Abstract
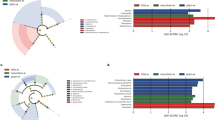
With the aim of learning more about functional bacterial communities in the tobacco rhizosphere and phyllosphere, a total of 96 nicotine-degrading (ND) bacterial strains isolated using nicotine as the sole carbon source—56 from the rhizosphere and 40 from the phyllosphere—were quantified and analyzed phylogenetically. The ND efficiency of 19 phyllosphere strains (47.5 %) and 39 rhizosphere strains (69.6 %) exceeded 90 %. Patterns of phylogenetic relationships based on 16S rRNA gene sequences revealed a high heterogeneity of community composition and suggested the existence of microenvironment-specific communities. The phyllospheric ND bacterial community was distributed over ten genera, of which Pseudomonas spp. was the dominant population. However, the rhizospheric ND bacterial community was composed of six genera, of which Arthrobacter spp. was the major group. This is the first report of members of genera Massilia, Erwinia, Brevundimonas and Paenibacillus capable of degrading nicotine. Diversity indices were calculated provisionally using sequence data obtained from each ND bacterial library. The species richness, diversity and dominance index of the ND bacterial community of the phyllosphere were higher than that of rhizosphere community, while the evenness index of the phyllopheric community was lower compared to rhizospheric ND bacteria. Metabolic intermediate detection showed that the Pseudomonads isolates possessed all four proposed metabolic pathways of nicotine degradation while the Arthrobacter strains all had only one pyridine pathway. These results greatly enhance our knowledge of the diversity of ND bacteria and demonstrate that the tobacco-associated micro-environment contain diverse and novel ND bacteria, which might be a valuable biotechnological resource for biodegradation of nicotine.


Similar content being viewed by others
References
Armstrong DW, Wang X, Ercal N (1998) Enantiomeric composition of nicotine in smokeless tobacco, medicinal products, and commercial reagents. Chiriality 10:587–591
Baitsch D, Sandu C, Brandsch R, Igloi GL (2001) Gene cluster on pAO1 of Arthrobacter nicotinovorans involved in degradation of the plant alkaloid nicotine: cloning, purification, and characterization of 2, 6-dihydroxypyridine 3-hydroxylase. J Bacteriol 183:5262–5267
Brandsch R (2006) Microbiology and biochemistry of nicotine degradation. Appl Microbiol Biotechnol 69:493–498
Chen CM, Li XM, Yang JK, Gong XW, Li X, Zhang KQ (2008) Isolation of nicotine-degrading bacterium Pseudomonas sp. Nic22, and its potential application in tobacco processing. Int Biodeterior Biodegrad 62:226–231
Chiribau CB, Mihasan M, Ganas P, Igloi GL, Artenie V, Brandsch R (2006) Final steps in the catabolism of nicotine. FEBS J 273:1528–1536
Coombs JT, Franco CM (2003) Isolation and identification of actinobacteria from surface-sterilized wheat roots. Appl Environ Microbiol 69:5603–5608
Gherna RL, Richardson SH, Rittenberg SC (1965) The bacterial oxidation of nicotine.VI. The metabolism of 2,6-dihydroxypseudooxynicotine. J Biol Chem 240:3669–3674
Gong XW, Yang JK, Duan YQ, Dong JY, Zhe W, Wang L, Li QH, Zhang KQ (2009) Isolation and characterization of Rhodococcus sp. Y22 and its potential application to tobacco processing. Res Microbiol 160:200–204
Hayat R, Ali S, Amara U, Khalid R, Ahmed I (2010) Soil beneficial bacteria and their role in plant growth promotion: a review. Ann Microbiol 60:579–598
Hill TC, Walsh KA, Harris JA, Moffett BF (2003) Using ecological diversity measures with bacterial communities. FEMS Microbiol Ecol 43:1–11
Hirano SS, Upper CD (2002) Bacteria in the leaf ecosystem with emphasis on Pseudomonas syringae, a pathogen, ice nucleus and epiphyte. Microbiol Mol Biol Rev 64:624–653
Hochstein LI, Rittenberg CS (1959) The bacterial oxidation of nicotine. II. The isolation of the first product and its identification as (l)-6-hydroxynicotine. J Biol Chem 234:156–162
Holmes PE, Rittenberg SC (1972) The bacterial oxidation of nicotine. VII. Partial purification and properties of 2,6-dihydroxypyridine oxidase. J Biol Chem 247:7622–7627
Huang J, Yang J, Duan Y, Gu W, Gong X, Zhe W, Su C, Zhang KQ (2010) Bacterial diversities on unaged and aging flue-cured tobacco leaves estimated by 16S rRNA sequence analysis. Appl Microbiol Biotechnol 88:553–562
Igloi GL, Brandsch R (2003) Sequence of the 165-kilobase catabolic plasmid pAO1 from Arthrobacter nicotinovorans and identification of a pAO1-dependent nicotine uptake system. J Bacteriol 185:1976–1989
Innerebner G (2011) Identification of indigenous phyllosphere isolates of the genus Sphingomonas as plant-protective bacteria. Dissertation, Leopold-Franzens-Universität Innsbruck
Jiang HJ, Ma Y, Qiu GJ, Wu FL, Chen SL (2011) Biodegradation of nicotine by a novel strain Shinella sp. HZN1 isolated from activated sludge. J Environ Sci Health B 48:703–708
Kimura M (1980) A simple method for estimating evolutionary rates of base substitutions through comparative studies of nucleotide sequences. J Mol Evol 16:111–120
Li HJ, Li XM, Duan YQ, Zhang KQ, Yang JK (2010) Biotransformation of nicotine by microorganism: the case of Pseudomonas spp. Appl Microbiol Biotechnol 86:11–17
Li HJ, Duan YQ, Ma GH, Lei LP, Zhang KQ, Yang JK (2011) Isolation and characterization of Acinetobacter sp. ND12 capable of degrading nicotine. Afr J Microbiol Res 5:1335–1341
Ma GH, Lei LP, Xia ZY, Gong XW, Zhou W, Yang JK (2012) Diversity and phylogenetic analyses of nicotine-degrading bacteria isolated from tobacco plantation soils. Afr J Microbiol Res 6:6392–6398
Novotny TE, Zhao F (1999) Consumption and production waste: another externality of tobacco use. Tob Control 8:75–80
Qiu J, Ma Y, Wen Y, Chen L, Wu L, Liu W (2012) Functional identification of two novel genes from Pseudomonas sp. strain HZN6 involved in the catabolism of nicotine. Appl Environ Microbiol 78:2154–2160
Raman G, Mohan K, Manohar V, Sakthivel N (2013) Biodegradation of nicotine by a novel nicotine-degrading bacterium, Pseudomonas plecoglossicida TND35 and its new biotransformation intermediates. Biodegradation 25(1):95–107. doi:10.1007/s10532-013-9643-4
Ruan AD, Min H, Peng X, Huang Z (2005) Isolation and characterization of Pseudomonas sp. strain HF-1, capable of degrading nicotine. Res Microbiol 156:700–706
Ruinen J (1956) Occurrence of Beijerinckia species in the phyllosphere. Nature (London) 177:220–221
Sachelaru P, Schiltz E, Igloi GL, Brandsch R (2005) An α/β-Fold C-C bond hydrolase is involved in a central step of nicotine catabolism by Arthrobacter nicotinovorans. J Bacteriol 187:8516–8519
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Schenk S, Hoelz A, Krauss B, Decker K (1998) Gene structures and properties of enzymes of the plasmid-encoded nicotine catabolism of Arthrobacter nicotinovorans. J Mol Biol 284:1323–1339
Sguros PL (1955) Microbial transformations of the tobacco alkaloids. I. Cultural and morphological characteristics of a nicotinophile. J Bacteriol 69:28–37
Smits THM, Rezzonico F, Kamber T, Blom J, Goesmann A, Frey JE, Duffy B (2010) Complete genome sequence of the fire blight pathogen Erwinia amylovora CFBP 1430 and comparison to other Erwinia spp. Mol Plant Microbe Interact 23:384–393
Su K, Gu W, Zhe W, Zhang KQ, Duan Y, Yang J (2011) Diversity and phylogeny of bacteria on Zimbabwe tobacco leaves estimated by 16S rRNA sequence analysis. Appl Microbiol Biotechnol 92:1033–1044
Su C, Lei L, Duan Y, Zhang KQ, Yang J (2012) Culture-independent methods for studying environmental microorganisms: methods, application, and perspective. Appl Microbiol Biotechnol 93:993–1003
Sun KD, Zhu CJ, Zhong WH, Chen JM, Ye ZJ, Liu PJ, Zhou Q (2008) Isolation and characterization of a high nicotine-degrading bacterium, Pseudomonas sp. strain ZUTSKD. Acta Sci Circum 28:1294–1301
Tang HZ, Wang SN, Ma LY, Meng XZ, Deng ZX, Zhang D, Ma CQ, Xu P (2008) A novel gene encoding 6-hydroxy-3-succinoylpyridine hydroxylase, involved in nicotine degradation by Pseudomonas putida strain S16. Appl Environ Microbiol 74:1567–1574
Thompson JD, Higgins DG, Gibson TJ (1994) CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res 22:4673–4680
Wada E, Yamasaki K (1954) Degradation of nicotine by soil bacteria. J Am Chem Soc 76:155–157
Wang SN, Liu Z, Tang HZ, Meng J, Xu P (2007) Characterization of environmentally friendly nicotine degradation by Pseudomonas putida biotype A strain S16. Microbiology 153:1556–1565
Wang SN, Liu Z, Xu P (2009) Biodegradation of nicotine by a newly isolated Agrobacterium sp. strain S33. J Appl Microbiol 107:838–847
Wang HH, Yin B, Peng XX, Wang Y, Xie ZH, Gao J, Tang XK (2012) Biodegradation of nicotine by newly isolated Pseudomonas sp. CS3 and its metabolites. J Appl Microbiol 112:258–268
Wei HL, Lei LP, Xia ZY, Liu XZ (2008) Characterization of a novel aerobic nicotine-biodegrading strain of Pseudomonas putida. Ann Microbiol 58:41–45
Xu X, Iba MM, Weisel CP (2004) Simultaneous and sensitive measurement of anabasine, nicotine, and nicotine metabolites in human urine by liquid chromatography–tandem mass spectrometry. Clin Chem 50:2323–2330
Yang CH, Crowley DE, Borneman J, Keen NT (2001) Microbial phyllosphere populations are more complex than previously realized. Proc Natl Acad Sci USA 98:3889–3894
Yuan YJ, Lu ZX, Wu N, Huang LJ, Lü FX, Bie XM (2005) Isolation and preliminary characterization of a novel nicotine-degrading bacterium, Ochrobactrum intermedium DN2. Int Biodeterior Biodegrad 56:45–50
Zhang W, Hu YG, Huang GH, Gao HW (2007) Soil microbial diversity of artificial peashrub plantation on North Loess Plateau of China. Acta Microbiol Sin 47:751–756
Acknowledgments
This work was supported by a grant from International Foundation for Science (IFS) (no. F/4583-1) to Hai-Lei Wei and two grants from Yunnan Tobacco Company (no. 2010YN16) and State Tobacco Monopoly Administration (no. 110201101035 JH-10).
Author information
Authors and Affiliations
Corresponding author
Additional information
Liping Lei and Zhenyuan Xia contributed equally to this work.
Rights and permissions
About this article
Cite this article
Lei, L., Xia, Z., Liu, X. et al. Occurrence and variability of tobacco rhizosphere and phyllosphere bacterial communities associated with nicotine biodegradation. Ann Microbiol 65, 163–173 (2015). https://doi.org/10.1007/s13213-014-0847-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13213-014-0847-6