Abstract
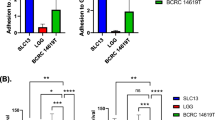
Helicobacter pylori (H. pylori) is one of most commonly found pathogen in the stomach. In spite of emergence of different treatment strategies, H. pylori infection remains difficult to treat. The bioengineered probiotic lactobacilli that could displace H. pylori and simultaneously present immunogenic peptides such as heparan sulphate binding protein (Hsbp) to elicit immune response could emerge as a potential therapeutic agent. The aim of this study was to discover the anti-H. pylori activities and faster exclusion of H. pylori from host cells by the recombinant strain of Lactobacillus expressing the immunogenic Hsbp protein. The results were promising and showed a 65% reduction in H. pylori adhesion after two hours of pre-incubation with recombinant-LGG and HeLa S3 cells, followed by the adhesion of H. pylori pathogen (P < 0.002). Additionally, 36% and 39% reduction were examined in co-incubation and post-incubation with recombinant-LGG, respectively. When challenged with H. pylori, the proinflammatory cytokine expression was also down regulated in recombinant-LGG treated HeLa S3 cells. This promising result provides a new insight of bioengineered probiotic lactobacilli which could displace H. pylori and simultaneously has immunogenic properties thereby may be useful to prevent H. pylori infection.



Similar content being viewed by others
Data availability
All data generated or analysed during this study are included in this published article (and its supplementary information files).
References
Anbazhagan K, Sasikumar P, Gomathi S, Priya HP, Selvam GS (2013) In vitro degradation of oxalate by recombinant Lactobacillus plantarum expressing heterologous oxalate decarboxylase. J Appl Microbiol 115(3):880–887. https://doi.org/10.1111/jam.12269
Åvall-Jääskeläinen S, Hynönen U, Ilk N, Pum D, Sleytr U, Palva A (2008) Identification and characterization of domains responsible for self-assembly and cell wall binding of the surface layer protein of Lactobacillus brevis ATCC 8287. BMC Microbiol 8:165. https://doi.org/10.1186/1471-2180-8-165
Axelsson L, Lindstad G, Naterstad K (2003) Development of an inducible gene expression system for Lactobacillus sakei. Lett Appl Microbiol 37(2):115–120. https://doi.org/10.1046/j.1472-765X.2003.01360.x
Bermúdez-Humarán LG, Kharrat P, Chatel J-M, Langella P (2011) Lactococci and lactobacilli as mucosal delivery vectors for therapeutic proteins and DNA vaccines. Microb Cell Fact 10(1):S4. https://doi.org/10.1186/1475-2859-10-S1-S4
Bezzerri V, Borgatti M, Finotti A, Tamanini A, Gambari R, Cabrini G (2011) Mapping the transcriptional machinery of the IL-8 gene in human bronchial epithelial cells. J Immunol 187(11):6069. https://doi.org/10.4049/jimmunol.1100821
Böhmer N, Lutz-Wahl S, Fischer L (2012) Recombinant production of hyperthermostable CelB from Pyrococcus furiosus in Lactobacillus sp. Appl Microbiol Biotechnol 96(4):903–912. https://doi.org/10.1007/s00253-012-4212-z
Bron P, Baarlen P, Kleerebezem M (2011) Emerging molecular insights into the interaction between probiotics and the host intestinal mucosa. Nat Rev Microbiol 10:66–78. https://doi.org/10.1038/nrmicro2690
Cano Garrido O, Seras J, Garcia-Fruitos E (2015) Lactic acid bacteria: reviewing the potential of a promising delivery live vector for biomedical purposes. Microb Cell Fact 14:137. https://doi.org/10.1186/s12934-015-0313-6
Carvalho RDDO, do Carmo FLR, de Oliveira Junior A, Langella P, Chatel J-M, Bermúdez-Humarán LG, Azevedo V, de Azevedo MS (2017) Use of wild type or recombinant lactic acid bacteria as an alternative treatment for gastrointestinal inflammatory diseases: a focus on inflammatory bowel diseases and mucositis. Front Microbiol 8:800–800. https://doi.org/10.3389/fmicb.2017.00800
Chen X, Xu J, Shuai J, Chen J, Zhang Z, Fang W (2007) The S-layer proteins of Lactobacillus crispatus strain ZJ001 is responsible for competitive exclusion against Escherichia coli O157:H7 and Salmonella typhimurium. Int J Food Microbiol 115(3):307–312. https://doi.org/10.1016/j.ijfoodmicro.2006.11.007
Duary RK, Rajput Y, Batish V, Grover S (2011) Assessing the adhesion of putative indigenous probiotic lactobacilli to human colonic epithelial cells. Indian J Med Res 134:664–671. https://doi.org/10.4103/0971-5916.90992
Fredriksen L, Mathiesen G, Sioud M, Eijsink VGH (2010) Cell wall anchoring of the 37-kilodalton oncofetal antigen by Lactobacillus plantarum for mucosal cancer vaccine delivery. Appl Environ Microbiol 76(21):7359–7362. https://doi.org/10.1128/AEM.01031-10
Fredriksen L, Kleiveland C, Olsen Hult L, Lea T, Nygaard C, Eijsink V, Mathiesen G (2012) Surface display of N-terminally anchored invasin by Lactobacillus plantarum activates NF- B in monocytes. Appl Environ Microbiol 78:5864–5871. https://doi.org/10.1128/AEM.01227-12
Jacobsen CN, Rosenfeldt V, Hayford AE, Møller PL, Michaelsen K, Paerregaard A, Sandström B, Tvede M, Jakobsen M (1999) Screening of probiotic activities of forty-seven strains of Lactobacillus spp. by in vitro techniques and evaluation of the colonization ability of five selected strains in humans. Appl Environ Microbiol 65:4949–4956. https://doi.org/10.1128/AEM.65.11.4949-4956.1999
Ji J, Yang H (2020) Using probiotics as supplementation for Helicobacter pylori antibiotic therapy. Int J Mol Sci 21(3):1136. https://doi.org/10.3390/ijms21031136
Ji D, Ma J, Xu M, Agyei D (2021) Cell-envelope proteinases from lactic acid bacteria: biochemical features and biotechnological applications. Compr Rev Food Sci Food Safety 20(1):369–400. https://doi.org/10.1111/1541-4337.12676
Johnson-Henry K, Hagen K, Gordonpour M, Tompkins T, Sherman P (2007) Surface-layer protein extracts from Lactobacillus helveticus inhibit enterohaemorrhagic Escherichia coli O157:H7 adhesion to epithelial cells. Cell Microbiol 9:356–367. https://doi.org/10.1111/j.1462-5822.2006.00791.x
Jørgensen CM, Vrang A, Madsen SM (2014) Recombinant protein expression in Lactococcus lactis using the P170 expression system. FEMS Microbiol Lett 351(2):170–178. https://doi.org/10.1111/1574-6968.12351
Keersmaecker S, Braeken K, Verhoeven T, Vélez M, Lebeer S, Vanderleyden J, Hols P (2006) Flow cytometric testing of green fluorescent protein-tagged Lactobacillus rhamnosus GG for response to defensins. Appl Environ Microbiol 72:4923–4930. https://doi.org/10.1128/AEM.02605-05
Kim MG, Kim CL, Kim YS, Jang JW, Lee GM (2021) Selective endocytosis of recombinant human BMPs through cell surface heparan sulfate proteoglycans in CHO cells: BMP-2 and BMP-7. Sci Rep 11(1):3378. https://doi.org/10.1038/s41598-021-82955-1
Kinoshita H, Hirata Y, Nakagawa H, Sakamoto K, Hayakawa Y, Takahashi R, Nakata W, Sakitani K, Serizawa T, Hikiba Y, Akanuma M, Shibata W, Maeda S, Koike K (2013) Interleukin-6 mediates epithelial-stromal interactions and promotes gastric tumorigenesis. PLoS One 8(4):e60914–e60914. https://doi.org/10.1371/journal.pone.0060914
Kobierecka P, Wyszyńska A, Maruszewska-Cheruiyot M, Wojtania A, Żylińska J, Bardowski J, Jagusztyn-Krynicka E (2015) Lactic acid bacteria as a surface display platform for Campylobacter jejuni antigens. J Mol Microbiol Biotechnol 25:1–10. https://doi.org/10.1159/000368780
Krachler AM, Orth K (2013) Targeting the bacteria–host interface: strategies in anti-adhesion therapy. Virulence 4:284–294. https://doi.org/10.4161/viru.24606
Kuo C-H, Wang SSW, Lu C-Y, Hu H-M, Kuo F-C, Weng B-C, Wu C-C, Liu C-J, Tsai P-Y, Lee T-C, Chen L-W, Cheng K-H, Chang L-L, Wu D-C (2013) Long-term use of probiotic-containing yogurts is a safe way to prevent <i>Helicobacter pylori</i>: based on a Mongolian Gerbil’s model. Biochem Res Int 2013:594561. https://doi.org/10.1155/2013/594561
Kusters J, van Vliet A, Kuipers E (2006) Pathogenesis of Helicobacter pylori infection. Clin Microbiol Rev 19:449–490. https://doi.org/10.1128/CMR.00054-05
Lesbros D, Corthesy-Theulaz I, Blum A (2007) Helicobacter pylori and probiotics. J Nutr 137:812S-818S. https://doi.org/10.1093/jn/137.3.812S
Li N, Russell WM, Douglas-Escobar M, Hauser N, Lopez M, Neu J (2009) Live and heat-killed Lactobacillus rhamnosus GG: effects on proinflammatory and anti-inflammatory cytokines/chemokines in gastrostomy-fed infant rats. Pediatr Res 66(2):203–207. https://doi.org/10.1203/PDR.0b013e3181aabd4f
Liu J-K, Wei C-H, Hou X-L, Yu L-Y (2014) Passive protection of mice pups through oral or intranasal immunization of dams with recombinant Lactobacillus casei vaccine against ETEC F41. Res Vet Sci 96(2):283–287. https://doi.org/10.1016/j.rvsc.2014.01.010
Majidzadeh Heravi R, Rouzbeh L, Sankian M, Kermanshahi H, Varasteh AR (2012) Optimization and comparison of two electrotansformation methods for lactobacilli. Biotechnology 11:50–54. https://doi.org/10.3923/biotech.2012.50.54
Martín C, Fernández-Vega I, Suárez J, Quirós L (2020) Adherence of Lactobacillus salivarius to hela cells promotes changes in the expression of the genes involved in biosynthesis of their ligands. Front Immunol 10:3019. https://doi.org/10.3389/fimmu.2019.03019
Mathiesen G, Namløs H, Risoen PA, Axelsson L, Eijsink V (2004) Use of bacteriocin promoters for gene expression in Lactobacillus plantarum C11. J Appl Microbiol 96:819–827. https://doi.org/10.1111/j.1365-2672.2004.02206.x
Miller JH (1972) Experiments in molecular genetics. Cold Spring Harbor Laboratory, Cold Spring Harbor, N.Y.
Pavan S, Hols P, Delcour J, Geoffroy M-C, Grangette C, Kleerebezem M, Mercenier A (2000) Adaptation of the Nisin-controlled expression system in Lactobacillus plantarum: a tool to study in vivo biological effects. Appl Environ Microbiol 66:4427–4432. https://doi.org/10.1128/AEM.66.10.4427-4432.2000
Pei H, Liu J, Cheng Y, Sun C, Wang C, Lu Y, Ding J, Zhou J, Xiang H (2005) Expression of SARS-coronavirus nucleocapsid protein in Escherichia coli and Lactococcus lactis for serodiagnosis and mucosal vaccination. Appl Microbiol Biotechnol 68(2):220–227. https://doi.org/10.1007/s00253-004-1869-y
Pontes D, Azevedo M, Chatel J-M, Langella P, Azevedo V, Miyoshi A (2011) Lactococcus lactis as live vector: heterologous protein production and DNA delivery systems. Protein Expr Purif 79:165–175. https://doi.org/10.1016/j.pep.2011.06.005
Pospiech A, Neumann B (1995) A versatile quick-prep of genomic DNA from gram-positive bacteria. Trends Genet 11(6):217–218. https://doi.org/10.1016/S0168-9525(00)89052-6
Reveneau N, Geoffroy M-C, Locht C, Chagnaud P, Mercenier A (2002) Comparison of the immune responses induced by local immunizations with recombinant Lactobacillus plantarum producing tetanus toxin fragment C in different cellular locations. Vaccine 20:1769–1777. https://doi.org/10.1016/S0264-410X(02)00027-0
Ruiz Rodríguez LG, Mohamed F, Bleckwedel J, Medina R, De Vuyst L, Hebert EM, Mozzi F (2019) Diversity and functional properties of lactic acid bacteria isolated from wild fruits and flowers present in Northern Argentina. Front Microbiol. https://doi.org/10.3389/fmicb.2019.01091
Ruiz-Bustos E, Sierra A, Romero MJ, Rodríguez-Jaramillo M, Ascencio F (2000) Protection of BALB/c mice against experimental Helicobacter pylori infection by oral immunisation with H. pylori heparan sulphate-binding proteins coupled to cholera toxin ??-subunit. J Med Microbiol 49:535–541. https://doi.org/10.1099/0022-1317-49-6-535
Sahoo A, Mandal AK, Dwivedi K, Kumar V (2020) A cross talk between the immunization and edible vaccine: current challenges and future prospects. Life Sci 261:118343–118343. https://doi.org/10.1016/j.lfs.2020.118343
Segers ME, Lebeer S (2014) Towards a better understanding of Lactobacillus rhamnosus GG - host interactions. Microb Cell Fact 13(1):S7. https://doi.org/10.1186/1475-2859-13-S1-S7
Selgrad M, Malfertheiner P (2008) New strategies for Helicobacter pylori eradication. Curr Opin Pharmacol 8:593–597. https://doi.org/10.1016/j.coph.2008.04.010
Song AA-L, In LLA, Lim SHE, Rahim RA (2017) A review on Lactococcus lactis: from food to factory. Microb Cell Fact 16(1):55. https://doi.org/10.1186/s12934-017-0669-x
Sun Y, Wilkinson B, Standiford T, Akinbi H, O’Riordan M (2012) Fatty acids regulate stress resistance and virulence factor production for listeria monocytogenes. J Bacteriol 194:5274–5284. https://doi.org/10.1128/JB.00045-12
Terpou A, Papadaki A, Lappa IK, Kachrimanidou V, Bosnea LA, Kopsahelis N (2019) Probiotics in food systems: significance and emerging strategies towards improved viability and delivery of enhanced beneficial value. Nutrients 11(7):1591. https://doi.org/10.3390/nu11071591
Tsuzaki M, Bynum D, Almekinders L, Yang X, Faber J, Banes AJ (2003) ATP modulates load-inducible IL-1β, COX 2, and MMP-3 gene expression in human tendon cells. J Cell Biochem 89(3):556–562. https://doi.org/10.1002/jcb.10534
Wyszyńska A, Kobierecka P, Bardowski J, Jagusztyn-Krynicka EK (2015) Lactic acid bacteria—20 years exploring their potential as live vectors for mucosal vaccination. Appl Microbiol Biotechnol 99(7):2967–2977. https://doi.org/10.1007/s00253-015-6498-0
Yang B, Chen H, Gu Z, Fengwei T, Ross R, Stanton C, Chen Y, Chen W, Zhang H (2014) Synthesis of conjugated linoleic acid by the linoleate isomerase complex in food-derived lactobacilli. J Appl Microbiol. https://doi.org/10.1111/jam.12524
Funding
This research work was financially supported by the Indian Council of Medical Research (ICMR), New Delhi, under the ICMR-PDF (3/1/3/PDF(7)/2013-HRD) program and Science and Engineering Research Board, Government of India, under Early Carrier Research Award (No. ECR/2017/000270) Grant.
Author information
Authors and Affiliations
Contributions
AKY received funds, conducted experiments, and wrote the manuscript. SVR conduct experiments of qPCR and analysed data. ND, AK and MK analysed data of the manuscript. RH thoroughly revised the manuscript for its intellectual content and gave final approval for the corrected version of the manuscript. All authors have read and agreed to the published version of the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Supplementary Information
Below is the link to the electronic supplementary material.
13205_2022_3428_MOESM1_ESM.pptx
Supplementary file1 Fig. S1 Growth curve of (A) recombinant-LGG and (B) non-recombinant LGG at different pH-2.5, 4.0 and 6.5. Growth pattern of recombinant-LGG and non-recombinant-LGG at different pH on various time intervals were non-significant (P>0.05). Fig. S2 Agarose gel showing PCR amplification of Signal and anchor sequences. (A) 1. Csp signal from genomic DNA of LP 21, 2. PrtR Anchor from Genomic DNA of LGG, M. 50 bp DNA Ladder, (B) Overlap PCR amplification fusion product using Csp_F_Nco1/PrtR_R_HindIII, M. 1kb DNA Ladder. Fig. S3 Expression study of recombinant LGG expressing green fluorescent protein. (A) Control: LGG cells harbouring pSIP503CMR (B) Fluorescence detection of recombinant-LGG cells harbouring pSIP503CMR-Gfp induced with 50 ng/ml of nisin using fluorescent microscope. Fig. S4 SDS-PAGE and western blot of Hsbp protein isolated from recombinant-LGG. SDS-PAGE showing the expression of Hsbp protein induced with nisin, Lane: 1. Un-induced LGG culture harbouring pSIP503CMR-Hsbp cell lysate, 2. LGG culture harbouring LGG-pSIP503CMR-Hsbp cell lysate induced with 25 ng/ml of nisin, 3. LGG culture harbouring LGG-pSIP503CMR-Hsbp cell lysate induced with 50 ng/ml of nisin. B. Western blot showing recombinant Hsbp protein using Novex ECL Chemiluminescent detection system. Fig. S5 Nucleotide sequence of the fusion construct. 1-6: NcoI, 7-71: Csp (signal sequence), 72-107 NEXT (NdeI, EcoRI, XhoI, Thrombin), 108-638: prtR (Anchor sequence) and 638-644: HindIII (PPTX 1272 KB)
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Yadav, A.K., Varikuti, S.R., Kumar, A. et al. Expression of heterologous heparan sulphate binding protein of Helicobacter pylori on the surface of Lactobacillus rhamnosus GG. 3 Biotech 13, 19 (2023). https://doi.org/10.1007/s13205-022-03428-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s13205-022-03428-4