Abstract
A brief survey towards an understanding of evolution of Basidiomycota in the last five decades is given. Between 1960–1990 studies were based mainly on morphology, subcellular structure, biochemistry, physiology and ecology. In the last 20 years, molecular methods have revolutionized the study of taxonomy, systematics, phylogeny, biogeography, population and microevolutionary processes in basidiomycetes. A fundamental improvement in the understanding of evolution of basidiomycetes has been achieved. It is anticipated that numerous new taxa at different taxonomic levels will be recognized based on integrated approaches, and new data will soon provide fascinating insights into the origin, evolution and biogeography of basidiomycetes in the coming years.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The phylum Basidiomycota is typically characterized by the presence of a basidium bearing sexual spores (i.e., basidiospores) in addition to several other characters (Oberwinkler 1978). A common informal term for all Basidiomycota is “basidiomycetes”. This is a very important group, being the second largest assemblage of the Kingdom Fungi, comprising approximately 31,000 described species (Kirk et al. 2008). The group is of almost cosmopolitan in distribution, encompassing numerous edible mushrooms, toadstools, pathogens, and endophytes besides numerous mycorrhizal partners and wood-rotting decomposers in forest ecosystems. The basidiomycetes have, as a result, drawn the attention of mycologists for a long time, since the very beginning of scientific mycology at the 18th century (e.g. Persoon 1801; Fries 1821; de Bary 1853, 1866; Brefeld 1888). Knowledge of the taxonomy, host range and distribution, phylogeny and evolution of this group of fungi has rapidly increased in the last 50 years. This is especially evident in the last 20 years with the development of molecular techniques.
The aim of paper is to summarize the last 50 years of research in the Basidiomycota, and also to review our present understanding of the phylum, emphasizing the highlights among selected groups and future perspectives. No attempt has been made to cite all of the relevant studies for the Basidiomycota, because studies on individual groups of basidiomycetes are too numerous to list.
The earlier thirty years: taxonomic and systematic researches
Between 1960–1990 gross phenotypic taxonomy was supplemented by microscopy and in vitro culturing (e.g. Miller 1971; Desjardin 1990). Many groups of basidiomycetes were intensively studied. At the same time important monographic or taxonomic works were published. A few of the most influential ones may be mentioned here; they are Corner (1966), Horak (1968), Cummins and Hiratsuka (1983), Pegler (1983), Vánky (1987), although there are many others.
In Europe, compilation and publication of a few important regional mycota, such as British Fungus Flora (1979–), and Flora Agaricina Neerlandica (1988–), have successfully been launched and promoted, with welcomed works by Moser (1983), Hjortstam et al. (1987, 1988), and Ryvarden and Gilbertson (1993, 1994). The monographic works “Fungi Europaei” (1984–) have been valuable references in the study on diversity of macromycetes both within and outside Europe. During the same period in North America, mycologists also were very active in studying basidiomycete diversity (e.g. Hesler and Smith 1979; Petersen 1981; Halling 1983; Mueller 1992). In East Asia, the mycota of Japan has been studied much more intensively than in any of the other countries in the region (e.g. Imazeki et al. 1988; Hiratsuka et al. 1992). The Flora Fungorum Sinicorum (1987–), covers such a diverse group of fungi that they can be finished only when specific groups have been intensively studied, and thus the publication will probably take several decades, although over 10 volumes on basidiomyctes in China are now available (e.g. Peng et al. 1992; Zhuang et al. 1998; Guo 2000; Yang 2005a; Zang 2006; Zhou 2007). Basidiomycetes in the Southern Hemisphere have also received much attention from a number of fungal taxonomists (e.g. Cunningham 1965; Dennis 1970; Heinemann 1972; Reid 1980; Garrido 1988).
With regard to the systematics and phylogeny of basidiomycetes, the works of Singer (1962, 1986), Donk (1964, 1971), Gäumann (1964), Kreisel (1969), Ainsworth et al. (1973), Oberwinkler (1977, 1978, 1982, 1985), Kühner (1980) and Jülich (1981) are probably among the most influential between 1960 and 1990. The gasteromycetes were often treated a single group, although some, such as the secotioid taxa, have anatomical similarities to certain agarics and boletes, and, as a result, were supposed to be related to agarics and boletes respectively. However, views were in conflict as regards to the direction of the evolutionary process (Singer and Smith 1960; Heim 1971; Thiers 1984; Singer 1986). Oberwinkler (1977, 1978), Thiers (1984) and others argued that it was more likely that sequestrate (secotioid or gasteroid) basidiomycetes were derived repeatedly and convergently, and should not be regarded as a single natural group.
In trying to elucidate the phylogeny of basidiomycetes, Oberwinkler (1982) exquisitely discussed the significance of the morphology of the basidium, together with the knowledge of the presence or absence of secondary spores, the host specificity and other aspects, and he pointed out that the evolution of the homobasidiomycetes from a phragmo- and/or holobasidial ancestral form was probably accompanied by the loss of the capacity to form secondary spores, and the formation of uniform basidium. Due to the unique basidial morphology, the connections of several groups of gasteromycetes with other basidiomycetes were unknown (Oberwinkler 1982).
Besides the morphology of basidia, spindle pole bodies (e.g. McLaughlin et al. 1995; Celio et al. 2006), and septa (e.g. Moore 1985, 1997; Khan and Kimbrough 1982; Oberwinkler and Bandoni 1982; Kimbrough 1994; Wells 1994; McLaughlin et al. 1995; Bauer et al. 1997; Müller et al. 2000; Hibbett and Thorn 2001; Van Driel et al. 2009) as well as physiological and biochemical characteristics (Bartnicki-Garcia 1968; Van der Walt and Yarrow 1984; Prillinger et al. 1993; Kurtzman and Fell 1998; Boekhout and Guého 2002) have significantly contributed to the systematics of basidiomycetes until the present day. The structural and biochemical database for fungi (Celio et al. 2006) aims to capture several of these characters in a comprehensive manner.
At the same time, for some groups of basidiomycetes that grow in culture, mating studies have been used to elucidate the specific or supraspecific consistency (Korhonen 1978a, b; Gordon and Petersen 1991; Petersen and Halling 1993; Petersen and Gordon 1994).
1990–2010: a transition from morphologically-based taxonomy and systematics to molecular-based phylogeny and evolution
Morphological and physiological studies left many questions and controversies unresolved, due in part to the paucity of characters, as well as the general absence of explicit phylogenetic analyses (Hibbett 2007). During the last 20 years, remarkable progress has made in the study of molecular evolution of basidiomycetes with the introduction of molecular methods.
The development of new statistical methods and advances in computational technology make the evaluation of evolution possible. In particular, with the invention and the development of the polymerase chain reaction (PCR) technique, phylogenetic analysis of DNA or protein sequences has become a powerful tool for studying molecular evolution in fungi (White et al. 1990; Bruns et al. 1992; Nei and Kumar 2000). Ribosomal DNA (rDNA) sequences have provided a wealth of information concerning phylogenetic relationships (Hillis and Dixon 1991), and studies of rDNA sequences have been used to infer phylogenetic history across a very broad spectrum, from studies among the basal lineages of life to relationships among closely related species and populations. Sequence data from ribosomal DNA (i.e. nSSU and nLSU rDNA), mtDNA and protein coding genes (e.g. tef1, rpb1, rpb2) have been used in fungal systematic studies (e.g. Swann and Taylor 1995; Fell et al. 2000; Lutzoni et al. 2004; Matheny et al. 2007a, b, c).
Classification in the basidiomycota
Before the molecular era, basidiomycetes were usually divided into Phragmobasidiomycetes and Holobasidiomycetes, or Heterobasidiomycetes and Homobasidiomycetes. Molecular phylogenetic data showed that a separation of heterobasidiomycetes from homobasidiomycetes is impossible, and, thus, such historical concepts have to be abandoned (Weiß et al. 2004a).
Molecular phylogenetic studies have led to significant advances in the understanding of the higher-level relationships of basidiomycetes, and consequently, the whole taxonomic hierarchy of the Basidiomycota, as in the remaining other groups of the Fungi, has been dramatically altered. Under the umbrella of the Deep Hypha Research Coordination Network and Assembling the Fungal Tree of Life project (Lutzoni et al. 2004; Blackwell et al. 2007), and additional projects, a few major publications elucidating relationships within the Fungi appeared in the last few years (Bauer et al. 2006; James et al. 2006; Liu et al. 2006; Aime et al. 2007). Within the Kingdom Fungi, molecular phylogenetic analyses support the monophyly of the Ascomycota and Basidiomycota, and these are regarded as the subkingdom Dikarya (James et al. 2006). A comprehensive classification of Fungi based on phylogenetic results was proposed (Hibbett et al. 2007) and adopted by the Dictionary of the Fungi (Kirk et al. 2008).
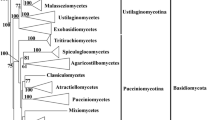
Recognized as a phylum, the phylogeny of Basidiomycota has been extensively revisited and many new taxa including diverse taxonomic ranks below the phylum level have been proposed in the last few years based on molecular phylogeny and reevaluation of morphological characters. The phylum Basidiomycota is generally regarded as having three major clades (Fig. 1; Swann and Taylor 1995; Lutzoni et al. 2004; Taylor et al. 2004; Bauer et al. 2006; Matheny et al. 2007a, b), the Pucciniomycotina (Urediniomycetes, Fig. 2a–d), the Ustilaginomycotina (Ustilaginomycetes, Fig. 2f–h), and the Agaricomycotina (Hymenomycetes, Fig. 2i–t), with the phylogenetic positions of additional two major lineages, the Entorrhizomycetes (Fig. 2e) and Wallemiomycetes yet unclear (Table 1; Zalar et al. 2005; Matheny et al. 2007c; Hibbett et al. 2007).
Diverse forms of spore-producing structures in Basidiomycota. a–d. Species of Pucciniomycotina. a. Puccinia recondita (Pucciniales, aecial stage) on Thalictrum rutifolium. b. Chrysomyxa succinea (Pucciniales, telial stage) on Rhododendron sp. c. Jola cf. javensis (Platygloeales) on moss. d. Sphacelotheca sp. (Microbotryales) on Polygonum sp. e. Entorrhiza casparyana (Entorrhizomycetes) on Juncus articulatus. f–h. Species of Ustilaginomycotina. f. Ustilago nuda (Ustilaginales) on Hordeum vulgare var. nudum. g. Anthracoidea filamentosae (Ustilaginales) on Carex crebra. h. Exobasidium deqinense (Exobasidiales) on Rhododendron sp. i–t. Species of Agaricomycotina. i. Dacrymyces yunnanensis (Dacrymycetales) on rotten wood. j. Auricularia auricula (Auriculariales) on rotten wood. k. Tremellodendropsis tuberosa (Auriculariales). l. Sebacina incrustans (Sebacinales). m. Multiclavula sinensis (Cantharellales, basidiolichen). n. Geastrum sacatum (Geastrales). o. Ramaria hemirubella (Gomphales). p. Phallus luteus (Phallales). q. Phallogaster saccatus (Hysterangiales). r. Agaricus bisporus (Agaricales). s. Crucibulum laeve (Agaricales). t. Boletus reticuloceps (Boletales)
It is worthy and interesting to note that Moncalvo et al. (2002) highlighted the complexity of the history of the Agaricomycotina. They found that many clades correspond to traditional taxonomic groups, many do not, and several clades are best supported by ecological, biochemical, or trophic habits rather than by morphological similarities. Such or similar phenomena may be revealed in many other groups of Fungi.
Gasteromycetation
Within the Basidiomycota, “gasteromycetes” (with spores that are not forcibly discharged, statismospores, see Figs. 1 and 3e, rather than forcibly discharged, ballistospores, see Fig. 3b) comprise a diverse, artificial assemblage of puffballs, earthstars, false earthstars, earthballs, bird’s nest and cannonball fungi, stinkhorns, secotioid agarics and boletes, and false truffles (Reijnders 1963; Heim 1971; Miller and Miller 1988). Molecular systematics studies have revealed that gasteromycetes have independently evolved many times within the basidiomycetes during the adaption of environmental selective pressures, such as arid conditions, dispersal vectors, and unknown mechanisms (Fig. 1; Bruns et al. 1989; Hibbett et al. 1997; Peintner et al. 2001; Binder and Bresinsky 2002; Binder et al. 2006; Henkel et al. 2010), as were suggested by Oberwinkler (1977, 1978, 1985), Thiers (1984) and many others. It was suggested that the evolution of the sequestrate state to be irreversible (Hibbett 2004, 2007).
A schema of gasteromycetation in Amanita (Agaricales). Torrendia (Fig. 3c, d) and Amarrendia (Fig. 3f) were regarded as genera independent from Amanita (Fig. 3a) by several authors (e.g. Bas 1975; Miller and Horak 1992; Bougher 1999; Bougher and Lebel 2002). Recent molecular phylogenetic analyses showed that species of these two genera just present gasteromycetations within Amanita (Justo et al. 2010)
The groups of the gasteromycetes whose connections with other basidiomycetes were unknown (Oberwinkler 1982) were revealed as either clades represented entirely by sequestrate taxa, i.e. Geastrales (Fig. 2n), Hysterangiales (Fig. 2q) and Phallales (Fig. 2p), or consisting of both sequestrate and non-sequestrate taxa, such as, Gomphales (Fig. 2o). The remaining groups, such as “Lycoperdales”, “Nidulariales”, and “Tulostomatales” have close relationships with Agaricaceae s.l. (Fig. 2r, s), while “Melanogastrales” and “Sclerodermatales” show phylogenetic affinity with Boletales (Hibbett et al. 1997; Vellinga 2004; Binder and Hibbett 2007; Hosaka et al. 2007; Fig. 2t).
Interestingly, some sequestrate fungi represent recent, divergent events that led to one or a few sequestrate species within a clade of non-sequestrate relatives (Fig. 3; e.g. Kretzer and Bruns 1997; Martin et al. 1999; Vellinga et al. 2003; Vellinga 2004; Albee-Scott 2007; Lebel and Catcheside 2009; Justo et al. 2010), while others of earlier origin have speciated and radiated across a wide spectrum of taxa (Fig. 1; e.g. Binder and Hibbett 2007; Hosaka et al. 2007). Newly most optimal models suggest that the net diversification rate of gasteroid forms exceeds that of non-gasteroid forms, and that gasteroid forms will eventually come to predominate over non-gasteroid forms in the clades in which they have arisen (Wilson et al. 2011).
Species criteria: challenge and opportunity
The basic rank in taxonomy of organisms is the species. Attempts to reach a consensus for a universal definition of species have been unsuccessful, and consequently over 20 different concepts have been used (Mayden 1997). For instance, the morphological species concept, the biological species concept, the ecological species concept, and the phylogenetic species concept virtually emphasize morphological divergence, reproductive isolation, adaptation to a particular ecological niche, and nucleotide divergence respectively (Giraud et al. 2008). However, these species criteria correspond to the different events that occur during lineage separation and divergence, rather than to fundamental differences of what is considered to represent a species (de Queiroz 1998, 2007; Giraud et al. 2008).
Morphological species concept is the classic approach used. However, exactly what different mycologists consider to be a species can vary widely, and there are different approaches for delineating them. In addition, many morphological characters are plastic or subtle, and difficult to assess. It has been repeatedly shown that similar characters can arise from evolutionary convergence or environmental constrains (Moncalvo 2005; Hibbett 2007), and, thus, morphological species concept is, in many cases, unsatisfactory for applications. The application of biological species concept or ecological species concept to fungi was favored between 1960–1990, and is still presently being used. However, there are still many limitations for its application (Taylor et al. 2000; Giraud et al. 2008).
Phylogenetic approaches and incorporation of molecular biological techniques, particularly the analysis of DNA nucleotide sequences have provided new information and the phylogenetic species concept is becoming a popular trend, particularly, when it is applied to asexual organisms, and connects the anamorph and teleomorphic stages of a single species (Guarro et al. 1999; Moncalvo 2005; Hyde et al. 2011). In fungi, the sequence data from the internal transcribed spacer region of the nuclear rDNA locus (ITS) have often been used to recognize fungal phylogenetic species and may well be the DNA barcoding locus used in barcoding (Seifert 2009; Begerow et al. 2010; Jargeat et al. 2010). However, it is better to use multigene genealogy concordance than to use a single gene to recognize species (Taylor et al. 2000). The current “gold standard” genealogical concordance phylogenetic species recognition criterion has proved very useful in fungi, because it is more finely discriminating than the other criteria in many cases. Genealogical concordance phylogenetic species recognition has been practiced recently in different groups of basidiomycetes (e.g. Kauserud et al. 2006; Jargeat et al. 2010; Van de Putte et al. 2010).
Prospects
Fungi play critically important roles in ecosystems, and their potential economic benefits as food and medicine are great. The importance of basidiomycetes in ecosystems as mycorrhizal partners, plant pathogens and decomposers cannot be overestimated. Although understanding of the origin and evolution of basidiomycetes has greatly been improved in recent years and has provided interesting new insights into the phylogeny and natural classification of Fungi, it is still far from satisfactory, as many issues relating to their taxonomy and phylogeny, ecology, and geographical distributions remain unclear. In the near future, the following aspects should be a few focal points of research interests:
-
1)
Accelerating the discovery and documentation of new taxa
It is generally accepted that only 5–10% of species on the earth have been discovered and named. An estimated 1.5 million fungal species exist and at most only about 5% of the fungal species on the Earth have been discovered (Hawksworth 1991, 2001). Major of the taxa of Fungi need to be uncovered (e.g. Jones et al. 2011). A recent estimation of worldwide diversity of macrofungi, including basidiomycetes and ascomycetes with large, easily observed spore-bearing structures that form above or below ground, calculated only 16–41% of macrofungi to be known to science and that endemism levels for macrofungi may be as high as 40–72% (Mueller et al. 2007). Bauer et al. (2006) pointed out that the ca. 8,000 described species of the simple-septate basidiomycetes may only represent the tip of the iceberg of this tremendous morphological and ecological diversified group.
On the other hand, it was assumed that Fungi are widely distributed, and consequently, for instance, many European or North American names were applied to morphologically similar Asian fungi. Recent data has shown that some species of Fungi, either saprotrophic or ectomycorrhizal or pathogenic, are indeed intercontinentally widely distributed, while many others are restricted in their range (Dai et al. 2003; Li et al. 2009; Liang et al. 2009; Dai 2010; Desprez-Loustau et al. 2011; O’Donnell et al. 2011). In consideration of global changes and dramatic deterioration of environments, largely due to human activities, acceleration of the inventory of fungi including basidiomycetes is an urgent task (Mueller et al. 2004; Piepenbring 2007).
Over the course of evolution, innumerous fungal taxa, such as plants and animals, have become extinct. Some unknown “living fossils” or unique taxa of basidiomycetes may be found in associated with plant living fossils. For instance, Bartheletia paradoxa, growing on leaf litter of Ginkgo biloba has a unique septal structure, and, like G. biloba, is a living fossil at the basal branching of the Agaricomycotina, which apparently used G. biloba as its Noah’s Ark (Scheuer et al. 2008). Taxa of significance in elucidating the phylogeny of Basidiomycota could well be harbored on living fossils of plants (e.g. Manchester et al. 2009). On the other hand, numerous previously unknown or little-known taxa in the tropics (Oberwinkler and Bauer 1990; Isaac et al. 1993; Watling et al. 2002) and on diverse substrates in other regions will contribute to a better understanding of the fungal diversity and evolution (e.g. Piepenbring 1996, 2007; Kirschner et al. 2001a, b; Kirschner and Chen 2004; Binder et al. 2006; Choeyklin et al. 2009; Coelho et al. 2009; Kirschner et al. 2010; Weiß et al. 2004b, 2011).
Molecular-data-based fungal systematics and phylogenetics have evolved very rapidly in the last two decades. However, morphological characters and ultrastructure, ecological traits, biochemical characters, chemical secondary metabolites as well as molecular phylogeny are all equally important in the understanding the evolution of the basidiomyctes. For instance, many hypotheses proposed in the last century based on morphology, ultrastructure, structure of pigments or metabolites have been verified by molecular approaches in the last two decades. To understand the megadiversity of basidimomycetes, multiple methodologies, thus, should be used (Bauer et al. 2001; Petersen and Hughes 2007; Wannathes et al. 2009; Hyde et al. 2010), although the shift from classical to molecular fungal taxonomy and systematics is becoming popular and inevitable (Seifert 2009). It may be worthy to mention that the integration of the on-going efforts of DNA barcoding into the inventory will accelerate the recovery and precise identification of a large number of unculturable, microscopic, and cryptic taxa of basidiomycetes (Moncalvo 2005; Begerow et al. 2010; Jargeat et al. 2010).
It is anticipated that numerous species, some monophyletic groups representing generic and suprageneric new taxa should be recognized within the Basidiomycota in the next few years (e.g. Binder et al. 2010). However, taxonomy, including fungal taxonomy, faces serious challenges (Agnarsson and Kuntner 2007), and thus, fungal taxonomists should consider adopting new modes of working (Hibbett et al. 2011), in order to accelerate the discovery and documentation of the world’s fungal heritage.
-
2)
Genome-based analyses of phylogeny and functional evolution
There has been a dramatic growth in multilocus fungal phylogenies in the last few years. Analyses of multigene sequences have resolved many major clades of Fungi, and have enabled development of a higher-level classification for the kingdom (e.g. Hibbett et al. 2007). Nevertheless, the framework is complete, but detailed information within the framework is largely absent, and there are some problematic deep nodes that are not well resolved, which limits our understanding of the evolutionary history of the Fungi (McLaughlin et al. 2009). Complete fungal genomes may reveal robust deep nodes of fungal tree of life (Fitzpatrick et al. 2006; Kuramae et al. 2006).
The use of high-throughput sequencing or next-generation sequencing technologies can produce dozens of gigabases per day. Future studies based on genomic data coupling efficient informatics approaches (e.g. Liu et al. 2009; Löytynoja and Goldman 2009) may contribute to the resolution of the major problematical nodes in the phylogeny of basidiomycetes and provide insight into its morphological, ecological and functional evolution. For instance, genome-based analyses may well resolve the backbone of the Agaricomycotina phylogeny and elucidate the diversity and evolution of the white rot and brown rot wood-decaying modes and shifts among hosts.
-
3)
Biogeographic inference
In comparison to plant or animal biogeography, biogeography of fungi is at its very young stages. For instance, understanding of the role of long distance dispersal of spores in the maintenance of fungal species cohesion is in its infancy. Some data suggest that fungal spores are seldom dispersed for distances greater than 100 m indicating that despite rare long distance dispersal events, significant gene flow via spore dispersal even between islands within Hawaii is quite unlikely (Bergemann and Miller 2002; Burnett 2003), while others suggests that a single fungal species can sustain appreciable gene flow across virtually global distributions (James et al. 2001; Petersen and Hughes 2007).
Biogeographic studies in fungi were impeded by the poor knowledge concerning the accurate distribution of fungal species. Up to now, biogeography of diverse groups of basidiomycetes is still very speculative and is only supported by fragmentary observations. Studies based only on morphological characters may provide a very incomplete and oversimplified picture of distribution patterns and associated historical events (Wu et al. 2000). Many intriguing morphological similarity based geographic distribution patterns, such as the well-known “Asa Gray disjunction” or a vicariance pattern in the Grayan distribution, and the Gondwanan distribution observed in the past (e.g. Horak 1983; Redhead 1989; Halling 2001; Mueller et al. 2001; Yang 2005b; Petersen and Hughes 2007), could well be inferred by molecular phylogenetic analyses in order to provide a much better understanding of their origin, historical biogeography and dispersal.
A more detailed and accurate understanding of the origin and evolution of a few selected groups of basidiomycetes have been revealed in the last few years, and are compelling areas for future research. For instance, through analyses of ITS and 26S rDNA sequences, and mt-ssu rDNA, Hibbett (2001) demonstrated that there are two main clades of the genus Lentinus, one in the New World, the other in the Old World. The Old World/New World disjunction could be due to fragmentation of an ancient Laurasian range. An alternative Gondwanan hypothesis is not supported by the molecular clock age estimates. Only one long distance dispersal event must be invoked in Lentinula, that being between Australia and New Zealand. Despite having airborne spores, long distance dispersal is rare in Lentinula. Aanen et al. (2002) showed that the symbiosis between fungus-growing termites and their fungal symbionts, Termitomyces, has a single African origin and that secondary domestication of other fungi or reversal of mutualistic fungi to a freeliving state has not occurred, and host switching has been frequent, especially at the lower taxonomic levels, and nests of single termite species can have different symbionts.
Recently, Hosaka et al. (2008) elucidated the biogeography of false truffles in the Hysterangiales. Their data are consistent with an Australian, or eastern Gondwanan origin of these fungi with subsequent range extensions into the Northern Hemisphere. A mosaic of vicariance and long distance events appears most plausible to explain the current distribution patterns in the false truffles. Using a relaxed molecular clock method, Matheny et al. (2009) reconstructed a phylogeny of the Inocybaceae with a geological timeline. Their data showed that the Inocybaceae initially diversified no later than the Cretaceous in Palaeotropical settings, in association with angiosperms. Diversification within major clades of the family accelerated during the Palaeogene in north and south temperate regions, whereas several relictual lineages persisted in the tropics. Both vicariance and dispersal patterns are detected. Species from Neotropical and south temperate regions are largely derived from immigrant ancestors from north temperate or Palaeotropical regions.
Without any doubt, more and more such studies on historical biogeography and evolution of different groups of basidiomycetes will soon appear.
-
4)
Study on species complex and cryptic species: to understand speciation and adaptation
Fungal speciation is one of the most fundamental issues of mycology (Kohn 2005; Giraud et al. 2008). The advent of molecular biology in the last 20 years has dramatically improved our ability to reveal cryptic diversity, speciation, and local adaption in basidiomycetes. Recent studies have shown that many morphospecies are complex or aggregates of taxa with distinct geographic, ecological or pathological traits, comprising several biological and/or phylogenetic species (e.g. Le Gac et al. 2007; Geml et al. 2008; Stubbe et al. 2010; O’Donnell et al. 2011).
It was found that there is often strong host specialization in basidiomycetes (e.g. Piepenbring et al. 1999; Begerow et al. 2004; Shefferson et al. 2007). However, high host specificity does not exclude possibilities for host shifts/host jumps, i.e., evolutionary lability (Parker and Gilbert 2004). Indeed, host jumps and host shifts are thought to be major driving forces in the evolution of basidiomycetes (Roy 2001; den Bakker et al. 2004; Refrégier et al. 2008; Li et al. 2009; Vercken et al. 2010; Li et al. 2011; Rochet et al. 2011). The next generation of high-throughput DNA sequencing methodologies, taking into account not only multilocus markers but also genome-wide studies, will provide enormous amounts of data for the study of phylogeny and population genetics, the reconstruction of patterns of origin and speciation in both time and space of basidiomycetes, leading to a more complete understanding of evolutionary and adaptation processes. The availability of molecular tools will prompt and yield a large number of new and highly interesting results in the near future.
References
Aanen DK, Eggleton P, Rouland-Lefèvre C et al (2002) The evolution of fungus-growing termites and their mutualistic fungal symbionts. Proc Natl Acad Sci USA 99:14887–14892
Agnarsson I, Kuntner M (2007) Taxonomy in a changing world: seeking solutions for a science in crisis. Syst Biol 56:531–539
Aime MC, Matheny PB, Henk D et al (2007) An overview of the higher-level classification of Pucciniomycotina based on combined analyses of nuclear large and small subunit rDNA sequences. Mycologia 98:896–905
Ainsworth GC, Sparrow FK, Sussman AS (eds) (1973) The fungi, an advanced treatise. Vol. IV B, a taxonomic review with keys: basidiomycetes and lower fungi. Academic, New York
Albee-Scott SR (2007) Does secotioid inertia drive the evolution of false-truffles? Mycol Res 111:1030–1039
Bartnicki-Garcia S (1968) Cell wall chemistry, morphogenesis, and taxonomy of fungi. Annu Rev Microbiol 22:87–108
Bas C (1975) A comparison of Torrendia(Gasteromycetes)with Amanita (Agaricales). In: Bigelow HE, Thiers HD (ed.) Studies on higher fungi. Nova Hedwigia Beiheft 51:53–60
Bauer R, Oberwinkler F, Vánky K (1997) Ultrastructural markers and systematics in smut fungi and allied taxa. Can J Bot 75:1273–1314
Bauer R, Begerow D, Oberwinkler F et al (2001) Ustilaginomycetes. In: McLaughlin DJ, McLaughlin EG, Lemke PA (eds) The Mycota. VII (B). Systematics and evolution. Springer, Berlin, pp 57–83
Bauer R, Begerow D, Sampaio JP et al (2006) The simple-septate basidiomycetes: a synopsis. Mycol Prog 5:41–66
Begerow D, Göker M, Lutz M et al (2004) On the evolution of smut fungi on their hosts. In: Agerer R, Piepenbring M, Blanz P (eds) Frontiers in basidiomyocte mycology. IHW-Verlag, Eching, pp 81–98
Begerow D, Nilsson H, Unterseher M et al (2010) Current state and perspectives of fungal DNA barcoding and rapid identification procedures. Appl Microbiol Biotechnol 87:99–108
Bergemann SE, Miller SL (2002) Size, distribution, and persistence of genets in local populations of the late-stage ectomycorrhizal basidiomycete, Russula brevipes. New Phytol 156:313–320
Binder M, Bresinsky A (2002) Derivation of a polymorphic lineage of gasteromycetes from boletoid ancestors. Mycologia 94:85–98
Binder M, Hibbett DS (2007) Molecular systematics and biological diversification of Boletales. Mycologia 98:971–981
Binder M, Hibbett DS, Wang Z et al (2006) Evolutionary relationships of Mycaureola diseae (Agaricales), a basidiomycete pathogen of a subtidal rhodophyte. Am J Bot 93:547–556
Binder M, Larsson K-H, Matheny PB et al (2010) Amylocorticiales ord. nov. and Jaapiales ord. nov.: early diverging clades of Agaricomycetidae dominated by corticioid forms. Mycologia 102:865–880
Blackwell M, Hibbett DS, Taylor JW et al (2007) Research coordination networks: a phylogeny for kingdom Fungi (Deep Hypha). Mycologia 98:829–837
Boekhout T, Guého E (2002) Basidiomycetous yeasts. In: Howard DH (ed) Pathogenic fungi in humans and animals. Marcel Dekker, New York, pp 535–564
Bougher NL (1999) New species of Torrendia (Fungi, Agaricales) from remnant woodlands in the wheatbelt region of Western Australia. Aust Syst Bot 12:145–156
Bougher NL, Lebel T (2002) Australasian sequestrate (truffle-like) fungi. XII. Amarrendia gen. nov.: an astipate, sequestrate relative of Torrendia and Amanita (Amanitaceae) from Australia. Aust Syst Bot 15:513–525
Brefeld O (1888) Basidiomyceten. II. Protobasidiomyceten. Untersuchungen aus dem Gesammtgebiete der Mykologie 7:1–178
Bruns TD, Fogel R, White TJ et al (1989) Accelerated evolution of a false truffle from a mushroom ancestor. Nature 339:140–142
Bruns TD, Vilgalys R, Barns SM et al (1992) Evolutionary relationships within the fungi: analyses of nuclear small subunit RNA sequences. Mol Phylogenet Evol 1:231–241
Burnett J (2003) Fungal populations and species. Oxford University Press, Oxfor
Celio GJ, Padamsee M, Dentinger BT et al (2006) Assembling the fungal tree of life: constructing the structural and biochemical database. Mycologia 98:850–859
Choeyklin R, Hattori T, Jaritkhuan S et al (2009) Bambusicolous polypores collected in Central Thailand. Fungal Divers 36:121–128
Coelho G, da Silveira RMB, Guerrero RT et al (2009) On poroid Hymenochaetales growing on bamboos in southern Brazil and NE Argentina. Fungal Divers 36:1–8
Corner EJH (1966) A monograph of cantharelloid fungi. Oxford University Press, Oxford
Cummins GB, Hiratsuka Y (1983) Illustrated genera of rust fungi. The American Phytopathological Society, St. Paul
Cunningham GH (1965) Polyporaceae of Australia and New Zealand. New Zeal Depart Sci Ind Res Bull 164:1–304
Dai Y-C (2010) Hymenochaetaceae (Basidiomycota) in China. Fungal Divers 45:131–343
Dai Y-C, Vainio EJ, Hantula J et al (2003) Investigations on Heterobasidion annosum s. lat. in central and eastern Asia with the aid of mating tests and DNA fingerprinting. For Pathol 33:269–286
de Bary HA (1853) Untersuchungen über die Brandpilze und die durch sie verursachten Krankheiten der Pflanzen mit Rücksicht auf das Getreide und andere Nutzpflanzen. Habilitationsschrift. Müller, Berlin
de Bary HA (1866) Morphologie und Physiologie der Pilze, Flechten und Myxomyceten. Engelmann, Leipzig
de Queiroz K (1998) The general lineage concept of species, species criteria, and the process of speciation. In: Howard DJ, Berlocher SH (eds) Endless forms: species and speciation. Oxford University Press, Oxford, pp 57–75
de Queiroz K (2007) Species concepts and species delimitation. Syst Biol 56:879–886
den Bakker HC, Zuccarello GC, Kuyper TW et al (2004) Evolution and host specificity in the ectomycorrhizal genus Leccinum. New Phytol 163:201–215
Dennis RWG (1970) Fungus flora of Venezuela and adjacent countries. Kew Bulletin Additional Series 3:1–531
Desjardin DE (1990) Culture morphology of Marasmius species. Sydowia 42:17–87
Desprez-Loustau M-L, Feau N, Mougou-Hamdane A et al (2011) Interspecific and intraspecific diversity in oak powdery mildews in Europe: coevolution history and adaptation to their hosts. Mycoscience 52:165–173
Donk MA (1964) A conspectus of the families of Aphyllophorales. Persoonia 3:199–324
Donk MA (1971) Progress in the study of the classification of the higher basidiomycetes. In: Petersen RH (ed) Evolution in the higher basidiomycetes. University of Tennessee Press, Knoxville, pp 3–25
Fell JW, Boekhout T, Fonseca A et al (2000) Biodiversity and systematics of basidiomycetous yeasts as determined by large-subunit rDNA D1/D2 domain sequence analysis. Int J Syst Evol Microbiol 50:1351–1371
Fitzpatrick DA, Logue ME, Stajich JE et al (2006) A fungal phylogeny based on 42 complete genomes derived from supertree and combined gene analysis. BMC Evol Biol 6:99
Fries E (1821) Systema mycologicum I. Gryphiswaldiae
Garrido N (1988) Agaricales s. l. und ihre Mykorrhizen in den Nothofagus-Wäldern Mittelchiles. Bibl Mycol 120:13–528
Gäumann E (1964) Die Pilze: Grundzüge ihrer Entwicklungsgeschichte und Morphologie. 2. Aufl. Birkhäuser Verlag, Basel
Geml J, Tulloss RE, Laursen GA et al (2008) Evidence for strong inter- and intracontinental phylogeographic structure in Amanita muscaria, a wind-dispersed ectomycorrhizal basidiomycete. Mol Phylogenet Evol 48:694–701
Giraud T, Refregier G, Le Gac M et al (2008) Speciation in fungi. Fungal Genet Biol 45:791–802
Gordon SA, Petersen RH (1991) Mating systems in Marasmius. Mycotaxon 41:371–385
Guarro J, Gene J, Stchigel AM (1999) Developments in fungal taxonomy. Clin Microbiol Rev 12:454–500
Guo L (2000) Flora fungorum sinicorum. Vol. 12. Ustilaginaceae. Science, Beijing
Halling RE (1983) The genus Collybia (Agaricales) in the northeastern United States and adjacent Canada. Mycol Mem 8:1–148
Halling RE (2001) Ectomycorrhizae: co-evolution, significance, and biogeography. Ann Mo Bot Gard 88:5–13
Hawksworth DL (1991) The fungal dimension of biodiversity: magnitude, significance, and conservation. Mycol Res 95:641–655
Hawksworth DL (2001) The magnitude of fungal diversity: the 1.5 million species estimate revisited. Mycol Res 105:1422–1432
Heim R (1971) The interrelationships between the Agaricales and gasteromycetes. In: Petersen RH (ed) Evolution in the higher basidiomycetes. The University of Tennessee Press, Knoxville, pp 505–534
Heinemann P (1972) Flore Iconographique des Champignons du Congo. Bruxelles
Henkel TW, Smith ME, Aime MC (2010) Guyanagaster, a new wood-decaying sequestrate fungal genus related to Armillaria (Physalacriaceae, Agaricales, Basidiomycota). Am J Bot 97:1–11
Hesler LR, Smith AH (1979) North American species of Lactarius. The University of Michigan Press, Ann Arbor
Hibbett DS (2001) Shiitake mushrooms and molecular clocks: historical biogeography of Lentinula. J Biogeogr 28:231–241
Hibbett DS (2004) Trends in morphological evolution in homobasidiomycetes inferred using maximum likelihood: a comparison of binary and multistate approaches. Syst Biol 53:889–903
Hibbett DS (2007) After the gold rush, or before the flood? Evolutionary morphology of mushroom-forming fungi (Agaricomycetes) in the early 21st century. Mycol Res 111:1001–1018
Hibbett DS, Thorn RG (2001) Basidiomycota: homobasidiomycetes. In: McLaughlin DJ, McLaughlin EG, Lemke PA (eds) The Mycota VII (B). Systematics and Evolution. Springer, Berlin, pp 121–168
Hibbett DS, Pine EM, Langer E et al (1997) Evolution of gilled mushrooms and puffballs inferred from ribosomal DNA sequences. Proc Natl Acad Sci USA 94:12002–12006
Hibbett DS, Binder M, Bischoff JF et al (2007) A higher-level phylogenetic classification of the Fungi. Mycol Res 111:509–547
Hibbett DS, Ohman A, Glotzer D et al (2011) Progress in molecular and morphological taxon discovery in Fungi and options for formal classification of environmental sequences. Fungal Biol Rev 25:38–47
Hillis DM, Dixon MT (1991) Ribosomal DNA: molecular evolution and phylogenetic inference. Q Rev Biol 66:411–453
Hiratsuka N, Sato S, Katsuya K et al (1992) The rust flora of Japan. Tsukuba Shuppankai, Tsukuba
Hjortstam K, Larsson K-H, Ryvarden L (1987) The Corticiaceae of North Europe, vol 1. Fungiflora, Oslo
Hjortstam K, Larsson K-H, Ryvarden L et al (1988) The Corticiaceae of North Europe, vol 8. Fungiflora, Oslo
Horak E (1968) Synopsis generum agaricalium (Die Gattungstypen der Agaricales). Beiträge zur Kryptogamenflora der Schweiz 13:1–741
Horak E (1983) Mycogeography in the South Pacific region: Agaricales, Boletales. Aust J Bot Suppl Ser 10:1–41
Hosaka K, Bates ST, Beever RE et al (2007) Molecular phylogenetics of the gomphoid-phalloid fungi with an establishment of the new subclass Phallomycetidae and two new orders. Mycologia 98:949–959
Hosaka K, Castellano MA, Spatafora JW (2008) Biogeography of Hysterangiales (Phallomycetidae, Basidiomycota). Mycol Res 112:448–462
Hyde KD, Abd-Elsalam K, Cai L (2010) Morphology: still essential in a molecular world. Mycotaxon 114:439–451
Hyde KD, McKenzie EHC, Ko TW (2011) Towards incorporating anamorphic fungi in a natural classification – checklist and notes for 2010. Mycosphere 2:1–88
Imazeki R, Otani Y, Hongo T (1988) Fungi of Japan. Yama-kei Publishers Co Ltd., Tokyo
Isaac S, Frankland JC, Watling R, Whalley AJS (1993) Aspects of tropical mycology. The University Press, Cambridge
James TY, Moncalvo J, Li S et al (2001) Polymorphism at the ribosomal DNA spacers and its relation to breeding structure of the widespread mushroom Schizophyllum commune. Genetics 157:149–161
James TY, Kauff F, Schoch C et al (2006) Reconstructing the early evolution of the fungi using a six gene phylogeny. Nature 443:818–822
Jargeat P, Martos F, Carriconde F et al (2010) Phylogenetic species delimitation in ectomycorrhizal fungi and implications for barcoding: the case of the Tricholoma scalpturatum complex (Basidiomycota). Mol Ecol 19:5216–5230
Jones MDM, Forn I, Gadelha C et al (2011) Discovery of novel intermediate forms redefines the fungal tree of life. Nature 474:200–203
Jülich W (1981) Higher taxa of basidiomycetes. Cramer, Lehre
Justo A, Morgenstern I, Hallen-Adams HE et al (2010) Convergent evolution of sequestrate forms in Amanita under Mediterranean climate conditions. Mycologia 102:675–688
Kauserud H, Stensrud O, Decock C et al (2006) Multiple gene genealogies and AFLPs suggest cryptic speciation and long-distance dispersal in the basidiomycete Serpula himantioides (Boletales). Mol Ecol 15:421–431
Khan SR, Kimbrough JW (1982) A reevaluation of the basidiomycetes based upon septal and basidial structures. Mycotaxon 15:103–210
Kimbrough JW (1994) Septal ultrastructure and ascomycete systematics. In: Hawksworth DL (ed) Ascomycete systematics: problems and perspectives in the nineties. Plenum, New York, pp 127–141
Kirk PM, Cannon PF, Minter DW et al (2008) Ainsworth & Bisby’s dictionary of the fungi, 10th edn. CABI, Wallingford
Kirschner R, Chen C-J (2004) Helicomyxa everhartioides, a new helicosporous sporodochial hyphomycete from Taiwan with relationships to the Hyaloriaceae (Auriculariales, Basidiomycota). Stud Mycol 50:337–342
Kirschner R, Bauer R, Oberwinkler F (2001a) Colacosiphon: a new genus described for a mycoparasitic fungus. Mycologia 93:634–644
Kirschner R, Sampaio JP, Gadanho M et al (2001b) Cuniculitrema polymorpha (Tremellales, gen. nov. and sp. nov.), a heterobasidiomycete vectored by bark beetles, which is the teleomorph of Sterigmatosporidium polymorphum. Antonie Van Leeuwenhoek 80:149–161
Kirschner R, Yang ZL, Zhao Q et al (2010) Ovipoculum album, a new anamorph with gelatinous cupulate bulbilliferous conidiomata from China and with affinities to the Auriculariales (Basidiomycota). Fungal Divers 43:55–65
Kohn LM (2005) Mechanisms of fungal speciation. Ann Rev Phytopathol 43:279–308
Korhonen K (1978a) Interfertility and clonal size in the Armillariella mellea complex. Karstenia 18:31–42
Korhonen K (1978b) Intersterility groups of Heterobasidion annosum. Communicationes Instituti Forestalis Fenniae 94:1–25
Kreisel H (1969) Grunndzüge eines natürlichen Systems der Pilze. J Cramer, Lehre
Kretzer A, Bruns TD (1997) Molecular revisitation of the genus Gastrosuillus. Mycologia 89:586–589
Kühner R (1980) Les Hyménomycètes Agaricoïdes (Agaricales, Tricholomatales, Pluteales, Russulales). Etude générale et classification. Numéro spécial du Bulletin Mensuel de la Société Linnéenne de Lyon 49e année:1–1027
Kuramae EE, Robert V, Snel B et al (2006) Phylogenomics reveal a robust fungal tree of life. FEMS Yeast Res 6:1213–1220
Kurtzman CP, Fell JW (eds) (1998) The yeasts: a taxonomic study, 4th edn. Amsterdam, Elsevier
Le Gac M, Hood ME, Fournier E et al (2007) Phylogenetic evidence of host-specific cryptic species in the anther smut fungus. Evolution 61:15–26
Lebel T, Catcheside PS (2009) The truffle genus Cribbea (Physalacriaceae, Agaricales) in Australia. Aust Syst Bot 22:39–55
Li YC, Yang ZL, Tolgor B (2009) Phylogenetic and biogeographic relationships of Chroogomphus species as inferred from molecular and morphological data. Fungal Divers 38:85–104
Li YC, Feng B, Yang ZL (2011) Zangia, a new genus of Boletaceae supported by molecular and morphological evidence. Fungal Divers. doi:10.1007/s13225-011-0096-y
Liang JF, Xu J, Yang ZL (2009) Divergence, dispersal and recombination in Lepiota cristata from China. Fungal Divers 38:105–124
Liu YJ, Hodson MC, Hall BD (2006) Loss of the flagellum happened only once in the fungal lineage: phylogenetic structure of Kingdom Fungi inferred from RNA polymerase II subunit genes. BMC Evol Biol 6:74. doi:10.1186/1471-2148-6-74
Liu K, Raghavan S, Nelesen S et al (2009) Rapid and accurate large-scale coestimation of sequence alignments and phylogenetic trees. Science 324:1561–1564
Löytynoja A, Goldman N (2009) Uniting alignments and trees. Science 324:1528–1529
Lutzoni F, Kauff F, Cox CJ et al (2004) Assembling the fungal tree of life: progress, classification, and evolution of subcellular traits. Am J Bot 91:1446–1480
Manchester SR, Chen Z-D, Lu A-M et al (2009) Eastern Asian endemic seed plant genera and their paleogeographic history throughout the Northern Hemisphere. J Syst Evol 47:1–42
Martin MP, Hogberg N, Listosella J (1999) Macowanites messapicoides, a hypogeous relative of Russula messapica. Mycol Res 103:203–208
Matheny PB, Curtis JM, Hofstetter V et al (2007a) Major clades of Agaricales: a multi-locus phylogenetic overview. Mycologia 98:982–995
Matheny PB, Wang Z, Binder M et al (2007b) Contributions of rpb2 and tef1 to the phylogeny of mushrooms and allies (Basidiomycota, Fungi). Mol Phylogenet Evol 43:430–451
Matheny PB, Gossman JA, Zalar P et al (2007c) Resolving the phylogenetic position of the Wallemiomycetes: an enigmatic major lineage of Basidiomycota. Can J Bot 84:1794–1805
Matheny PB, Aime MC, Bougher NL et al (2009) Out of the Paleotropics? Historical biogeography and diversification of the cosmopolitan ectomycorrhizal mushroom family Inocybaceae. J Biogeogr 36:577–592
Mayden RL (1997) A hierarchy of species concepts: the denoument in the saga of the species problem. In: Claridge MF, Dawah HA, Wilson MR (eds) Species: the units of diversity. Chapman and Hall, London, pp 381–423
McLaughlin DJ, Frieders EM, Lü H (1995) A microscopist’s view of heterobasidiomycete phylogeny. Stud Mycol 38:91–109
McLaughlin DJ, Hibbett DS, Lutzoni F et al (2009) The search for the fungal tree of life. Trends Microbiol 17:488–497
Miller OK Jr (1971) The relationship of cultural characters to the taxonomy of the agarics. In: Petersen RH (ed) Evolution in higher basidiomycetes. The University of Tennessee Press, Knoxville, pp 197–215
Miller OK Jr, Horak E (1992) Observations on the genus Torrendia and a new species from Australia. Mycologia 84:64–71
Miller OK Jr, Miller HH (1988) Gasteromycetes – Morphological and developmental features with keys to the orders, families, and genera. Mad River, Eureka
Moncalvo J-M (2005) Molecular Systematics: major fungal phylogenetic groups and fungal species concepts. In: Xu J (ed) Evolutionary genetics of fungi. Horizon Bioscience, Norfolk, pp 1–33
Moncalvo J-M, Vilgalys R, Redhead SA et al (2002) One hundred and seventeen clades of euagarics. Mol Phylogenet Evol 23:357–400
Moore RT (1985) The challenge of the dolipore/parenthosome septum. In: Moore D, Casselton LA, Wood DA et al (eds) Developmental biology of higher fungi. Cambridge University Press, Cambridge, pp 175–212
Moore RT (1997) Evolutionary advances in the higher fungi. Antonie Van Leeuwenhoek 72:209–218
Moser M (1983) Die Röhrlinge und Blätterpilze (Polyporales, Boletales, Agaricales, Russulales). In: Gams H (Hrg.) Kleine Kryptogamenflora, Band II b/2. Basidiomyceten, 2. Teil, 5. Aufl. Gustav Fischer Verlag, Stuttgart, pp 1–532
Mueller GM (1992) Systematics of Laccaria (Agaricales) in the continental United States and Canada, with discussions on extralimital taxa and descriptions of extant types. Fieldiana Botany New Series 30:1–158
Mueller GM, Wu QX, Huang YQ et al (2001) Assessing biogeographic relationships between North American and Chinese macrofungi. J Biogeogr 28:271–281
Mueller GM, Bills GF, Foster MS (2004) Biodiversity of fungi, inventory and monitoring methods. Elsevier Academic Press, Amsterdam
Mueller GM, Schmit JP, Leacock PR et al (2007) Global diversity and distribution of macrofungi. Biodivers Conserv 16:37–48
Müller WH, Stalpers JA, Van Aelst AC et al (2000) The taxonomic position of Asterodon, Asterostroma and Coltricia inferred from the septal pore cap ultrastructure. Mycol Res 104:1485–1491
Nei M, Kumar S (2000) Molecular evolution and phylogenetics. Oxford University Press, Oxford
O’Donnell K, Rooney AP, Mills GL et al (2011) Phylogeny and historical biogeography of true morels (Morchella) reveals an early Cretaceous origin and high continental endemism and provincialism in the Holarctic. Fungal Genet Biol 48:252–265
Oberwinkler F (1977) Das neue System der Basidiomyceten. In: Frey W, Hurka H, Oberwinkler F (eds) Beiträge zur Biologie der niedrigen Pflanzen. Gstav Fischer Verlag, Stuttgart, pp 59–105
Oberwinkler F (1978) Was ist ein Basidiomycet? Zeitschrift für Mykologie 44:13–29
Oberwinkler F (1982) The significance of the morpholgy of the basidium in the phylogeny of basidiomycetes. In: Wells K, Wells EK (eds) Basidium and basidiocarp. Evolution, cytology, function, and development. Springer Verlag, New York, pp 9–35
Oberwinkler F (1985) Anmerkungen zur Evolution und Systematik der Basidiomyceten. Botanische Jahrbücher für Systematik. Pflanzengeschichte und Pflanzengeographie 107:541–580
Oberwinkler F, Bandoni RJ (1982) A taxonomic survey of the gasteroid, auricularioid Heterobasidiomycetes. Can J Bot 60:1726–1750
Oberwinkler F, Bauer R (1990) Cryptomycocolax: a new mycoparasitic heterobasidiomycete. Mycologia 82:671–692
Parker IM, Gilbert GS (2004) The evolutionary ecology of novel plant-pathogen interactions. Annu Rev Ecol Evol Syst 35:675–700
Pegler DN (1983) The genus Lentinus, a world monograph. HMSO, London
Peintner U, Bougher NL, Castellano MA et al (2001) Multiple origins of sequestrate fungi related to Cortinarius (Cortinariaceae). Am J Bot 88:2168–2179
Peng YB, Liu B, Fan L (1992) Flora fungorum sinicorum. Vol. 2. Tremellales et Dacrymycetales. Science, Beijing
Persoon CH (1801) Synopsis methodica fungorum. H Dieterich, Gottingae
Petersen RH (1981) Ramaria subgenus Echinoramaria. Bibl Mycol 79:1–261
Petersen RH, Gordon SA (1994) Mating systems in hymenomycetes: new reports and new species. Mycologia 86:743–757
Petersen RH, Halling RE (1993) Mating systems in the Xerulaceae: Oudemansiella. Trans Mycol Soc Jpn 34:409–422
Petersen RH, Hughes KW (2007) Some agaric distribution patterns involving Pacific landmasses and Pacific Rim. Mycoscience 48:1–14
Piepenbring M (1996) Smut fungi (Ustilginales and Tilletiales) in Costa Roca. Nova Hedwigia Beiheft 113:1–155
Piepenbring M (2007) Inventoring the fungi of Panama. Biodivers Conserv 16:73–84
Piepenbring M, Begerow D, Oberwinkler F (1999) Molecular sequence data assess the value of morphological characteristics for a phylogenetic classification of species of Cintractia. Mycologia 91:485–498
Prillinger H, Oberwinkler F, Umile C et al (1993) Analysis of cell wall carbohydrates (neutral sugars) from ascomycetous and basidiomycetous yeasts with and without derivatization. J Gen Appl Microbiol 39:1–34
Redhead SA (1989) A biogeographical overview of the Canadian mushroom flora. Can J Bot 67:3003–3062
Refrégier G, Le Gac M, Jabbour F et al (2008) Cophylogeny of the anther smut fungi and their caryophyllaceous hosts: prevalence of host shifts and importance of delimiting parasite species. BMC Evol Biol 8:100. doi:10.1186/1471-2148-8-100
Reid DA (1980) A monograph of the Australian species of Amanita Pers. ex Hook. (fungi). Aust J BotSuppl Ser 8:1–97
Reijnders AFM (1963) Les problemes du developement des carpophores des Agaricales et de quelques groups voisins. W. Junk, The Hague
Rochet J, Moreau P-A, Manzi S et al (2011) Comparative phylogenies and host specialization in the alder ectomycorrhizal fungi Alnicola, Alpova and Lactarius (Basidiomycota) in Europe. BMC Evol Biol 11:40. doi:10.1186/1471-2148-11-40
Roy BA (2001) Patterns of association between crucifers and their flower-mimic pathogens: host jumps are more frequent than coevolution or cospeciation. Evolution 55:41–53
Ryvarden L, Gilbertson RL (1993) European polypores 1. Synop Fungorum 6:1–387
Ryvarden L, Gilbertson RL (1994) European polypores 2. Meripilus–Tyromyces. Synop Fungorum 6:389–743
Scheuer C, Bauer R, Lutz M et al (2008) Bartheletia paradoxa is a living fossil on Ginkgo leaf litter with a unique septal structure in the Basidiomycota. Mycol Res 112:1265–1279
Seifert KA (2009) Progress towards DNA barcoding of fungi. Mol Ecol Resour 9(Suppl 1):83–89
Shefferson RP, Taylor DL, Weiß M et al (2007) The evolutionary history of mycorrhizal specificity among lady’s slipper orchids. Evolution 61:1380–1390
Singer R (1962) The Agaricales in modern taxonomy, 2nd edn. J. Cramer, Weinheim
Singer R (1986) The Agaricales in modern taxonomy, 4th edn. Koeltz Scientific Books, Koenigstein
Singer R, Smith AH (1960) Studies on secotiaceous fungi IX. Memoirs of the Torrey Botanical Club 21:94–103
Stubbe D, Nuytinck J, Verbeken A (2010) Critical assessment of the Lactarius gerardii species complex (Russulales). Fungal Biol 114:271–283
Swann EC, Taylor JW (1995) Phylogenetic perspectives on basidiomycete systematics: evidence from the 18S rRNA gene. Can J Bot 73(Suppl):S862–S868
Taylor JW, Jacobson D, Kroken S et al (2000) Phylogenetic species recognition and species concepts in fungi. Fungal Genet Biol 31:21–32
Taylor JW, Spatafora J, O’Donnell K et al (2004) The fungi. In: Cracraft J, Donoghue MJ (eds) Assembling the fungal tree of life. Oxford University Press, Oxford. pp 171–194
Thiers HD (1984) The secotioid syndrome. Mycologia 76:1–8
Van de Putte K, Nuytinck J, Stubbe D et al (2010) Lactarius volemus sensu lato (Russulales) from northern Thailand: morphological and phylogenetic species concepts explored. Fungal Divers 45:99–130
Van der Walt JP, Yarrow D (1984) Methods for isolation, maintenance, classification, and identification of yeasts. In: Kregervan Rij NJW (ed) The yeasts, a taxonomic study. Elsevier Science Publishers, Amsterdam, pp 45–104
Van Driel KGA, Humbel BM, Verkleij AJ et al (2009) Septal pore complex morphology in the Agaricomycotina (Basidiomycota) with emphasis on the Cantharellales and Hymenochaetales. Mycol Res 113:559–576
Vánky K (1987) Illustrated genera of smut fungi. Gustav Fischer Verlag, Stuttgart
Vellinga EC (2004) Genera in the family Agaricaceae: evidence from nrITS and nrLSU sequences. Mycol Res 108:354–377
Vellinga EC, De Kok RPJ, Bruns TD (2003) Phylogeny and taxonomy of Macrolepiota (Agaricaceae). Mycologia 95:442–456
Vercken E, Fontaine MC, Gladieux P et al (2010) Glacial refugia in pathogens: European genetic structure of anther smut pathogens on Silene latifolia and Silene dioica. PLoS Pathog 6:e1001229. doi:10.1371/journal.ppat.1001229
Wannathes N, Desjardin DE, Hyde KD et al (2009) A monograph of Marasmius (Basidiomycota) from Northern Thailand based on morphological and molecular (ITS sequences) data. Fungal Divers 37:209–306
Watling R, Frankland JC, Ainsworth M et al (2002) Tropical mycology, Volume 1: macromycetes. CABI, Wallingford
Weiß M, Bauer R, Begerow D (2004a) Spotlights on heterobasidiomycetes. In: Agerer R, Piepenbring M, Blanz P (eds) Frontiers in basidiomyocte mycology. IHW-Verlag, Eching, pp 7–48
Weiß M, Selosse MA, Rexer KH et al (2004b) Sebacinales: a hitherto overlooked cosm of heterobasidiomycetes with a broad mycorrhizal potential. Mycol Res 108:1003–1010
Weiß M, Sýkorová Z, Garnica S et al (2011) Sebacinales everywhere: previously overlooked ubiquitous fungal endophytes. PLoS One 6:e16793. doi:10.1371/journal.pone.0016793
Wells K (1994) Jelly fungi, then and now. Mycologia 86:18–48
White TJ, Bruns T, Lee S et al (1990) Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In: Innis MA, Gelfand DH, Sninsky JJ, White TJ (eds) PCR protocols: a guide to methods and applications. Academic, San Diego, pp 315–322
Wilson AW, Binder M, Hibbett DS (2011) Effects of gasteroid fruiting body morphology on diversification rates in three independent clades of fungi estimated using binary state speciation and extinct analysis. Evolution 65:1305–1322
Wu QX, Mueller GM, Lutzoni FM et al (2000) Phylogenetic and biogeographic relationships of eastern Asian and eastern North American disjunct Suillus species (Fungi) as inferred from nuclear ribosomal RNA ITS sequences. Mol Phylogenet Evol 17:37–47
Yang ZL (2005a) Flora fungorum sinicorum. Vol. 27. Amanitaceae, vol 27. Science, Beijing
Yang ZL (2005b) Diversity and biogeography of higher fungi in China. In: Xu J (ed) Evolutionary genetics of fungi. Horizon Bioscience, Norfolk, pp 35–62
Zalar P, de Hoog GS, Schroers HJ et al (2005) Taxonomy and phylogeny of the xerophilic genus Wallemia (Wallemiomycetes and Wallemiales, cl. et ord. nov.). Antonie Leeuwenhoek 87:311–328
Zang M (2006) Flora fungorum sinicorum. Vol. 22. Boletales (I). Science, Beijing
Zhou TX (2007) Flora fungorum sinicorum. Vol. 36. Geastraceae and Nidulariaceae, vol 36. Science, Beijing
Zhuang JY, Wei SX, Wang YC (1998) Flora fungorum sinicorum. Vol. 10. Uredinales (I). Science, Beijing
Acknowledgements
I am very grateful to Prof. Dr. K. Hyde (Mae Fah Luang University, Thailand) for initiating this review and for his suggestions to improve the manuscript. Prof. Dr. J.Y. Zhuang and Prof. Dr. L. Guo (Institute of Microbiology, Chinese Academy of Sciences, China) are acknowledged for the identification of my collections of rusts and smuts respectively. Prof. Dr. M. Piepenbring (J. W. Goethe-Universität Frankfurt, Frankfurt/Main, Germany) is acknowledged for providing an image of Entorrhiza casparyana. This study was supported by the Joint Funds of the National Natural Science Foundation of China and Yunnan Provincial Government (No. U0836604), the National Basic Research Program of China (No. 2009CB522300), and the Hundred Talents Program of the Chinese Academy of Sciences.
Open Access
This article is distributed under the terms of the Creative Commons Attribution Noncommercial License which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
Open Access This is an open access article distributed under the terms of the Creative Commons Attribution Noncommercial License (https://creativecommons.org/licenses/by-nc/2.0), which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
About this article
Cite this article
Yang, Z.L. Molecular techniques revolutionize knowledge of basidiomycete evolution. Fungal Diversity 50, 47–58 (2011). https://doi.org/10.1007/s13225-011-0121-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13225-011-0121-1