Abstract
The well-known secondary metabolite-producing bacterium Streptomyces coelicolor is a natural choice for the development of super-hosts optimized for the heterologous expression of antibiotic biosynthetic gene clusters (BGCs). In this study, we used S. coelicolor M145 and its derivative strain M1146 where all active BGCs have been deleted and generated high-resolution quantitative time series metabolite profiles under two cultivation conditions (phosphate and nitrogen limitation to cease growth and trigger secondary metabolism). Five targeted LC–MS/MS-based methods were used to quantify intracellular primary metabolites covering phosphorylated metabolites, amino acids, organic acids, (deoxy) nucleoside/sugar phosphates, Nicotinamide adenine dinucleotide (NAD), and Coenzyme A (CoA). The nitrogen limitation resulted in a sharp decline in respiration and an immediate drop in the cell mass concentration. Intracellularly, a reduction in the level of the metabolites next to α-ketoglutarate in the tricarboxylic acid cycle and a decrease in the NADH pool were among the most prominent adaptation to this nutrient limitation. Phosphate limitation evoked a different adaptation of the metabolite pools as most of the phosphorylated metabolite pools except 6-phosphogluconic acid (6PG) pool were downregulated. 13C-isotope-labeling experiments revealed the simultaneous activity of both glycolysis and gluconeogenesis during the co-utilization of glucose and glutamate. The S. coelicolor M1146 strain had similar time-series metabolite profile dynamics as the parent M145 strain, except for a visibly increased 6PG pool in the stationary phase. In general, the nutrient limitation had a larger effect on the metabolite pool levels than the absence of secondary metabolite production in M1146. This study provides new insight into the primary carbon metabolism and its link to the secondary metabolism which is needed for further optimization of both super-host genotype and cultivation conditions.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Antibiotic resistance is dramatically increasing requiring enhanced efforts to search for new antibiotics to combat the emerging clinical crisis (Hawkey and Jones 2009; Hwang et al. 2014). The rediscovery of the same bioactive biomolecules is one of the main bottlenecks undermining global efforts. Actinobacteria contribute to over half of the identified bacterial bioactive molecules and among these, around 70% of the antibiotics are produced by the Streptomyces genus (Mahajan and Balachandran 2012; Ranjani et al. 2016). Streptomyces coelicolor A3(2) is a well-studied model system producing different types of secondary metabolites (antibiotics, pigments) e.g. actinorhodin, undecylprodigiosin, calcium-dependent antibiotic (CDA) (Rioseras et al. 2014; Bystrykh et al. 1996). It was the first Streptomyces sp. to be fully genome sequenced and contains almost thirty cryptic tentative secondary metabolite gene clusters (BGCs) (Bentley et al. 2002). The genetic toolbox is quite advanced which has made it the most frequently used model strain to investigate the regulation of antibiotic biosynthesis (Kieser et al. 2000). Thus, S. coelicolor is a natural choice for the development of optimized super-hosts for the heterologous expression of BGCs from different native sources to produce new bioactive compounds.
The central carbon metabolites (CCMs) carry most of the metabolic flux and provide precursors, reductive power, and energy used for biomass generation and secondary metabolite synthesis and their analysis can give real-time status of phenotype (Coze et al. 2013; Kumar et al. 2021; Rost et al. 2020a; Kumar and Bruheim 2021a, b). Therefore, information about the intracellular metabolite concentrations provided by metabolome analysis will further enhance our understanding of the genotype–phenotype interactions both by direct single interpretation of metabolomics data but also when integrated with multi omics-data such as genomics (Xu et al. 2017; Nguyen et al. 2012), transcriptomics (Kim and Kim 2016; Yague et al. 2014; Nieselt et al. 2010) and proteomics (Thomas et al. 2012). Mathematical modeling, such as genome-scale metabolic models will also become important in this workflow (Amara et al. 2018; Borodina et al. 2005; Sulheim et al. 2020).
The success of comprehensive and absolute quantitative metabolite profiling depends upon robust and reproducible cultivation conditions combined with optimized workflow consisting of several steps such as sampling methods, sample processing, and the efficiency of the LC–MS/MS system to generate high-resolution metabolite data (Villas-Boas and Bruheim 2007). The filamentous growth of Streptomyces makes it challenging for homogeneous and consistent sampling (Yague et al. 2010). Previously, cultivation conditions of S.coelicolor A3(2) were improved to obtain dispersed and reproducible growth by subjecting it to a high agitation rate causing high shear stress (Wentzel et al. 2012a). In the same study, a media was reported consisting of two carbon sources: D-glucose and L-glutamate; that at optimized concentrations delay the onset of secondary metabolism and provide enough biomass for sampling during the distinct growth, transition and antibiotic production phases. Several studies have reported metabolome data based on the analysis of secondary metabolites (Cihak et al. 2017; Goodwin et al. 2015; Lee et al. 2015; Morgenstern et al. 2015; Traxler et al. 2013), but only a few studies are reported for understanding the physiology of S.coelicolor by diving into comprehensive coverage of primary metabolites (Nitta et al. 2020; Wentzel et al. 2012b). Comparing the metabolite data sets from different laboratories is also challenging because they are generated using different continuously improved analytical methodologies, under different cultivation set-ups and physiological conditions. Most data is also only reported as abundances and compared on a relative scale.
Primary metabolites have both different physicochemical properties and roles, based on which they are classified into different metabolite classes [i.e. amino acids, organic acids, phosphorylated metabolites (deoxy) nucleoside/sugar phosphates, and cofactors like Nicotinamide adenine dinucleotide (NAD) and Coenzyme A (CoA) metabolites]). This is the reason to adopt several metabolite class-specific sampling and analytical protocols for comprehensive and complete quantitative coverage of the CCMs. We have developed a set of LC–MS/MS methods for the absolute quantitative metabolite profiling including the 13C-isotope dilution approach for enhanced analytical accuracy (Supplementary Fig. S1) (Bartosova et al. 2021; Kumar et al. 2021; Rost et al. 2020a; Kumar and Bruheim 2021b). The current study aimed to apply these methods for the quantification of CCMs of S. coelicolor M145 during the growth and antibiotic production phases in two nutrient-limited conditions (L-glutamate and phosphate limited). S coelicolor M145 is a plasmid cured derivative strain of the wild type strain A3(2) (Kieser et al. 2000) and a reference strain of a series of new derivative strains (Gomez-Escribano and Bibb 2011). Recently, we have published a comprehensive coverage of primary metabolome data by comparing super host strain S. coelicolor M1152 and its derivative S. coelicolor M1581 heterologously expressing a chloramphenicol BGC (Kumar and Bruheim 2021b). The current study was designed to further decipher the distribution and participation of carbon from two substrates (D-glucose and L-glutamate) in primary metabolic pathways and their role in the growth and support of antibiotic biosynthesis. The M145 reference strain and the derivative strain S. coelicolor M1146 where the four native BGCs being expressed (actinorhodin, undecylprodigiosin, calcium-dependent antibiotics, methylenomycin) have been inactivated were selected for this study as this mutant strain provides a cleaner background for heterologous expression of new and potentially bioactive secondary metabolites. Importantly, this is probably the first report on LC–MS/MS-based absolute quantification of intracellular concentrations of NAD, NADH, NADP, and NADPH during growth, and secondary metabolite production phases. From a super-host development perspective, it is especially critical to verify that sufficient NADPH supplies are available throughout all cultivation stages.
Materials and methods
Strains and cultivation conditions
Two strains of Streptomyces coelicolor were used in this study: S. coelicolor M145 and S. coelicolor M1146. The protocols for spore preparation, pre-germination, media preparation, and setting up the experiments were adopted from earlier reports (Wentzel et al. 2012a; Kumar and Bruheim 2021b). The cultivations were conducted in 3 L bioreactors (Eppendorf Bioflo 320) with an initial working volume of 1.8 L and the inoculum was pre-germinated spores having ~ 1 × 109 CFU L−1. The study was conducted in phosphate (SSBM-P) and nitrogen (L-glutamate) limited (SSBM-E) media at 30 °C, pH 7.0. SSBM media consisted of two carbon sources: D-glucose and L-glutamate. SSBM-P media consisted of D-glucose (40 g L−1), Sodium L-glutamate monohydrate (61.1 g L−1), Phosphate (4.6 mM), Magnesium sulfate (2.0 mM), and previously reported minimal medium trace element solution (Wentzel et al. 2012a). SSBM-E media had a similar composition as SSBM-P media except for a reduced amount of Sodium L-glutamate monohydrate (16.6 g L−1), and an increased amount of phosphate (9.2 mM). The 13C isotope labeling experiments were conducted at 30 °C, pH 7.0, in customized mini bioreactors (working volume of 200 mL) where the agitation was provided using a magnetic stirrer. The SSBM-P media used in the study had a similar composition as stated above except the D-glucose was 13C labeled at all carbon positions (U-13C6, 99%, Cambridge Isotope Laboratories, Inc.) and its concentration was reduced to 20 g L−1. Time-series sampling was analyzed for better understating and the direct interpretation of dynamic 13C labeling patterns (Buescher et al. 2015).
Quantification of exometabolites
The fermentation broth was sampled into 15 mL centrifuge tubes and biomass pelleted using centrifugation at 4500 × g, 4 °C and 5 min. The pellets were washed three times with NaCl solution (9 g L−1) followed by three times with ultrapure water using centrifugation. Dry cell weight (DCW) was quantified by transferring biomass pellets into a pre-dried and pre-weighed aluminum pan and dried at 110 °C till constant weight was achieved.
The supernatant of the fermentation broth was used for the analysis of D-glucose and excreted organic acids using high-performance liquid chromatography (HPLC). Before analysis, the supernatant samples were filtered using syringe filters with a 0.2 μm polyethersulfone membrane (VWR). The Waters Alliance HPLC was equipped with a refractive index (RI) and a UV/VIS detector to measure the carbohydrates and organic acids, respectively. The Hi-Plex column of dimension 300 × 7.7 mm was used to separate the sugars and organic acids. The column and the sample compartment were set at 45 °C and 10 °C, respectively. The mobile phase was 0.05 M H2SO4 in ultrapure water at the flow rate of 0.8 mL min−1.
The extracellular L-glutamate was analyzed separately using LC–MS/MS following a similar protocol (described later) as for the intracellular amino acid analysis method except using a shorter C18 column (1.7 µm, 2.1 × 75 mm) to reduce the analysis run time.
The intracellular (actinorhodin, Act) and extracellular (γ Act) blue pigment were extracted from biomass pellets and supernatants, respectively. The protocol was adopted from the previous report (Coze et al. 2013). The Act was analyzed by resuspending the pellet in 1.0 mL KOH (1.0 M), and with thorough mixing for 20 min at 4 °C and centrifuged at 20,817 × g, 4 °C, and 5 min. After this step, Act and γ Act were extracted following similar steps. The supernatant was collected and 0.5 mL HCl (3.0 M) was added to precipitate the Act. The Act was pelleted by incubating the sample at 4 °C for 15 min, followed by centrifugation at 20,817 × g, 4 °C for 5 min. The pellet was resuspended in 1.0 mL KOH (1.0 M) and the absorbance of the samples was measured at 644 nm using a plate reader. The Act concentration was calculated using the molar extinction coefficient of 25,320 L mol−1 cm−1 for pure Act in 1.0 M KOH.
The extraction of intracellular red pigment (Undecylprodigiosin, Red) was adopted from the previous report (Sheng-wan Tsao et al. 1985). The biomass pellet obtained from 1.0 mL fermentation broth was resuspended in 1.0 mL methanol, and with thorough mixing for 30 min at 4 °C, and centrifuged at 20,817 × g, 4 °C, and 5 min. The supernatant was collected and 0.5 mL HCl was added, and the sample was incubated at room temperature for 5 min and followed by centrifugation at 20,817 × g for 5 min at room temperature. The absorbance of the samples was measured at 530 nm using a plate reader. The Red concentration was calculated using the molar extinction coefficient of 100,500 L mol−1 cm−1 for the pure Red in an equal volume of methanol/HCl (1 M).
Quantification of endometabolites
Sampling and sample preparation
Time-series sampling for intracellular metabolite profiling was conducted in the exponential and the nutrient depletion phase. The fermentation cell suspension was pipetted from the bioreactors and immediately filtered using a Durapore® 0.65 μm polyvinylidene fluoride filter (Merck) through a funnel connected to a vacuum (200 bar), and rinsed with 10 mL, ice-cooled 2.3% NaCl solution followed by 10 mL ice-cooled, ultrapure water. The filter containing the biomass was transferred to 13 mL ice-cooled 55:45 ACN: H2O solution in 50 mL tubes, followed by immediate quenching in liquid nitrogen, N2(l), and stored at -80 °C until further processing.
The intracellular metabolite from the samples was extracted following three times freezing and thawing and with an intermittent brief vertexing after each thawing. The thawing step was conducted at 0 °C in an ethanol bath and freezing was done using liquid nitrogen, N2(l). This was followed by centrifugation (4500 × g, 4 °C, 10 min) and aliquoting 3.5 mL of sample into three 15 mL tubes. The aliquoted samples were dried for ~ 24 h at − 105 °C and 12 Pa in a freeze dryer. The dried metabolites were re-constituted in 550 μL HPLC-grade water and centrifuged (4500 × g, 4 °C, 5 min) to remove large cell debris. The supernatant was transferred into 3-kDa-molecular-weight spin cut-off filters (516-0228, VWR) and centrifuged at 20,817 × g, 4 °C, 5 min to get the sample for further analysis.
Amino acids, organic acids, and phosphorylated metabolites analysis
For amino acids and organic acids, the samples were first derivatized before quantification using LC–MS/MS. The derivatization protocol and LC–MS/MS analysis applied for amino acids and organic acids were adopted as previously reported (Kumar et al. 2021). All phosphorylated metabolites and some tricarboxylic acid (TCA) cycle metabolites were quantified using capillary ion chromatography and tandem mass spectroscopy (CapIC-MS/MS) as described in previous reports (Kvitvang et al. 2014; Kumar et al. 2021; Stafsnes et al. 2018).
Nicotinamide adenine dinucleotide (NAD) and Coenzyme A (CoA) metabolites analysis
A different sampling protocol had to be developed for the quantitation of CoAs and NADs. First, pellets were collected by centrifugation at 4 °C, for one minute at a maximum speed: 4500 × g for 15 mL centrifuge tubes (for exponential phase samples with less biomass) and 20,817 × g for 2 mL centrifuge tubes (for stationary phase samples). Supernatants were discarded, and pellets were quickly quenched in N2(l) and stored at − 80 °C until further analysis. NAD and CoA metabolites extraction were optimized using S. coelicolor as a biological model system and then extracted following cold extraction and quantified as described previously (Bartosova et al. 2021; Kumar and Bruheim 2021a; Rost et al. 2020b). Cold extractions in 50 mM NH4Ac having a composition of acetonitrile:buffer:CH3OH (7:2:1) including 5% of a 13C-labelled biological extract (see below), at pH 9 and 5, respectively, for NADs and CoAs. The extracts were incubated at 4 °C for 3 min while agitating at 1500 rpm. 13C-labeled biological extract (E.coli, 10 mL, OD 2.3) was prepared similarly. Samples were centrifuged at 4 °C, 2 min, 20,817 × g for clearing cell debris. It was followed by transferring supernatants into 3-kDa-molecular-weight spin cut-off filters (516-0228, VWR) and centrifugation at 4 °C, 5 min, 20,817 × g. The standards used to prepare the calibration curves were prepared in a matrix containing 5%, 13C-labelled E.coli biological extract. Chromatographic separation was adopted from a previous report (Rost et al. 2020b) and performed using hydrophilic interaction liquid chromatography (HILIC) coupled with tandem mass spectroscopy. More about the development and assessment of the NAD and CoA quantitation can be found in Supplementary Figs. S2 and S3, respectively.
Data analysis and calculation
The specific growth rate, μ was calculated in the exponential phase, which is equal to the slope of ln[x] plotted as a function of time. The specific rates (specific D-glucose uptake rate, qgluc; specific L-glutamate uptake rate, qglut; specific Act, γAct and Red production rate were pAct, pγAct, pred, respectively) were calculated by dividing the volumetric rate by the average biomass between the two points. The carbon recovery percentage was measured between the first and the last point of analysis, which was calculated based on the amount of carbon (on a mole basis) recovered from the biomass, organic acids, CO2, and products (pigments), and the amount of carbon supplied in the form of substrates. The molecular weight of biomass was assumed as 27.0 g dry weight/mol (Kumar and Bruheim 2021b). The respiratory quotient (RQ) was calculated from the ratio of CO2 evolution to oxygen consumption.
The MS data were processed using the TargetLynx 4.2 software. Integration was performed automatically but manually checked to ensure quality. All metabolite concentrations were calculated based on an individual linear standard calibration curve. All metabolites concentrations were normalized using biomass. All intracellular metabolites pools were reported as an average of two to three independent replicas and/or supported by another independent batch cultivation. The CCMs are visualized using the Omix software (Droste et al. 2011). The principal component analysis (PCA) scores plot was plotted for S. coelicolor M145 metabolite time series data in SSBM-E and SSBM-P media using MetaboAnalyst software (Xia et al. 2009). Missing values were replaced by feature mean and autoscaled before PCA.
The equation below was used to calculate the Energy charge (Atkinson 1977):
In the 13C labeled experiment, the relative summed fractional labeling (Rel SFL) of each metabolite was calculated (scale 0 to 100) as shown below, where In is the signal area of metabolite with n number of labeled carbon (Isotopologues) (Gombert et al. 2001).
Results and discussion
In this study, we first present the cultivation data of the reference strain S. coelicolor M145 in the two different nutrients limited mediums (glutamate limited/N-lim: SSBM-E, and phosphate limited/P-lim: SSBM-P) and the derivative strain M1146 in SSBM-P conditions. Besides being an additional source of carbon, L-glutamate was the sole nitrogen source in the SSBM media. Therefore, the depletion of L-glutamate in the media corresponded to nitrogen limitation. No cultures are carbon limited at any state since glucose is in high abundance (never below 10 gL−1). Next, time series of primary metabolite pool profiles are presented and followed by the 13C-isotope labeling results to further enhance our understanding of the distribution and interaction of substrate carbon in the different central metabolic pathways. Finally, the metabolite pool composition of the derivative strain S. coelicolor M1146 is compared to the reference S. coelicolor M145 under P lim conditions.
Growth kinetics of S. coelicolor M145
L-glutamate/ N lim and P lim caused characteristic different CO2 evolution profiles (Fig. 1A and C). N lim resulted in a sharp decline in the CO2 evolution at the time of L-glutamate depletion, whereas a gradual decrease in the CO2 evolution was observed in P lim. The slope of the respiratory coefficient (RQ – ratio of CO2 production vs. O2 consumption) can be used to assess the respiratory state of the two cultivation conditions. In both cultivation conditions, RQ increased in the exponential phase till close to 1.1, followed by a sharp decline in N lim and a gradual decline in P lim (Fig. 1A and C). SSBM-E supported a slightly higher growth rate compared to the SSBM-P media (0.21 ± 0.04 vs. 0.16 ± 0.01 h−1) (Table 1), most likely due to the lower initial concentration of phosphate in P lim. There is a slight (statistically non-significant) trend that the specific L-glutamate uptake rate, qglut was higher compared to the D-glucose uptake rate. The qglut and qgluc were higher in P lim compared to the N lim media even though the growth rate was lower. The depletion of either L-glutamate or phosphate at a defined time resulted in a switch to the secondary production phase from the growth phase with a large decrease in the substrate consumption rates. Three antibiotics/pigments were quantified: intracellular actinorhodin (Act), extracellular actinorhodin (γ Act), and undecylprodigiosin (Red), which were produced after nutrient limitation in both media (Fig. 1B and D). It was observed that the specific production rates (pAct, pγAct, pred) of all three antibiotics/pigments were around 1.3, 2.0, 2.6 times higher in the P lim as compared to the N lim, respectively (Table 1). The M1146 strain cultivated in P lim conditions grew faster than M145 but the substrate consumption rates were comparable (Table 1 and Supplementary Fig. S7).
The online and offline cultivation data of S. coelicolor M145 in L-glutamate limited (A and B), and phosphate limited (C and D) media. The molecular weight of biomass is assumed as 27.0 g dry weight per mole. Offline data are the average of at least duplicate samples, with their respective standard deviation
Mass spectrometric profiling of central intracellular metabolite pools
More than 80 metabolites, with almost complete coverage in the pentose phosphate pathway (PPP), glycolytic, deoxy-/nucleoside, and sugar phosphates, amino acids, organic acids, CoA, and NAD metabolite classes, were quantified by applying five different LC–MS/MS methods (Supplementary Fig. S1). The time-series metabolite analysis revealed the dynamics in the S. coelicolor M145 metabolite pool composition during growth and stationary/secondary metabolism phases. Preliminary assessment of the global data set by PCA revealed a stronger dependence on cultivation conditions (N lim vs. P lim) since sampling points before the nutrient depletion clustered with respective later samples rather than a time-dependent growth phase clustering (Supplementary Fig. S4).
The intracellular metabolite concentrations varied over 104 orders of magnitude as shown in the time-course heat-map, where metabolites were grouped over seven color concentration intervals (Fig. 2). The relative standard deviation for most of the metabolites was below 30% (Supplementary Table S2 and Fig. S6). Several amino acids (Glutamate—Glu, Glutamine—Gln, Aspartate—Asp, Alanine—Ala) and organic acids (Pyruvate—Pyr and α-ketoglutarate—αKG) were among the highest abundant ones in both cultivation conditions, while there were many low abundant among the glycolytic, deoxynucleoside phosphate, NAD, and CoA metabolites. In general, many metabolite pools decreased in the stationary phase. Of particular interest to note is that the concentration of NAD was much higher than the other NAD metabolites and dropped dramatically in the P lim stationary phase. The culture still maintained the same energy charge levels (ECR). The average ECR for the S. coelicolor M145 was 0.48 ± 0.02 (in SSBM-E media), and 0.44 ± 0.07 (in SSBM-P media) (Supplementary Table S1).
Heat map representation of the absolute concentration of intracellular metabolite pools of S.coelicolor M145. White indicates that the metabolites were not analyzed/not included/ trace amount (< 1 nmol/gDCW). R5P is the combination of X5P, R5P, and RL5P. Concentrations and standard deviations can be found in Supplementary Table S2. Supplementary Fig. S6 also shows the heat-map representation of relative standard deviation for two to three samples. Expo and Stat stand for the sample points analyzed from the exponential and the stationary phase, respectively
Next, intracellular metabolites were grouped in metabolite classes to draw a better understanding of the effect of cultivation conditions and growth phases. The intracellular Glu level was not included when calculating the percentage composition because of two reasons: first, Glu contributes to most of the concentration of the total metabolites, thus overshadowing the visualization of the pool contribution of the other metabolites, and second, a possible overestimated quantification of Glu due to its contamination from the L-glutamate containing media. Similar to our previous report on S. coelicolor M1152 and M1581 strains (Kumar and Bruheim 2021b), a relative downregulation of all the phosphorylated metabolites and upregulation of amino acids were observed after phosphate depletion (Fig. 3). In contrast, L-glutamate depletion did not compromise the phosphorylated metabolite pools, but most noteworthy was the reduction in the TCA cycle metabolite pools succeeding aKG.
The stacked bar diagram shows the intracellular metabolites pool (excluding glutamate) in the exponential phase and the stationary phase when S. coelicolor M145 was cultivated in (A) L-glutamate limited and B phosphate limited media. The pointed arrow on the X-axis separates the exponential (Expo) and the stationary (Stat) phase
The relative change in individual metabolite levels across different time points and cultivation conditions was visualized through another time series heat-map representation. For this purpose, the ratio of the metabolite concentration at individual time points to the average concentration across all sampling points was log2 transformed for each cultivation. In N lim medium, most of the metabolites had a high level at the early exponential phase, decreased around glutamate depletion but recovered at the end of the cultivation (Supplementary Fig. S5). Contrary to N lim, in general, metabolite pools peaked in the mid-exponential phase in P lim medium except for the amino acids that were low in the growth phase but accumulated in the P lim induced stationary phase. Most of the phosphorylated metabolites were decreased after experiencing phosphate depletion. The Fig. S5 plot highlights the dynamics of the central metabolome during growth, transition, and finally secondary metabolism phase.
Inspection and interpretation of such large data sets are challenging, and therefore the highest metabolite levels from each of the exponential and nutrient depletion phases were selected for further assessment of the results. The log2 transformed ratio between the nutrient depletion and exponential phase is presented in the metabolic network perspective in Fig. 4. The overall picture is quite remarkable: upper glycolytic and PPP pools accumulate while lower glycolytic, TCA, and amino acids pools decrease in the N lim cultivation, and the opposite adaption in the metabolite pools are observed in the P lim cultivation (Fig. 4A vs B). It is only the nucleoside phosphate pools that change in the same direction (downregulation) for both cultivations. In N lim of S. coelicolor M145, the depletion of L-glutamate in the cultivation broth is not counteracted by upregulation of glucose consumption (Table 1). This led to a downregulation of TCA activity resulting in decreased TCA metabolites and NADH levels, as well as the lower glycolytic metabolites (Fig. 4A). Both carbon sources are available throughout the whole P lim cultivation but a larger decrease in the stationary phase is observed in the glucose consumption rate than the glutamate consumption rate (Table 1). The twice as high (on mole basis) consumption of glutamate might be reflected in the accumulation of TCA and lower glycolytic metabolites.
Log2 fold change of central carbon metabolites (CCMs) of S.coelicolor M145 corresponding to the maximum levels of metabolites in the stationary phase relative to the exponential phase in (A) L-glutamate limited, B phosphate limited media. Grey indicates that the metabolites were not analyzed/not included. All metabolites were visualized using the Omix software (Droste et al. 2011)
Industrial Streptomyces bioprocesses are mostly designed with phosphate limitations since this supports higher secondary metabolite production rates, as also observed in the current study with defined medium conditions. Therefore, it is interesting to observe that many primary metabolite pools are downregulated in P lim (Fig. 4B) but there is not a single pronounced metabolite that stands out as the sole bottleneck/ limiting agent, neither among carbon precursors, reducing power (NADPH) nor energetic level (i.e. NADH and ATP). The current model system is not a high yield system and the next study should be to apply the same quantitative metabolite profiling methodology on a set of high yield strains with a much larger (e.g. > 10–20%) carbon flux from the substrates to the secondary metabolites. The current study is focused on validating the analytical methodology and verifying that the BGCs deletions of M1146 did not cause negative effects on the physiological level (see final paragraph).
13C-isotope tracing
A 13C-isotope labeling experiment was performed on S. coelicolor M145 in the P lim media to investigate how the carbons of D-glucose and L-glutamate were distributed among the CCMs in the growth and the antibiotics production phase. This study could potentially aid in the better understanding of the physiology of Streptomyces sp. and in the interpretation of the metabolite profile data described above, as also performed earlier by others, e.g. Obanye and co-workers used radiorespiroometry and reported a correlation between carbon flux through the PPP and production of the antibiotic (Obanye et al. 1996) and Gunnarson and co-workers applied GC–MS-based 13C labeled experimentation and discovered a switch from glycolysis to Entner-Doudoroff pathway during the transition from growth to secondary metabolism in the Actinomycete Nonomuria gerenzanensis (Gunnarson et al. 2004).
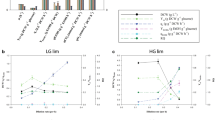
Isotopologues of selected metabolites from central metabolic pathways (upper and lower glycolysis, PPP, TCA cycle, purine, and pyrimidine nucleoside phosphate) showing 13C-label enrichment from the U-13C6-glucose are displayed in Fig. 5. Glucose-6-phosphate, (G6P), Fructose-6-phosphate (F6P), and Xylulose-5-phosphate/Ribose-5-phosphate/Ribulose-5-phosphate (X5P/R5P/RL5P) were almost completely 13C-labeled in the growth phase but there was a slight reduction in the stationary phase, which is due to the introduction of 12C-carbons from L-glutamate. Interestingly, this was not due to general 12C enrichment in all isotopologues but limited to M + 3 for the hexose-phosphate and M + 2 the pentose phosphates which indicates the simultaneous glycolysis and gluconeogenesis (bi-directionality) in these central metabolic pathways where complete 12C-C3 carbon skeletons (e.g. PEP) from L-glutamate is supplied to glycolysis through anaplerotic reactions connecting the glycolytic pathway and TCA (Coze et al. 2013; Sauer and Eikmanns 2005). Different 13C labeling patterns in isotopologues of Citrate (Cit) depended upon the reaction between the labeling pattern of oxaloacetate (OAA) and Acetyl coenzyme A (AcCoA). The cyclic nature of TCA and the simultaneous glycolysis and gluconeogenesis can potentially yield all isotopologues of Cit. However, it could be observed that there was an abundance of isotopologues of M + 0, M + 1, and M + 2. Succinate (Suc) mostly consisted of isotopologue of M + 0 carbon, signifying those metabolites downstream of the α-KG receiving most of the carbons from 12C carbon of L-glutamate. Purine (ATP and GTP) had a higher Rel SFL as compared to pyrimidine (CDP) nucleoside phosphates.
The relative abundance of the isotopologues of the selected metabolites from central carbon metabolites (CCMs) shows the distribution of 13C carbon from D-glucose and 12C carbon from L-glutamate of S.coelicolor M145 in phosphate limited media. A Glucose-6-phosphate, G6P; B fructose-6-phosphate, F6P; C xylulose-5-phosphate/ribose-5-phosphate/ribulose-5-phosphate, X5P/R5P/RL5P; D phosphoenolpyruvate, PEP; E citrate; F succinate; G ATP; H GTP; I CDP. The square dotes in each of the figures show the relative Summed Fractional Labeling (Rel SFL), corresponding to each time point. M + 0, M + 1, M + 2…M + n represent the molecular weight of metabolite isotopologues formed due to incorporation of 13C labeled carbons, where n is the maximum number of carbon present in that metabolite
In summary, upper glycolytic intermediates and PPP were rich with carbons from D-glucose, and lower glycolytic intermediates and Cit (upstream to α-KG) were rich with carbons from both D-glucose and L-glutamate, whereas intermediates downstream to α-KG were mostly composed of carbons from L-glutamate. The simultaneous bidirectionality of the glycolytic pathway was observed with 12C carbons of L-glutamate the following gluconeogenesis whereas 13C carbons of D-glucose were directed towards glycolysis. The bidirectionality of the glycolytic pathway also implied that some of the 12C carbons reentered into the TCA cycle as revealed by isotopologues of Cit. A progressive enrichment of 12C labeled carbon into CCMs coincided with a relative increase of L-glutamate over D-glucose consumption rate with the time of cultivation, especially in the antibiotics production phase.
Derivative strain M1146 vs reference M145 strain during Plim conditions
The derivative strain M1146 had a similar characteristic CO2 evolution profile but grew slightly faster than the parent M145 strain (Table 1 and Supplementary Fig. S7A). As expected, this BGCs deleted strain did not produce the pigments corresponding to the deleted BGCs. A higher specific growth rate of S. coelicolor M1146 might be because of a lower and simpler genetic and metabolic burden due to the deletion of four BGCs from this strain (Coze et al. 2013).
Phosphate limitation had nearly a similar effect on metabolite pools on both strains (Supplementary Fig. S7B–E). The metabolite pools were high in the exponential phase and most of the phosphorylated metabolites were sharply downregulated after experiencing the phosphate limitation. Similar to the wild/reference strain in P lim media, αKG and Pyruvate levels were increased in S. coelicolor M1146. The phosphate limitation in both the strains resulted in a decrease in the Suc pool in the growth phase and its reverse trend in the production phase. Interestingly, in the phosphate-depleted phase, a higher level of 6PG was observed in M1146 which might be linked to less NADPH consumption in absence of antibiotics production. M1146 had a slightly higher ECR (0.58 ± 0.11) compared to M145 (0.44 ± 0.07) which is in resonance with previous studies where an elevated level of ATP/ADP ratios correlated with improved growth performance (Esnault et al. 2017; Coze et al. 2013).
Conclusion
High-resolution quantitative metabolite profiles of Streptomyces coelicolor have been obtained in the nutrient-limited batch cultivation. The nutrient limitation had a larger effect on the metabolite pool composition compared to the deletion of BGCs. A 13C based study revealed the cultivation phase-dependent interplay of D-glucose and L-glutamate carbons into the central carbon metabolism.
Abbreviations
- 6PGL:
-
6-Phosphogluconolactone
- 6PG:
-
Phosphogluconic acid
- RL5P:
-
Ribulose-5-phosphate
- R5P:
-
Ribose-5-phosphate
- X5P:
-
Xylulose-5-phosphate
- E4P:
-
Erythrose 4-phosphate
- S7P:
-
Sedoheptulose-7-phosphate
- G6P:
-
Glucose-6-phosphate
- F6P:
-
Fructose-6-phosphate
- F1,6BP:
-
Fructose-1,6-bisphosphate
- DHAP:
-
Dihydroxyacetone phosphate
- G3P:
-
D-Glyceraldehyde 3-phosphate
- 3PG:
-
3-Phosphoglyceric acid
- 2PG:
-
2-Phosphoglyceric acid
- PEP:
-
Phosphoenolpyruvic acid
- Pyr:
-
Pyruvic acid
- 2-HG/OHGlu:
-
2-Hydroxyglutaric acid
- Cit:
-
Citric acid
- ICit:
-
Isocitric acid
- αKG:
-
α-Ketoglutaric acid
- Suc:
-
Succinic acid
- Fum:
-
Fumaric acid
- Mal:
-
Malic acid
- OAA:
-
Oxalacetic acid
- Ala:
-
Alanine
- Arg:
-
Arginine
- Asn:
-
Asparagine
- Asp:
-
Aspartic acid
- Gln:
-
Glutamine
- Glu:
-
Glutamic acid
- Gly:
-
Glycine
- His:
-
Histidine
- Leu:
-
Leucine,
- Ile:
-
Isoleucine
- Lys:
-
Lysine
- Met:
-
Methionine
- Phe:
-
Phenylalanine
- Pro:
-
Proline
- Ser:
-
Serine
- Thr:
-
Threonine
- Trp:
-
Tryptophane
- Tyr:
-
Tyrosine
- Val:
-
Valine
- AMP:
-
Adenosine monophosphate
- ADP:
-
Adenosine diphosphate
- ATP:
-
Adenosine triphosphate
- GMP:
-
Guanosine monophosphate
- GDP:
-
Guanosine diphosphate
- GTP:
-
Guanosine triphosphate
- CMP:
-
Cytidine monophosphate,
- CDP:
-
Cytidine diphosphate
- CTP:
-
Cytidine triphosphate
- UMP:
-
Uridine monophosphate
- UDP:
-
Uridine diphosphate
- UTP:
-
Uridine triphosphate
- TMP:
-
Thiamine monophosphate
- TDP:
-
Thymidine diphosphate
- TTP:
-
Thymidine triphosphate
- IMP:
-
Inosine monophosphate
- ITP:
-
Inosine triphosphate
- dAMP:
-
Deoxyadenosine monophosphate
- dADP:
-
Deoxyadenosine diphosphate
- dATP:
-
Deoxyadenosine triphosphate
- dGMP:
-
Deoxyguanosine monophosphate
- dGDP:
-
Deoxyguanosine diphosphate
- dGTP:
-
Deoxyguanosine triphosphate
- dCTP:
-
Deoxycytosine triphosphate
- dUMP:
-
Deoxyuridine monophosphate
- dUTP:
-
Deoxyuridine triphosphate
- dTTP:
-
Deoxythymidine triphosphate
- dIMP:
-
Deoxyinosine monophosphate
- cAMP:
-
Cyclic monophosphate
- PRPP:
-
Phosphoribosyl-pyrophosphate
- GAL1P:
-
Galactose-1-phosphate
- GAL6P:
-
Galactose-6-phosphate
- G1P:
-
Glucose-1-phosphate, G1P
- M1P:
-
Mannose-1-phosphate
- F1P:
-
Fructose-1-phosphate
- M6P:
-
Mannose-6-phosphate
- GL3P:
-
Glycerol-3-phosphate
- UDP-glu:
-
UDP-glucose
- UDP-Glc-Nac:
-
UDP-acetylglucosamine
- DOXP:
-
Deoxyxylulose 5-Phosphate
- 2-IPML:
-
2-Isopropylmalic acid
- IPP:
-
Isopentenyl pyrophosphate
- GA6P:
-
Glucosamine 6-phosphate
- GlcN6P:
-
N-Acetyl-glucosamine 6-phosphate
- Lac:
-
L-Lactic acid
- T6P:
-
Trehalose 6-phosphate
- CoA:
-
Coenzyme A
- AcCoA:
-
Acetyl coenzyme A
- Mal CoA:
-
Malonyl coenzyme A
- Suc CoA:
-
Succinyl coenzyme A
- NAD:
-
Nicotinamide adenine dinucleotide (oxidised)
- NADH:
-
Nicotinamide adenine dinucleotide (reduced)
- NADP:
-
Nicotinamide adenine dinucleotide phosphate (oxidised)
- NADPH:
-
Nicotinamide adenine dinucleotide phosphate (reduced)
- FAD:
-
Flavin adenine dinucleotide
References
Amara A, Takano E, Breitling R (2018) Development and validation of an updated computational model of Streptomyces coelicolor primary and secondary metabolism. BMC Genomics 19(519):1–9. https://doi.org/10.1186/s12864-018-4905-5
Atkinson DE (1977) Cellular energy control – adenylate energy-charge is a key factor. Trends Biochem Sci 2(9):198–200. https://doi.org/10.1016/0968-0004(77)90190-6
Bartosova Z, Ertesvag H, Nyflot EL, Kampe K, Aasen IM, Bruheim P (2021) Combined metabolome and lipidome analyses for in-depth characterization of lipid accumulation in the DHA producing Aurantiochytrium sp. T66. Metabolites 11(135):1–19. https://doi.org/10.3390/metabo11030135
Bentley SD, Chater KF, Cerdeno-Tarraga AM, Challis GL, Thomson NR, James KD, Harris DE, Quail MA, Kieser H, Harper D, Bateman A, Brown S, Chandra G, Chen CW, Collins M, Cronin A, Fraser A, Goble A, Hidalgo J, Hornsby T, Howarth S, Huang CH, Kieser T, Larke L, Murphy L, Oliver K, O’Neil S, Rabbinowitsch E, Rajandream MA, Rutherford K, Rutter S, Seeger K, Saunders D, Sharp S, Squares R, Squares S, Taylor K, Warren T, Wietzorrek A, Woodward J, Barrell BG, Parkhill J, Hopwood DA (2002) Complete genome sequence of the model actinomycete Streptomyces coelicolor A3(2). Nature 417(6885):141–147. https://doi.org/10.1038/417141a
Borodina I, Krabben P, Nielsen J (2005) Genome-scale analysis of Streptomyces coelicolor A3(2) metabolism. Genome Res 15(6):820–9. http://www.genome.org/cgi/doi/10.1101/gr.3364705
Buescher JM, Antoniewicz MR, Boros LG, Burgess SC, Brunengraber H, Clish CB, DeBerardinis RJ, Feron O, Frezza C, Ghesquiere B, Gottlieb E, Hiller K, Jones RG, Kamphorst JJ, Kibbey RG, Kimmelman AC, Locasale JW, Lunt SY, Maddocks OD, Malloy C, Metallo CM, Meuillet EJ, Munger J, Noh K, Rabinowitz JD, Ralser M, Sauer U, Stephanopoulos G, St-Pierre J, Tennant DA, Wittmann C, Vander Heiden MG, Vazquez A, Vousden K, Young JD, Zamboni N, Fendt SM (2015) A roadmap for interpreting (13)C metabolite labeling patterns from cells. Curr Opin Biotechnol 34:189–201. https://doi.org/10.1016/j.copbio.2015.02.003
Bystrykh LV, FernandezMoreno MA, Herrema JK, Malpartida F, Hopwood DA, Dijkhuizen L (1996) Production of actinorhodin-related “blue pigments” by Streptomyces coelicolor A3(2). J Bacteriol 178(8):2238–2244. https://doi.org/10.1128/jb.178.8.2238-2244.1996
Cihak M, Kamenik Z, Smidova K, Bergman N, Benada O, Kofronova O, Petrickova K, Bobek J (2017) Secondary metabolites produced during the germination of Streptomyces coelicolor. Front Microbiol 8(2495):1–13. https://doi.org/10.3389/fmicb.2017.02495
Coze F, Gilard F, Tcherkez G, Virolle MJ, Guyonvarch A (2013) Carbon-flux distribution within Streptomyces coelicolor metabolism: a comparison between the actinorhodin-producing strain M145 and Its non-producing derivative M1146. PLoS ONE 8(12):e84151. https://doi.org/10.1371/journal.pone.0084151
Droste P, Miebach S, Niedenfuhr S, Wiechert W, Noh K (2011) Visualizing multi-omics data in metabolic networks with the software Omix—a case study. Biosystems 105(2):154–161. https://doi.org/10.1016/j.biosystems.2011.04.003
Esnault C, Dulermo T, Smirnov A, Askora A, David M, Deniset-Besseau A, Holland I-B, Virolle M-J (2017) Strong antibiotic production is correlated with highly active oxidative metabolism in Streptomyces coelicolor M145. Sci Rep 7:200. https://doi.org/10.1038/s41598-017-00259-9
Gombert AK, Moreira dos Santos M, Christensen B, Nielsen J (2001) Network identification and flux quantification in the central metabolism of Saccharomyces cerevisiae under different conditions of glucose repression. J Bacteriol 183(4):1441–1451. https://doi.org/10.1128/JB.183.4.1441-1451.2001
Gomez-Escribano JP, Bibb MJ (2011) Engineering Streptomyces coelicolor for heterologous expression of secondary metabolite gene clusters. Microb Biotechnol 4(2):207–215. https://doi.org/10.1111/j.1751-7915.2010.00219.x
Goodwin CR, Covington BC, Derewacz DK, McNees CR, Wikswo JP, McLean JA, Bachmann BO (2015) Structuring microbial metabolic responses to multiplexed stimuli via self-organizing metabolomics maps. Chem Biol 22(5):661–670. https://doi.org/10.1016/j.chembiol.2015.03.020
Gunnarson N, Bruheim P, Nielsen J (2004) Glucose metabolism in the antibiotic producing Actinomycete Nonomuria sp. ATCC 39727. Biotechnol Bioeng 88(5):652–663. https://doi.org/10.1002/bit.20279
Hawkey PM, Jones AM (2009) The changing epidemiology of resistance. J Antimicrob Chemother 64(Suppl 1):i3-10. https://doi.org/10.1093/jac/dkp256
Hwang KS, Kim HU, Charusanti P, Palsson BO, Lee SY (2014) Systems biology and biotechnology of Streptomyces species for the production of secondary metabolites. Biotechnol Adv 32(2):255–268. https://doi.org/10.1016/j.biotechadv.2013.10.008
Kieser T, Bibb MJ, Buttner MJ, Chater KF, Hopwood D (2000) Practical streptomyces genetics. John Innes Foundation, Norwich, UK
Kim SH, Kim BG (2016) NAD(+)-specific glutamate dehydrogenase (EC.1.4.1.2) in Streptomyces coelicolor; in vivo characterization and the implication for nutrient-dependent secondary metabolism. Appl Microbiol Biot 100(12):5527–5536. https://doi.org/10.1007/s00253-016-7433-8
Kumar K, Bruheim P (2021a) Large dependency of intracellular NAD and CoA pools on cultivation conditions in Saccharomyces cerevisiae. BMC Res Notes 14(372):1–5. https://doi.org/10.1186/s13104-021-05783-6
Kumar K, Bruheim P (2021b) A comparative study at bioprocess and metabolite levels of superhost strain Streptomyces coelicolor M1152 and its heterologously expressed chloramphenicol BGC derivative M1581. Biotech Bioeng. https://doi.org/10.1002/bit.27958
Kumar K, Venkatraman V, Bruheim P (2021) Adaptation of central metabolite pools to variations in growth rate and cultivation conditions in Saccharomyces cerevisiae. Microb Cell Fact. https://doi.org/10.1186/s12934-021-01557-8
Kvitvang HFN, Kristiansen KA, Bruheim P (2014) Assessment of capillary anion exchange ion chromatography tandem mass spectrometry for the quantitative profiling of the phosphometabolome and organic acids in biological extracts. J Chromatogr A 28(1370):70–79. https://doi.org/10.1016/j.chroma.2014.10.029
Lee MY, Kim HY, Lee S, Kim JG, Suh JW, Lee CH (2015) Metabolomics-based chemotaxonomic classification of Streptomyces spp and its correlation with antibacterial activity. J Microbiol Biotechn 25(8):1265–1274. https://doi.org/10.4014/jmb.1503.03005
Mahajan GB, Balachandran L (2012) Antibacterial agents from actinomycetes—a review. Front Biosci (elite Ed) 1(4):240–253. https://doi.org/10.2741/373
Morgenstern A, Paetz C, Behrend A, Spiteller D (2015) Divalent transition-metal-ion stress induces prodigiosin biosynthesis in Streptomyces coelicolor M145: formation of coeligiosins. Chem-Eur J 21(16):6027–6032. https://doi.org/10.1002/chem.201405733
Ngugen Q-T, Merlo ME, Medema MH, Jankevics A, Breitling R, Takano E (2012) Metabolomics methods for the synthetic biology of secondary metabolism. FEBS Lett 586:2177–2183. https://doi.org/10.1016/j.febslet.2012.02.008
Nieselt K, Battke F, Herbig A, Bruheim P, Wentzel A, Jakobsen OM, Sletta H, Alam MT, Merlo ME, Moore J, Omara WAM, Morrissey ER, Juarez-Hermosillo MA, Rodriguez-Garcia A, Nentwich M, Thomas L, Iqbal M, Legaie R, Gaze WH, Challis GL, Jansen RC, Dijkhuizen L, Rand DA, Wild DL, Bonin M, Reuther J, Wohlleben W, Smith MCM, Burroughs NJ, Martin JF, Hodgson DA, Takano E, Breitling R, Ellingsen TE, Wellington EMH (2010) The dynamic architecture of the metabolic switch in Streptomyces coelicolor. BMC Genomics 11(10):1–9. https://doi.org/10.1021/bi9722251
Nitta K, Del Carratore F, Breitling R, Takano E, Putri SP, Fukusaki E (2020) Multi-omics analysis of the effect of cAMP on actinorhodin production in Streptomyces coelicolor. Front Bioeng Biotech 8(595552):1–14. https://doi.org/10.3389/fbioe.2020.595552
Obanye AIC, Hobbs G, Gardner DCJ, Oliver SG (1996) Correlation between carbon flux through the pentose phosphate pathway and production of the antibiotic methylenomycin in Streptomyces coelicolor A3(2). Microbiol 142(1):133–137. https://doi.org/10.1099/13500872-142-1-133
Ranjani A, Dhanasekaran D, Gopinath PM (2016) An introduction to actinobacteria. In: Dhanasekaran D, Jiang Y (eds) Actinobacteria—basics and biotechnological applications. IntechOpen, London, pp 1–37. https://doi.org/10.5772/62329
Rioseras B, Lopez-Garcia MT, Yague P, Sanchez J, Manteca A (2014) Mycelium differentiation and development of Streptomyces coelicolor in lab-scale bioreactors: programmed cell death, differentiation, and lysis are closely linked to undecylprodigiosin and actinorhodin production. Bioresour Technol 151:191–198. https://doi.org/10.1016/j.biortech.2013.10.068
Rost LM, Brekke Thorfinnsdottir L, Kumar K, Fuchino K, Eide Langorgen I, Bartosova Z, Kristiansen KA, Bruheim P (2020a) Absolute quantification of the central carbon metabolome in eight commonly applied prokaryotic and eukaryotic model systems. Metabolites 10(2):74. https://doi.org/10.3390/metabo10020074
Rost LM, Shafaei A, Fuchino K, Bruheim P (2020b) Zwitterionic HILIC tandem mass spectrometry with isotope dilution for rapid, sensitive and robust quantification of pyridine nucleotides in biological extracts. J Chromatogr B Analyt Technol Biomed Life Sci 1144:122078. https://doi.org/10.1016/j.jchromb.2020.122078
Sauer U, Eikmanns BJ (2005) The PEP-pyruvate-oxaloacetate node as the switch point for carbon flux distribution in bacteria. Fems Microbiol Rev 29(4):765–794. https://doi.org/10.1016/j.femsre.2004.11.002
Sheng-wan Tsao BAMR, Xian-Guo HE, Chang C-J, Floss HG (1985) Identification of a red pigment from Streptomyces coelicolor A3 (2) as a mixture of prodigiosin derivatives. J Antibiotics 38(1):128–131. https://doi.org/10.7164/antibiotics.38.128
Stafsnes MH, Rost LM, Bruheim P (2018) Improved phosphometabolome profiling applying isotope dilution strategy and capillary ion chromatography-tandem mass spectrometry. J Chromatogr B Analyt Technol Biomed Life Sci 1083:278–283. https://doi.org/10.1016/j.jchromb.2018.02.004
Sulheim S, Kumelj T, van Dissel D, Salehzadeh-Yazdi A, Du C, van Wezel GP, Nieselt K, Almaas E, Wentzel A, Kerkhoven EJ (2020) Enzyme-constrained models and omics analysis of Streptomyces coelicolor reveal metabolic changes that enhance heterologous production. Iscience 23(9):101525. https://doi.org/10.1016/j.isci.2020.101525
Thomas L, Hodgson DA, Wentzel A, Nieselt K, Ellingsen TE, Moore J, Morrissey ER, Legaie R, Wohlleben W, Rodriguez-Garcia A, Martin JF, Burroughs NJ, Wellington EMH, Smith MCM, Consortium S (2012) Metabolic switches and adaptations deduced from the proteomes of Streptomyces coelicolor wild type and phoP mutant grown in batch culture. Mol Cell Proteomics 11(2):M111.013797. https://doi.org/10.1074/mcp.M111.013797
Traxler MF, Watrous JD, Alexandrov T, Dorrestein PC, Kolter R (2013) Interspecies interactions stimulate diversification of the Streptomyces coelicolor secreted metabolome. Mbio 4(4):e00459-e513. https://doi.org/10.1128/mBio.00459-13
Villas-Boas SG, Bruheim P (2007) Cold glycerol-saline: the promising quenching solution for accurate intracellular metabolite analysis of microbial cells. Anal Biochem 370(1):87–97. https://doi.org/10.1016/j.ab.2007.06.028
Wentzel A, Bruheim P, Overby A, Jakobsen OM, Sletta H, Omara WAM, Hodgson DA, Ellingsen TE (2012a) Optimized submerged batch fermentation strategy for systems scale studies of metabolic switching in Streptomyces coelicolor A3(2). BMC Syst Biol 6(59):1–16. https://doi.org/10.1186/1752-0509-6-59
Wentzel A, Sletta H, Stream C, Ellingsen TE, Bruheim P (2012b) Intracellular metabolite pool changes in response to nutrient depletion induced metabolic switching in Streptomyces coelicolor. Metabolites 2(1):178–194. https://doi.org/10.3390/metabo2010178
Xia J, Psychogios N, Young N, Wishart DS (2009) MetaboAnalyst: a web server for metabolomic data analysis and interpretation. Nucl Acids Res 37:W652-660. https://doi.org/10.1093/nar/gkp356
Xu Z, Wang YM, Chater KF, Ou HY, Xu HH, Deng ZX, Tao MF (2017) Large-scale transposition mutagenesis of Streptomyces coelicolor identifies hundreds of genes influencing antibiotic biosynthesis. Appl Environ Microb. https://doi.org/10.1128/AEM.02889-16
Yague P, Manteca A, Simon A, Diaz-Garcia ME, Sanchez J (2010) New method for monitoring programmed cell death and differentiation in submerged Streptomyces cultures. Appl Environ Microbiol 76(10):3401–3404. https://doi.org/10.1128/AEM.00120-10
Yague P, Rodriguez-Garcia A, Lopez-Garcia MT, Rioseras B, Martin JF, Sanchez J, Manteca A (2014) Transcriptomic analysis of liquid non-sporulating Streptomyces coelicolor cultures demonstrates the existence of a complex differentiation comparable to that occurring in solid sporulating cultures. PLoS One 9(1):e86296. https://doi.org/10.1371/journal.pone.0086296
Acknowledgements
The NTNU NV-faculty Mass Spectrometry facility is thanked for support during all MS analysis, and technical help from Kåre Andre Kristiansen and Amanda Holstad Singleton is highly appreciated. The INBioPharm project team and the Centre for Digital Life Norway are thanked for administrative support during the study (https://www.ntnu.edu/dln/centre-for-digital-life-norway).
Funding
Open access funding provided by NTNU Norwegian University of Science and Technology (incl St. Olavs Hospital - Trondheim University Hospital). This work was supported by the Centre for Digital Life Norway project INBioPharm (Research Council of Norway) (grant number 248885).
Author information
Authors and Affiliations
Contributions
Conceptualization, KK and PB; methodology, KK and PB; software, KK and PB; validation, KK and PB; formal analysis, KK and PB; investigation, KK and PB; resources, PB; data curation, KK and PB; writing—original draft preparation, KK and PB; writing—review and editing, KK and PB; visualization, KK and PB; supervision, KK and PB; project administration, PB; funding acquisition, PB.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests regarding the publication of this study.
Research involved with human or animals participants
The research does not involve human participants or animals. Both authors consent submission of this study.
Supplementary Information
Below is the link to the electronic supplementary material.
13205_2022_3146_MOESM1_ESM.pdf
Supplementary file1 Figure 1: The LC–MS/MS methods covering CCM, Figure S2: NADs optimization graphs, Figure S3: CoAs optimization graphs, Figure S4: PCA plot of S. coelicolor) M145 in N and P lim, Figure S5: Heat maps on intracellular metabolite pools, Figure S6: Heat-map of relative standard deviation (%), Figure S7: Growth and metabolites profiling of S. coelicolor M1146, Table S1: Energy charge ratio, Table S2: metabolite concentrations of M145 and M146 cultivations (PDF 987 KB)
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Kumar, K., Bruheim, P. Nutrient-depended metabolic switching during batch cultivation of Streptomyces coelicolor explored with absolute quantitative mass spectrometry-based metabolite profiling. 3 Biotech 12, 80 (2022). https://doi.org/10.1007/s13205-022-03146-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s13205-022-03146-x