Abstract
COVID-19 is caused by severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2, has become a global pandemic resulting in significant morbidity and mortality. This study presents 12 new peptidomimetic fullerene-based derivatives in three groups that are investigated theoretically as SARS-CoV-2 Mpro inhibitors to increase the chance of treating COVID-19. Studied compounds are designed and optimized at B88-LYP/DZVP method. Molecular descriptors results show the stability and reactivity of the compounds with Mpro, especially in the 3rd group (Ser compounds). However, Lipinski's Rule of Five values indicates that the compounds are not suitable as oral drugs. Furthermore, molecular docking simulations are carried out to investigate the binding affinity and interaction modes of the top five compounds (compounds 1, 9, 11, 2, and 10) with the Mpro protein, which have the lowest binding energy. Molecular dynamics simulations are also performed to evaluate the stability of the protein–ligand complexes with compounds 1 and 9 and compare them with natural substrate interaction. The analysis of RMSD, H-bonds, Rg, and SASA indicates that both compounds 1 (Gly-α acid) and 9 (Ser-α acid) have good stability and strong binding affinity with the Mpro protein. However, compound 9 shows slightly better stability and binding affinity compared to compound 1.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
On December 31, 2019, infections of unknown etiology with pneumonia first appeared in Wuhan City, located in central China (Lu et al. 2020; Wang et al. 2020). Initially, WHO classified the virus that causes this pneumonia infection as 2019-nCoV (Hui et al. 2020) and then SARS-CoV-2 (Gorbalenya et al. 2020a). This infectious disease has been named Coronavirus disease 2019 (COVID-19) by WHO (Gorbalenya et al. 2020b). Since this time, our lives have changed upside down. Worldwide, there are > 663,000,000 confirmed cases and > 6,000,000 confirmed deaths as of July 2022 (Shrinivasan 2021; World Health Organization 2021). Despite the widespread use of vaccination, the need for effective treatments is urgent for those who do not respond adequately to the Coronavirus vaccination (Yan et al. 2021; Herman et al. 2022).
Coronavirinae is subfamily of Coronaviridae and is divided into 4 main genera: α, β, γ and δ-coronavirus. α and β-coronavirus are responsible for strains of Human coronaviruses (HCoVs). α-coronavirus has two strains: 229E & NL63, while β-coronavirus has five strains: OC43, HKU1, MERS-CoV, SARS-HCoV and recently SARS-CoV-2 (Hui et al. 2020; Singhal 2020; Harapan et al. 2020). Both SARS-CoV and MERS-CoV are associated with a high level of pathogenicity and mortality (Báez-Santos et al. 2014; Kramer et al. 2006; Hemida and Alnaeem 2019; Hakomori and itiroh 1954).
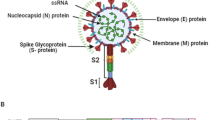
SARS-CoV-2, a positive and enveloped RNA virus, has a genome of 30,000 nucleotides. Its genome contains two overlapping open reading frames, ORF1a and ORF1b, which encode polyprotein precursors (NSPs; non-structural proteins & SPs; structural proteins). The SPs are (N) nucleocapsid protein, (E) envelope protein, (S) spike protein and (M) membrane protein while the NSPs are sixteen NSPs that regulate virus replication (Chen et al. 2020; Kim et al. 2020; Naqvi et al. 2020). The viral proteases are responsible for these proteolytic cleavages. SARS-CoV-2 has two protease proteins: NSP5 is (Mpro) main protease or 3 (3CLpro) chymotrypsin-like cysteine protease, which cleaves eleven polyprotein sites and NSP3 is (PLpro) papain-like protease which cleaves three polyproteins’ sites (Ullrich and Nitsche 2020; Morse et al. 2020; Vuong et al. 2020; Bakhshandeh et al. 2021; Raj 2021). The three main domains of SARS-CoV-2 Mpro are as follows: I (with 8–101 residues), II (with 102–184 residues) and III (with 201–303 residues). An antiparallel β-barrel structure is formed by domains I and II. Five α-helices is formed by domain III that are arranged in a cluster of antiparallel globular with (185–200 residues) a loop connecting domain II to III. SARS-CoV-2 Mpro active site contains a unique catalytic dyad Cys145 & His41 which cleaves polyproteins typically at Leu-Gln ↓ Ser, Ala or Gly sequence (Jin et al. 2020; Hsu et al. 2005; Pillaiyar et al. 2016). A key role is played by SARS-CoV-2 Mpro in the replication and transcription of Coronaviruses, making it an attractive drug target (Singh et al. 2022, 2021; Kashyap et al. 2022; Chauhan et al. 2022; Patel et al. 2022).
Many peptidomimetic inhibitors have been studied as protease inhibitors for various viruses, including SARS-CoV-2 (Vankadara et al. 2021; Paul et al. 2021; Somboon et al. 2021; Wang et al. 2022). Peptidomimetic inhibitors are capable of interacting with biological targets and producing the same therapeutic effects as native peptides. Using peptidomimetic inhibitors can eliminate some of the drawbacks of using native peptides, including proteolysis and low bioavailability (Vankadara et al. 2021; Paul et al. 2021; Somboon et al. 2021; Wang et al. 2022; Phoo et al. 2018; Colarusso et al. 2022; Ezat et al. 2015; Mostafa et al. 2014; Ibrahim et al. 2013; Saleh and Elshemey 2017; Calugi et al. 2014; Frecer et al. 2008).
This study introduces novel tri-peptidomimetic inhibitors against SARS-CoV-2 Mpro that have been significantly improved by adding this tripeptide to fulleropyrrolidine and attaching it with different ketone groups to form three groups of twelve suggested compounds. A computer-aided drug design approach is performed to investigate their inhibition activity through molecular descriptors calculation (ligand-based drug design) and molecular docking and MDS (structure-based drug design) (Marshall 1987; Yu and Mackerell 2017). According to the results, the suggested compounds are stable and produce good binding affinity with the active site pocket of SARS-CoV-2 Mpro especially, compounds 1, 9 11, 2, and 10.
Materials and methods
Building structures
The present author suggested the investigated compounds which are designed and built using SCIGRESS (version FJ 2.9.1) software simulations (Marchand et al. 2014). Many fullerene-based (C60) derivatives and nanotubes have been studied computationally as potential COVID-19 inhibitors (Sengupta and Hussain 2021; Rad et al. 2021; Gawad et al. 2019; Bagheri Novir and Aram 2020; Skariyachan et al. 2021; Zhang et al. 2021; Riley and Narayan 2021). In this study (Fig. 1), the main cap group for the studied inhibitors is water-soluble fulleropyrrolidine, which has demonstrated numerous antiviral and biomedical activities (Pochkaeva et al. 2020; Kazakova et al. 2019; Marchesan et al. 2005; Ibrahim et al. 2010a, b; Ibrahim et al. 2012; Saleh 2015). The natural substrate sequence for Mpro is Leu-Gln↓(Ser, Ala, Gly), which serves as the recognition sequence at most cleavage sites for Mpro, with P2-P1-P1ʹ notation. These tripeptide sequences are added to fulleropyrrolidine to form peptidomimetic inhibitors. A carbonyl group is added between the peptide sequences and fulleropyrrolidine to mimic peptide bond behaviour and decrease steric hindrance between the two nitrogen (N) atoms (one N atom from the pyrrolidine ring and one N atom from the peptide sequences). Both peptide sequence and carbonyl group are considered as a linker for the studied compounds. At the end of the suggested compounds, a head group is attached. This head group is the ketone group which can be α-ketoacid, α-ketoamide, β-ketoacid or β-ketoamide. The effectiveness of the ketone group as an anti-SARS-CoV-2 agent, as well as an anticancer agent, has been demonstrated in several studies (Wang et al. 2021; Zhang et al. 2020a, b).
As presented in Table 1, twelve compounds have been studied and are categorized into three groups: the first group consists of compounds where the peptide sequence is Leu-Gln-Gly and the head group is either α-ketoacid (compound 1: Gly-α acid), α-ketoamide (compound 2: Gly-α amide), β-ketoacid (compound 3: Gly-β acid) or β-ketoamide (compound 4: Gly-β amide). The second group includes compounds where the peptide sequence is Leu-Gln-Ala and the head group is either α-ketoacid (compound 5: Ala-α acid), α-ketoamide (compound 6: Ala-α amide), β-ketoacid (compound 7: Ala-β acid) or β-ketoamide (compound 8: Ala-β amide). The third group contains compounds where the peptide sequence is Leu-Gln-Ser and the head group is either α-ketoacid (compound 9: Ser-α acid), α-ketoamide (compound 10: Ser-α amide), β-ketoacid (compound 11: Ser-β acid) or β-ketoamide (compound 12: Ser-β amide).
Geometric optimization of structures
To begin, SCIGRESS software (Stewart 2008–2022) is used to optimize the geometrical structures of studied compounds at semi-empirical method (PM3) (Stewart 1989). The structures were then confirmed using FTIR calculation to ensure that they are not imaginary. To prepare the suggested compounds for molecular descriptors calculation, docking and molecular dynamics simulation, the geometrical structural compounds (Fig. 2) are re-optimized at the density functional theory (DFT) level using D-Gauss at B88-LYP GGA [Becke’s B88 (Becke 1988) Lee–Yang–Parr (Lee et al. 1988) functional generalized gradient approximation (Perdew et al. 1996)] functional with the DZVP (double-zeta valence polarized) (Chiodo et al. 2005; Schuchardt et al. 2007) basis sets. D-Gauss at B88-LYP method uses the exchange (B88) and correlation (LYP) functional which expresses the correlation energy density in terms of the electron density. The GAA improves the description of atom and molecule spin densities by accounting for the exchange–correlation energy. For the proposed molecules containing O, H, C and N atoms, the implemented method B88-LYP with DZVP basis set is suitable for these calculations (Pritchard et al. 2019).
The geometrical structure of optimized compounds at B88-LYP/ DZVP method. Compounds 1, 2, 3 and 4 form 1st group (Gly-compounds). Compounds 5, 6, 7 and 8 form 2nd group (Ala-compounds). Compounds 9, 10, 11 and 12 form 3rd group (Ser-compounds) The grey, white, blue and red balls represent the carbon, hydrogen, nitrogen and oxygen atoms, respectively
Molecular descriptors
For the optimized compounds using the B88-LYP method, several molecular descriptors are calculated use SCIGRESS software (Stewart 2008–2022) to assess their potential as viral inhibitors. These descriptors include the total optimization energy, ionization energy, electron affinity, lowest unoccupied molecular orbital (LUMO), highest occupied molecular orbital (HOMO), frontier band gap energy (ΔE), dipole moment, heat of formation and solvent accessible surface area (SASA). The heat of formation is calculated using the MO-G PM3 (Stewart 1989) method, while the solvent accessible surface area is calculated using the MO-G PM3/H2O method. All other molecular descriptors are calculated using the B88-LYP GGA functional with DZVP basis sets and the D-Gauss method.
Toxicity by Lipinski’s rule of five (RoF)
To prove the inhibitory activity of investigated compounds, the RoF is also calculated for the studied compounds. This rule assesses the ability of a compound to be used as drug, especially as an orally active compound (Lipinski 2004). The parameters used in this study to determine the Rule of Five are logP, molecular weight, (HBD) hydrogen-bond donors and (HBA) hydrogen-bond acceptors. These parameters are calculated using SCIGRESS software (Stewart 2008–2022).
Molecular docking
Molecular docking is performed for the investigated compounds against SARS-CoV-2 Mpro active site to provide an overview of the binding modes between the suggested compounds and active site amino acids residues. The SARS-CoV-2 Mpro is retrieved from www.rcsb.org with PDB code (7L0D) (Lockbaum et al. 2021) and prepared by SCIGRESS software (Stewart 2008–2022). Hydrogen atoms are added to SARS-CoV-2 Mpro and any other molecules or duplicated protein are removed. During molecular docking calculations, the active site residues (conserved catalytic dyad Cys145 and His41 in addition to conserved residues Tyr161, Met162 and His163 (Anand et al. 2003)) of SARS-CoV-2 Mpro are selected as a group to simulate and calculate the binding energy for the best docking score. In SCIGRESS software, The FASTDOCK genetic algorithm is performed using the scoring function PMF04 (Muegge 2000, 2006; Xue et al. 2010; Shen et al. 2011). The calculation of the binding energy is done using the following equation:
Molecular dynamics simulation
The two poses with the best interactions in molecular docking are with compounds 1 and 9, along with the complex with the natural substrate (Leu-Gln-Ser). These were selected to perform Molecular Dynamics Simulation (MDS) for 100 ns using NAMD. (Phillips et al. 2020). The CHARMM36m force field parameters are used for the simulation. The topology files of both SARS-CoV-2 Mpro and ligands is created using the CHARMM-GUI server for the simulation (Jo et al. 2008; Lee et al. 2016; Brooks et al. 2009). The SARS-CoV-2 Mpro used in this Molecular Dynamics (MD) simulation is retrieved from www.rcsb.org with PDB code (7K3T) with 1.20 Å resolution (Andi et al. 2022). The complexes are solvated in a cubic water box with an edge distance of 10 Å. Additionally, Monte-Carlo method is used to add KCl ions at a concentration of 0.15 M. The steepest descent is performed for energy minimization. The two complex structures are equilibrated at 300 K for 5000 ps with NVT boundary conditions. The molecular dynamics simulation is produced at NPT boundary conditions for 100 ns at 300 K with a fixed temperature and at 1 atm with a fixed pressure and a time step of integration (2 fs). Electrostatic interactions are specified by the particle-mesh Ewald (PME) algorithm with a C cutoff value of 1.2 nm (Darden et al. 1993). During the simulations, all the bond distances containing hydrogen atoms are fixed by the Lincs algorithm (Hess et al. 1997).
VMD program (Humphrey et al. 1996) is used to retrieve the resulting simulation data, which includes root mean square deviation (RMSD), hydrogen bonds, radius of gyration (Rg) and solvent accessible surface area (SASA).
Results and discussion
The novel suggested compounds are designed as competitive inhibitors to inhibit SARS-CoV-2 Mpro, which is one of the vital target proteins in SARS-CoV-2 virus. As shown in Fig. 1, the studied compounds have three main parts: (1) Fulleropyrrolidine-based system that provides the antiviral activity, (2) Natural substrate sequences Leu-Gln↓(Ser, Ala, Gly) as (P2-P1-P1ʹ) for SARS-CoV-2 Mpro to form peptidomimetic behaviour and decrease drug resistance effect and (3) A ketone group in different forms (α or β, acid or amide) to enhance the inhibition activity.
In this study, these essential parts are used to build 12 novel peptidomimetic inhibitors in three groups against main protease of SARS-CoV-2. Table 1 lists the chemical structures of these suggested compounds. The first group has Gly residue with α ketoacid, α ketoamide, β ketoacid or β ketoamide (compounds 1, 2, 3 or 4, respectively). The second group has Ala residue with α ketoacid, α ketoamide, β ketoacid or β ketoamide (compounds 5, 6, 7 or 8, respectively). Finally, the third group has Ser residue with α ketoacid, α ketoamide, β ketoacid or β ketoamide (compounds 9, 10, 11 or 12, respectively). The geometric structures for these compounds are optimized at DFT level using D-Gauss at B88-LYP GGA functional with the DZVP basis sets to investigate their molecular descriptors, Lipinski’s rule of five, molecular docking and molecular dynamic simulation. The geometric optimization of investigated compounds is represented in Fig. 2.
Molecular descriptors
Some molecular descriptors are calculated for studied compounds to investigate their physical and chemical properties, as well as to estimate the inhibition activity. Table 2 shows the calculated molecular descriptors at B88-LYP level, except for Heat of formation and SASA. The first parameter in Table 2 is the total energy for optimized geometry. It varies from − 2,377,836.01 to − 2,340,701.30 kcal/mol for the first group (Gly compounds) and from − 2,390,020.97 to − 2,365,360.67 kcal/mol for the second group (Ala compounds). For the third group (Ser compounds), it varies from − 2,449,693.47 to − 2,412,566.48 kcal/mol. The third group with serine residue has the lowest total energy, which indicates that this group has the most stable compounds in this study.
The ability of suggested compounds to lose an electron and to react with a surrounding medium is investigated by calculating ionization energy. The ionization energy for studied compounds is negative, indicating that it is easier to remove an electron and therefore more reactive. The ionization energy values for the three groups are close to each other and range from − 5.37 to − 5.27 eV. The variation in these values is due to the nature of the ketoacid and ketoamide groups in the compounds.
The exothermic energy released from neutral studied compounds when an electron is added is calculated as electron affinity. The compounds in the three groups with α-ketoacid head group have electron affinity values ranging from (4.37 eV for the first group, 4.21 eV for the second group and 4.58 eV for the third group). The other head groups result in lower electron affinity values for the studied compounds, as shown in Table 2.
HOMO and LUMO energies for the introduced compounds are listed in Table 2 to calculate the frontier energy gap (ΔE). Once again, the α-ketoacid group causes an increase in reactivity by decreasing ΔE value in each group. In the first group, compound 1 (Gly-α acid) has a ΔE value of 0.97 eV, in the second group, compound 5 (Ala-α acid) has a ΔE value of 1.16 eV and in the third group, compound 9 (Ser-α acid) has a ΔE value of 0.76 eV. Compounds 1, 5 and 9 with α-ketoacid group in each group have the lowest dipole moment value (10.66 debye, 8.99 debye and 10.85 debye, respectively). The rest of the compounds with α-ketoamide, β-ketoacid or β-ketoamide have high dipole moment values (Table 2).
During the formation of a mole of a compound, the amount of heat absorbed or raised is measured as the heat of formation. Heat of formation of the suggested compounds is calculated in Table 2. Both compounds with head groups, α-ketoacid and α-ketoamide, for each group have low heat of formation values. This explains their higher stability especially, for the α-ketoacid and α-ketoamide compounds in the third group (Ser compounds) which have the lowest heat of formation. Compounds 1 (Gly-α acid) and 2 (Gly-α amide) in the first group have 491.45 kcal/mol and 481.18 kcal/mol, respectively. Compounds 5 (Ala-α acid) and 6 (Ala-α amide) in the second group have 485.22 kcal/mol and 474.58 kcal/mol, respectively. Compounds 9 (Ser-α acid) and 10 (Ser-α amide) in the third group have 446.84 kcal/mol and 435.71 kcal/mol, respectively. In contrast, the compounds with β-ketoacid and β-ketoamide head groups have a high heat of formation. As a result, it is worth mentioning that the compounds with β-ketoacid and β-ketoamide in the third group also have a low heat of formation compared to their counterparts in other groups.
The surface area of the introduced compounds accessible to a solvent (SASA) is shown in Table 2. Due to the higher polarity of amide group compared to carboxylic acids, the compounds with ketoamide are more polar than ketoacid compounds. Therefore, the compounds with α or β-ketoamide have a larger solvent accessible surface area (SASA) than α or β-ketoacid compounds in each group and are more reactive in biological systems.
Lipinski’s rule of five (RoF) for toxicity
It is essential to investigate the susceptibility of studied compounds as oral drugs. The rule of five (RoF) for drug-like (oral bioavailability) compounds are calculated in Table 3. This rule recommends that the molecular weight should be equal to or less than 500, log P parameter should be equal to or less than 5, H-bond donors number (HBD) should be equal to or less than 5 and H-bond acceptors number (HBA) should be equal to or less than 10.
Based on the chemical composition of the suggested compounds, the compounds with Gly residue (first group) have a lower molecular weight than Ala compounds (second group) and Ser compounds (third group) as shown in Table 3. The molecular weight of studied compounds is more than 500, ranging from 1133.083 au for compound 3 (Gly-β acid) to 1178.12 au for compound 10 (Ser-α amide).
Log P values of investigated compounds range from 5.115 for compound 11 (Ser-β acid) to 7.635 for compound 6 (Ala-α amide), which is more than 5. So it does not obey the rule of five.
As listed in Table 3, the H-bond donors (HBD) number in the studied compounds is more than 5. The compounds in the first and second groups have identical numbers of HBD, while the third group has slightly higher numbers. The compounds with α-ketoacid and α-ketoamide have lower HBD number than that of β-ketoacid and β-ketoamide compounds. HBA number is 12 for the first and second groups, and 13 for the third group.
The calculated RoF of suggested compounds indicates that they are not likely to be suitable as oral drugs.
Binding interaction analysis
The molecular docking simulation aims to investigate the intermolecular interactions between the suggested compounds and SARS-CoV-2 Mpro. The study employs SARS-CoV-2 Mpro, which is identified by PDB code 7L0D, as the target protein. The active site of the protein is composed of catalytic dyad residues; Cys145 and His41, and conserved residues; Tyr161, Met162, and His163. These amino acids are selected as the active site residues for the docking analysis.
Table 4 presents the general properties of SARS-CoV-2 Mpro as determined by the MM3 classical mechanics method. The table shows the chemical formula, molecular weight, total energy and the total number of atoms, heavy atoms and bonds for the main protease protein. Table 5 provides a detailed analysis of binding energy for each docking system, including the amino acid residues involved and the number of hydrogen bonds formed. An asterisk next to an amino acid's name in Table 5 indicates that it is one of the selected active site residues.
According to the results in Table 5, it can be observed that compound 1 (Gly-α acid) in the first group has the lowest binding energy value (− 281.21 kcal/mol), indicating the most stable interaction with SARS-CoV-2 Mpro active site. Conversely, compound 7 (Ala-β acid) in the second group has the highest binding energy value (− 145.26 kcal/mol), suggesting the weakest interaction. The results also suggest that hydrogen bonding plays a significant role in stabilizing the interaction between SARS-CoV-2 Mpro and the studied compounds. The top five compounds for interaction with SARS-CoV-2 Mpro, as shown in Table 5, come from the first and third groups. These include compound 1 (Gly-α acid), compound 9 (Ser-α acid), compound 11 (Ser-β acid), compound 2 (Gly-α amide) and compound 10 (Ser-α amide) with the binding energies of − 281.21 kcal/mol, − 280.50 kcal/mol, − 218.66 kcal/mol, − 210.25 kcal/mol, − 206.14 kcal/mol, respectively. Figure 3 illustrates the mode of interaction simulation of molecular docking between these five compounds and the active site of SARS-CoV-2 Mpro.
The best docking poses interactions between spike protein of SARS-CoV-2 Mpro and compounds 1, 9, 11, 2 and 10. These compounds are compound 1 (Gly-α acid), compound 9 (Ser-α acid), compound 11 (Ser-β acid), compound 2 (Gly-α amide) and compound 10 (Ser-α amide). The studied compounds are shown as cylinder and protein’s amino acids are shown as ball and cylinder. The blue dashed lines are hydrogen bonds between SARS-CoV-2 Mpro and studied compounds in docking complex
As seen in Table 5 and Fig. 3, compound 1 (Gly-α acid) produces only 5 H-bonds with three amino acids. One H-bond is between Gly146 and amino group of P1, one H-bond is between Ser147 and OH group of ketoacid. Gly149 forms two H-bonds with NH2 residue of P1 and one H-bond with C=O of ketoacid.
Compound 9 (Ser-α acid) forms 7 hydrogen bonds with four active site residues: Val114, Tyr126, Cys128, and His163. His163 is an active site residue that forms one H-bond with the amino group of P2. Cys128 and Val114 Each form one H-bond with C=O of ketoacid. Tyr126 forms 4 H-bonds, one with OH group of ketoacid, one with C = O of P1 and finally two H-bonds with the amino group of P1.
Ten H-bonds are created in docking system with compound 11 (Ser-β acid). The active site residues His41, Cys145, and His163 are involved in the hydrogen bonding interactions with compound 11 (Ser-β acid), along with other residues. His41 forms two H-bonds with amino group of P2 for ligand compound while His163 forms three H-bonds: one with C=O residue of P1, one with NH2 residue of P1 and one with OH residue of P1ʹ. Cys145 has two H-bonds one with amino group of P1 and another with amino group of P1ʹ. The ketoacid group of compound 11 forms one H-bond with Ala173. One H-bond is formed between Ser144 residue and amino group of P1ʹ. Finally, one H-bond is formed between Gly146 and OH residue of P1ʹ.
In the first group, compound 2 (Gly-α amide) forms 9 H-bonds with active site of main protease for SARS-CoV-2. Four of these H-bonds form with two active site conserved residues (Cys145 and His163). Cys145 forms one H-bond with C=O of ketoacid. His163 generates two H-bonds with NH2 group of ketoacid and one H-bond with amino group of P1ʹ. Each one of Asn142 and Leu141 forms H-bond with NH2 residue of P1. Ser144 produces two H-bonds: one of them with C=O residue of P1 and the other with C=O of ketoamide. C=O of peptide bond of P1 forms one H-bond with His172.
For compound 10 (Ser-α amide), amino group P1ʹ forms three H-bonds with His163, His164 and Met165 while OH residue of P1ʹ forms two H-bonds with Ala173. amino group P2 for compound 10 forms two H-bonds with His164 and His41.
The remaining suggested compounds, as listed in Table 5, have binding energies greater than − 206.14 kcal/mol. Compounds 3 and 4 in the first group form three and thirteen H-bonds, respectively. Compounds 5, 6, 7 and 8 in the second group form two, six, seven and ten H-bonds, respectively. Compound 12 in the third group produces ten H-bonds. Table 5 lists the amino acids involved in the hydrogen bonding interactions for each compound.
Some earlier studied in-silico analyses on SARS-CoV-2 Mpro are compared with the suggested compounds to classify their potency as Mpro inhibitors. The intermolecular interaction binding energy of the suggested compounds is compared with that of other studies. Some compounds in the flavonoid family have been investigated as Mpro inhibitors. Silibinin, Tomentin A, Tomentin B, 4′ -O-methyldiplacone, Hesperidin Amentoflavone and Bilobetin compounds have effective binding energies with SARS-CoV-2 Mpro (ranging from − 9.24 kcal/mol to − 12.43 kcal/mol) (Hadni et al. 2022). In another study, 7809 natural compounds from different databases were collected to study their inhibition activities against SARS-CoV-2 Mpro. Based on docking and dynamic simulation, the top five natural compounds (theaflavin, ginkgetin, hesperidin, withanolide D and psoralidin) were found to generate bingeing energy with Mpro ranging from 10.04 to 7.55 kcal/mol (Patel et al. 2022). As a result of the inclusion complex of the antiviral drug acyclovir and methyl-b-cyclodextrin (Mb-CD:AVR), SARS-CoV-2 Mpro also showed higher stability by generating a binding energy of − 21.67 kcal/mol (Mohandoss et al. 2022). The investigated compounds in this study confirm that they produce higher SARS-CoV-2 Mpro stability and lower binding energies compare to previous in-silico studies.
Post-dynamics trajectories analysis
The dynamic aspects of the docking complex involving the two top compounds 1 and 9 are simulated for 100 ns. MDSs are applied to investigate the stability of the protein and ligand during their interaction. This is done by calculating and analysing various parameters such as RMSD, H-bonds, Rg, and SASA.
Figure 4a illustrates the RMSD of Cα atoms of the backbone of the SARS-CoV-2 Mpro protein complexes with compounds 1 and 9 during 100 ns of simulation. The RMSD for Mpro with compound 1 fluctuates between 0.11 and 0.39 nm, while with compound 9, it fluctuates between 0.14 and 0.34 nm. The average values of RMSD for Mpro with compounds 1 and 9 are 0.23 nm and 0.25 nm, respectively. The RMSD of Mpro with compounds 1 and 9 increases from the start of the simulation to around 65 ns, after which it remains relatively stable with very low variation until the end of the simulation. The maximum value as well as the average value is still very low, indicates the stability of the Mpro protein in the complex interaction.
The root mean square deviation (RMSD) graph of a SARS-CoV-2 Mpro in complexes with compound 1 (Gly-α acid) and compound 9 (Ser-α acid), b compound 1 (Gly-α acid) and compound 9 (Ser-α acid) in complexes with SARS-CoV-2 Mpro. The blue curve represents the SARS-CoV-2 Mpro complex with compound 1 (Gly-α acid). The red curve represents the SARS-CoV-2 Mpro complex with compound 9 (Ser-α acid)
As shown in Fig. 4b, the RMSD is also calculated for compounds 1 and 9 in their complexes with Mpro. For compound 1, the RMSD fluctuates between 0.07 and 0.69 nm (average value = 0.32 nm). For compound 9, the RMSD varies between 0.31 and 0.83 nm (average value = 0.5 nm). These values indicate the stability of the docking interaction poses. The change in RMSD values for compounds 1 and 9 is large from the start of the simulation until around 40 ns, then it becomes small until the end of the simulation.
Hydrogen bonds (H-bonds) are considered one of the most crucial bonds in biological systems. The total number of H-bonds in the two interaction systems involving compounds 1 and 9 are analysed in molecular dynamics, as represented in Fig. 5a. Both compounds 1 and 9 generated 4 H-bonds with the active site pocket of SARS-CoV-2 Mpro. The probability of forming four H-bonds with Mpro is higher for compound 9 than compound 1. This is because the production of four H-bonds with compound 9 is generated eight times during the simulation, while for compound 1 only generated four H-bonds once in the simulation. A snapshot of the complexes, represented in Fig. 5b, shows the trajectory of compounds 1 and 9 in the active site pocket of Mpro.
a The number of hydrogen bonds in complexes between SARS-CoV-2 Mpro and compound 1 (Gly-α acid) and compound 9 (Ser-α acid). The blue curve represents the SARS-CoV-2 Mpro complex with compound 1. The red curve represents the SARS-CoV-2 Mpro complex with compound 9. b The snapshots of the SARS-CoV-2 Mpro complex with compound 1 (Gly-α acid) and compound 9 (Ser-α acid). The compound is shown in a stick (blue) representation
The radius of gyration (Rg) provides an indication of the compactness of the protein–ligand interaction and the degree of protein folding in the interaction complex. Figure 6a shows that the average values of Rg for protein–ligand complexes with compounds 1 and 9 are almost the same, 2.25 nm and 2.22 nm, respectively. During the first part of the simulation (0–50 ns), there is no significant variation between Rg values for complexes with compounds 1 and 9. However, during the second part of the simulation (50–100 ns), the complex with compound 9 has lower Rg values than the complex with compound 1. This means that the interaction between Mpro and compound 9 is slightly more compact and the complex is slightly more stable than that with compound 1. Additionally, Mpro is more folded in the complex with compound 9 than in the complex with compound 1.
a Radius of gyration (Rg) of complexes with compounds 1 (Gly-α acid) and compound 9 (Ser-α acid). b solvent‐accessible surface area (SASA) analysis for complexes with compounds 1(Gly-α acid) and compound 9 (Ser-α acid). The blue curve represents the SARS-CoV-2 Mpro complex with compound 1 (Gly-α acid). The red curve represents the SARS-CoV-2 Mpro complex with compound 9 (Ser-α acid)
SASA for the two complexes (with compounds 1 and 9) is presented in Fig. 6b. The average value of SASA for the complex containing compound 1 is 156.11 nm2 and for the complex containing compound 9 is 155.25 nm2. Throughout the majority of the simulation, the SASA for the interaction with compound 9 is less than that with compound 1. This indicates that the interaction between Mpro and compound 9 is slightly more stable than the interaction with compound 1.
As compound 9 (Ser-α acid) has shown to cause more stability in complex with SARS-CoV-2 Mpro, the complex with the natural substrate (Leu-Gln-Ser) is compared using MDS to investigate the effect of modified tri-peptidomimetic inhibitors on the stability of complex with Mpro. Figure 7 depicts the MDS analysis of natural substrate interaction. Figure 7a illustrates the RMSD of both Cα atoms of the backbone of Mpro protein and natural substrate in complex interaction during 100 ns of simulation. The RMSD for Mpro fluctuates between 0.11 and 2.55 nm with an average value of 1.54 nm, while for natural substrate, it fluctuates between 0.14 and 9.98 nm with an average value of 4.07 nm. These large variations show that neither Mpro protein nor natural substrate are stable during interaction. Although the Mpro protein become semi-stable from 60 to 100 ns, the natural substrate remains unstable. The number of H-bonds in complex with natural substrate is represented in Fig. 7b. There are five H-bonds that are generated four times during the simulation, and there are many times interval during the simulation that don’t have any H-bonds. This is further evidence of the instability of the interaction.
The MDS results of natural substrate (Leu-Gln-Ser) interaction. a The Root Mean Square Deviation (RMSD) of SARS-CoV-2 Mpro and natural substrate in complex. b The number of hydrogen bonds in complex. c Radius of gyration Rg (blue curve) and solvent‐accessible surface area SASA (red curve) analysis of complex. d The snapshots of the SARS-CoV-2 Mpro complex with natural substrate (Leu-Gln-Ser)
The radius of gyration (Rg) and SASA results for the complex between Mpro and natural substrate are shown in Fig. 7c. The blue curve represents the Rg of complex with average 2.89 nm. Compared with compound 9, there are large fluctuations in Rg values from 2.21 to 3.6 nm. The red curve represents SASA of complex with natural substrate, which also shows large fluctuations in SASA values from 151.11 nm2 to 360.15 nm. The complex with natural substrate has a high value of Rg and SASA compared to the complexes with compounds 1 and 9.
Conclusion
In conclusion, this study presents 12 novel peptidomimetic inhibitors against the main protease of SARS-CoV-2 virus. The compounds are designed to inhibit the Mpro by mimicking the natural substrate sequence and incorporating a fulleropyrrolidine-based system for antiviral activity. The compounds are optimized at the DFT level and evaluated for their physical and chemical properties, as well as their potential as oral drugs using the rule of five. The results show that group three Ser-containing compounds among the studied compounds are the most stable and interact with the surrounding systems, especially the compounds containing α-ketoacid in each group. The suggested compounds have lower binding energies and higher stability compared to previously studied compounds, but their potential as oral drugs is limited by their high molecular weight and logP values. Molecular docking and dynamics simulations have been performed to investigate the binding modes and stability of the compounds with the protease. The results of the docking systems show high stability and binding affinity of the complexes with the studied compounds, particularly compounds 1, 9, 11, 2, and 10. The MDS results for the complexes with compounds 1 and 9 have been analysed, and they indicate the stability of the complexes, particularly with compound 9. By comparing these results with the MDS for natural substrate complex, it can be concluded that the modification of the suggested compounds by fulleropyrrolidine, α or β-ketoacid, and α or β-ketoamide has resulted in increased stability and improved binding with the active site of SARS-CoV-2 Mpro. Overall, this study provides promising candidates for the development of new drugs for the treatment of COVID-19. Future studies will further investigate the ability of the studied compounds to inhibit SARS-CoV-2 Mpro variants using computer-aided drug design approaches. This will help to identify the most promising compounds for further development as potential inhibitors of SARS-CoV-2 Mpro.
Data availability
On request, the corresponding author can provide optimization structures, molecular descriptors, molecular docking and molecular dynamic simulation.
References
Anand K, Ziebuhr J, Wadhwani P et al (2003) Coronavirus main proteinase (3CLpro) structure: basis for design of anti-SARS drugs. Science (80-) 300:1763–1767. https://doi.org/10.1126/science.1085658
Andi B, Kumaran D, Kreitler DF et al (2022) Hepatitis C virus NS3/4A inhibitors and other drug-like compounds as covalent binders of SARS-CoV-2 main protease. Sci Rep 12:12197. https://doi.org/10.1038/s41598-022-15930-z
Báez-Santos YM, Mielech AM, Deng X et al (2014) Catalytic function and substrate specificity of the papain-like protease domain of nsp3 from the Middle East respiratory syndrome coronavirus. J Virol 88:12511–12527. https://doi.org/10.1128/jvi.01294-14
Bagheri Novir S, Aram MR (2020) Quantum mechanical simulation of Chloroquine drug interaction with C60 fullerene for treatment of COVID-19. Chem Phys Lett 757:137869. https://doi.org/10.1016/j.cplett.2020.137869
Bakhshandeh B, Jahanafrooz Z, Abbasi A et al (2021) Mutations in SARS-CoV-2; consequences in structure, function, and pathogenicity of the virus. Microb Pathog 154:104831. https://doi.org/10.1016/j.micpath.2021.104831
Becke AD (1988) Density-functional exchange-energy approximation with correct asymptotic behavior. Phys Rev A 38:3098–3100. https://doi.org/10.1103/PhysRevA.38.3098
Brooks BR, Brooks CL, Mackerell AD et al (2009) CHARMM: the biomolecular simulation program. J Comput Chem 30:1545–1614. https://doi.org/10.1002/jcc.21287
Calugi C, Guarna A, Trabocchi A (2014) Identification of constrained peptidomimetic chemotypes as HIV protease inhibitors. Eur J Med Chem 84:444–453. https://doi.org/10.1016/j.ejmech.2014.07.049
Chauhan M, Bhardwaj VK, Kumar A et al (2022) Theaflavin 3-gallate inhibits the main protease (Mpro) of SARS-CoV-2 and reduces its count in vitro. Sci Rep 12:13146. https://doi.org/10.1038/s41598-022-17558-5
Chen Y, Liu Q, Guo D (2020) Emerging coronaviruses: genome structure, replication, and pathogenesis. J Med Virol 92:418–423
Chiodo S, Russo N, Sicilia E (2005) Newly developed basis sets for density functional calculations. J Comput Chem 26:175–184. https://doi.org/10.1002/jcc.20144
Colarusso S, Ferrigno F, Ponzi S et al (2022) SAR evolution towards potent C-terminal carboxamide peptide inhibitors of Zika virus NS2B-NS3 protease. Bioorgan Med Chem 57:116631. https://doi.org/10.1016/j.bmc.2022.116631
Darden T, York D, Pedersen L (1993) Particle mesh Ewald: an N-log (N) method for Ewald sums in large systems. J Chem Phys 98:10089–10092
Ezat AA, Mostafa HI, El-Bialy NS et al (2015) Computational approaches to study peptidomimetic and macrocyclic hepatitis C virus NS3 protease inhibitors. J Comput Theor Nanosci 12:52–59. https://doi.org/10.1166/jctn.2015.3697
Frecer V, Berti F, Benedetti F, Miertus S (2008) Design of peptidomimetic inhibitors of aspartic protease of HIV-1 containing—PheΨPro—core and displaying favourable ADME-related properties. J Mol Graph Model 27:376–387. https://doi.org/10.1016/j.jmgm.2008.06.006
Gawad AE, Bayoumy AM, Ibrahim MA (2019) Molecular interactions of fullerene-based derivatives and SARS-CoV-2. Res Sq 1–11
Gorbalenya A, Baker S, Baric R et al (2020a) Severe acute respiratory syndrome-related coronavirus—The species and its viruses, a state-ment of the Coronavirus Study Group. bioRxiv (preprint). https://doi.org/10.1101/2020.02.07.937862. Bio RXIV 1–15
Gorbalenya AE et al (2020b) Coronaviridae Study Group of the International Committee on Taxonomy of Viruses. The species severe acute respiratory syndrome related coronavirus. Nat Microbiol 5:536–544
Hadni H, Fitri A, Benjelloun AT et al (2022) Evaluation of flavonoids as potential inhibitors of the SARS-CoV-2 main protease and spike RBD: molecular docking, ADMET evaluation and molecular dynamics simulations. J Indian Chem Soc 99:100697. https://doi.org/10.1016/j.jics.2022.100697
Hakomori S (1954) Chemistry and biology of lipids. Tohoku J Exp Med 60:331–345. https://doi.org/10.1620/tjem.60.331
Harapan H, Itoh N, Yufika A et al (2020) Coronavirus disease 2019 (COVID-19): a literature review. J Infect Public Health 13:667–673. https://doi.org/10.1016/j.jiph.2020.03.019
Hemida MG, Alnaeem A (2019) Some one health based control strategies for the Middle East respiratory syndrome coronavirus. One Health 8:100102. https://doi.org/10.1016/j.onehlt.2019.100102
Herman GA, O’Brien MP, Forleo-Neto E et al (2022) Efficacy and safety of a single dose of casirivimab and imdevimab for the prevention of COVID-19 over an 8-month period: a randomised, double-blind, placebo-controlled trial. Lancet Infect Dis 22:1444–1454. https://doi.org/10.1016/S1473-3099(22)00416-9
Hess B, Bekker H, Berendsen HJC, Fraaije JGEM (1997) LINCS: A Linear Constraint Solver for molecular simulations. J Comput Chem 18:1463–1472. https://doi.org/10.1002/(SICI)1096-987X(199709)18:12%3c1463::AID-JCC4%3e3.0.CO;2-H
Hsu MF, Kuo CJ, Chang KT et al (2005) Mechanism of the maturation process of SARS-CoV 3CL protease. J Biol Chem 280:31257–31266. https://doi.org/10.1074/jbc.M502577200
Hui DS, Azhar I, E, Madani TA, et al (2020) The continuing 2019-nCoV epidemic threat of novel coronaviruses to global health—the latest 2019 novel coronavirus outbreak in Wuhan, China. Int J Infect Dis 91:264–266. https://doi.org/10.1016/j.ijid.2020.01.009
Humphrey W, Dalke A, Schulten K (1996) VMD: visual molecular dynamics. J Mol Graph 14:33–38
Ibrahim M, Saleh NA, Hameed AJ et al (2010a) Structural and electronic properties of new fullerene derivatives and their possible application as HIV-1 protease inhibitors. Spectrochim Acta Part A Mol Biomol Spectrosc 75:702–709. https://doi.org/10.1016/j.saa.2009.11.042
Ibrahim M, Saleh NA, Elshemey WM, Elsayed AA (2010b) Computational notes on fullerene based system as HIV-1 protease inhibitors. J Comput Theor Nanosci 7:224–227. https://doi.org/10.1166/jctn.2010.1350
Ibrahim M, Saleh NA, Elshemey WM, Elsayed AA (2012) Fullerene derivative as Anti-HIV protease inhibitor: molecular modeling and QSAR approaches. Mini Rev Med Chem 12:447–451. https://doi.org/10.2174/138955712800493762
Ibrahim M, Saleh NA, Elshemey WM, Elsayed AA (2013) QSAR properties of novel peptidomimetic NS3 protease inhibitors. J Comput Theor Nanosci 10:785–788. https://doi.org/10.1166/jctn.2013.2771
Jin Z, Du X, Xu Y et al (2020) Structure of Mpro from SARS-CoV-2 and discovery of its inhibitors. Nature 582:289–293. https://doi.org/10.1038/s41586-020-2223-y
Jo S, Kim T, Iyer VG, Im W (2008) CHARMM-GUI: a web-based graphical user interface for CHARMM. J Comput Chem 29:1859–1865. https://doi.org/10.1002/jcc.20945
Kashyap P, Bhardwaj VK, Chauhan M et al (2022) A ricin-based peptide BRIP from Hordeum vulgare inhibits Mpro of SARS-CoV-2. Sci Rep 12:1–11. https://doi.org/10.1038/s41598-022-15977-y
Kazakova AV, Konev AS, Zorin IM et al (2019) PEG-modified aziridines for stereoselective synthesis of water-soluble fulleropyrrolidines. Org Biomol Chem 17:9864–9873. https://doi.org/10.1039/c9ob01949a
Kim D, Lee J-Y, Yang J-S et al (2020) The architecture of SARS-CoV-2 transcriptome. Cell 181:914–921
Kramer A, Schwebke I, Kampf G (2006) How long do nosocomial pathogens persist on inanimate surfaces? A systematic review. BMC Infect Dis 6:130. https://doi.org/10.1186/1471-2334-6-130
Lee C, Yang W, Parr RG (1988) Development of the Colle–Salvetti correlation-energy formula into a functional of the electron density. Phys Rev B 37:785–789. https://doi.org/10.1103/PhysRevB.37.785
Lee J, Cheng X, Swails JM et al (2016) CHARMM-GUI input generator for NAMD, GROMACS, AMBER, OpenMM, and CHARMM/OpenMM simulations using the CHARMM36 additive force field. J Chem Theory Comput 12:405–413. https://doi.org/10.1021/acs.jctc.5b00935
Lipinski CA (2004) Lead-and drug-like compounds: the rule-of-five revolution. Drug Discov Today Technol 1:337–341
Lockbaum GJ, Reyes AC, Lee JM et al (2021) Crystal structure of sars-cov-2 main protease in complex with the non-covalent inhibitor ml188. Viruses 13:174. https://doi.org/10.3390/v13020174
Lu H, Stratton CW, Tang YW (2020) Outbreak of pneumonia of unknown etiology in Wuhan, China: the mystery and the miracle. J Med Virol 92:401–402. https://doi.org/10.1002/jmv.25678
Marchesan S, Da Ros T, Spalluto G et al (2005) Anti-HIV properties of cationic fullerene derivatives. Bioorgan Med Chem Lett 15:3615–3618. https://doi.org/10.1016/j.bmcl.2005.05.069
Marshall GR (1987) Computer-aided drug design. Annu Rev Pharmacol Toxicol 27:193–213
Mohandoss S, Sukanya R, Ganesan S et al (2022) SARS-CoV-2 main protease (3CLpro) interaction with acyclovir antiviral drug/methyl-β-cyclodextrin complex: Physiochemical characterization and molecular docking. J Mol Liq 366:120292. https://doi.org/10.1016/j.molliq.2022.120292
Morse JS, Lalonde T, Xu S, Liu WR (2020) Learning from the past: possible urgent prevention and treatment options for severe acute respiratory infections caused by 2019-nCoV. ChemBioChem 21:730–738. https://doi.org/10.1002/cbic.202000047
Mostafa H, El-bialy N, Ezat A et al (2014) QSAR analysis and molecular docking simulation of suggested peptidomimetic NS3 protease inhibitors. Curr Comput Aided Drug Des 10:28–40. https://doi.org/10.2174/15734099113096660048
Muegge I (2000) A knowledge-based scoring function for protein–ligand interactions: probing the reference state. Perspect Drug Discov Des 20:99–114. https://doi.org/10.1023/A:1008729005958
Muegge I (2006) PMF scoring revisited. J Med Chem 49:5895–5902. https://doi.org/10.1021/jm050038s
Naqvi AAT, Fatima K, Mohammad T et al (2020) Insights into SARS-CoV-2 genome, structure, evolution, pathogenesis and therapies: structural genomics approach. Biochim Biophys Acta Mol Basis Dis 1866:165878. https://doi.org/10.1016/j.bbadis.2020.165878
Patel CN, Jani SP, Prasanth Kumar S et al (2022) Computational investigation of natural compounds as potential main protease (Mpro) inhibitors for SARS-CoV-2 virus. Comput Biol Med 151:106318. https://doi.org/10.1016/j.compbiomed.2022.106318
Paul A, Sarkar A, Saha S et al (2021) Synthetic and computational efforts towards the development of peptidomimetics and small-molecule SARS-CoV 3CLpro inhibitors. Bioorgan Med Chem 46:116301. https://doi.org/10.1016/j.bmc.2021.116301
Perdew JP, Burke K, Ernzerhof M (1996) Generalized gradient approximation made simple. Phys Rev Lett 77:3865–3868. https://doi.org/10.1103/PhysRevLett.77.3865
Phillips JC, Hardy DJ, Maia JDC et al (2020) Scalable molecular dynamics on CPU and GPU architectures with NAMD. J Chem Phys 153:44130. https://doi.org/10.1063/5.0014475
Phoo WW, Zhang Z, Wirawan M et al (2018) Structures of Zika virus NS2B-NS3 protease in complex with peptidomimetic inhibitors. Antiviral Res 160:17–24. https://doi.org/10.1016/j.antiviral.2018.10.006
Pillaiyar T, Manickam M, Namasivayam V et al (2016) An overview of severe acute respiratory syndrome-coronavirus (SARS-CoV) 3CL protease inhibitors: peptidomimetics and small molecule chemotherapy. J Med Chem 59:6595–6628. https://doi.org/10.1021/acs.jmedchem.5b01461
Pochkaeva EI, Podolsky NE, Zakusilo DN et al (2020) Fullerene derivatives with amino acids, peptides and proteins: from synthesis to biomedical application. Prog Solid State Chem 57:100255. https://doi.org/10.1016/j.progsolidstchem.2019.100255
Pritchard BP, Altarawy D, Didier B et al (2019) New basis set exchange: an open, up-to-date resource for the molecular sciences community. J Chem Inf Model 59:4814–4820. https://doi.org/10.1021/acs.jcim.9b00725
Rad AS, Ardjmand M, Esfahani MR, Khodashenas B (2021) DFT calculations towards the geometry optimization, electronic structure, infrared spectroscopy and UV–Vis analyses of Favipiravir adsorption on the first-row transition metals doped fullerenes; a new strategy for COVID-19 therapy. Spectrochim Acta Part A Mol Biomol Spectrosc 247:119082. https://doi.org/10.1016/j.saa.2020.119082
Raj R (2021) Analysis of non-structural proteins, NSPs of SARS-CoV-2 as targets for computational drug designing. Biochem Biophys Reports 25:100847. https://doi.org/10.1016/j.bbrep.2020.100847
Riley PR, Narayan RJ (2021) Recent advances in carbon nanomaterials for biomedical applications: a review. Curr Opin Biomed Eng 17:100262. https://doi.org/10.1016/j.cobme.2021.100262
Saleh NA (2015) The QSAR and docking calculations of fullerene derivatives as HIV-1 protease inhibitors. Spectrochim Acta Part A Mol Biomol Spectrosc 136:1523–1529. https://doi.org/10.1016/j.saa.2014.10.045
Saleh NA, Elshemey WM (2017) Structure-based drug design of novel peptidomimetic cellulose derivatives as HCV-NS3 protease inhibitors. Life Sci 187:58–63. https://doi.org/10.1016/j.lfs.2017.08.021
Schuchardt KL, Didier BT, Elsethagen T et al (2007) Basis set exchange: a community database for computational sciences. J Chem Inf Model 47:1045–1052. https://doi.org/10.1021/ci600510j
Sengupta J, Hussain CM (2021) Carbon nanomaterials to combat virus: a perspective in view of COVID-19. Carbon Trends 2:100019. https://doi.org/10.1016/j.cartre.2020.100019
Shen Q, Xiong B, Zheng M et al (2011) Knowledge-based scoring functions in drug design: 2. can the knowledge base be enriched? J Chem Inf Model 51:386–397. https://doi.org/10.1021/ci100343j
Shrinivasan R (2021) COVID-19 data explorer—our world in data. In: Scroll.in. https://ourworldindata.org/. Accessed 22 Jan 2023
Singh R, Bhardwaj VK, Sharma J et al (2021) Identification of potential plant bioactive as SARS-CoV-2 Spike protein and human ACE2 fusion inhibitors. Comput Biol Med 136:104631. https://doi.org/10.1016/j.compbiomed.2021.104631
Singh R, Bhardwaj VK, Das P et al (2022) Benchmarking the ability of novel compounds to inhibit SARS-CoV-2 main protease using steered molecular dynamics simulations. Comput Biol Med 146:105572. https://doi.org/10.1016/j.compbiomed.2022.105572
Singhal T (2020) A review of coronavirus disease-2019 (COVID-19). Indian J Pediatr 87:281–286. https://doi.org/10.1007/s12098-020-03263-6
Skariyachan S, Gopal D, Deshpande D et al (2021) Carbon fullerene and nanotube are probable binders to multiple targets of SARS-CoV-2: insights from computational modeling and molecular dynamic simulation studies. Infect Genet Evol 96:105155. https://doi.org/10.1016/j.meegid.2021.105155
Somboon T, Mahalapbutr P, Sanachai K et al (2021) Computational study on peptidomimetic inhibitors against SARS-CoV-2 main protease. J Mol Liq 322:114999. https://doi.org/10.1016/j.molliq.2020.114999
Stewart JJP (1989) Optimization of parameters for semiempirical methods I. Method. J Comput Chem 10:209–220. https://doi.org/10.1002/jcc.540100208
Marchand N, Lienard P, Siehl HU et al (2014) Applications of molecular simulation software SCIGRESS in industry and university. Fujitsu Sci Tech J 50:46–51
Ullrich S, Nitsche C (2020) The SARS-CoV-2 main protease as drug target. Bioorgan Med Chem Lett 30:127377. https://doi.org/10.1016/j.bmcl.2020.127377
Vankadara S, Wong YX, Liu B et al (2021) A head-to-head comparison of the inhibitory activities of 15 peptidomimetic SARS-CoV-2 3CLpro inhibitors. Bioorgan Med Chem Lett 48:128263. https://doi.org/10.1016/j.bmcl.2021.128263
Vuong W, Khan MB, Fischer C et al (2020) Feline coronavirus drug inhibits the main protease of SARS-CoV-2 and blocks virus replication. Nat Commun 11:1–8. https://doi.org/10.1038/s41467-020-18096-2
Wang C, Horby PW, Hayden FG, Gao GF (2020) A novel coronavirus outbreak of global health concern. Lancet 395:470–473. https://doi.org/10.1016/S0140-6736(20)30185-9
Wang J, Liang B, Chen Y et al (2021) A new class of α-ketoamide derivatives with potent anticancer and anti-SARS-CoV-2 activities. Eur J Med Chem 215:113267. https://doi.org/10.1016/j.ejmech.2021.113267
Wang H, Pei R, Li X et al (2022) The structure-based design of peptidomimetic inhibitors against SARS-CoV-2 3C like protease as Potent anti-viral drug candidate. Eur J Med Chem 238:114458. https://doi.org/10.1016/j.ejmech.2022.114458
World Health Organization (2021) WHO Coronavirus Disease (COVID-19) Dashboard With Vaccination Data|WHO Coronavirus (COVID-19) Dashboard With Vaccination Data. In: World Heal. Organ. https://covid19.who.int/. Accessed 20 Jan 2023
Xue M, Zheng M, Xiong B et al (2010) Knowledge-based scoring functions in drug design. 1. Developing a target-specific method for kinase–ligand interactions. J Chem Inf Model 50:1378–1386. https://doi.org/10.1021/ci100182c
Yan Z, Yang M, Lai C-L (2021) COVID-19 vaccinations: a comprehensive review of their safety and efficacy in special populations. Vaccines 9:1097
Yu W, Mackerell AD (2017) Computer-aided drug design methods. In: Methods in molecular biology. Springer, Humana Press, New York, NY, pp 85–106
Zhang L, Lin D, Sun X et al (2020a) Crystal structure of SARS-CoV-2 main protease provides a basis for design of improved a-ketoamide inhibitors. Science (80-) 368:409–412. https://doi.org/10.1126/science.abb3405
Zhang L, Lin D, Kusov Y et al (2020b) α-ketoamides as broad-spectrum inhibitors of coronavirus and enterovirus replication: structure-based design, synthesis, and activity assessment. J Med Chem 63:4562–4578. https://doi.org/10.1021/acs.jmedchem.9b01828
Zhang F, Wang Z, Vijver MG, Peijnenburg WJGM (2021) Probing nano-QSAR to assess the interactions between carbon nanoparticles and a SARS-CoV-2 RNA fragment. Ecotoxicol Environ Saf 219:112357. https://doi.org/10.1016/j.ecoenv.2021.112357
Acknowledgements
A grateful acknowledgment is expressed to the Department of Biophysics of the Faculty of Science at Cairo University for providing access to SCIGRESS software facilities.
Funding
The author declares that this research did not receive any specific grant from funding agencies in the public, commercial, or not-for-profit sectors.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The author declares that there is no competing interest in this work.
Research involving human participants and/or animals
Not applicable.
Informed consent
The author gives her consent to publish details of the manuscript in 3 Biotech journal.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Saleh, N.A. In-silico study: docking simulation and molecular dynamics of peptidomimetic fullerene-based derivatives against SARS-CoV-2 Mpro. 3 Biotech 13, 185 (2023). https://doi.org/10.1007/s13205-023-03608-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s13205-023-03608-w