Abstract
The family Ajellomycetaceae (Onygenales) includes mammal-associated pathogens within the genera Blastomyces, Emmonsia, Histoplasma and Paracoccidioides, as well as the recently described genera, Emergomyces that causes disease in immunocompromised hosts, and Emmonsiellopsis, known only from soil. To further assess the phylogenetic relationships among and between members of these genera and several previously undescribed species, we sequenced and analyzed the DNA-directed RNA polymerase II (rPB2), translation elongation factor 3-α (TEF3), β-tubulin (TUB2), 28S large subunit rDNA (LSU) and the internal transcribed spacer regions (ITS) in 68 strains, in addition to morphological and physiological investigations. To better understand the thermal dimorphism among these fungi, the dynamic process of transformation from mycelial to yeast-like or adiaspore-like forms was also assessed over a range of temperatures (6–42 °C). Molecular data resolved the relationships and recognized five major well-supported lineages that correspond largely to the genus level. Emmonsia, typified by Emmonsia parva, is a synonym of Blastomyces that also accommodates Blastomyces helicus (formerly Emmonsia helica). Emmonsia crescens is phylogenetically distinct, and found closely related to a single strain from soil without known etiology. Blastomyces silverae, Emergomyces canadensis, Emergomyces europaeus and Emmonsia sola are newly described. Almost all of the taxa are associated with human and animal disease. Emmonsia crescens, B. dermatitidis and B. parvus are prevalently associated with pulmonary disease in humans or animals. Blastomyces helicus, B. percursus, Emergomyces africanus, Es. canadensis, Es. europaeus, Es. orientalis and Es. pasteurianus (formerly Emmonsia pasteuriana) are predominantly found in human hosts with immune disorders; no animal hosts are known for these species except B. helicus.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The majority of known systemic, thermally dimorphic fungi are classified in a single family of the order Onygenales, the Ajellomycetaceae. Members of the family differ in host predilection and virulence, but in general the family is unique in the fungal kingdom for its large number of species exhibiting specialized invasive phases adapted to survival and replication in tissues of mammal hosts. For the most part, the course of infection by these fungi shares the following key features: inhalation of infectious propagules from the environment, pulmonary and extrapulmonary disease, a significant role of the acquired cellular immune system in the control of the infection, and, in some cases, endogenous reactivation upon severe immune impairment of the host (de Hoog et al. 2016b). Molecular findings have suggested that fungi with the above life cycle and described in the genera Blastomyces, Emmonsia, Histoplasma, and Paracoccidioides are phylogenetically coherent. Lacazia loboi, the cause of lobomycosis (which follows subcutaneous inoculation), shares a common ancestor with Paracoccidioides (Taborda et al. 1999; Vilela and Mendoza 2018). Genera recently added to the Ajellomycetaceae include Emmonsiellopsis comprising only geophilic species (Marin-Felix et al. 2015) and Emergomyces that comprises systemic pathogens primarily of immunocompromised human hosts (Dukik et al. 2017b). Nearly all species described within the Ajellomycetaceae share morphological and ecological features of thermal dimorphism (conversion from filamentous to yeast-like or enlarged adiaspore-like morphotypes, expressed under saprobic and pathogenic conditions, respectively), a highly similar conidial apparatus, often restricted geographic distribution and absence of keratinolytic activity (Baumgardner and Paretsky 1999; Restrepo et al. 2000; Untereiner et al. 2004; Sigler 2005; Bagagli et al. 2006, 2008). Sexual phases (previously known as Ajellomyces states) are known within the genera Emmonsia, Blastomyces and Histoplasma and comprise heterothallism, complex ascomata (gymnothecia) with coiled appendages, evanescent asci and oblate ascospores (McDonough and Lewis 1968; Kwon-Chung 1973; Sigler 1996).
Several species of Ajellomycetaceae are etiologic agents of well-known diseases in otherwise healthy mammals. The endemic mycoses, i.e. blastomycosis, histoplasmosis, and paracoccidioidomycosis, have considerable impact on public health with a combined estimated burden of 32,000 cases of life-threatening disease each year (Colombo et al. 2011). Disease caused by Emmonsia species, on the other hand, is less well understood. Until recently, it was considered to be primarily a disease of mammals other than humans. Emmonsia parva and Ea. crescens are associated with adiaspiromycosis, a localized lung infection mainly in wild rodents and occasionally in humans (Emmons and Jellison 1960; Hubálek et al. 1998; Sigler 2005). The association of Emmonsia species with invasive human disease received little attention until Emmonsia pasteuriana (recently renamed Emergomyces pasteurianus; Dukik et al. 2017b) was described as the cause of disseminated disease associated with advanced HIV infection in an individual from Europe (Drouhet et al. 1998). Recently Kenyon et al. (2013) reported a disease among HIV-positive individuals in South Africa caused by another emmonsia-like fungus, since described as Emergomyces africanus (Dukik et al. 2017b). To date, over 80 human cases of disease caused by Es. africanus have been diagnosed in South Africa (Maphanga et al. 2017). In a retrospective review of the first 52 cases from South Africa, all patients were immunocompromised, cutaneous disease occurred in 95% of patients and the case-fatality rate of infections was 50% (Schwartz et al. 2015b).
Several other emmonsia-like agents have been reported sporadically but with increasing frequency from human hosts (Schwartz et al. 2015b; Dukik et al. 2017b; Yang et al. 2017). However, little is known about the phylogeny of these organisms and several have not been formally described. To further assess relationships of these emerging fungi with pathogens in the Ajellomycetaceae, a phylogenetic study was undertaken. Prior molecular studies revealed that ribosomal DNA sequences generated well-supported clades, but species definitions were insufficiently clarified (Untereiner et al. 2004; Peterson and Sigler 1998). In this study, in addition to the analysis of the traditional loci, ITS and LSU, sequences of partial protein-coding genes were obtained, i.e. rPB2, TEF3 and TUB2. We combined multilocus phylogenetic analysis with extensive analysis of morphogenetic processes over a series of temperatures to better understand the process of conversion from saprobic (mycelial) to invasive (yeast-like or other) stages. Thermodependent conversion has been studied extensively in species of Histoplasma, Blastomyces, and Paracoccidioides, but data are lacking for lesser known and novel members of the Ajellomycetaceae.
Materials and methods
Morphology and physiology
Sixty-eight emmonsia-like strains were analyzed (Table 1). These were obtained from the Centraalbureau voor Schimmelcultures (CBS, housed at Westerdijk Fungal Biodiversity Institute, Utrecht, The Netherlands), the University of Alberta Microfungus Collection and Herbarium (UAMH), Edmonton, Canada; now UAMH Centre for Global Microfungual Biodiversity, University of Toronto, Toronto, Canada) and from the National Collection of Pathogenic Fungi (NCPF), Mycology Reference Laboratory, Bristol, England), supplemented by kind donations of individual researchers.
Conditions for conidial formation and thermal conversion were pre-tested on several commercially available media; Malt Extract Agar (MEA, Oxoid) was found to be optimal for both sporulation and conversion. Morphology and growth rate of colonies were determined on MEA during 4 weeks in the dark at temperatures of 6, 15, 21, 24, 27, 30, 33, 36, 37, 40 and 42 °C. Colony characters were assessed on MEA after 3 weeks at 24 °C (saprobic phase), 33 °C (intermediary phase) and 37 °C (thermotolerant phase). Production of urease was determined in Christensen’s urea broth (Oxoid) after incubation for 8 h, 24 h and 7 days at 24 and 37 °C, a positive response being a color change from yellow to fuchsia. Cycloheximide tolerance was evaluated by comparing growth on plain Sabouraud glucose agar (SGA) (Oxoid) with that on SGA supplemented with 0.2% cycloheximide (Sigma-Aldrich, Zwijndrecht, The Netherlands) at 24 and 37 °C for 21 days. Isolates were grown on blood agar (BioMérieux, Marcy-l’Étoile, France) for 14 days at 24 and 37 °C to assess hemolysis (clearing) according to Kane et al. (1997). Slide cultures, applied to examine microscopic features and to assess the process of thermal conversion, were prepared using the method of Dukik et al. (2017b), incubated at 21, 24, 27, 30, 33 and 37 °C and examined after 7, 14, 21 and 28 days. Slides were made using Shear’s mounting medium without pigments (Samson et al. 2010). Micrographs were taken using a Nikon Eclipse 80i microscope and DS Camera Head DS-Fi1/DS-5m/DS-2Mv/DS-2MBW using NIS-Element freeware package (Nikon Europe, Badhoevedorp, The Netherlands). Measurements of conidia and other structures were determined using the Nikon Eclipse 80i measurement module.
At saprobic temperature strains were filamentous. Lateral cells bearing conidial structures are referred to as conidiophores, which have or have not a septum at the base, and may bear short lateral branches (secondary conidiophores) arising at right angles. Conidia are sessile or borne on minute pedicels. Incubation at 33–37 °C led to conversion to another growth form (thermotolerant phase) in nearly all strains, resembling the forms known in host tissues. This was initiated by moderate or pronounced swelling, then leading to giant cells. Large, spherical cells in vitro measuring over 30 μm (up to 140 µm) without budding or any other type of reproduction (terminal phase) and with very thick walls are known as adiaspores. Cells between 10–30 μm in diam, mostly produced at or beyond intermediate temperatures (33 °C) and with thinner walls, often more irregular in shape due to occasional broad-based budding are referred to as adiaspore-like cells. Cells measuring 5–10 μm in diam and producing daughter cells from a broad base are large yeast, and cells below 5 μm in diam budding at narrow bases are small yeast. Budding in any cell type may be unipolar, bipolar or multilateral. A diagrammatic overview of the main morphological features is given in Fig. 1.
Molecular analysis
DNA was obtained from 68 strains grown for 7–14 days on MEA at 24 °C in a class II biological safety cabinet. DNA extracts of 12 reference strains belonging to Blastomyces (2), Histoplasma (7) and Paracoccidioides (3) were obtained from the Centraalbureau voor Schimmelcultures (housed at Westerdijk Fungal Biodiversity Institute, Utrecht, The Netherlands) because of their hazard level. About 10 mm2 of material was added to a screw-capped tube containing 490 μL CTAB-buffer (2% cetyltrimethyl ammonium bromide, 100 mM Tris–HCl, 20 mM EDTA, 1.4 M NaCl) and 6–10 acid-washed glass beads (~ 1.5–2 mm diam). Ten microliters proteinase K were added and the mixture was vortexed for 10 min. Tubes were incubated at 60 °C for 1 h, again vortexed and 500 μL of chloroform: isoamylalcohol (24:1) were added followed by shaking for 2 min. Tubes were spun at 14,000 r.p.m. in a microfuge for 10 min and the upper layer was collected in new sterile tubes with 0.55 volume ice-cold iso-propanol and spun again. Finally, pellets were washed with 70% ethanol, air-dried and suspended in 100 μL TE buffer.
Five genomic regions were amplified using primers using standard PCR conditions except for partial LSU with a cycle extension of 90 s (Liu et al. 1999; Stielow et al. 2015; Dukik et al. 2017b). PCR products were visualized on 1% agarose gels and sequenced using ABI big dye terminator v3.1. Sequencing reaction was at 95 °C for 1 min, 30 cycles of 95 °C for 10 s, 50 °C for 5 s and 60 °C for 4 min. A capillary electrophoresis system (Life Technologies 3730XL DNA analyser) was used for bidirectional sequencing. Sequences were those listed in a preceding study (Dukik et al. 2017b), retrieved from GenBank, or were newly generated (291, including 12 ITS, 46 LSU, 80 rPB2, 50 TEF3 and 49 TUB2) and deposited in GenBank (Table 1); in six of eight Es. africanaus TEF3 could not be obtained. Consensus sequences for each locus were obtained and edited using SeqMan in the Lasergene software (DNAStar, WI, U.S.A.). Alignment was done with Muscle (Edgar 2004) using Mega v6.0 (Tamura et al. 2013) with minor manual editing. Missing data for partial or complete sequences were recoded. A concatenated alignment was made with DataConvert1 for ITS, partial LSU, TUB2, TEF3, and rPB2.
Maximum likelihood (ML) and Bayesian inference analyses (BI) were performed using RAxML (Stamatakis 2014) and MrBayes v3.1.2 (Ronquist and Huelsenbeck 2003). Robustness of tree topology for each ML analysis was evaluated by 1000 bootstrap replicates. Suitable substitution models of ML trees were determined using Mega v6.0. ML in the Cipres web server (https://www.phylo.org). Bayesian analyses were performed with two sets of four chains (one cold and three heated) and the Stoprule option, stopping analyses at an average standard deviation of split frequencies of 0.01. The combined dataset was partitioned per locus and the analysis was done in MrBayes v3.1.2. Two parallel runs with four Markov chain Monte Carlo (MCMC) simulations for each run were set for 2,000,000 generations and stopped when average standard deviation of split frequencies fell below 0.01. The sample frequency was set to 100 and the first 25% of trees were removed as burn-in. The different loci within the combined data sets were analysed as separate partitions. Arthroderma flavescens, a member of family Arthrodermataceae (Onygenales) was used as outgroup for all analyses. Phylograms are shown using Figtree v1.3.1 (Shapiro et al. 2010). Bootstrap values ≥ 80% and posterior probabilities ≥ 0.99 were considered as statistically supported and were indicated above thickened branches (Figs. 2, 3, 4).
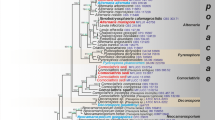
Phylogram (50% majority rule) resulting from ML analysis of concatenated rPB2, TUB2, TEF3, ITS and LSU sequences with confidence values of bootstrap proportions from the ML analysis and posterior probabilities from the Bayesian analysis above branches (> 50% for BS from ML analyses, > 0.95 for PP from Bayesian). Branches with full statistical support (ML-BS ≥ 99%; PP ≥ 0.99) are highlighted by thickened branches. Genus and species clades are discriminated with boxes of different colours. The scale bar shows the expected number of changes per site
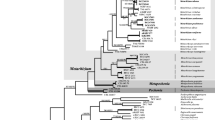
Phylogram (50% majority rule) resulting from a ML of the partial rPB2 gene alignment, with the confidence values BS and PP analysis above branches (> 50% for BS from ML analyses, > 0.95 for PP from Bayesian). Branches with full statistical support (ML-BS ≥ 99%; PP ≥ 0.99) are highlighted by thickened branches. Figures in the column at the right demonstrate the main morphologic features at 24, 33 and 37 °C
APE and SPIDER and their associated R statistical package were also used to evaluate sequence data (https://cran.r-project.org/; http://spider.r-forge.r-project.org/; Brown et al. 2012; Popescu et al. 2012). A sliding window and barcoding gap analysis was conducted to identify the gene section that provides the highest information content in the identification of members in Ajellomycetaceae. The sliding window was set to a length of 100 nucleotides as a prior and the distance metric was set as Tamura’s ‘K2P’ (Popescu et al. 2012; Van den Brink et al. 2015).
Results
Morphology and physiology
Colony morphologies are similar among below-described species of Emmonsia, Blastomyces and Emergomyces. On MEA at 24 °C the colonies have slow to moderate growth and are yellowish-white to buff colored, rather compact, sometimes with grooves (sulcate) or slightly zonate and lack exudates. Colony types are similar to those distinguished by Carmichael (1951), dependent on medium and incubation temperature. Granular colonies are powdery, with low aerial mycelium and with sometimes folded surface. This colony type is common among species of Emmonsia, Emergomyces and Blastomyces and is associated with heavy conidial sporulation. Cottony colonies are dense with felty aerial mycelium, while floccose colonies have loose aerial mycelium giving the appearance of sheep’s wool. The cottony colony type may be associated with lower sporulation or individual strain degeneration as is also the case with glabrous colonies that are smooth to slightly hairy, flat or slightly elevated, sometimes with tufts of hyphae centrally. Glabrous colonies are often produced when isolates are grown at temperatures above 30–33 °C and occur commonly among Es. africanus strains at 24 °C.
Among species of Emmonsia, Blastomyces and Emergomyces, vegetative hyphae are hyaline, narrow (2.5–4.5 μm wide), regularly septate and branched. Conidiophores often have a septum at the base, are slightly to moderately swollen near the tip, and often bear short secondary conidiophores arising at right angles (Fig. 1). Conidia are formed by holothallic conidiogenesis and are sessile or borne on minute pedicels at the tip or on the sides of the conidiophores. Conidia are solitary or occur in chains of two to three, are single-celled, often subspherical, smooth to finely roughened and have rhexolytic dehiscence. Conidial formation was absent in B. helicus and in three other strains: B. percursus (NCPF4091) and B. parvus (CBS 178.60 and CBS 205.48). Thin- to thicker-walled helically coiled hyphae are sometimes present.
Growth rates and the dynamic process of conversion over a range of temperatures were evaluated for all species with particular attention to type strains (Fig. 5, Table 2). The temperature of 33 °C was determined as optimum for elaboration of the intermediate transition stages for species that were able to grow at 37 °C or higher. Swollen, adiaspore-like cells between 10–30 μm wide were produced at or beyond 33 °C, were somewhat irregular in shape and had thinner walls. Conversion to non-filamentous growth forms in vitro occurred in nearly all strains incubated at 37 °C. In most cases, this thermotolerant phase produced in culture resembled forms that occur in infected host tissues and can be hypothesized to represent a pathogenic phase, although pathogenicity is unknown in some species (Emmonsia sola, Emmonsiellopsis species). Blastomyces dermatitidis is unable to produce conidia beyond 33 °C. In Ea. sola NCPF 4289 conidia were produced until 33 °C, leaving a narrower temperature range of the thermotolerant phase, and conversion was more difficult.
Summarized cladogram of the Ajellomycetaceae, compared with growth and conversion characteristics. Bold branches represent bootstrap support (> 80%) in the original phylogram based on multilocus sequencing. Left panel represents growth at temperature ranges after 3 weeks incubation on MEA. Mean diameters were determined by 3 biological replicates, each with 3 technical replicates. Right panel summarizes conversion at different temperatures after 4 weeks incubation on MEA. 0 = saprobic phase, 1–3 = Thermotolerant phase, at three levels of conversion. Yellow circles represent maximum temperatures of conidiation
Physiological characteristics were analyzed for all species with a subset of isolates (Table 2). All tested isolates were tolerant of cycloheximide at 24 °C, but the degree of tolerance differed among species. Blastomyces species varied in tolerance with B. dermatitidis having lowest tolerance. Emmonsia crescens was less tolerant than B. parvus as noted previously (Sigler 1996). Species of Emergomyces had the lowest tolerance to cycloheximide, showing relative inhibition over 35%. Both Emmonsiellopsis species were strongly tolerant with Emms. terrestris even growing better on SGA with cycloheximide. Positive urease activity occurred within 24 h at 24 and 37 °C in Emmonsia species and some Emergomyces species but not in Blastomyces or Emmonsiellopsis species. After 7 days incubation at these temperatures, two Emergomyces species (Es. canadensis and Es. orientalis) and Emmonsiellopsis species were urease negative and urease activity was variable among some Blastomyces strains. Almost all strains showed hemolysis on blood agar after 2 weeks at 37 °C; results varied among Emergomyces species at 24 °C.
Phylogeny and barcoding
In the combined phylogenetic analysis of five gene loci, the family Ajellomycetaceae formed a well-supported lineage (ML/BI 100/1.00), with the geophilic genus Emmonsiellopsis in ancestral position (Clade V) (Fig. 2). Five clades were strongly supported as follows: Clade I (ML/BI 95/0.98), Clade II (ML/BI 100/1.00), Clade III (ML/BI 100/0.99), Clade IV (ML/BI 100/1.00) and CladeV (ML/BI 100/1.00). The topology of the tree largely corresponds with those of Marin-Felix et al. (2015), Dukik et al. (2017b) and Muñoz et al. (2015). No topological conflicts were found when comparing the reciprocal tree topologies based on the rPB2, TUB2 and TEF3 datasets. The concatenated alignment consisted of 3302 characters (including alignment gaps) with 628, 663, 1088, 370 and 536 characters used in the ITS, LSU, rPB2, TEF3 and TUB2 partitions, respectively. For Bayesian inference, a SYM+I+G model was selected for rPB2, GTR+I+G model for ITS, TrN +I+G model for LSU, a K81uf +I+G model for TUB2 and TEF3. Analyses of the rPB2 gene region demonstrated the same five well-supported clades as revealed with the combined data, except that Clade I had lower bootstrap support (Fig. 3). Most species were resolved in the rPB2 and ITS trees (Fig. 4). TEF3 and TUB demonstrated species delimitation but with lower bootstap support (not shown). Generic relationships were better resolved in the rPB2 and combined analyses, but these genes individually were too variable to show relationships over the entire family. In the multigene phylogram, Histoplasma (Clade III) was paraphyletic to Blastomyces (Clade II) (ML/BI 100/0.99). Paracoccidioides brasiliensis (Clade IV) was distant from other genera (ML/BI 99/0.99) with strong statistical support (ML/BI 100/1.00).
Clade I comprises two strongly-supported subclades including strains formerly classified as Emmonsia crescens (Clade I-1; ML/BI 100/1.00) and the other Emergomyces species (Clade I-3; ML/BI 100/1.00) (Figs. 2, 3). While a single strain CBS 142607 (= NCPF 4289) (Clade I-2) is close to Ea. crescens, there is no statistical support for the sister relationship and the strain deviated from its nearest neighbors in at least 20 nucleotide positions in ITS, and 23 and 16 positions in rPB2 and TUB2, respectively. Most strains in the Ea. crescens clade are derived from lungs of terrestrial animal hosts except for two soil-associated strains (Table 1). The clade is comprised of two closely related lineages as was previously noted by Peterson and Sigler (1998) on the basis of ITS-partial LSU data. Their Eurasian lineage, shown here with high support (Clade I-1a; ML/BI 100/0.99) includes the ex-type strain of Ea. crescens and ten other strains, while their North American group comprising strains from Canada and the United States, is less well supported (Clade I-1b; ML/BI 65/0.97) (Fig. 2). The Eurasian and North American groups differed from each other at several positions both in protein-coding genes and the rDNA operon: 1 position in rPB2, 2 in TUB2, 1–2 in TEF3, ITS and LSU. One strain CBS 191.55 deviated from other strains at five positions in rPB2 and two positions in TUB2 and phenotypically in lacking sporulation and a lower maximum growth temperature of 33 °C. The fungi in the Ea. crescens subclades differ from those in the Emergomyces clade in host associations, pathogenicity and pathogenic cell types.
The Emergomyces subclade (Clade I-3; ML/BI 100/1.00) includes five well-supported lineages encompassing species having small yeast-like cells as invasive forms and associated only with human infections (Dukik et al. 2017b). Included are the previously described species Es. pasteurianus from Europe (Clade I-3c; ML/BI 100/1.00) (Drouhet et al. 1998) and Es. africanus from southern Africa (Clade I-3e; ML/BI 100/1.00) (Dukik et al. 2017b), each of which received strong support. Another strain (CBS 102426) from Germany (Clade I-3d), here described as Es. europaeus, has a close relationship with these two species but differs from Es. pasteurianus and Es. africanus in 10 positions in ITS and in 14 vs. 19 and 29 vs. 30 positions in rPB2 and TUB2, respectively. Emergomyces orientalis (Clade I-3b) (Wang et al. 2017) is closely related to two strains from Canada here designated Es. canadensis (Clade I-3a). Comparison of sequences reveals differences in 7 positions in ITS, 8 in rPB2, 17 in TUB2 and 7 in TEF3. Compared to other species in the Ajellomycetaceae these differences exceed the maintained species limit and we therefore keep them as separate species pending analysis of additional strains.
Clade II is strongly supported (ML/BI 100/1.00) and comprises five highly supported monophyletic lineages (Figs. 2, 3). One lineage includes B. dermatitidis (Clade II-3; ML/BI 100/1.00), the type species of Blastomyces together with its relative B. gilchristii (Fig. 4). A second lineage (Clade II-1; ML/BI 100/1.00) comprises the generic type species of Emmonsia, Ea. parva represented by the type strain (CBS 139881 = UAMH 130) which is designated herein as B. parvus. The B. parvus lineage contains subgroups each containing a few strains that show differences in growth rates or morphologies. The lower subclade includes four sublineages having moderate bootstrap support (ML/BI 77/0.91). One with strong support (Clade II-2; ML/BI 98/1.00) includes seven strains representing a new species, B. silverae. A small intra-lineage difference was noted for B. silverae strain (CBS 509.78 = ATCC 32539) that was sister to the main bootstrap-supported cluster and unusual in producing conidia over a wide temperature range (21–37 °C). The other three sublineages are sister to B. dermatitidis (ML/BI 93/0.99) and represent the recently described species B. percursus (Dukik et al. 2017b) (Clade II-4; ML/BI 100/1.00) and B. helicus (formerly Emmonsia helica Sigler, Clade II-5) comprising eight strains that varied at no more than 2 nucleotide positions in ITS, and fewer than 5 in rPB2 and TUB2.
The genus Emmonsiellopsis (Clade V; ML/BI 100/1.00) occurs in an ancestral position within the Ajellomycetaceae and is phylogenetically distant to other genera. The genus comprises two environmental species initially suggested as closely related to the pathogenic dimorphic fungi in the family (Marin-Felix et al. 2015). Emmonsiellopsis terrestris (ML/BI 100/0.99) is represented by the type and two other strains (CBS 273.77 = UAMH 2304; CBS 139889 = UAMH 141; CBS 137499 = FMR 4023) while Emms. coralliformis comprises a single isolate (CBS 137500 = FMR 4024). The thermodependent form is produced in culture with difficulty and resembles that of some Blastomyces species.
We assessed information density of the different loci individually to infer: (a) which marker shows the best potential alone to recognize the described taxa, thus maximizing affiliation to an existing taxon e.g. in the case of an unknown clinical sample, and (b) which sections of the here studied genes are most informative and render taxa with the highest probability as monophyletic. The R packages SPIDER and APE were used to assess these density metrics over all loci. Figure 6 outlines subsequently the following plots for each locus: (a) ‘congruence of NJ (neighbor joining) trees’, (b) ‘proportion of species that are monophyletic’ and (c) ‘sum of diagnostic nucleotides’. The first metric describes the proportion/quantity of neighbor joining trees being congruent (identical) to each other for a given sliding window, and thus is a consistency metric for statistical resampling. It describes the robustness of the given section for a given locus to retrieve the same species affiliations. The second metric describes in a similar way to the first metric, how consistent taxa could be rendered as monophyletic entities over the statistical resampling process. The third metric is quantitatively describing how many nucleotides per window position are informative/diagnostic, thus describing the information content numerically. Values for the first and second metric range from 0 to 1, with 0 no NJ trees/no species being congruent/monophyletic, respectively, while 1 is interpreted as the opposite, and the third metric range is from 0 to infinitive. Secondly, Fig. 7 shows the non-conspecific K2P distances over the intra-specific K2P distances over each window, in other words the ‘classical’ barcoding gap (Meyer and Paulay 2005).
Sliding window analysis conducted with the R package SPIDER, covering a 100 bp sliding-window for the analyzed partial gene sequences (multiple alignment) respectively. A Congruency of neighbor-joining (NJ) trees, when re-sampled 1000 times over each window. B Proportion of species that are monophyletic, indicating quantity of discrete species entities derived from NJ re-sampling process and as defined in this study. C Sum of diagnostic nucleotides at each window position corresponding to the degree of informativeness given a particular alignment position. Note: scaling of Y-axis is not proportional, values range from 0 (lowest) to 1 (highest) for (A) and (B) respectively, for C, number indicates quantities of diagnostic nucleotides for a given window
Pairwise distance sliding window analysis of five genetic loci alignments (analysed individually) showing closest inter-specific (orange whiskers) and intra-specific (blue whiskers) distances (A) and proportion of monophyletic species over a 100 bp sliding window (B). Hypervariable alignment sections were automatically excluded, as indicated by ‘gaps’ for the plot ‘proportion of species that are monophyletic’ per gene (section w140e200 bp in BT2 and w150e230 bp in ITS region)
The informativeness of the analysed loci was relatively even between the individual loci, thus none of the genes exposed a higher non-conspecific distance (inter-specific distance) over a lower intra-specific distance between the analysed taxa. However, clear qualitative differences where observable with particular attention to (a) the informativeness of the individual sequence section analysed, and (b) information density over a particular section analysed. When referring to the informativeness of the individual sections (NJ tree congruency/Proportion of monophyletic species), the ranking is the following: TUB2 > rPB2 > ITS/TEF3 (approximately equal) > LSU. TUB2 here provides the highest consistency over the entire sequence analysed, as revealed in Fig. 6 and the corresponding plots for ‘NJ tree congruency/Proportion of monophyletic species’, particularly since the latter values are consistently > 0.8 from ~ 90–450 bp of the sequence matrix. A similar result was obtained for rPB2, which scores high in its information content. Complete rPB2 was determined as slightly superior over any other of the other four partial gene sections for recognition of the here studied fungi. In contrast to these positive results, surprisingly only the ITS-1 of the gold standard DNA barcode renders NJ-trees/taxa proportionally consistent, while the ITS-2 section performs poorly and indicates almost a complete drop-out as is revealed in Fig. 6, and in the corresponding plot ‘Sum of diagnostic nucleotides’ (section ~ 250–550 bp). While the fungal specific TEF3 also exposes a relatively high consistency, it overall ranks lower (or similarly equal to ITS) due to a lower ratio of diagnostic nucleotides, e.g. when compared to TUB2. The most uninformative locus, at least at the species level, was partial LSU; all sliding window plots revealed that only a small section, approximately between 250 and 350 bp contains a sufficiently variable number of nucleotides to recognize the taxa studied here. However, maximum consistency between NJ trees/monophyletic taxa is approximately half of the best two loci TUB2/rPB2.
Taxonomy
Emmonsia Cif. & Montemartini – Mycobank MB8151
Colonies yellowish-white to ochraceous. Hyphae hyaline. Conidiophores arising at right angles from vegetative hyphae, with secondary conidiophores arising from a swollen tip, Conidia forming at the tip or on the sides of conidiophores, or sessile on vegetative hyphae. Conidia holothallic, with rhexolitic dehiscence. Adiaspores usually produced at 37 °C, spherical with thickened walls, over 100 µm in diam. No growth at 40 °C.
Comments: The genus Emmonsia was proposed with Haplosporangium parvum as type species (Ciferri and Montemartini 1959). The type species of Emmonsia, Ea. parva, is shown here and in prior reports to be a species of Blastomyces (de Hoog et al. 2016a; Dukik et al. 2017b). A second species, Ea. crescens, did not group in Blastomyces and its status is insufficiently resolved in the phylogenetic trees generated by the used barcoding genes. Ea. crescens requires transfer to a different genus or, potentially, conservation of the genus Emmonsia with Ea. crescens as type species. We therefore postpone a name change until investigation of the most suitable option is completed.
Emmonsia crescens C.W. Emmons & Jellison—Ann. N. Y. Acad. Sci. 89: 98. 1960. MycoBank MB330349 (Fig. 8)
= Ajellomyces crescens Sigler—J. Med. Vet. Mycol. 34(5): 305. 1996.
Emmonsia crescens (CBS 177.60). A–C Colonies on MEA 3 weeks at 24, 33 and 37 °C. D–F 24 °C. Slightly swollen conidiophore; secondary conidiophore present. G–I 33 °C. Conidiophores and hyphae swelling and disarticulating to form giant cells. J–M 37 °C. Swollen hyphae leading to adiaspores. Scale bar = 10 μm
Type culture: CBS 177.60 = ATCC 13704 = UAMH 3008, Hamar, Norway, isolated from lungs of rodent Arvicola terrestris by C.W. Emmons and W.L. Jellison, January 1959.
Saprobic phase (24 °C, MEA, 3 weeks): Colonies 35 mm diam, dense, granular to felty, yellowish-white. Conidiophores about 1.5–3.7 (2.2 ± 0.5) μm wide at the base, with secondary conidiophores. Conidia solitary or rarely in chains of two, subspherical, 2.1–4.8 × 2.4–4.9 μm (3.0 ± 0.5 × 3.7 ± 0.5, n = 44), smooth to finely roughened. Intermediate phase (33 °C, MEA, 3 weeks): Colonies 2 mm diam, glabrous, cream, without aerial tufts. Conidia absent. Hyphae densely septate and somewhat swollen, 2.9–7.1 μm wide. Swollen hyphal cells and conidiophores sometimes becoming giant cells, 5.9–25 μm (12.6 ± 3.9, n = 44) or adiaspore-like cells. Thermotolerant phase (37 °C, MEA, 3 weeks): Colonies 4–5 mm, glabrous to slightly hairy, yellowish white. Adiaspores abundant, ovoidal to spherical, abundant, 16–117 μm (69.0 ± 39.2, n = 30) in diam, with smooth walls about 10 μm thick.
Comments: The species is the only one that produces large thick walled adiaspores (Sigler 2005; this study) requiring about 2 weeks at 37 °C. Intermediate morphology includes giant cells without budding; Optimal sporulation occurs at 21–24 °C; conidia absent at 30 °C. Grows at 6 °C, no growth at 40 °C.
Emmonsia sola Y. Jiang, Borman & de Hoog, sp. nov.—MycoBank MB821087 (Fig. 9)
Etymology: referring to soil as the species’ habitat.
Holotype: Arizona, U.S.A., specimen of culture CBS 142607 (preserved in metabolically inactive condition in liquid nitrogen) from soil, sent by A. Borman, December 2012; living strain CBS 142607 = NCPF 4289.
Saprobic phase (24 °C, MEA, 3 weeks): Colonies 64 mm diam, floccose, buff to pale buff at periphery, somewhat sulcate centrally, margin thin. Coiled hyphae sometimes present. Conidiophores 1.1–1.7 μm (1.3 ± 0.2) wide with a septum at the base, sometimes multiseptate, erect, rather long (18–74 μm); short secondary conidiophores arising at right angles. Conidia single or in chains of two to four, subspherical, 1.3–2.8 × 1.7–2.8 μm (1.9 ± 0.4 × 2.2 ± 0.3, n = 46), smooth-walled to finely roughened. Intermediate phase (33 °C, MEA, 3 weeks): Colonies 57 mm diam, dense, flat, cottony, buff centrally, white peripherally, radially sulcate with numerous fissures. Conidia and conidiophores similar to those at 24 °C; conidiophores swelling near the middle and distally, becoming chlamydospore-like; no transformation to other forms. Thermotolerant phase (37 °C, MEA, 3 weeks): Colonies 6 mm diam, compact, white, moist to slightly hairy. Hyphal elements moniliform, variably shaped, 2.0–5.7 μm wide. Conidiophores mostly swollen, ampulliform; septa absent or located at the base, forming giant cells 6. 2–17.7 μm (11.3 ± 2.4; n = 44) in diam with thin walls; sometimes liberated with broad-based budding.
Comments: Emmonsia sola, known only from the type strain NCPF 4289, was identified originally as Emmonsia parva. It was isolated from soil in the U.S.A., but precise strain data are lacking. Our multilocus analysis demonstrated that this strain was close to Ea. crescens with low bootstrap support (ML/BI 68/–). It deviated from all other strains in this study in at least 20 positions in ITS, and in 23 and 16 positions in rPB2 and TUB2, respectively. The species differs from Ea. crescens in its habitat from soil and from all other species by (1) producing giant cells less than 20 μm diam with thin walls and occasional broad-based budding and requiring 4 weeks of incubation at 37 °C to develop; (2) having smaller conidia and longer primary conidiophores (up to 74 μm in length); (3) producing conidia at 33 °C; (4) having strong cycloheximide tolerance and the strongest hemolytic reaction at 24 °C. The single strain of this species had the strongest hemolytic reaction at 24 °C. Due to insufficient taxon sampling, with only a single strain available, phylogenetic resolution remained poor. Strain NCPF 4289 is preliminarily preserved in Emmonsia as Ea. sola. Optimal sporulation occurs at 24–27 °C with conidia produced up to 33 °C. Conversion is slow, requiring about 4 weeks to produce giant cells at 37 °C. Grows at 6 °C; no growth at 40 °C.
Blastomyces Gilchrist & W.R. Stokes—J. Exp. Med. 3: 76. 1898; conserved name, non Blastomyces Costantin & Rolland—Bull. Soc. Mycol. Fr. 4: 153. 1889. MycoBank MB7389.
Colonies yellowish-white to ochraceous. Hyphae hyaline, sometimes helically twisted. Conidiophores arising at right angles from the vegetative hyphae, narrow, unbranched, sometimes with secondary conidiophores arising from a swollen tip, Conidia forming at the tip or on the sides of conidiophores, or sessile on vegetative hyphae, holothallic with rhexolytic dehiscence. At 37 °C producing large yeast-like cells with budding at a broad base or giant cells that may proliferate. Growth at 37 °C, some species up to 40 °C.
Type species: Blastomyces dermatitidis Gilchrist & W.R. Stokes.
Comments: The genus Blastomyces Gilchrist and W.R. Stokes has recently been conserved (de Hoog et al. 2016a) with B. dermatitidis as type species and CBS 674.68 (=ATCC 18187 = UAMH 3539) as epitype. Our multilocus and rPB2 analyses (Figs. 2, 3) provided strong support for Clade II including Ea. parva, type species of Emmonsia, Ea. helica (Sigler 2015), the recently described species B. percursus (Dukik et al. 2017b) and a new species, B. silverae. B. dermatitidis and B. percursus produce large yeast cells with broad-based budding at 37 °C. Blastomyces silverae and B. parvus mainly produce giant cells which could proliferate, as well as occasional large yeast cells with broad-based budding. Species of Blastomyces show a tolerance to higher temperature and to cycloheximide but urease activity is delayed and variably positive among strains of B. parvus and B. silverae (Table 2). No strains of the recently described species B. gilchristii were available for study.
Blastomyces dermatitidis Gilchrist & W.R. Stokes—J. Exp. Med. 3: 76. 1898. MycoBank MB361754 (Fig. 10)
Blastomyces dermatitidis (CBS 642.72). A–C Colonies on MEA 3 weeks at 24, 33 and 37 °C. D–F 24 °C. Swollen, flexuose conidiophore without secondary conidiophores. G–I 33 °C. Swollen, flexuose, hyphae producing terminal yeast-like cells and giant cells which may show multilateral budding. J–M 37 °C. Large yeast-like cells produced multilaterally from irregularly swollen giant cells, large yeast cells abundant, showing uni- or bipolar budding at a broad base. Scale bars: D–M = 10 μm
= Ajellomyces dermatitidis McDonough & A.L. Lewis—Mycologia 60: 77. 1968.
Lectotype: plate VI, Gilchrist & W.R. Stokes – J. Exp. Med. 3: 53–78, pls. IV–VIII. 1898; epitype culture: CBS 674.68 (= ATCC 18188 = UAMH 3539), U.S.A., isolated from human patient, by E.S. McDonough, 1968 (de Hoog et al. 2016a).
Saprobic phase (24 °C, MEA, 3 weeks): Colonies 41 mm diam, cottony to felty, white to buff at the centre, radially sulcate, margin thin, reverse pale buff. Hyphae narrow with some spirally twisted hyphae present. Conidiophores septate at the base and at conidial insertion, 1.6–4.1 (2.2 ± 0.5) μm wide, unbranched slightly bent apically (Fig. 10E). Conidia borne singly, subspherical, 1.8–5.5 × 2.4–5.7 μm (3.5 ± 0.7 × 3.6 ± 0.7, n = 45), finely roughened sometimes adherent to the conidiophore. Intermediate phase (33 °C, MEA, 3 weeks): Colonies 12 mm diam, cerebriform, slightly raised, yellowish white, dense, felty. Swollen, bent, conidiophore-like structures emerge terminally producing inflated yeast-like cells with mulitlateral daughter cells. Some cells become giant cells with thin walls 10.6–29.2 μm (16.9 ± 4.7, n = 40) diam with some multipolar budding. Yeast-like cells with broad-based budding present (Fig. 10I) measuring 6.8–10.0 × 5.4–8.7 μm (9.1 ± 0.5 × 6.6 ± 0.7, n = 45). Thermotolerant phase (37 °C, MEA, 3 weeks): Colonies 6 mm diam, glabrous, smooth, yellowish white. Swollen, bent, conidiophore-like structures producing large giant cells, 10.6–29.2 μm (16.9 ± 4.7, n = 40) in diam (Fig. 10L). Yeast cells measuring 5.0–14.2 × 3.5–8.9 μm (8.0 ± 1.6 × 5.8 ± 1.1, n = 40) with unipoloar, rarely bipolar budding at a broad base.
Comments: B. dermatitidis is distinguished by its simple conidiophores without secondary conidiophores in contrast to other Blastomyces species, Ea. crescens and Emergomyces species. Giant cells are produced as initial stages, while at 37 °C abundant broad-based budding cells are formed within 1 week. Optimal conidiation occurs at 24–27 °C. Growth occurs at 9 °C, optimum 24 °C, no growth at 42 °C.
Blastomyces helicus (Sigler) Y. Jiang, Sigler, Schwartz & de Hoog, comb. nov.—MycoBank MB821083 (Fig. 11)
Basionym: Emmonsia helica Sigler—Index Fungorum 237: 1. 2015.
Blastomyces helicus (CBS 139874). A–C Colonies on MEA 3 weeks at 24, 33 and 37 °C. D–F 24 °C. Helically coiled and swollen hyphae present, conidia absent. G–I 33 °C. Structures similar and giant cells with thin walls and multilateral budding sometimes present. J–M 37 °C. Giant cells bearing spherical daughter cells over the surface, with spindle-shaped daughter cells. Yeast-like cells with multilateral budding at a broad base. Scale bars: D–M = 10 μm
= Emmonsia sp. 1, Schwartz et al. – PLoS Pathog. 11: e1005198, p. 2. 2015.
Type culture: CBS 140056 = UAMH 7101, Edmonton, Canada, specimen of culture from sputum, blood from a male chronic leukemia with disseminated infection, P. Kibsey, December 1991.
Saprobic phase (24 °C, MEA, 3 weeks): Colonies 37 mm diam, yellowish white, cottony to felty, radially sulcate; reverse ochraceous-buff to warm buff peripherally. Hyphae 1.0–2.0 μm wide, hyaline, septate, branched; yellowish helically coiled hyphae often present (Fig. 11D). Conidiophore-like stalks arising at right angles from hyphae, having a basal septum and sometimes with several septa (Fig. 11E, F) but no conidia are produced (Fig. 11E, F). Intermediate phase (33 °C, MEA, 3 weeks): Colonies 45 mm diam, flat, glabrous with radial fissures, tufts of hyphae occurring centrally. Hyphae somewhat swollen, 2–4 μm, hyaline, slightly pigmented, septate at short intervals, constricted at the septa. Conidiophore-like structures swollen; giant cells 4.8–12.9 μm (7.5 ± 1.4, n = 40) with multilateral budding, producing chains of thin-walled variably shaped yeast-like cells budding at a broad base. Thermotolerant phase (37 °C, MEA, 3 weeks): Colonies 20 mm diam, folded, yeast-like, yellowish-tan. Large giant cells bearing spindle-shaped daughter cells 4.2–9.8 × 1.3–3.2 μm (6.5 ± 1.7 × 2.1 ± 0.5, n = 40) (Fig. 11L). Yeast cells varying in shape, budding multilaterally, occurring in short branched chains, sometimes liberating and measuring, 2.4–7.0 × 2.6–6.2 μm (5.3 ± 0.8 × 4.0 ± 0.7, n = 50).
Comments: Blastomyces helicus is differentiated from all here described ajellomycetaceous fungi by absence of conidia and by formation of variably shaped yeast-like cells in short chains. The temperature for conversion is low, starting at 33 °C, and conversion time is short, taking about 1–2 weeks to produce the yeast-like stage at 37 °C. Grows at 9 °C, optimum 30 °C, grows at 40 °C.
All strains of this species (n = 8) originate from North America. Our phylogenetic data show that B. helicus is sister to B. dermatitidis and the species have a number of features in common. Both are thermophilic. All strains of B. helicus grow better at 33 °C than at 24 °C, and at 37 °C, show the most prolific growth of all species studied attaining 20 mm in diam on MEA in 3 weeks. Strains CBS 140058 (= UAMH 11034), CBS 139874 (= UAMH 3398) and CBS 139875 (= UAMH 11294) were able to grow at 40 °C and beyond. Blastomyces helicus is characterized by absence of conidia in the filamentous phase, only spirally coiled hyphae with some brownish pigmentation being present, and in the thermotolerant phase strains producing yeast-like cells (Fig. 11L). After prolonged incubation (4–5 weeks) at 24–27 °C some dumbbell-shaped, spherical or ovoidal conidium-like cells were produced on MEA (Fig. 11E), consistent with observations of Sekhon et al. (1982) who mentioned dumbbell-shaped conidia after several weeks on cereal agar at 25 °C. The latter author reported the infection caused by strain CBS 139874 as a case of blastomycosis. The strain CBS 139874 showed neurotropism, being isolated from spinal fluid; also a second isolate, CBS 139877 (= UAMH 11718) originated from human CSF from a patient with brain infection. Central nervous system involvement is known to occur in B. dermatitidis (Bariola et al. 2010) and B. percursus (Dukik et al, 2017b). A further clinical similarity is the occurrence of disease in animals. While B. dermatitidis disease predominates in dogs (Baumgardner et al. 1995), canine infection with B. helicus also seems to occur: CBS 140058 and CBS 140059 of B. helicus were isolated from lungs of dogs. Feline infections caused by B. helicus have also been reported (Schwartz et al. 2017).
Blastomyces parvus (Emmons & Ashburn) Y. Jiang, Sigler & de Hoog, comb. nov. – MycoBank MB821085 (Fig. 12)
Basionym: Haplosporangium parvum Emmons & Ashburn – Publ. Health Rep., Wash. 57: 1719. 1942 ≡ Emmonsia parva (Emmons & Ashburn) Cif. & Montemartini – Mycopath. Mycol. Appl. 10: 314. 1959 ≡ Chrysosporium parvum (Emmons & Ashburn) Carmichael var. parvum (Emmons & Ashburn) Carmichael – Can. J. Bot. 46: 1164. 1962.
Blastomyces parvus (CBS 139881). A–C Colonies on MEA 3 weeks at 24, 33 and 37 °C. D–F 24 °C. Slightly swollen conidiophore with secondary conidiophore. G–I 33 °C. Conidiophores and conidia. J–M 37 °C. Swollen hyphae; some cells becoming giant cells with thin walls; large yeast-like cells with thick walls present, rarely showing broad-based budding. Scale bars: D–M = 10 μm
Lectotype: Fig. 1 in C.W. Emmons & L.L. Ashburn – Publ. Health Rep. 57: 1720. No. 46; epitype: Arizona, U.S.A., specimen of culture CBS 139881 (preserved in metabolically inactive condition in liquid nitrogen), isolated from lungs of rodent, C.W. Emmons, November 1946; living strain CBS 139881 = UAMH 130.
Saprobic phase (24 °C, MEA, 3 weeks): Colonies 50 mm diam, white, cottony to glabrous with hyphae tufts in centre, radially sulcate, sometimes cracking the agar, margin flat, thin, reverse warm-buff to light buff peripherally. Hyphae 0.9–3.0 μm wide, with few spirally twisted hyphae. Conidiophores arising at right angles from vegetative hyphae, with septa at the base; conidiophores less than 1.5 μm in width, slightly swollen at the tip, occasionally with secondary conidiophores on which conidia are formed at the tip or on the sides. Conidia solitary, rarely in short chains. subspherical, 1.6–3.7 × 1.9–4.1 μm (2.7 ± 0.42 × 2.8 ± 0.5, n = 44), smooth to finely roughened, sometimes adherent to the conidiophore. Intermediate phase (33 °C, MEA, 3 weeks): Colonies 39 mm diam, in macro- and microscopic details nearly identical to 24 °C. Conidia 1.5–3.8 × 1.5–4.0 μm (2.4 ± 0.6 × 2.6 ± 0.6, n = 50). Thermotolerant phase (37 °C, MEA, 3 weeks): Colonies 8 mm diam, tan, compact, moist to slightly hairy. Hyphal elements short, swollen, 2.3–4.1 μm wide, hyaline, septate at short intervals, constricted at the septa; some cells enlarging to become giant cells with thin walls, 4.7–15.7 μm (9.3 ± 2.3, n = 44) wide or forming occasional yeast-like cells with broad-based buds (Fig. 12M).
Comments: No type material was designated by Emmons and Ashburn so a lectotype and epitype are proposed herein. The B. parvus lineage contains subgroups each containing a few strains that show differences in growth rates or morphologies. Strains in one subcluster (CBS 139880, CBS 139883, CBS 204.48) produce conidia abundantly at 21–33 °C, while strains of a second (CBS 178.60, CBS 205.48) did not sporulate and grew faster at 37 °C. Strains demonstrate high tolerance to cycloheximide, with growth at 37 °C, while other species of Blastomyces were inhibited at this temperature. The thermotolerant phase of B. parvus has traditionally been interpreted as small adiaspores (Dowding 1947; Emmons and Jellison 1960; Dvořak et al. 1973; Sigler 2005), but the cells produced at 37 °C have occasional broad-based budding. In contrast, the thick-walled adiaspores of Ea. crescens represent a terminal phase without further reproduction. Transfer temperature is high, at least 36 °C, conversion time is long, taking about 4 weeks. Optimal sporulation temperature: 24–27 °C; grows at 6 and 40 °C.
Blastomyces percursus Dukik, Muñoz, Sigler & de Hoog – Mycoses 60: 306. 2017. MycoBank MB817662 (Fig. 13)
Type culture: CBS 139878 = Kemna 408-93 = UAMH 7425 = UAMH 7426, Israel, specimen of culture from granulomatous lesion on lip of otherwise healthy patient with disseminated infection, isolated by I. Polachek, November 1993.
Blastomyces percursus (CBS 139878). A–C Colonies on MEA 3 weeks at 24, 33 and 37 °C. D–F 24 °C. Conidiophores and secondary conidiophores, some yeast cells arising from vegetative hyphae. G–I 33 °C. Swollen, ampulliform conidiophores and conidia present, some becoming giant cells with thin or thick walls. J–M 37 °C. Swollen hyphae, septate at short intervals, constricted at the septa, fragmenting to become large yeast-like cells with uni- or bipolar budding at a broad base. Scale bars: D–M = 10 μm
Saprobic phase (24 °C, MEA, 3 weeks): Colonies 51 mm diam, white, sectoring, glabrous to floccose, with tufts of hyphae centrally, radially sulcate, reverse warm-buff to light buff periperally. Hyphae 1.1–2.8 μm wide, with few spirally twisted hyphae. Conidiophores septate near the base and below the conidium, often with an intercalary swelling and measuring 1.6–4.1 μm (2.2 ± 0.6) in width; secondary conidiophores present. Conidia solitary, forming in groups of 3–4 on swollen conidiophores, subspherical, 1.5–4.4 × 1.7–4.6 μm (2.7 ± 0.6 × 2.6 ± 0.5, n = 45), smooth-walled to slightly roughened, adherent to the conidiophore. Yeast-like cells arising from vegetative hyphae (Fig. 13F), sessile, spherical, 3.0–8.0 μm (5.1 ± 1.1, n = 45), budding from narrow base. Intermediate phase (33 °C, MEA, 3 weeks): Colonies 20 mm diam, velvety, cerebriform, tan to greyish-white. Hyphal elements moniliform 3.5–5.5 μm wide. Conidiophore-like structures (Fig. 13G, H) composed of swollen cells and arising from intercalary swollen cells 3.7–6.6 μm, sometimes producing conidia multilaterally; conidia 1.9–4.7 × 2.2–4.4 μm (3.2 ± 0.6 × 2.6 ± 0.4, n = 45), smooth-walled to finely roughened, sessile. Some giant cells present, 7.2–24.0 μm (12.8 ± 3.2, n = 40) diam. Thermotolerant phase (37 °C, MEA, 3 weeks): Colonies 6 mm, smooth, yeast-like, yellowish-white. Hyphal elements swollen producing giant cells 7.2–24.0 μm (12.8 ± 3.2, n = 40) diam and hyphae fragmenting to become large subspherical yeast cells 5.2–12.2 × 2.4–6.5 μm (8.1 ± 1.7 × 4.8 ± 0.9, n = 45), with uni- or bipolar budding at a broad base.
Comments: Dukik et al. (2017b) considered B. percursus as being closely related to B. dermatitidis and that is verified in our multilocus analyses. The two available strains, both from human sources, grouped with high support in all analyses. One strain failed to sporulate (CBS 142605 = NCPF 4091). The species is differentiated by producing large yeasts (greater than 5 μm in length) mostly from fragmenting hyphae within 2 weeks of incubation at 37 °C. At lower temperatures (24–33 °C), characteristic, spherical, sessile yeast-like cells are present along hyphae. The optimal sporulation temperature is 21–24 °C, conidia being absent at 30 °C. Growth occurs at 9 °C, optimum 27 °C, no growth at 40 °C.
Blastomyces silverae Y. Jiang, Sigler & de Hoog, sp. nov.—MycoBank MB821084 (Fig. 14)
Etymology: named after Canadian mycologist Eleanor Silver Keeping (née Dowding).
Blastomyces silverae (CBS 139879). A–C Colonies on MEA 3 weeks at 24, 33 and 37 °C. D–F 24 °C. Slightly swollen conidiophore with secondary conidiophore. G–I 33 °C. Swollen conidiophores and conidia. J–M 37 °C. Conidia, giant cells with thin or thick walls, large yeast-like cells with thick walls and broad-based budding. Scale bars: D–M = 10 μm
Holotype: Alberta, Canada, specimen of culture CBS 139885 (preserved in metabolically inactive condition under liquid nitrogen) from coyote dung, isolated by R.S. Currah, 3 May 1983; living strain CBS 139885 = UAMH 4470.
Saprobic phase (24 °C, MEA, 3 weeks): Colonies 68 mm diam, yellowish-white, floccose, radially sulcate; margin thin 3–5 mm wide, reverse warm-buff to light buff peripherally. Few spirally twisted hyphae present. Conidiophores cylindrical, about 1 μm wide at the base, with basal septum, often swollen near the tip, secondary conidiophores present. Conidia sessile, occasionally on pedicels when alongside hyphae, spherical, 2.2–5.6 × 1.8–5.6 μm (3.9 ± 0.6 × 3.8 ± 0.7, n = 50), smooth or slightly roughened, often adherent. Some broad-based budding cells may be present. Intermediate phase (33 °C, MEA, 3 weeks): Colonies restricted 16 mm diam, glabrous with central tufts of hyphae centrally; margin fimbriate. Hyphal elements moniliform, 3.7–4.5 μm wide. Conidiophores cylindrical, sometimes swollen, 2.0–4.6 μm wide. Conidia larger than at 24 °C, subspherical, 3.0–5.6 × 2.4–5.7 μm (4.4 μm ± 0.7 × 3.8 ± 0.7, n = 45), smooth-walled to slightly roughened, adherent to supporting structures. Some broad-based budding or giant cells may be present. Thermotolerant phase (37 °C, MEA, 3 weeks): Colonies restricted, 4–5 mm, cerebriform, glabrous to slightly hairy, yellowish-white to buff; hyphal elements partly swollen, 2.3–4.1 μm wide, densely septate. Conidiophores simple or absent. Conidia identical to those at 33 °C. Some thick-walled, broad-based budding cells or giant cells with thin walls measuring 9.2–28.7 μm (13.3 ± 3.3, n = 44) may be present.
Comments: Some strains in B. silverae were identified initially as Ea. parva (Sigler 1996) but analysis of partial ITS and LSU sequences (Peterson and Sigler 1998) and results of our study determined that they cluster separately from Ea. parva. Blastomyces silverae produces conidia over a temperature range of 21–37 °C; giant cells and yeast-like cells are produced at 36–37 °C but conversion time is slow, requiring about 3–4 weeks. One strain (CBS 139871 = UAMH 4489) deviated by the producing diffusing brown pigment when grown at 24 °C. A seventh strain (CBS 509.78 = ATCC 32539) was sister to the main bootstrap-supported cluster, and it showed some phenotypic differences. The strain did not grow at 6 °C and formed giant cells more quickly at 37 °C. Four strains come from human and animal sources but pathogenicity has not been demonstrated for this species. Optimum sporulation temperature is 24–27 °C, with conidia present until 37 °C. Growth occurs at 6 °C, no growth at 40 °C.
Emergomyces Dukik, Sigler & de Hoog—Mycoses 60: 304. 2017. MycoBank MB818569.
Colonies yellowish-white. Hyphae hyaline. Conidiophores short, unbranched, arising at right angles from vegetative hyphae, slightly swollen at the top. Conidia holothallic, on short stalks. At 37 °C small yeast cells are produced, less than 5 µm in diam, budding at narrow bases; large, broad-based budding cells occasionally present. No growth at 40 °C.
Type species: Emergomyces pasteurianus (Drouhet, Guého & Gori) Dukik, Sigler & de Hoog
Comments: Emergomyces was described recently for Emmonsia pasteurianus and Es. africanus (Dukik et al. 2017b). Our data show these species grouping with two novel species in a well-supported lineage (Clade I-3, Fig. 2) (ML/BI 100/1.00). Strains of all species come from human specimens (Table 1) and produce small yeast cells as the thermotolerant phase. A second strongly supported subclade within Clade I comprises Emmonsia crescens (Figs. 2, 3). These subclades are supported by ecological and phenotypic differences distinguishing Emergomyces from Emmonsia crescens, i.e. prevalence of human rather than animal (largely rodent) hosts and conversion to yeasts rather than adiaspores. Emergomyces species are associated with disease in immunocompromised hosts especially among those with HIV infection (Schwartz et al. 2015a, b).
Emergomyces africanus Dukik, Kenyon, Schwartz, Govender & de Hoog—Mycoses 60: 305, 2017. MycoBank MB818571 (Fig. 15)
= Emmonsia sp., Kenyon et al. – N. Engl. J. Med. 369: 1416. 2013.
Emergomyces africanus (CBS 139543). A–C Colonies on MEA 3 weeks at 24, 33 and 37 °C. D–F 24 °C. Slightly swollen conidiophore without secondary conidiophores. G–I 33 °C. Hyphae short and swollen, disarticulating to liberate giant cells. J–M 37 °C. Scant, swollen hyphae, abundant small yeast cells with unipolar budding at narrow base. Scale bars: D–M = 10 μm
= Emmonsia sp. 5, Schwartz et al. – PLoS Pathog. 11: e1005198, p. 2. 2015.
Type culture: CBS 136260, New Somerset Hospital, Cape Town, South Africa, isolated from skin biopsy of an HIV-infected male, collected by N.P. Govender, 11 June 2010.
Saprobic phase (24 °C, MEA, 3 weeks): Colonies 26 mm diam, glabrous to floccose centrally, yellowish white, radially sulcate, reverse warm-buff to light buff peripherally. Few helical hyphae present. Conidiophores 0.9–3.3 (1.6 ± 0.3) μm wide with a septum at the base and at conidial insertion; moderately swollen at the tip with 4–8 conidia borne on narrow pedicels. Conidia single or in short chains (2–4), subspherical, 1.2–3.2 × 1.7–3.8 μm (2.2 ± 0.5 × 2.7 ± 0.5, n = 45), smooth to finely roughened. Intermediate phase (33 °C, MEA, 3 weeks): Colonies 14 mm, yeast-like, smooth, yellowish-white. Hyphae densely sepate, somewhat moniliform, 1.6–3.0 μm wide. Conidia absent. Conidiophore-like cells arising from hyphae, 5–15 × 1.5–2.2 μm (Fig. 15G). Small ovoidal yeast-like cells produced from irregularly shaped adiaspore-like cells, 3.9–10.0 μm (5.7 ± 1.3, n = 44) (Fig. 15H, I) or from tip of conidiophore-like structures, ovoidal, 1.7–5.3 × 0.9–2.2 μm (2.9 ± 0.7 × 1.6 ± 0.3, n = 45). Thermotolerant phase (37 °C, MEA, 3 weeks): Colonies 7 mm diam, yeast-like, cerebriform, yellowish-white. Hyphae scant. Yeast cells abundant, ovoidal to subspherical, 1.7–5.3 × 0.9–2.2 μm (2.9 ± 0.7 × 1.6 ± 0.3, n = 45) with unipolar budding at a narrow base.
Comments: The species is distinguished by development of secondary conidiophores which lead to a complex cluster of four to eight conidia and production of small-celled yeasts at 37 °C within 1 week. Optimal sporulation temperature is 21–24 °C; conidia are absent at 30 °C. Grows at 6 °C, optimum 24 °C, no growth at 40 °C.
Emergomyces canadensis Y. Jiang, Sigler & de Hoog, sp. nov.—MycoBank MB821102 (Fig. 16)
= Emmonsia sp., Sigler et al. – J. Clin. Microbiol. 36: 2920. 1998.
Emergomyces canadensis (CBS 139872). A–C Colonies on MEA 3 weeks at 24, 33 and 37 °C. D–F 24 °C. Conidiophores, conidia and secondary conidiophores. G–I 33 °C. Swollen conidiophores and subspherical conidia; hyphal cells and conidia in part swell and become giant cells with thin walls. J–M 37 °C. Scant, swollen hyphae, abundant yeast cells with budding at narrow base. Scale bars: D–M = 10 μm
= Emmonsia sp. 2, Schwartz et al. – PLoS Pathog. 11: e1005198, p. 2. 2015.
Etymology: referring to the country of origin of the type specimen.
Holotype: Saskatchewan, Canada, specimen of culture CBS 139872 (preserved in metabolically inactive condition under liquid nitrogen) from skin lesions of HIV-positive patient, H. Congly, June 1992; living strain CBS 139872 = UAMH 7172.
Saprobic phase (24 °C, MEA, 3 weeks): Colonies 30 mm diam, yellowish white, cottony to glabrous, with tufts of hyphae centrally, radially sulcate, reverse ochraceous-buff to warm-buff peripherally. Few helical hyphae present. Conidiophores 1.4–2.5 μm (1.7 ± 0.3) wide with septum at the base, cylindrical or slightly swollen in the middle and at the tip, bearing 1–2 conidia on narrow pedicels (< 1 μm long). Conidia subspherical, smooth to slightly roughened 2.1–3.8 × 1.8–3.4 μm (2.8 ± 0.4 × 2.7 ± 0.3, n = 45). Intermediate phase (33°C, MEA, 3 weeks): Colonies 9 mm diam, cerebriform, felty, yellowish-white. Hyphae broader 2.8–5.8 μm wide. Conidiophores mostly swollen, ampulliform; septa absent or present at the base, 1.7–4.6 μm wide. Conidia spherical, 2.3–7.1 μm (4.4 ± 1.1, n = 45). Hyphal cells and conidia in part swelling and becoming giant cells, 5.3–11.3 μm (7.8 ± 1.6, n = 40). Small yeast cells present, some originating by budding from giant cells. Thermotolerant phase (37 °C, MEA, 3 weeks): Colonies restricted, about 3 mm, yeast-like, smooth, yellowish-white. Yeast cells abundant, spherical, 2.2–4.8 μm (3.5 ± 0.63, n = 45) with uni- or bipolar budding at narrow base. Few short, swollen hyphal elements and giant cells present.
Comments: Emergomyces canadensis is closely related to Es. orientalis (Wang et al. 2017; Figs. 2, 3) and both species come from human sources (in Canada and China). At 37 °C they both produce small spherical yeast cells below 5 μm in diam. Urease broth test remained negative even after a week of incubation, and a red pigment is produced on BHI and TSA at 37 °C. Their distinction as separate species is supported by differences between their sequences, including 7 nucleotide positions in ITS, 8 in rPB2 and 17 in TUB2, 7 in TEF3, and by phenotypic differences. Optimal sporulation temperature for Es. canadensis is 24–27 °C, while Es. orientalis produced conidia only at 21 °C. Emergomyces canadensis produces small budding yeast cells at 33 °C and time for conversion at 37 °C is fast (1 week), while Es. orientalis budding occurs at lower temperature (> 30 °C) and conversion at 37 °C is slow (2 weeks). Compared to remaining species in the Ajellomycetaceae these differences exceed the maintained species limit. Despite the near-absence of phenotypic characters we therefore keep them as separate species pending availability of more material.
Emergomyces europaeus Y. Jiang, Sigler & de Hoog, sp. nov. – MycoBank MB821103 (Fig. 17)
= Emmonsia sp., Wellinghausen et al. – Int. J. Med. Microbiol. 293: 441–445. 2003.
Emergomyces europaeus (CBS 102456). A–C Colonies on MEA 3 weeks at 24, 33 and 37 °C. D–F 24 °C. Slightly swollen conidiophore without secondary conidiophores; conidia slightly roughened. G–I 33 °C. Swollen hyphae disarticulating to liberate giant cells; small yeasts produced. J–M 37 °C. Scant, irregularly swollen hyphae, small yeast cells with budding at narrow base. Scale bars: D–M = 10 μm
= Emmonsia sp. 6, Schwartz et al. – PLoS Pathog. 11: e1005198, p. 2. 2015.
Etymology: referring to the origin of the type specimen.
Holotype: Ulm, Germany, specimen of culture CBS 102456 (preserved in metabolically inactive condition under liquid nitrogen) from transbronchial biopsy of 64-year-old male farmer with chronic granulomatous lung infection, G. Haase, 2003; living culture CBS 102456 = UAMH 10427.
Saprobic phase (24 °C, MEA, 3 weeks): Colonies 30 mm diam, dense, white, felty to floccose, radially sulcate, glabrous at the margin; reverse warm-buff to light buff periphally. Few helical hyphae present. Conidiophores unbranched, 0.9–2.3 μm (1.6 ± 0.2) wide, with septum at the base, cylindrical to slightly swollen at the tip, bearing one or two conidia. Conidia subspherical 2.9–5.7 × 3.0–5.7 μm (3.9 ± 0.6 × 4.2 ± 0.7, n = 45) slightly roughened. Intermediate phase (33 °C, MEA, 3 weeks): Colonies 10 mm diam, yeast-like, pasty, cerebriform, tan. Hyphae scant, densely septated and somewhat inflated, 1.8–4.8 μm wide, forming slightly swollen conidiophore-like cells 1.2–3.8 μm wide. Conidia absent. Small yeast cells present with uni- or bipolar budding from narrow base, ovoidal, 2.6–5.9 × 1.7–3.8 μm (3.7 ± 0.6 × 2.6 ± 0.5, n = 45), developing from swollen conidiophore-like cells or from giant cells 4–10 μm (7.2 ± 1.3, n = 44). Thermotolerant phase (37 °C, MEA, 3 weeks): Colonies strongly inhibited, 4 mm diam, with features similar to those at 33 °C. Swollen hyphae and giant cells present; yeast cells ovoidal to subspherical, 2.6–5.9 × 17–3.8 μm (3.7 ± 0.6 × 2.6 ± 0.5, n = 45), with uni- or bipolar budding at a narrow base.
Comments: This species is known from a single strain from a human source and the isolate was thought initially to be Es. pasteurianus but its conidial phenotype was different (Wellinghausen et al. 2003). In our multilocus analyses, Es. europaeus differs from Es. pasteurianus and Es. africanus in 10 positions in ITS, and in (14, 19) and (29, 30) positions in rPB2 and TUB2. The three species are similar at 37 °C in producing ovoidal to subspherical yeast cells of less than 5 μm in diam, but the saprobic phases differ. Emergomyces europaeus has conidiophores without secondary branches and conidia are larger and more roughened than those of other species. Yeast transformation occurs at a lower temperature (33 °C). The Es. europaeus strain is the only one among all Emergomyces species in its source from an immunocompetent individual, although the patient had been on prolonged low-dose corticosteroid therapy. This strain was isolated from a lung lesion rather than from skin, which is more common in the other two species. Transfer time is short, only 1 week being needed for total conversion at 37 °C .The optimal sporulation temperature is 21–24 °C; conidia are absent at 30 °C. Grows at 6 °C, optimum 24 °C, no growth at 40 °C.
Emergomyces orientalis P. Wang, Y.C. Xu & H.W. Fan—Mycoses 60: 316. 2017. MycoBank MB517245 (Fig. 18)
= Emmonsia sp. 7, Schwartz et al. – PLoS Pathog. 11: e1005198, p. 2. 2015.
Emergomyces orientalis (CBS 124587). A–C Colonies on MEA 3 weeks at 24, 33 and 37 °C. D–F 21 °C. Slightly swollen conidiophores and coiled hyphae; conidia present (absent at 24 °C); G–I 33 °C. Swollen conidiophores and hyphae forming giant cells with thin walls. J–M 37 °C. Hyphae scant, swollen; small spherical yeast cells with budding at narrow base. Scale bars: D–M = 10 μm
Type culture: CBS 124587 = Peng 5Z489 = CGMCC2.4011, Shanxi, China, isolated from sputum of a 64-year-old male with type-2 diabetes mellitus by P. Wang, 2005.
Saprobic phase (21 °C, MEA, 3 weeks): Colonies 23 mm diam, yellowish white, felty with hyphal tufts centrally, radially sulcate, reverse ochraceous-buff to warm-buff peripherally. Coiled hyphae frequently present (Fig. 18F). Conidiophores cylindrical or slightly swollen in the middle, with a septum at the base, 1.7–3.7 (2.8 ± 0.7) μm, with 1–2 thin secondary conidiophores. Conidia subsphaerical, 1.1–2.8 × 1.7–4.8 μm (1.9 ± 0.4 × 2.8 ± 0.6, n = 40), smooth to slightly roughened; absent at 24 °C. Intermediate phase (33 °C, MEA, 3 weeks): Colonies 12 mm diam, yeast-like, cerebriform, yellowish-white. Hyphae somewhat inflated, 2.7–5.0 μm wide, constricted at septa. Part of inflated hyphal cells and conidiophores swelling to become giant cells, 5.7–12.7 μm (8.5 ± 1.7, n = 26) diam, sometimes with narrow-based budding. Yeast cells spherical, 2.0–4.5 (3.4 ± 0.6, n = 45) diam. Thermotolerant phase (37 °C, MEA, 3 weeks): Colonies restricted, 5 mm diam, yeast-like, cerebriform, yellowish white. Hyphal elements scant. Yeast cells spherical, 2.0–4.5 (3.4 ± 0.64, n = 45) diam, with uni- or bipolar budding at a narrow base. Few giant cells present.
Comments: This species, known only from the type strain (CBS 124587), produced conidia at 21 °C on MEA but not at 24 °C. Conversion to yeast stage occurs at lower temperature (33 °C) and time for transformation to yeast at 37 °C is short (1 week). See comments under Es. canadensis for comparisons between these two species. Grows at 6 °C, optimum 24 °C, no growth at 40 °C.
Emergomyces pasteurianus (Drouhet, Guého & Gori) Dukik, Sigler & de Hoog – Mycoses 60: 304. 2017. MycoBank MB517245 (Fig. 19)
≡ Emmonsia pasteuriana Drouhet, Guého & Gori – J. Mycol. Méd. 8: 70. 1998.
Emergomyces pasteurianus (CBS 101426). A–C Colonies on MEA 3 weeks at 24, 33 and 37 °C. D–F 24 °C. Slightly swollen conidiophores and secondary conidiophores. G–I 33 °C. Conidiophores and hyphal fragments becoming swollen and disarticulating. J–M 37 °C. Hyphae producing giant cells; small and large yeast cells with uni- or bipolar budding. Scale bars: D–M = 10 μm
= Emmonsia sp. 4, Schwartz et al. – PLoS Pathog. 11: e1005198, p. 2. 2015.
Type culture: CBS 101426 = UAM H9510 = NCPF 4236 = IP 2310.95, Italy, isolated from disseminated cutaneous disease in a 40-year-old HIV+ female by E. Guého, April 1999.
Saprobic phase (24 °C, MEA, 3 weeks): Colonies 38 mm diam, yellowish white, dense, felty to floccose, radially sulcate, with thin margin, reverse ochraceous-buff to warm buff peripherally. Few helical hyphae present. Conidiophores with septa at the base and at conidial insertion, at base 0.6–1.7 μm (1.2 ± 0.3) wide, cylindrical or moderately swollen at the tip. Conidia formed singleton narrow pedicels (ca. 1 μm long) or in short chains (2–4), subspherical, 0.9–2.8 × 1.8–3.2 μm (2.0 ± 0.4 × 2.5 ± 0.4, n = 45), smooth-walled to finely roughened. Some chlamydospore-like cells arising terminally on short lateral branches, with thickened walls and often with a median septum. Intermediate phase (33 °C, MEA, 3 weeks): Colonies 21 mm diam, cerebriform, yellowish white. Hyphae scant and ampulliform conidiophore-like cells becoming somewhat inflated and disarticulating. Conidia absent. Thermotolerant phase (37 °C, MEA, 3 weeks): Colonies restricted, 8 mm diam, yeast-like, cerebriform, yellowish white. Hyphae scant, moniliform, some cells becoming giant cells, 5.4–12.0 μm (8.4 ± 1.6, n = 42) wide (Fig. 19K). Yeast cells arising from giant cells or from fragments of swollen conidiophores or hyphae; small yeasts with narrow-based budding, 2.1–5.1 × 1.6–4.2 μm (3.7 ± 0.7 × 2.9 ± 0.6, n = 44); larger yeasts 5.0–11.2 × 2.4–6.3 μm (8.0 ± 1.7× 4.7 ± 0.80, n = 40), with uni- or bipolar budding from narrow or broad bases,
Comments: Emergomyces pasteurianus was first reported from a case of disseminated infection with secondary cutaneous lesions in a female with advanced HIV disease and a history of injection drug use in Italy in 1994 (Gori et al. 1998). The species produces small yeasts with narrow based budding and large yeasts with broad-based budding at 37 °C, characteristic of Emergomyces species and Blastomyces species, respectively. Conversion to yeast is slower (2–3 weeks) and occurs at higher temperature than in Es. africanus and Es. europaeus. Conidia occur at 21–24 °C but are absent at 30 °C, except strain CBS 140361 that deviates by conidiation until 33 °C and formation of a brown pigment. Growth at 6 °C, optimum 24 °C, no growth at 40 °C.
Emmonsiellopsis Marín, Stchigel, Guarro & Cano – Mycoses 58: 455. 2015. MycoBank MB811334
Type species: Emmonsiellopsis terrestris Marín, Stchigel, Guarro & Cano.
Colonies whitish to cream, pale brownish near the centre. Hyphae hyaline, sometimes contorted to dendritic. Conidia ovoidal to clavate, smooth walled and verrucose or spinulose born laterally or terminally on conidiophores, sessile on vegetative hyphae or on pedicels.
Comments: The genus Emmonsiellopsis was differentiated from other members of Ajellomycetaceae in part by absence of thermal dimorphism (Marin-Felix et al. 2015). Its distinction as a separate genus is supported in our multilocus analysis (ML/BI 100/0.99). In the present study both species were found to transform to large yeast-like cells.
Emmonsiellopsis coralliformis Marín, Stchigel, Guarro & Cano – Mycoses 58: 455. 2015. MycoBank MB811335 (Fig. 20)
Type culture: CBS 137500 = FMR 4024 (holotype: CBS H-21624), Girona, Spain, ex fluvial sediment, P. Vidal, June 1991.
Emmonsiellopsis coralliformis (CBS 139500). A–C Colonies on MEA 3 weeks at 24, 30 and 33 °C. D–F 24 °C. Conidiophores lacking secondary conidiophores; conidia roughened. G–I (30 °C). Conidia, swollen cells andbudding cells present. J–M 33 °C. Swollen hyphae and conidia becoming giant cells with rather thick walls, some showing broad-based budding. Scale bars: D–M = 10 μm
Saprobic phase (24 °C, MEA, 3 weeks): Colonies 35 mm diam, floccose, somewhat cerebriform to sulcate, yellowish white, reverse warm-buff. Conidiophores cylindrical, about 1 μm wide, somewhat flexuose (20F), lacking secondary conidophores. Conidia solitary, mostly sessile, subspherical, 2.9–4.9 × 2.3–4.6 μm (3.9 ± 0.5 × 3.6 ± 0.60, n = 45), roughened to spinulose. Intermediate phase (30 °C, MEA, 3 weeks): Colonies 21 mm diam, yellowish white, yeast-like to slightly hairy, cerebriform. Conidia-like cells swollen, 3.1–6.5 × 2.5–5.4 μm (4.6 ± 0.8 × 3.7 ± 0.7, n = 45); some with broad-based budding. Thermotolerant phase (33 °C, MEA, 3 weeks): Colonies restricted 5 mm diam, yellowish white, yeast-like, cerebriform. Hyphal cells and conidia becoming swollen, forming giant cells, 7.1–17.3 μm (10.7 ± 2.2, n = 45) diam, with some broad-based budding, uni- or bipolar, occasionally multilateral. Conidia spherical, 2.0–3.8 μm (3.0 ± 0.4, n = 45) diam often sessile on supporting structures (Fig. 20K).
Comments: This species is known only from the type strain which has a lower transformation temperature, starting at 30 °C with formation of giant cells, which develop into thicker-walled adiaspore-like cells (less than 20 μm diam) that may bud at a broad base (33 °C). Conversion time is slow (4–5 weeks). Grows at 9 °C, optimum 24 °C, no growth above 33 °C.
Emmonsiellopsis terrestris Marín, Stchigel, Guarro & Cano – Mycoses 58: 456. 2015. MycoBank MB811598 (Fig. 21)
Type culture: CBS 273.77 = FMR 13882 = UAMH 2304 (holotype CBS H-22118), Phillips County, Kansas, U.S.A., ex barnyard, C.W. Emmons, Jan. 1965.
Emmonsiellopsis terrestris (CBS 273.77). A–C Colonies on MEA 3 weeks at 24, 33 and 37 °C. D–F 24 °C. Conidiophores short, slightly swollen, without secondary conidiophores conidia sometimes sessile, roughened. G–I 33 °C. Conidia. J–M 37 °C. Conidia, some becoming giant cells with occasional budding at a broad base. Scale bars: D–M = 10 μm
Saprobic phase (24 °C, MEA, 3 weeks): Colonies 58 mm diam, cottony-floccose, yellowish white; reverse warm-buff to pale buff peripherally. Conidiophores cylindrical, about 1 μm wide at the base, sometimes slightly swollen or flexuous, producing 1–4 (up to 7) solitary conidia; secondary conidiophores lacking. Conidia spherical, 2.8–7.3 × 2.2–4.9 μm (4.9 ± 0.9 × 3.8 ± 0.70, n = 45), roughened to spinulose, sessile, often adherent to the conidiophore, some budding at a broad base. Intermediate phase (33 °C, MEA, 3 weeks): Colonies 30 mm diam, cottony-floccose, yellowish white, sulcate. Conidiation moderate; conidia slightly larger than at 24 °C. Thermotolerant phase (37 °C, MEA, 3 weeks): Colonies restricted 7 mm, cerebriform, glabrous to hairy, greyish-white. Hyphal elements moniliform. Conidia 3.0–7.7 × 2.8–6.9 μm (5.3 ± 1.1 × 4.3 ± 1.0, n = 45), enlarging to form giant cells 5.7–23.0 μm (14.4 ± 4.3, n = 44) diam; with uni- or bipolar, sometimes multilateral, budding at a broad base. Conversion reluctant, taking almost 4–5 weeks on MEA.
Comments: Emmonsiellopsis species are similar in lacking secondary conidiophores. Emmonsiellopsis terrestris differs from Emms. coralliformis in conversion temperature at or beyond 37 °C but time of conversion is slow in both species (4–5 weeks). Optimal sporulation temperature: 24–30 °C. Growth occurs at 9 °C, optimum 21 °C, no growth at 40 °C.
Discussion
Over the last two decades, several new genera and species have been added to the family Ajellomycetaceae suggesting the need for a comprehensive review of phylogenetic relationships and morphological features among taxa in this important family of vertebrate pathogens. In particular, several fungi having emmonsia-like morphologies have emerged as causative agents of human disease in case reports globally (Schwartz et al. 2015a, b; Dukik et al. 2017b) (Fig. 22). In the present paper we resolved phylogenetic relationships among Emmonsia and emmonsia-like fungi, which proved to be closely similar to Blastomyces and Emergomyces and to a less extent to other members of Ajellomycetaceae, using concatenated sequence data of the loci LSU, ITS, TUB2, TEF3, and rPB2.
As shown in earlier phylogenetic overviews (Peterson and Sigler 1998; Brown et al. 2013, Schwartz et al. 2015b), the generic type species Emmonsia parva, with epitype strain CBS 139881 (= UAMH 130) clusters in the Blastomyces clade (Clade II). The genus Blastomyces has recently been validated (de Hoog et al. 2016a) with B. dermatitidis as type species and strain CBS 674.68 (= ATCC 18188 = UAMH 3539) as epitype of B. dermatitidis. Emmonsia is now a younger synonym of this genus. As circumscribed in the present study, Blastomyces now comprises five species having strong support: B. dermatitidis/B. gilchristii (ML/BI 100/1.00), B. helicus (ML/BI 100/1.00), B. parvus (ML/BI 100/1.00), B. percursus (ML/BI 100/1.00) and B. silverae (ML/BI 98/1.00). While B. gilchristii (ML/BI 100/1.00) could not be distinguished from B. dermatitidis in our study with standard barcoding by ITS (Fig. 4), the species has been segregated on the basis of genealogical concordance phylogenetic species recognition (GCPSR; Brown et al. 2013) using seven partial genes, some of which are not commonly used in phylogenetic studies (e.g. histidine kinase, orotidine 5’-phosphate decarboxylase). Blastomyces gilchristii was found to be reproductively and genetically isolated from B. dermatitidis and had a smaller, overlapping geographic distribution. Similar diversity had been noted by Meece et al. (2011). Whole genome sequencing of strains of B. dermatitidis (ER-3, ATCC 18188 and ATCC 26199) and B. gilchristii (SLH14081) provided additional evidence for separation of these species (Muñoz et al. 2015).
Species of the genus Blastomyces are associated with human and animal disease. Blastomyces dermatitidis and B. gilchristii cause disease pulmonary or disseminated disease in immunocompetent humans, companion animals and other mammals (Baumgardner et al. 1995; Bariola et al. 2010; Saccente and Woods 2010; Brown et al. 2013). B. helicus is associated with disease in predominantly immunocompromised humans and in companion animals (Kappagoda et al. 2017; Rofael et al. 2018; Schwartz et al. 2017) whereas known cases of B. percursus infection include immunocompetent and immunocompromised hosts (Kemna et al. 1994; Heys et al. 2014; Dukik et al. 2017b). B. parvus causes pulmonary disease only in rodents and terrestrial mammals; no human cases are substantiated (Sigler 2005; Anstead et al. 2012). All species are shown here to produce moderate to large yeast cells at higher temperatures in culture that replicate by broad-based budding, including, to a lesser extent, B. parvus in which giant cells may propagate as broad based yeast cells.
With the transfer of the type species, Ea. parva to Blastomyces, as proposed herein, the disposition of Ea. crescens is not resolved. Strains of Ea. cresens clustered with strong support in a single, moderately variable clade in both ML and rPB2 analyses. The clade was paraphyletic to clade I-3 comprising the genus Emergomyces but Ea. crescens differs from all members of the Emergomyces clade in not demonstrating budding, in host associations and pathogenicity. Herein we continue to use the current name to avoid nomenclatural instability. Emmonsia crescens has been recorded from all five continents, yet our study confirms data of Peterson and Sigler (1998) in showing two molecular subgroups within the species: a panboreal Eurasian population, and a North American population. Sequence differences are small, and there are no significant phenotypic differences between the two populations and Sigler (1996) showed interfertility among several strains of the two populations. The observed structuring of the two populations may indicate that the vector of dispersal of Ea. crescens is slow and local, in line with mammal vectoring and leading to a gradual separation of Eurasian and American genotypes.
Emmonsia crescens occurs in animal and, less commonly, in human hosts in which it forms large non-replicating, thick-walled adiaspores in lung and occasionally other tissues. Almost all data on this species’ host and geographic distribution come from animal surveys in which lungs and other tissues of trapped animals have been dissected and examined for presence of the large adiaspores (up to 400 µm diam) characteristic of this species (Dvořak et al. 1973; Emmons and Jellison 1960; Hubalek 1999; Hubálek et al. 1995, 1998; Jellison 1969; Sigler 2005; Borman et al. 2009, 2018). These adiaspores are pathognomonic and can be recognized in published reports even in the absence of preserved material. Over 120 species of mammal hosts have been identified and include members of the orders Insectivora, Edentata, Lagomorpha, Rodentia and Carnivora. Terrestrial rodents and burrowing animals are most commonly represented. Rare reports concern domestic or farm animals including two dogs (Al-Doory et al. 1971; Koller et al. 1976), a goat (Koller and Helfer 1978), a horse (Pusterla et al. 2002) and a deer (Matsuda et al. 2015), although no voucher material is available to confirm the diagnoses. Approximately 50 cases of human pulmonary adiaspiromycosis have been reported from the Americas, Europe, and North Africa (England and Hochholzer 1993; Sigler 1998; Anstead et al. 2012). Dvořak et al. (1973) suggested that growth of the saprobic phase is more important as determinant of the life cycle than the host itself. They suggested that localization of Ea. crescens in the biotope of the animal burrow in temperate to subboreal climates is a significant ecological prerequisite. The limited number of conidia produced may contaminate the air inside the burrow sufficiently to provide an effective inoculum, thus enabling the animal to transfer conidia to another site having the same ecological parameters. The species has a wide range of temperature tolerance, from 6 °C which is comparable to winter temperatures of burrows at around 3–5 °C, and up to 40 °C at which it grows restrictedly (Fig. 5). At higher temperatures, fast-growing saprobes may suppress the growth of Ea. crescens, which explains that highest incidence of infection occurs in winter and early spring (Dvořak et al. 1973). In accordance with this hypothesis, we found that conidiation in Ea. crescens occurred at 21 °C and below.
The recently-described genus Emergomyces (Clade I-3) is here confirmed to include taxa exclusively associated with human disease, including the previously described species Es. pasteurianus [formerly Ea. pasteuriana] (Gori et al. 1998; Dukik et al. 2017b)], Es. africanus (Dukik et al. 2017b), Es. orientalis (Wang et al. 2017) and the new species Es. canadensis and Es. europaeus. The species differ slightly with respect to maximum growth temperatures and sizes of yeast cells as well geographic distribution. Both Es. canadensis (Clade I-3a) and Es. orientalis (Clade 1-3b; Wang et al. 2017) have been isolated from human patients (North America and China respectively), produce small, spherical yeast cells at 37 °C and a unique red pigment on BHI and TSA at 37 °C. Differences occur with respect to mold-to-yeast transitions: in Es. canadensis small yeasts are produced at 33 °C which reproduce by budding (Fig. 16H), while in Es.orientalis beyond 30 °C yeast-like cells originate from swollen hyphal cells (Fig. 18I). Additional isolates are needed to gain a better understanding of the relationship between these two species.
According to the above, a range is observed from the adiaspore of Ea. crescens to the small yeast of Emergomyces, and from reluctant budding in Ea. sola (NCPF 4289) to abundant budding in Emergomyces. Although we could not satisfactorily reconstruct the evolutionary process, in line with the different forms of the parasitic phase, the species seem to acquire certain ecologically relevant capacities, e.g. to infect and colonize mammal hosts; this situation seems to open new opportunities in terms of expanding its traditional ecological niche and exerting pressures for speciation (Coyne and Orr 2004). The significance of Ea. crescens colonizing lungs of burrowing animals is unknown. From a longer evolutionary perspective, what is its role in the fungus’s adaptation and speciation? The adiaspore is so unique that it is unlikely to be coincidental. Ea. crescens might be an interesting model for study that may serve as a useful comparator to other dimorphic fungi.
The morphological features of thermal transformation among members of the Ajellomycetaceae have been central to the taxonomy and ecology of the family. Among the species examined here, those having a long saprobic phase, i.e. expressing conidiation over a broad temperature range and having a short thermotolerant phase take more time to convert (Fig. 5, Table 2). It is unknown whether this can be linked to virulence (Medoff et al. 1986). Strains with relatively long saprobic phases almost invariably originated from rodents and their burrows. Species that were regularly found in human hosts were more often able to grow at 40 °C or higher, had a broader temperature range of the thermotolerant phase, and required less time to convert. For example, B. helicus CBS 139874, from CSF of a patient with fungal meningitis, tolerated 40 °C, required 1 week to convert at 37 °C, and was noted to have already converted at 33 °C. Mammals have different body temperatures, ranging from 35 to 40 °C, and significant differences are observed between species e.g. armadillo, rodent and feline hosts (Fig. 23). Emmonsia crescens and B. parvus are both associated with rodents. In Ea. crescens, these are small terrestrial animals and rodents: moles (Talpa europaea), dark polecats (Trichosurus vulpecula), European water voles (Arvicola amphibius), common voles (Microtus arvalis), red-backed voles (Myodes sp.), field mice (Apodemus sp.) and deer mice (Peromyscus sp.) (Dvořak et al. 1973), among others, which have relatively low body temperatures of 35.0–37.5 °C. Hosts of B. parvus have somewhat higher body temperatures, e.g. Neotama micropus (CBS 139882 = UAMH 134) and pocket gopher (CBS 178.60 = ATCC14051), that are 36–40 °C. Similarly B. silverae (CBS 139879 = UAMH 139), originated from a weasel with a body temperature of 37.8–40 °C. No infections caused by members of the Ajellomycetaceae are known to occur in cold-blooded hosts.
Approximate relationships between optimal conversion temperatures of ajellomycetaceous species and their prevalent host animal, with average body temperatures listed in the right column. Several novel species are represented by very few isolates so that only scant information on host ranges is available
Inferring the evolutionary histories on traits of thermal conversion, fungal pathogens could have evolved with their specific animal and human hosts. Emmonsiellopsis, found in an ancestral position, has no known animal hosts but has a predominantly filamentous life cycle, and reluctant conversion. The prime host of Paracoccidioides is the armadillo, a terrestrial mammal with a primitive immune system and a low body temperature of 34 °C (Bagagli et al. 2006, 2008; Hrycyk et al. 2018). The prime host of Histoplasma is the bat, with a very low winter body temperature which rises to 37.8–40.6 °C during flight. Most rodents that are hosts for Ea. crescens have temperatures ca. 35–37 °C. Blastomyces dermatitidis is commonly found in felines and canines, which have slightly higher body temperatures (38–40 °C) and a more advanced immune system. Emergomyces clade comprises species in humans (body temperature 36.1–37.2 °C) and as yet no known animal or environmental alternative cycle.
It should be noted that diverse types of giant cells occur at elevated temperatures across most taxa in Ajellomycetaceae, even the genus Emmonsiellopsis described by Marin-Felix et al. (2015). Two strains of Emms. terrestris (CBS 139889 = UAMH 141; CBS 273.77 = UAMH 2304) examined by Sigler (1996) and reexamined in the present study were able to produce giant cells at 37 °C. Already in 1947, Dowding suggested that Emmonsia parva was more close to Blastomyces than to Ea. crescens, as was substantiated with molecular data by Perterson and Sigler (1998) and in the present paper. In line with this, the giant cells of B. parvus (Emmonsia parva) appeared not to be a terminal (i.e., non-replicating) structure in the fungus’ pathogenic phase, as they are in Ea. crescens, but are able to propagate as broad-based yeast cells, very similar to those of classical Blastomyces species (Fig. 3). Wu et al. (2005) reviewed five cases of blastomycosis that presented histopathologically with very large budding cells (20–40 μm diam). Paracoccidioides brasiliensis appears as spherical budding cells with multiple buds; younger cells measure 2–10 μm in diam, and mature cells often > 30 μm, or even reach 60 μm in diam (Brummer et al. 1993). The biological coherence of taxa in Ajellomycetaceae is underlined by teleomorphs. Ajellomyces-type sexuality has been obtained after matings in H. capsulatum, in B. dermatitidis, and in Ea. crescens and the morphologies of the ascomata and ascospores is relatively homogeneous over the species. All these features show significant overlap and apparent atavisms, suggesting that all species might be attributed to a single evolutionary series, i.e. classified in a single genus. However, such a taxonomic decision resulting in many name changes of clinically important pathogens is undesirable because it could sow confusion among clinicians, clinical laboratories and patients about these fungi and the diseases that they cause.
With sequence data of the loci LSU, ITS, TUB2, TEF3, and rPB2, 80 strains were analyzed, of which 44 clustered in Blastomyces/Emergomyces, and with representative isolates from classical genera of dimorphic fungi, i.e. Histoplasma (n = 7) and Paracoccidioides (n = 3). Despite our best effort to obtain a complete sequence dataset for all the genes and strains employed, the sequence of some genes, especially the nuclear protein-coding gene TEF3, could not be determined for a small percentage of strains because of failure in the PCR amplification or sequencing reactions. Interestingly, all unsuccessful reactions in TEF3 originated from Es. africanus. Specifically, 2.5, 1.25, 2.5, 7.5 and 5.0% of the total 80 strains employed failed in the sequence determination of the LSU, ITS, rPB2, TEF3 and TUB2 genes, respectively. This problem is known from all groups of fungi (Schoch et al. 2012). Another study has shown that an inferred phylogeny is not sensitive to 25% or even 50% missing data for sufficiently large alignments (e.g., ~ 30,000 positions and 36 species; Philippe et al. 2004). Though the length of the five-gene alignment in this study comprises only about 3302 positions, the amount of missing data is much less. Thus, we assume that the relative minor amount of missing data in our study will not significantly influence the reliability of the resulting phylogeny. The phylogenies of the taxa compared in this study were inferred from analyses using different data sets and algorithms. The topologies of the trees constructed using different algorithms performed on different data sets were largely congruent, which make the delimitation of major lineages and clades more clear and confident. A recent study of fewer representatives by whole genome sequencing data constructed a tree of similar topology to our multilocus data (Dukik et al. 2017b), confirming that the main topological traits of the present study are robust.
Bayesian analysis is usually believed to be more reliable compared to parsimony and neighbour-joining methods, especially for an extensive sampling with a high divergence occurring among the sequences (Alfaro et al. 2003; Holder and Lewis 2003; James et al. 2006). In line with expectations, the Bayesian analysis of the five-gene data set showed a robust phylogeny. Bayesian and ML supports showed significant correlation of PP and BP values. In Eurotiomycetes, ribosomal ITS is often considered to be an inadequate marker for species distinction (Skouboe et al. 1999; Seifert et al. 2007; Visagie et al. 2014). Based on previous and present phylogenetic studies, it was demonstrated that the rDNA operon is sufficiently variable to distinguish most species of Ajellomycetaceae (Peterson and Kurtzman 1991; Berres et al. 1995; Kretzer et al. 1996; Peterson and Sigler 1998; Untereiner et al. 2004; this study; Fig. 4), but phylogenetic relationships among genera remained largely unresolved (Untereiner et al. 2004). In our ITS analysis (Fig. 4) the genera Blastomyces, Emmonsia, Emmonsiellopsis, Emergomyces, Helicocarpus, Histoplasma, Lacazia and Paracoccidioides were included in the Ajellomycetaceae, but the backbone of the tree showed low support. Histoplasma was nested in Blastomyces, and some taxa lacked strong bootstrap support, especially B. parvus (ML/BI 10/–) and B. silverae (ML/BI 65/–). The family Ajellomycetaceae had the geophilic genus Emmonsiellopsis in an ancestral position (Clade V). Paracoccidioides (Clade IV) was remote from remaining genera, and Histoplasma (Clade III) was paraphyletic to Blastomyces (Clade II). Emergomyces (Clade I-3) was paraphyletic to Ea. crescens (Clade I-1). With multilocus analysis, strains CBS 178.60 and CBS 205.48 formed a bootstrap-supported subclade (ML/BI 100%/1.00), paraphyletic to B. parvus, matching with a difference of 4 bases in ITS, 4 in TUB2, 0 in TEF3 and 14 in rPB2. The strains originated from lungs of rodents and were indistinguishable from B. parvus strains in all phenotypic and ecological markers. We suggest that at this level of molecular diversity of the currently used markers, strains may still belong to the same species.
Markers may have different functions. In addition to barcodes for species distinction, less variable genes are needed for the development of pan-family diagnostic primers. Also, markers for phylogenetic purposes may be different from those used for diagnostic purposes, where barcoding gaps should preferably be present. TUB2 has successfully been used species and variety delimitation in Histoplasma, Blastomyces and Paracoccidioides (Matute et al. 2006; de Teixeira et al. 2009, 2016); this gene was sequenced over our entire set of strains. TUB2 is easy to amplify, with a success rate in our strains of 95%. The gene has been reported to vary in the number of introns, and sometimes PCR results in the amplification of paralogous genes (Peterson 2008; Hubka and Kolarik 2012), but these problems were not encountered in our data set. Other options for secondary markers include TEF3, which was recently introduced by Stielow et al. (2015) as an excellent marker, but the success rate of sequencing for each strain is lower (92.5%). rPB2 was shown to have similar discriminatory power as TUB2 (Liu et al. 1999). PCR amplification in our dataset was significant (97.5%). In this study, the TEF3 and TUB2 sequences showed less parsimony-informative characters for the inference of phylogenetic relationships in different taxa compared to rPB2. The TEF3 and TUB2 data sets generated lower resolution across the Bayesian and ML trees, in which only 3 strongly supported clades were resolved with high BP and PP values, respectively. Among the three protein-coding genes, rPB2-based phylogeny drawn from Bayesian and ML analyses not only resolved the same number of clades in the family Ajellomycetaceae, but also had an equivalent resolution power, and topologies of trees were comparable to multilocus trees in Blastomyces (Clade II) (ML/BI 100/0.99), Histoplasma (Clade III) (ML/BI 100/0.99), Paracoccidioides (Clade IV) (ML/BI 100/0.99), Emmonsiellopsis (Clade V) (ML/BI 100/0.99). Emergomyces (Clade I-3) (ML/BI 99/0.99), leaving only the relationship between genera Emmonsia and Emergomyces unresolved with lower bootstrap support (ML/BI 46/–).
Barcoding analysis confirms that rPB2 and TUB2 with the current primer pairs allow recognition of entities. There is still room for improvement of primer design considering that not all parts of the sequences are equally informative. This aspect is particularly useful for the design of pan-Ajellomycetaceae primers or genus-specific barcode identifiers (Heinrichs et al. 2012). For example, the ITS-1 section is useful, as it provides essential information to delimit entities, while the ITS-2 region contributes less. Usually shorter fragments coincide with higher PCR fidelity, shorter PCR cycles (= shorter time-to-result) and higher capillary sequencing success, a key criterion to successful application in the clinics, and faster patient treatment. We consequently conclude that (1) rPB2 and TUB2 can both be recommended for routine identification, (2) TEF3, in combination with ITS, TUB2, and rPB2 yield stable topologies for multilocus sequence typing and GCPSR, and (3) for the development of a pan-Ajellomycetaceae primer to detect hitherto unknown species, from animal burrows or in archived lung specimens of terrestrial animals, rDNA ITS is probably sufficient. Whole genome trees usually show excellent resolution (Muñoz et al. 2015; Dukik et al. 2017b), but as yet relatively low numbers of genomes are available and trees suffer from significant strain and taxon sampling effects.
Biosafety considerations
Data presented in this paper including detailed species descriptions and reports of associated human and animal infections provides a foundation for assessment of biosafety consideration. Risk assessment should include the capacity of the organism to cause human disease, severity of the disease, and availability of effective treatment (Anon. 2009). Formal biosafety risk assessments by national level committees exist for only some members of the Ajellomycetacaeae, described in this report i.e. B. dermatitidis, B. parvus, Ea. crescens, and Es. pasteurianus,
Emergomyces species have only been reported to cause disease among persons with immunocompromising conditions (Gori et al. 1998, Pelegrin et al. 2011, Feng et al. 2015, Malik et al. 2016, Dukik et al. 2017b, Schwartz et al. 2018). In vitro antifungal susceptibility testing results suggested susceptible to available antifungals, including polyenes and most triazoles (Maphanga et al. 2017). Belgium’s Scientific Institute of Public Health (www.biosafety.be), the Swiss Agency for the Environment, Forests and Landscape (http://www.uab.cat/doc/fongs), and the American Type Culture Collection have classified Es. pasteurianus as a biosafety level 2 organism. We propose that other species in the genus be handled similarly.
Emmonsia crescens is a rodent pathogen and has occasionally been implicated in pulmonary disease in immune competent persons based on size of adiapores in lung (Anstead et al. 2012). In vitro antifungal susceptibility results showed susceptibility to amphotericin B, itraconazole, voriconazole and caspofungin (Borman et al. 2009; Dukik et al. 2017a). However, the role of antifungal therapy is controversial, because in vivo replication does not occur (Anstead et al. 2012). The species has been classified as a hazard group 2 pathogen by the Advisory Committee for Dangerous Pathogens in the United Kingdom (ACDP 2013).
Blastomyces dermatitidis can cause severe disease in immune competent hosts. In vitro susceptibility studies have reported susceptibility to amphotericin B, itraconazole, and other triazoles (Dukik et al. 2017a). The Biosafety in Microbiological and Biomedical Laboratories 5th edition classifies the yeast phase of B. dermatitidis as a biosafety level 2 pathogen and the mycelial phase as a biosafety level 3 pathogen (Anon. 2009). In contrast, ACDP considers both phase as hazard group 3 pathogens (ACDP 2013).
Blastomyces parvus has been stated as the cause of human disease in two immune compromised patients (Echavarria et al. 1993; Turner et al. 1999), but the identification of the putative pathogen is disputed (Sigler 2005; Anstead et al. 2012). There is no reliable report of B. parvus human infection. In vitro antifungal susceptibilities suggest susceptibility to commonly used antifungals. Health risks of this fungus for humans are low, but its close affinity to B. dermatitidis, as established in this paper, support its classification as a hazard group 2 pathogen by ACDP (ACDP 2013).
Blastomyces helicus infection appears to primarily cause disease in immune compromised persons (Sekhon et al. 1982; Kappagoda et al. 2017; Schwartz et al. 2017; Rofael et al. 2018). Conidia are not produced in vitro, and thus the risk of occupational inhalational exposure is considered low, although primary inoculation disease, may be possible. In vitro antifungal susceptibility results suggested susceptibility to amphotericin B and extended azoles (Dukik et al. 2017a). National level biosafety risk assessments for B. helicus do not yet exist. We propose that B. helicus be handled as a biosafety level 2 pathogen.
References
ACDP (2013) Approved list of biological agents. Advisory Committee on Dangerous Pathogens (ACDP). Health and Safety Executive (HSE), London. http://www.hse.gov.uk/pubns/misc208.pdf
Al-Doory Y, Vice TE, Mainster ME (1971) Adiaspiromycosis in a dog. J Am Vet Med Assoc 159(1):87–90
Alfaro ME, Zoller S, Lutzoni F (2003) Bayes or bootstrap? A simulation study comparing the performance of Bayesian Markov chain Monte Carlo sampling and bootstrapping in assessing phylogenetic confidence. Mol Biol Evol 20:255–266. https://doi.org/10.1093/molbev/msg028
Anon (2009) Biosafety in microbiological and biomedical laboratories, 5th edn. US Department of Health and Human Services, Centers for Disease Control and Prevention & National Institutes of Health, Atlanta
Anstead GM, Sutton DA, Graybill JR (2012) Adiaspiromycosis causing respiratory failure and a review of human infections due to Emmonsia and Chrysosporium spp. J Clin Microbiol 50(4):1346–1354. https://doi.org/10.1128/JCM.00226-11
Bagagli E, Bosco SMG, Theodoro RC, Franco M (2006) Phylogenetic and evolutionary aspects of Paracoccidioides brasiliensis reveal a long coexistence with animal hosts that explain several biological features of the pathogen. Infect Genet Evol 6(5):344–351. https://doi.org/10.1016/j.meegid.2005.12.002
Bagagli E, Theodoro RC, Bosco SM, McEwen JG (2008) Paracoccidioides brasiliensis: phylogenetic and ecological aspects. Mycopathologia 165(4–5):197–207. https://doi.org/10.1007/s11046-007-9050-7
Bariola J, Perry P, Pappas PG et al (2010) Blastomycosis of the central nervous system: a multicenter review of diagnosis and treatment in the modern era. Clin Infect Dis 50(6):797–804. https://doi.org/10.1086/650579
Baumgardner DJ, Paretsky DP (1999) The in vitro isolation of Blastomyces dermatitidis from a woodpile in north central Wisconsin, USA. Med Mycol 37(3):163–168
Baumgardner DJ, Paretsky DP, Yopp AC (1995) The epidemiology of blastomycosis in dogs: north 473 central Wisconsin, USA. J Med Vet Mycol 33:171–176
Berres ME, Szabo LJ, McLaughlin DJ (1995) Phylogenetic relationships in auriculariaceous basidiomycetes based on 25S ribosomal DNA sequences. Mycologia 87:821–840
Borman AM, Simpson VR, Palmer MD et al (2009) Adiaspiromycosis due to Emmonsia crescens is widespread in native British mammals. Mycopathologia 168(4):153–163. https://doi.org/10.1007/s11046-009-9216-6
Borman AM, Jiang Y, Dukik K, Sigler L, Schwartz IS, de Hoog S. (2018) Adiaspiromycosis and diseases caused by related fungi in Ajellomycetaceae. Springer International Publishing AG (in press)
Brown SD, Collins RA, Boyer S et al (2012) Spider: an R package for the analysis of species identity and evolution, with particular reference to DNA barcoding. Molec Ecol Res 12(3):562–565. https://doi.org/10.1111/j.1755-0998.2011.03108.x
Brown EM, McTaggart LR, Zhang SX et al (2013) Phylogenetic analysis reveals a cryptic species Blastomyces gilchristii, sp. nov. within the human pathogenic fungus Blastomyces dermatitidis. PLoS ONE 8(3):e59237. https://doi.org/10.1371/journal.pone.0059237
Brummer E, Castaneda E, Restrepo A (1993) Paracoccidioidomycosis: an update. Clin Microbiol Rev 6(2):89–117
Carmichael JW (1951) The pulmonary fungus Haplosporangium parvum II. Strain and generic relationships. Mycologia 43:605–624
Ciferri R, Montemartini A (1959) Taxonomy of Haplosporangium parvum. Mycopath Mycol Appl 10:303–316
Colombo AL, Tobón A, Restrepo A et al (2011) Epidemiology of endemic systemic fungal infections in Latin America. Med Mycol 49(8):785–798. https://doi.org/10.3109/13693786.2011.577821
Coyne JA, Orr HA (2004) Speciation. Sinauer Associates, Sunderland
De Hoog GS, Redhead SA, Feng P et al (2016a) Proposals to conserve Blastomyces Gilchrist & W.R. Stokes against Blastomyces Costantin & Rolland and Ajellomycetaceae against Paracoccidioidaceae (Ascomycota: Onygenales). Taxon 65:1167–1169
De Hoog GS, Guarro J, Gené J, Figueras MJ (2016b). Atlas of clinical fungi, E-version 4.1.4. CBS-KNAW Fungal Biodiversity Centre, Utrecht
de Teixeira M, Theodoro RC, de Carvalho MJ et al (2009) Phylogenetic analysis reveals a high level of speciation in the Paracoccidioides genus. Mol Phylogenet Evol 52(2):273–283. https://doi.org/10.1016/j.ympev.2009.04.005
de Teixeira M, Patané JS, Taylor ML et al (2016) Worldwide phylogenetic distributions and population dynamics of the genus Histoplasma. PLoS Neglect Trop Dis 10(6):e0004732. https://doi.org/10.1371/journal.pntd.0004732
Dowding ES (1947) The pulmonary fungus Haplosporangium parvum, and its relationship with some human pathogens. Can J Res Sect E Med Sci 25:195–206
Drouhet E, Guého E, Gori S et al (1998) Mycological, ultrastructural and experimental aspects of a new dimorphic fungus Emmonsia pasteuriana sp. nov. isolated from a cutaneous disseminated mycosis in AIDS. J Mycol Méd 8:64–77
Dukik K, Al-Hatmi AMS, Curfs-Breuker I et al (2017a) Antifungal susceptibility of emerging dimorphic pathogens in the family Ajellomycetaceae. Antimicrob Agents Chemother. https://doi.org/10.1128/AAC.01886-17
Dukik K, Muñoz JF, Jiang Y et al (2017b) Novel taxa of thermally dimorphic systemic pathogens in the Ajellomycetaceae (Onygenales). Mycoses 60(5):296–309. https://doi.org/10.1111/myc.12601
Dvořak J, Otčenašek M, Rosicky B (1973) Adiaspiromycosis caused by Emmonsia crescens, Emmons and Jellison 1960. Studie ČSAV (Acad Praha) 14:1–120
Echavarria E, Cano EL, Restrepo A (1993) Disseminated adiaspiromycosis in a patient with AIDS. J Med Vet Mycol 31(1):91–97
Edgar RC (2004) MUSCLE: Multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res 32(5):1792–1797
Emmons CW, Ashburn LL (1942) The isolation of Haplosporangium parvum n. sp. and Coccidioides immitis from wild rodents. Publ Health Rep 57:1715–1727
Emmons CW, Jellison WL (1960) Emmonsia crescens sp. n. and adiaspiromycosis (haplomycosis in mammals). Ann NY Acad Sci 89:91–101
England DM, Hochholzer L (1993) Adiaspiromycosis: an unusual fungal infection of the lung. Am J Surg Pathol 17(9):876–886
Feng P, Yin S, Zhu G et al (2015) Disseminated infection caused by Emmonsia pasteuriana in a renal transplant recipient. J Dermatol 42(12):1179–1182. https://doi.org/10.1111/1346-8138.12975
Gori S, Drouhet E, Guého E et al (1998) Cutaneous disseminated mycosis in a patient with AIDS due to a new dimorphic fungus. J Mycol Méd 8:57–63
Heinrichs G, de Hoog GS, Haase G (2012) Barcode identifiers as a practical tool for reliable species assignment of medically important black yeast species. J Clin Microbiol 50(9):3023–3030. https://doi.org/10.1128/JCM.00574-12
Heys I, Taljaard J, Orth H (2014) An Emmonsia species causing disseminated infection in South Africa. N Engl J Med 370(3):283–284. https://doi.org/10.1056/NEJMc1314277#SA1
Holder M, Lewis PO (2003) Phylogeny estimation: traditional and Bayesian approaches. Nat Rev 4(4):275–284. https://doi.org/10.1038/nrg1044
Hrycyk MF, Garcia Garces H, Bosco SMG et al (2018) Ecology of Paracoccidioides brasiliensis, P. lutzii and related species: infection in armadillos, soil occurrence and mycological aspects. Med Mycol. https://doi.org/10.1093/mmy/myx142
Hubalek Z (1999) Emmonsiosis of wild rodents and insectivores in Czechland. J Wildl Dis 35:243–249
Hubálek Z, Nesvadbová J, Rychnovsky B et al (1995) A heterogeneous distribution of Emmonsia parva var. crescens in an agro-ecosystem. J Med Vet Mycol 33(3):197–200
Hubálek Z, Nesvadbová J, Halouzka J (1998) Emmonsiosis of rodents in an agroecosystem. Med Mycol 36(6):387–390
Hubka V, Kolarik M (2012) β-Tubulin paralogue tubC is frequently misidentified as the benA gene in Aspergillus section Nigri taxonomy: primer specificity testing and taxonomic consequences. Persoonia 29:1–10. https://doi.org/10.3767/003158512X658123
James TY, Kauff F, Schoch CL et al (2006) Reconstructing the early evolution of fungi using a six-gene phylogeny. Nature 443:818–822. https://doi.org/10.1038/nature05110
Jellison WL (1969) Adiaspiromycosis (=haplomycosis). Mountain Press Publishers, Montana
Kane J, Summerbell RC, Sigler L et al (1997) Laboratory handbook of dermatophytes: a clinical guide and laboratory manual of dermatophytes and other filamentous fungi from skin, hair, and nails. Star Publishing Co., Belmont
Kappagoda S, Adams JY, Luo R et al (2017) Fatal Emmonsia sp. infection and fungemia after orthotopic liver transplantation. Emerg Infect Dis 23(2):346–349. https://doi.org/10.3201/eid2302.160799
Kemna ME, Weinberger M, Sigler L et al (1994) A primary oral blastomycosis-like infection in Israel. In: 94th General Meeting of the American Society for Microbiology, Washington, DC, Abstract F-75, p 601
Kenyon C, Bonorchis K, Corcoran C et al (2013) A dimorphic fungus causing disseminated infection in South Africa. N Engl J Med 369(15):1416–1424. https://doi.org/10.1056/NEJMoa1215460
Koller LD, Helfer DH (1978) Adiaspiromycosis in the lungs of a goat. J Am Vet Med Assoc 173:80–81
Koller LD, Patton NM, Whitsett DK (1976) Adiaspiromycosis in the lungs of a dog. J Am Vet Med Assoc 169:1316–1317
Kretzer A, Li Y, Szaro T et al (1996) Internal transcribed spacer sequences from 38 recognized species of Suillus sensu lato: phylogenetic and taxonomic implications. Mycologia 88:776–785
Kwon-Chung KJ (1973) Studies on Emmonsiella capsulata I. Heterothallism and development of the ascocarp. Mycologia 65(1):109–121
Liu YJ, Whelen S, Hall BD (1999) Phylogenetic relationships among ascomycetes: evidence from an RNA polymerse II subunit. Mol Biol Evol 16(12):1799–1808. https://doi.org/10.1093/oxfordjournals.molbev.a026092
Malik R, Capoor MR, Vanidassane I et al (2016) Disseminated Emmonsia pasteuriana infection in India: a case report and a review. Mycoses 59(2):127–132. https://doi.org/10.1111/myc.12437
Maphanga TG, Britz E, Zulu TG et al (2017) In vitro antifungal susceptibility of yeast and mold phases of isolates of dimorphic fungal pathogen Emergomyces africanus (formerly Emmonsia sp.) from HIV-infected South African patients. J Clin Microbiol 55(6):1812–1820. https://doi.org/10.1128/JCM.02524-16
Marin-Felix Y, Stchigel AM, Cano-Lira JF et al (2015) Emmonsiellopsis, a new genus related to the thermally dimorphic fungi of the family Ajellomycetaceae. Mycoses 58(8):451–460. https://doi.org/10.1111/myc.12336
Matsuda K, Niki H, Yukawa A et al (2015) First detection of adiaspiromycosis in the lungs of a deer. J Vet Med Sci 77(8):981–983. https://doi.org/10.1292/jvms.15-0083
Matute DR, McEwen JG, Puccia R et al (2006) Cryptic speciation and recombination in the fungus Paracoccidioides brasiliensis as revealed by gene genealogies. Mol Biol Evol 23(1):65–73. https://doi.org/10.1093/molbev/msj008
McDonough ES, Lewis AL (1968) The ascigerous stage of Blastomyces dermatitidis. Mycologia 60:76–83
Medoff G, Maresca B, Lambowitz AM et al (1986) Correlation between pathogenicity and temperature sensitivity in different strains of Histoplasma capsulatum. J Clin Invest 78:16381–16647
Meece JK, Anderson JL, Fisher MC et al (2011) Population genetic structure of clinical and environmental isolates of Blastomyces dermatitidis, based on 27 polymorphic microsatellite markers. Appl Environ Microbiol 77(15):5123–5131. https://doi.org/10.1128/AEM.00258-11
Meyer CP, Paulay G (2005) DNA barcoding: error rates based on comprehensive sampling. PLoS Biol 3(12):e422. https://doi.org/10.1371/journal.pbio.0030422
Muñoz JF, Gauthier GM, Desjardins CA et al (2015) The dynamic genome and transcriptome of the human fungal pathogen Blastomyces and close relative Emmonsia. PLoS Genet 11(10):e1005493. https://doi.org/10.1371/journal.pgen.1005493
Pelegrin I, Ayats J, Xiol X et al (2011) Disseminated adiaspiromycosis: case report of a liver transplant patient with human immunodeficiency infection, and literature review. Transpl Infect Dis 13(5):507–514. https://doi.org/10.1111/j.1399-3062.2011.00611.x
Peterson SW (2008) Phylogenetic analyses of Aspergillus species using DNA sequences from four loci. Mycologia 100(2):205–226
Peterson SW, Kurtzman CP (1991) Ribosomal RNA sequence divergence among sibling species of yeasts. Syst Appl Microbiol 14:124–129
Peterson SW, Sigler L (1998) Molecular genetic variation in Emmonsia crescens and Emmonsia parva, etiologic agents of adiaspiromycosis, and their phylogenetic relationship to Blastomyces dermatitidis (Ajellomyces dermatitidis) and other systemic fungal pathogens. J Clin Microbiol 36(10):2918–2925
Philippe H, Snell EA, Bapteste E et al (2004) Phylogenomics of Eukaryotes: impact of missing data on large alignments. Mol Biol Evol 21(9):1740–1752. https://doi.org/10.1093/molbev/msh182
Popescu AA, Huber KT, Paradis E (2012) ape 3.0: new tools for distance-based phylogenetics and evolutionary analysis in R. Bioinformatics 28(11):1536–1537. https://doi.org/10.1093/bioinformatics/bts184
Pusterla N, Pesavento PA, Leutenegger CM et al (2002) Disseminated pulmonary adiaspiromycosis caused by Emmonsia crescens in a horse. Equine Vet J 34(7):749–752
Restrepo A, Baumgardner DJ, Bagagli E et al (2000) Clues to the presence of pathogenic fungi in certain environments. Med Mycol 38:67–77
Rofael M, Schwartz IS, Sigler L et al (2018) Emmonsia helica infection in HIV-infected man, California, USA. Emerg Infect Dis 24(1):166–168. https://doi.org/10.3201/eid2401.170558
Ronquist F, Huelsenbeck JP (2003) MrBayes 3: Bayesian phylogenetic inference under mixed models. Bioinformatics 19(12):1572–1574
Saccente M, Woods GL (2010) Clinical and laboratory update on blastomycosis. Clin Microbiol Rev 23(2):367–381. https://doi.org/10.1128/CMR.00056-09
Samson RA, Houbraken J, Thrane U et al (2010) Food and indoor fungi. CBS KNAW Biodiversity Center, Utrecht
Schoch CL, Seifert KA, Huhndorf S et al (2012) Nuclear ribosomal internal transcribed spacer (ITS) region as a universal DNA barcode marker for Fungi. Proc Natl Acad Sci USA 109(16):6241–6246. https://doi.org/10.1073/pnas.1117018109
Schwartz IS, Govender NP, Corcoran C et al (2015a) Clinical characteristics, diagnosis, management and outcomes of disseminated emmonsiosis: a retrospective case series. Clin Infect Dis 61(6):1004–1012. https://doi.org/10.1093/cid/civ439
Schwartz IS, Kenyon C, Feng P et al (2015b) 50 Years of Emmonsia disease in humans: the dramatic emergence of a cluster of novel fungal pathogens. PLoS Pathog 11(11):e1005198. https://doi.org/10.1371/journal.ppat.1005198
Schwartz IS, Wiederhold NP, Patterson TF, Sigler L (2017) Blastomyces helicus, an emerging dimorphic fungal pathogen causing fatal pulmonary and disseminated disease in humans and animals in western Canada and United States. IDWeek, San Diego, CA, 4–8 October 2017
Schwartz IS, Sanche S, Wiederhold NP, Patterson TF, Sigler L (2018) Emergomyces canadensis, a dimorphic fungus causing fatal systemic human disease in North America. Emerg Infect Dis 24(4):758–761. https://doi.org/10.3201/eid2404.171765
Seifert KA, Hughes SJ, Boulay H et al (2007) Taxonomy, nomenclature and phylogeny of three cladosporium-like hyphomycetes, Sorocybe resinae, Seifertia azaleae and the Hormoconis anamorph of Amorphotheca resinae. Stud Mycol 58:235–245. https://doi.org/10.3114/sim
Sekhon AS, Jackson FL, Jacobs HJ (1982) Blastomycosis: report of the first case from Alberta, Canada. Mycopathologia 79:65–69
Shapiro B, Ho SY, Drummond AJ et al (2010) A Bayesian phylogenetic method to estimate unknown sequence ages. Mol Biol Evol 28(2):879–887. https://doi.org/10.1093/molbev/msq262
Sigler L (1996) Ajellomyces crescens sp. nov., taxonomy of Emmonsia spp., and relatedness with Blastomyces dermatitidis (teleomorph Ajellomyces dermatitidis). J Med Vet Mycol 34(5):303–314
Sigler L (1998) Agents of adiaspiromycosis. In: Ajello L, Hay R (eds) Topley & Wilson’s microbiology and microbial infections, vol 4, 9th edn. Arnold, London, pp 571–583
Sigler L (2005) Adiaspiromycosis and other infections caused by Emmonsia species. In: Hodder A (ed) Topley & Wilson’s microbiology and microbial infections, 10th edn. Wiley, London, pp 809–824
Sigler L (2015) Emmonsia helica Sigler. Index Fungorum 237:1. http://www.speciesfungorum.org/Names/SynSpecies.asp?RecordID=551157
Skouboe P, Frisvad JC, Taylor JW et al (1999) Phylogenetic analysis of nucleotide sequences from the ITS region of terverticillate Penicillium species. Mycol Res 103:873–881
Stamatakis A (2014) RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30(9):1312–1313. https://doi.org/10.1093/bioinformatics/btu033
Stielow JB, Levesque CA, Seifert KA et al (2015) One fungus, which genes? Development and assessment of universal primers for potential secondary fungal DNA barcodes. Persoonia 35:242–263. https://doi.org/10.3767/003158515X689135
Taborda PR, Taborda VA, McGinnis MR (1999) Lacazia loboi gen. nov., comb. nov., the etiologic agent of lobomycosis. J Clin Microbiol 37(6):2031–2033
Tamura K, Stecher G, Peterson D et al (2013) MEGA6: Molecular Evolutionary Genetics Analysis version 6.0. Mol Biol Evol 30(12):2725–2729. https://doi.org/10.1093/molbev/mst197
Turner D, Burke M, Bashe E et al (1999) Pulmonary adiaspiromycosis in a patient with acquired immunodeficiency syndrome. Eur J Clin Microbiol Infect Dis 18(12):893–895
Untereiner WA, Scott JA, Naveau FA et al (2004) The Ajellomycetaceae, a new family of vertebrate-associated Onygenales. Mycologia 96(4):812–821
Van den Brink J, Facun K, de Vries M et al (2015) Thermophilic growth and enzymatic thermostability are polyphyletic traits within Chaetomiaceae. Fung Biol 119(12):1255–1266. https://doi.org/10.1016/j.funbio.2015.09.011
Vilela R, Mendoza L (2018) Paracoccidioidomycosis ceti (lacaziosis/lobomycosis) in dolphins. In Seyedmousavi S et al. (eds) Emerging and epizootic fungal infections in animals (in press)
Visagie CM, Hirooka Y, Tanney JB et al (2014) Aspergillus, Penicillium and Talaromyces isolated from in house dust samples collected around the world. Stud Mycol 78:63–139. https://doi.org/10.1016/j.simyco.2014.07.002
Wang P, Kenyon C, de Hoog GS et al (2017) A novel dimorphic pathogen, Emergomyces orientalis (Onygenales), agent of disseminated infection. Mycoses 60(5):310–319. https://doi.org/10.1111/myc.12583
Wellinghausen N, Kern WV, Haase G et al (2003) Chronic granulomatous lung infection caused by the dimorphic fungus Emmonsia sp. Int J Med Microbiol 293(6):441–445. https://doi.org/10.1078/1438-4221-00281
Wu SJ, Valyi-Nagy T, Engelhard HH et al (2005) Secondary intracerebral blastomycosis with giant yeast forms. Mycopathologia 160(3):253–257
Yang Y, Ye Q, Li K et al (2017) Genomics and comparative genomic analyses provide insight into the taxonomy and pathogenic potential of novel Emmonsia pathogens. Front Cell Infect Microbiol 7:105. https://doi.org/10.3389/fcimb.2017.00105
Acknowledgements
Yanping Jiang is indebted to Prof. Li Zuo, director of Department of Immunology, Basic Medical School, Guizhou Medical University and director of the Department of Dermatology, The Affiliated Hospital, Guizhou Medical University for supporting her studies performed at Westerdijk Institute. Yanping also gratefully acknowledges the China Scholarship Council (CSC) Fund support for the Advancement of Scholarship. Andy Borman is acknowledged for kindly making strains available for study.
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Jiang, Y., Dukik, K., Muñoz, J.F. et al. Phylogeny, ecology and taxonomy of systemic pathogens and their relatives in Ajellomycetaceae (Onygenales): Blastomyces, Emergomyces, Emmonsia, Emmonsiellopsis. Fungal Diversity 90, 245–291 (2018). https://doi.org/10.1007/s13225-018-0403-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13225-018-0403-y