Abstract
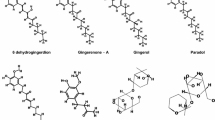
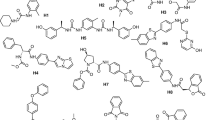
Staphylococcus aureus is a lethal pathogen that can cause various bacterial infections. This study targets the CrtM enzyme of S. aureus, which is crucial for synthesizing golden carotenoid pigment: staphyloxanthin, which provides anti-oxidant activity to this bacterium for combating antimicrobial resistance inside the host cell. The present investigation quests for human SQS inhibitors against the CrtM enzyme by employing structure-based drug design approaches including induced fit docking (IFD), molecular dynamic (MD) simulations, and binding free energy calculations. Depending upon the docking scores, two compounds, lapaquistat acetate and squalestatin analog 20, were identified as the lead molecules exhibit higher affinity toward the CrtM enzyme. These docked complexes were further subjected to 100 ns MD simulation and several thermodynamics parameters were analyzed. Further, the binding free energies (ΔG) were calculated for each simulated protein–ligand complex to study the stability of molecular contacts using the MM-GBSA approach. Pre-ADMET analysis was conducted for systematic evaluation of physicochemical and medicinal chemistry properties of these compounds. The above study suggested that lapaquistat acetate and squalestatin analog 20 can be selected as potential lead candidates with promising binding affinity for the S. aureus CrtM enzyme. This study might provide insights into the discovery of potential drug candidates for S. aureus with a high therapeutic index.


Similar content being viewed by others
Data availability
Data will be made available on request.
References
Ahmad B, Saeed A, Castrosanto MA et al (2022) Identification of natural marine compounds as potential inhibitors of CDK2 using molecular docking and molecular dynamics simulation approach. J Biomol Struct Dyn. https://doi.org/10.1080/07391102.2022.2135594
Aribisala JO, Sabiu S (2022) Cheminformatics identification of phenolics as modulators of penicillin-binding protein 2a of Staphylococcus aureus: a structure–activity-relationship-based study. Pharmaceutics 14:1818. https://doi.org/10.3390/pharmaceutics14091818
Banu S, Bollu R, Naseema M et al (2018) A novel templates of piperazinyl-1,2-dihydroquinoline-3-carboxylates: synthesis, anti-microbial evaluation and molecular docking studies. Bioorg Med Chem Lett 28:1166–1170. https://doi.org/10.1016/j.bmcl.2018.03.007
Berman HM (2000) The protein data bank. Nucleic Acids Res 28:235–242. https://doi.org/10.1093/nar/28.1.235
Che Omar MT (2020) Data analysis of molecular dynamics simulation trajectories of β-sitosterol, sonidegib and cholesterol in smoothened protein with the CHARMM36 force field. Data Brief 33:106350. https://doi.org/10.1016/j.dib.2020.106350
Choudhary MI, Shaikh M, Tul-Wahab A, Ur-Rahman A (2020) In silico identification of potential inhibitors of key SARS-CoV-2 3CL hydrolase (Mpro) via molecular docking, MMGBSA predictive binding energy calculations, and molecular dynamics simulation. PLOS ONE 15:e0235030. https://doi.org/10.1371/journal.pone.0235030
Dalal V, Dhankhar P, Singh V et al (2021) Structure-based identification of potential drugs against FmtA of Staphylococcus aureus: virtual screening, molecular dynamics, MM-GBSA, and QM/MM. Protein J 40:148–165. https://doi.org/10.1007/s10930-020-09953-6
Desmond Molecular Dynamics System, D. E. Shaw Research, New York, NY, 2021. Maestro-Desmond Interoperability Tools, Schrödinger, New York, NY, 2021
Elmesseri RA, Saleh SE, Elsherif HM et al (2022) Staphyloxanthin as a potential novel target for deciphering promising anti-Staphylococcus aureus agents. Antibiot Basel Switz 11:298. https://doi.org/10.3390/antibiotics11030298
Gao P, Davies J, Kao RYT (2017) Dehydrosqualene desaturase as a novel target for anti-virulence therapy against Staphylococcus aureus. mBio 8:e01224-17. https://doi.org/10.1128/mBio.01224-17
Gupta A, Chaudhary N, Aparoy P (2018) MM-PBSA and per-residue decomposition energy studies on 7-Phenyl-imidazoquinolin-4(5H)-one derivatives: Identification of crucial site points at microsomal prostaglandin E synthase-1 (mPGES-1) active site. Int J Biol Macromol 119:352–359. https://doi.org/10.1016/j.ijbiomac.2018.07.050
Husain A, Ahmad A, Khan SA et al (2016) Synthesis, molecular properties, toxicity and biological evaluation of some new substituted imidazolidine derivatives in search of potent anti-inflammatory agents. Saudi Pharm J 24:104–114. https://doi.org/10.1016/j.jsps.2015.02.008
Kahlon AK, Roy S, Sharma A (2010) Molecular docking studies to map the binding site of squalene synthase inhibitors on dehydrosqualene synthase of Staphylococcus Aureus. J Biomol Struct Dyn 28:201–210. https://doi.org/10.1080/07391102.2010.10507353
Kim S, Thiessen PA, Bolton EE et al (2016) PubChem substance and compound databases. Nucleic Acids Res 44:D1202–D1213. https://doi.org/10.1093/nar/gkv951
Kumari R, Rathi R, Pathak SR, Dalal V (2022) Structural-based virtual screening and identification of novel potent antimicrobial compounds against YsxC of Staphylococcus aureus. J Mol Struct 1255:132476. https://doi.org/10.1016/j.molstruc.2022.132476
Lee SK, Lee IH, Kim HJ et al (2003) The PreADME approach: web-based program for rapid prediction of physico-chemical, drug absorption and drug-like properties, EuroQSAR 2002 designing drugs and crop protectants: processes, problems and solutions. Blackwell Publishing, Massachusetts, pp 418–420
Liu C-I, Liu GY, Song Y et al (2008) A cholesterol biosynthesis inhibitor blocks Staphylococcus aureus virulence. Science 319:1391–1394. https://doi.org/10.1126/science.1153018
Marques SM, Bednar D, Damborsky J (2019) Computational study of protein-ligand unbinding for enzyme engineering. Front Chem 6:650. https://doi.org/10.3389/fchem.2018.00650
Metwaly AM, Elwan A, El-Attar A-AMM et al (2022) Structure-based virtual screening, docking, ADMET, molecular dynamics, and MM-PBSA calculations for the discovery of potential natural SARS-CoV-2 helicase inhibitors from the Traditional Chinese Medicine. J Chem 2022:1–23. https://doi.org/10.1155/2022/7270094
Musil M, Jezik A, Jankujova M et al (2022) Fully automated virtual screening pipeline of FDA-approved drugs using Caver Web. Comput Struct Biotechnol J 20:6512–6518. https://doi.org/10.1016/j.csbj.2022.11.031
Nayak C, Singh SK (2021) In silico identification of natural product inhibitors against Octamer-binding transcription factor 4 (Oct4) to impede the mechanism of glioma stem cells. PLoS ONE 16:e0255803. https://doi.org/10.1371/journal.pone.0255803
Pinto GP, Hendrikse NM, Stourac J et al (2022) Virtual screening of potential anticancer drugs based on microbial products. Semin Cancer Biol 86:1207–1217. https://doi.org/10.1016/j.semcancer.2021.07.012
Schrödinger Release 2022–4: LigPrep, Schrödinger, LLC, New York, NY, 2022. https://www.schrodinger.com/
Schrödinger Release 2022–4: Maestro, Schrödinger, LLC, New York, NY, 2022. https://www.schrodinger.com/
Schrödinger Release 2022–4: Prime, Schrödinger, LLC, New York, NY, 2022. https://www.schrodinger.com/
Singh G, Soni H, Tandon S et al (2022) Identification of natural DHFR inhibitors in MRSA strains: structure-based drug design study. Results Chem 4:100292. https://doi.org/10.1016/j.rechem.2022.100292
Stourac J, Vavra O, Kokkonen P et al (2019) Caver Web 1.0: identification of tunnels and channels in proteins and analysis of ligand transport. Nucleic Acids Res 47:W414–W422. https://doi.org/10.1093/nar/gkz378
Sullivan LE, Rice KC (2021) Measurement of Staphylococcus aureus pigmentation by methanol extraction. Methods Mol Biol Clifton NJ 2341:1–7. https://doi.org/10.1007/978-1-0716-1550-8_1
Thapa B, Raghavachari K (2019) Energy decomposition analysis of protein-ligand interactions using molecules-in-molecules fragmentation-based method. J Chem Inf Model 59:3474–3484. https://doi.org/10.1021/acs.jcim.9b00432
Xiong G, Wu Z, Yi J et al (2021) ADMETlab 2.0: an integrated online platform for accurate and comprehensive predictions of ADMET properties. Nucleic Acids Res 49:W5–W14. https://doi.org/10.1093/nar/gkab255
Author information
Authors and Affiliations
Contributions
SR and IB conceptualized and designed the research study; IB and VP performed data curation and experiments. IB and VP performed in-silico data analysis; IB and SR wrote the manuscript; VPR and SR critically reviewed and edited the manuscript. All authors approved the final version of the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors hereby declare that they have no conflict of interest in the publication of the manuscript.
Ethical approval
There is no involvement of any potential conflicts of interest, funding, or research involving human participation and/or animals.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Bhogal, I., Pankaj, V., Provaznik, V. et al. In silico investigation of cholesterol-lowering drugs to find potential inhibitors of dehydrosqualene synthase in Staphylococcus aureus. 3 Biotech 14, 39 (2024). https://doi.org/10.1007/s13205-023-03862-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s13205-023-03862-y