Abstract
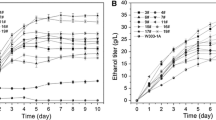
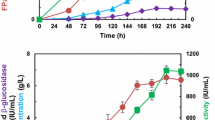
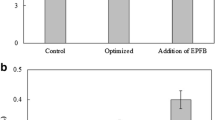
To improve the efficiency of enzymatic saccharification for lignocellulose, an efficient crude multi-enzyme was produced by Trichoderma reesei using corncob, a low cost inducer. Expression of cbh1, bgl1, egl1, xyn1 and positive regulator xyr1 induced by corncob increased significantly compared to that by cellulose. After 120 h induction by corncob, enzymatic activities on filter, CMC, β-glucose and xylan increased 86.5, 46.9, 120.9 and 291.2% compared to those induced by cellulose, and the concentration of secreted protein increased by 120.8%. FPase:β-glucosidase and FPase:xylanase values in crude multi-enzyme I (ECI, induced by corncob) were higher than that in crude multi-enzyme II (ECII, induced by cellulose). Under the same hydrolysis conditions, the volume dosage of ECI was only half of ECII, but ECI still showed a maximum of 12.5 and 33.4% higher than ECII in the total reducing sugar and glucose yield in lignocellulose hydrolysis. Corncob could be a candidate for low cost production of multi-enzyme for efficient lignocellulose degradation, and this work could guide the genetic modification of T. reesei to obtain efficient multi-enzyme for lignocellulose hydrolysis.




Similar content being viewed by others
References
Adav SS, Ravindran A, Sze SK (2012) Quantitative proteomic analysis of lignocellulolytic enzymes by Phanerochaete chrysosporium on different lignocellulosic biomass. J Proteomics 75:1493–1504
Alriksson B, Rose SH, van Zyl WH, Sjöde A, Nilvebrant N, Jönsson LJ (2009) Cellulase production from spent lignocellulose hydrolysates by recombinant Aspergillus niger. Appl Environ Microbiol 75:2366–2374
Bailey MJ, Biely P, Poutanen K (1992) Interlaboratory testing of methods for assay of xylanase activity. J Biotechnol 23:257–270
Bradford MM (1976) A rapid and sensitive method for quantification of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem 72:248–254
Ghose TK (1987) Measurement of cellulase activities. Pure Appl Chem 59:257–268
He J, Wu A, Chen D, Yu B, Mao X, Zheng P, Yu J, Tian G (2014) Cost-effective lignocellulolytic enzyme production by Trichoderma reesei on a cane molasses medium. Biotechnol Biofuels 7:43–52
Herold S, Bischof R, Metz B, Seiboth B, Kubicek CP (2013) Xylanase gene transcription in Trichoderma reesei is triggered by different inducers representing different hemicellulosic pentose polymers. Eukaryot Cell 12:390–398
Herpoël-Gimbert I, Margeot A, Dolla A, Jan G, Mollé D, Lignon S, Mathis H, Sigoillot JC, Monot F, Asther M (2008) Comparative secretome analyses of two Trichoderma reesei RUT-C30 and CL847 hypersecretory strains. Biotechnol Biofuels 1:18–30
Jurgens G, Survase S, Berezina O, Sklavounos E, Linnekoski J, Kurkijärvi A, Väkevä M, van Heiningen A, Granström T (2012) Butanol production from lignocellulosics. Biotechnol Lett 34:1415–1434
Karaffa L, Fekete E, Gamauf C, Szentirmai A, Kubicek CP, Seiboth B (2006) d-Galactose induces cellulase gene expression in Hypocrea jecorina at low growth rates. Microbiology 152:1507–1514
Laemmli UK (1970) Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 227:680–685
Li Q, Gao Y, Wang H, Li B, Liu C, Yu G, Mu X (2012) Comparison of different alkali-based pretreatments of corn stover for improving enzymatic saccharification. Bioresour Technol 125:193–199
Liao H, Li S, Wei Z, Shen Q, Xu Y (2014) Insights into high-efficiency lignocellulolytic enzyme production by Penicillium oxalicum GZ-2 induced by a complex substrate. Biotechnol Biofuels 7:162–178
Ling M, QinY Li N, Liang Z (2009) Binding of two transcriptional factors, Xyr1 and ACEI, in the promoter region of cellulase cbh1 gene. Biotechnol Lett 31:227–231
Liu D, Li J, Zhao S, Zhang R, Wang M, Miao Y, Shen Y, Shen Q (2013) Secretome diversity and quantitative analysis of cellulolytic Aspergillus fumigatus Z5 in the presence of different carbon sources. Biotechnol Biofuels 6:149–165
Mandels M, Reese ET (1960) Induction of cellulase in fungi by cellobiose. J Bacteriol 79:816–826
Merino ST, Cherry J (2007) Progress and challenges in enzyme development for biomass utilization. Adv Biochem Eng Biotechnol 108:95–120
Miller G (1959) Use of dinitrosalicylic acid reagent for determination of reducing sugar. Anal Chem 31:426–428
Ng IS, Tsai SW, Ju YM, Yu SM, Ho TD (2011) Dynamic synergistic effect on Trichoderma reesei cellulases by novel β-glucosidases from Taiwanese fungi. Bioresour Technol 102:6073–6081
Ouyang J, Yan M, Kong D, Xu L (2006) A complete protein pattern of cellulase and hemicellulase genes in the filamentous fungus Trichoderma reesei. Biotechnol J 1:266–1274
Payne CM, Knott BC, Mayes HB, Hansson H, Himmel ME, Sandgren M, Ståhlberg J, Beckham GT (2015) Fungal cellulases. Chem Rev 115:1308–1448
Peterson R, Nevalainen H (2012) Trichoderma reesei RUT-C30—30 years of strain improvement. Microbiology 158:58–68
Portnoy T, Margeot A, Seidl-Seiboth V, Le Crom S, Ben Chaabane F, Linke R, Seiboth B, Kubicek CP (2011) Differential regulation of the cellulase transcription factors XYR1, ACE2, and ACE1 in Trichoderma reesei strains producing high and low levels of cellulase. Eukaryot Cell 10:262–271
Pribowo A, Arantes V, Saddler JN (2012) The adsorption and enzyme activity profiles of specific Trichoderma reesei cellulase/xylanase components when hydrolyzing steam pretreated corn stover. Enzyme Microb Technol 50:195–203
Sluiter A (2008) Determination of structural carbohydrates and lignin in biomass. In: Laboratory Analytical Procedure. National renewable energy laboratory, Golden, Colardo
Sun Y, Cheng J (2002) Hydrolysis of lignocellulosic materials for ethanol production: a review. Bioresour Technol 83:1–11
Xia L, Shen X (2004) High-yield cellulase production by Trichoderma reesei ZU-02 on corncob residue. Bioresour Technol 91:259–262
Acknowledgements
The authors acknowledge the Key Project of Natural Science Foundation of Tianjin (16JCZDJC31800), and the National Natural Science Fund of China (31770625).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Ethical approval
This study was focused on analyzing the efficient crude multi-enzyme produced by Trichoderma reesei using corncob for the hydrolysis of lignocellulose. Every parts of the research did not involve human participants and other animals. Our manuscript complies with the Ethical Rules applicable for 3 Biotech.
Conflict of interest
The authors have declared that no conflicts of interest exist.
Rights and permissions
About this article
Cite this article
Jiang, F., Ma, L., Cai, R. et al. Efficient crude multi-enzyme produced by Trichoderma reesei using corncob for hydrolysis of lignocellulose. 3 Biotech 7, 339 (2017). https://doi.org/10.1007/s13205-017-0982-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s13205-017-0982-4