peiris JS (2011) Amino acid residues 253 and 591 of the PB2 protein of avian influenza virus A H9N2 contribute to mammalian pathogenesis. J Virol 85:9641–9645
Okamatsu M, Sakoda Y, Kishida N, Isoda N, Kida H (2008) Antigenic structure of the hemagglutinin of H9N2 influenza viruses. Archiv Virol 153:2189–2195
Pan Q, Liu A, Zhang F, Ling Y, Ou C, Hou N, He C (2012) Co-infection of broilers with Ornithobacterium rhinotracheale and H9N2 avian influenza virus. BMC vet Res 8:104
Peiris JS, Guan Y, Markwell D, Ghose P, Webster RG, Shortridge KF (2001) Cocirculation of avian H9N2 and contemporary “human” H3N2 influenza A viruses in pigs in southeastern China: potential for genetic reassortment? J Virol 75:9679–9686
Saito T, Lim W, Suzuki T, Suzuki Y, Kida H, Nishimura SI, Tashiro M (2001) Characterization of a human H9N2 influenza virus isolated in Hong Kong. Vaccine 20:125–133
Shanmuganatham K, Feeroz MM, Jones-Engel L, Smith GJ, Fourment M, Walker D, McClenaghan L, Alam SM, Hasan MK, Seiler P et al. (2013) Antigenic and molecular characterization of avian influenza A(H9N2) viruses, Bangladesh. Emerg Infect Dis 19: doi:10.3201/eid1909.130336
Shi W, Li W, Li X, Haywood J, Ma J, Gao GF, Liu D (2014) Phylogenetics of varied subtypes of avian influenza viruses in China: potential threat to humans. Protein Cell 5:253–257
Sorrell EM, Wan H, Araya Y, Song H, Perez DR (2009) Minimal molecular constraints for respiratory droplet transmission of an avian-human H9N2 influenza A virus. Proc Natl Acad Sci U S A 106:7565–7570
Subbarao K, Katz J (2000) Avian influenza viruses infecting humans. Cell Mol Life Sci CMLS 57:1770–1784
Sun Y, Bi Y, Pu J, Hu Y, Wang J, Gao H, Liu L, Xu Q, Tan Y, Liu M et al (2010a) Guinea pig model for evaluating the potential public health risk of swine and avian influenza viruses. PLoS One 5:e15537
Sun Y, Pu J, Jiang Z, Guan T, Xia Y, Xu Q, Liu L, Ma B, Tian F, Brown EG et al (2010b) Genotypic evolution and antigenic drift of H9N2 influenza viruses in China from 1994 to 2008. Vet Microbiol 146:215–225
Sun Y, Qin K, Wang J, Pu J, Tang Q, Hu Y, Bi Y, Zhao X, Yang H, Shu Y et al (2011) High genetic compatibility and increased pathogenicity of reassortants derived from avian H9N2 and pandemic H1N1/2009 influenza viruses. Proc Natl Acad Sci U S A 108:4164–4169
Sun Y, Pu J, Fan L, Sun H, Wang J, Zhang Y, Liu L, Liu J (2012) Evaluation of the protective efficacy of a commercial vaccine against different antigenic groups of H9N2 influenza viruses in chickens. Vet Microbiol 156:193–199
Sun X, Xu X, Liu Q, Liang D, Li C, He Q, Jiang J, Cui Y, Li J, Zheng L et al (2013a) Evidence of avian-like H9N2 influenza A virus among dogs in Guangxi, China. Infect Genet Evol: J Mol Epidemiol Evol Genet Infect Dis 20:471–475
Sun Y, Tan Y, Wei K, Sun H, Shi Y, Pu J, Yang H, Gao GF, Yin Y, Feng W et al (2013b) Amino acid 316 of hemagglutinin and the neuraminidase stalk length influence virulence of H9N2 influenza virus in chickens and mice. J Virol 87:2963–2968
Uyeki TM, Chong YH, Katz JM, Lim W, Ho YY, Wang SS, Tsang TH, Au WW, Chan SC, Rowe T et al (2002) Lack of evidence for human-to-human transmission of avian influenza A (H9N2) viruses in Hong Kong, China 1999. Emerg Infect Dis 8:154–159
Wan H, Perez DR (2007) Amino acid 226 in the hemagglutinin of H9N2 influenza viruses determines cell tropism and replication in human airway epithelial cells. J Virol 81:5181–5191
Wan H, Sorrell EM, Song H, Hossain MJ, Ramirez-Nieto G, Monne I, Stevens J, Cattoli G, Capua I, Chen LM et al (2008) Replication and transmission of H9N2 influenza viruses in ferrets: evaluation of pandemic potential. PLoS One 3:e2923
Wan Z, Ye J, Xu L, Shao H, Jin W, Qian K, Wan H, Qin A (2014) Antigenic mapping of the hemagglutinin of an H9N2 avian influenza virus reveals novel critical amino acid positions in antigenic sites. J Virol 88:3898–3901
Wang B, Chen Q, Chen Z (2012a) Complete genome sequence of an H9N2 avian influenza virus isolated from egret in Lake Dongting wetland. J Virol 86:11939
Wang J, Sun Y, Xu Q, Tan Y, Pu J, Yang H, Brown EG, Liu J (2012b) Mouse-adapted H9N2 influenza A virus PB2 protein M147L and E627K mutations are critical for high virulence. PLoS One 7:e40752
Wang Q, Ju L, Liu P, Zhou J, Lv X, Li L, Shen H, Su H, Jiang L, Jiang Q (2014) Serological and virological surveillance of avian influenza A virus H9N2 subtype in humans and poultry in Shanghai, China, between 2008 and 2010. Zoonoses Public Health. doi:10.1111/zph.12133
Wu R, Zhang H, Yang K, Liang W, Xiong Z, Liu Z, Yang X, Shao H, Zheng X, Chen M et al (2009) Multiple amino acid substitutions are involved in the adaptation of H9N2 avian influenza virus to mice. Vet Microbiol 138:85–91
Wu A, Su C, Wang D, Peng Y, Liu M, Hua S, Li T, Gao GF, Tang H, Chen J et al (2013) Sequential reassortments underlie diverse influenza H7N9 genotypes in China. Cell Host Microbe 14:446–452
Xu KM, Smith GJ, Bahl J, Duan L, Tai H, Vijaykrishna D, Wang J, Zhang JX, Li KS, Fan XH et al (2007) The genesis and evolution of H9N2 influenza viruses in poultry from southern China, 2000 to 2005. J Virol 81:10389–10401
Yu Q, Liu L, Pu J, Zhao J, Sun Y, Shen G, Wei H, Zhu J, Zheng R, Xiong D et al (2013) Risk perceptions for avian influenza virus infection among poultry workers, China. Emerg Infect Dis 19:313–316
Zhang GZJ (2013) The analysis of epidemic diseases in poultry in 2013 China Poultry 35:56–59
Zhang ZJ, Chen B, Chen WB (1994) The study of avian influenza: II. The incidence and serological survey of avian influenza. Chin J Vet Med 20:6–7
Zhang P, Tang Y, Liu X, Peng D, Liu W, Liu H, Lu S, Liu X (2008) Characterization of H9N2 influenza viruses isolated from vaccinated flocks in an integrated broiler chicken operation in eastern China during a 5 year period (1998–2002). J Gen Virol 89:3102–3112
Zhang K, Xu W, Zhang Z, Wang T, Sang X, Cheng K, Yu Z, Zheng X, Wang H, Zhao Y et al (2013a) Experimental infection of non-human primates with avian influenza virus (H9N2). Archiv Virol 158:2127–2134
Zhang K, Zhang Z, Yu Z, Li L, Cheng K, Wang T, Huang G, Yang S, Zhao Y, Feng N et al (2013b) Domestic cats and dogs are susceptible to H9N2 avian influenza virus. Virus Res 175:52–57
Zhao M, Liu Q, Sun Q, Zhang W, Zhao G, Gu M, Wang X, Hu S, Liu X, Liu X (2013) Full genome sequence of a natural reassortant H9N2 Avian influenza virus isolated from domestic ducks in Jiangsu Province, China. Genome Announc 1:e00463-13
ACKNOWLEDGMENTS
This work was supported by the National Natural Science Foundation of China (Grant Nos. 31430086 and 31302102), the National Basic Research Program (973 Program) (No. 2011CB504702), grants from the Chang Jiang Scholars Program, and the National Science Fund for Distinguished Young Scholars (31025029).
COMPLIANCE WITH ETHICS GUIDELINES
Yipeng Sun and Jinhua Liu declare that they have no conflict of interest.
This article does not contain any studies with human or animal subjects performed by the any of the authors.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution License which permits any use, distribution, and reproduction in any medium, provided the original author(s) and the source are credited.
About this article
Cite this article
Sun, Y., Liu, J. H9N2 influenza virus in China: a cause of concern. Protein Cell 6, 18–25 (2015). https://doi.org/10.1007/s13238-014-0111-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13238-014-0111-7