Abstract
Oilseed rape is one of the important oil plants. Pod shattering is one of the problems in oilseed rape production especially in regions with dry conditions. One of the important genes in Brassica pod opening is SHATTERPROOF1 (SHP1). Down-regulation of BnSHP1 expression by RNAi can increase resistance to pod shattering. A 470 bp of the BnSHP1 cDNA sequence constructed in an RNAi-silencing vector was transferred to oilseed rape cv. SLM046. Molecular analysis of T2 transgenic plants by RT-PCR and Real-time PCR showed that expression of the BnSHP alleles was highly decreased in comparison with control plants. Morphologically, transgenic plants were normal and produced seeds at greenhouse conditions. At ripening, stage pods failed to shatter, and a finger pressure was needed for pod opening.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Oilseed rape (Brassica napus L.) is the third most important oilseed crop in the world (Basalma 2008). Seeds have about 40–48 % oil with a high amount oleic acid and low linolenic acid suitable for frying applications and cooking. Dehiscence of pods causes significant yield loss (Raman et al. 2011). Ordinary yield losses are in the range of 10–25 % (Price et al. 1996). Seed losses have been reported as much as 50 % of the expected yield when adverse climatic conditions delayed harvesting (Macleod 1981; Child and Evans 1989). The process of pod shatter begins with degradation and separation of cell walls along a layer of few cells, termed the dehiscence zone (Meakin and Roberts 1990). Resistance to shattering is an important and necessary trait for oilseed rape improvement (Kadkol 2009). Attempts to solve this problem by interspecific hybridization using related species such as B. nigra, B. juncea and B. rapa have been faced with some difficulties as other undesirable traits will be integrated too (Prakash and Chopra 1990; Kadkol 2009). In Arabidopsis, which is in the same family of brassicaceae, several genes including the ALCATRAZ (ALC), INDEHISCENT (IND), SHATTERPROOF1 (SHP1) and SHATTERPROOF2 (SHP2) and FRUITFUL (FUL) have been shown to be involved in pod dehiscence (Raman et al. 2011). Genes for a number of hydrolytic enzymes, such as endopolygalacturonases, have also roles in dehiscence (Petersen et al. 1996). In Arabidopsis, SHP genes are specifically expressed in flowers with strong expression in the outer replum (Savidge et al. 1995; Flanagan et al. 1996). SHP gene also has mainly effect in the ripening of strawberries (Daminato et al. 2013). In B. napus, three BnSHP alleles (BnSHP1, BnSHP2a and BnSHP2b) have been identified (Tan et al. 2009). BnSHP1 and BnSHP2 show 80 % identity at nucleotide sequence. The expression of BnSHP2a and BnSHP2b (Two alleles of BnSHP gene which differ only in downstream sequences) are mainly in root, floral buds and pods, and most strongly in floral buds (Tan et al. 2009). It is suggested that less severe phenotype of indehiscence will be better, and the SHP, IND and ALC genes are ideal candidates for research and application in breeding new lines suitable for mechanized harvest (Liljegren et al. 2000; Tan et al. 2009). Recent advances about the role of MADS-box genes in dehiscence zone development have been reviewed (Ferrándiz and Fourquin 2014). In this study, we report the effect of the silencing cassette on expression of BnSHP alleles in transgenic oilseed rape plants using RNAi approach.
Materials and methods
Nucleic acid isolation
DNA was isolated from 100 mg leaf tissues using the procedure of Dellaporta et al. (1983). For RNA isolation, total RNA was extracted from floral buds using RNeasy Mini Kit (Qiagene Co.). The quantity and quality of RNA samples were checked using nano spectrophotometry and agarose gel electrophoresis. First-strand cDNA was synthesized using 2 µg of total RNA with iScript Select cDNA synthesis kit (Bio-rad Co.) in a 20-µl reaction using oligo-dT’s according to manufacturer’s instructions.
Construction of RNAi cassette
A 470-bp fragment of the BnSHP1 cDNA (Accession, AY036062) without MADS-box region was amplified by PCR using specific primers; F: 5′-ATACTAGTGGCGCGCCCCGTTAACCCTCCACTG-3′ and R: 5′-GCCTTAATTAAATTTAAATTTGAAGAGGAGGTTGGTC-3′ containing restriction enzyme digestion sites for Asc1, Aws1, Spe1 and Pac1 (underlined), for cloning the sense and antisense fragments in the above sites in pGSA1252 behind the CaMV35S promoter. The RNAi cassette was removed with Pst1 digestion and sub-cloned in the Pst1 site in pCAMBIA3301 to make pCAMRNAi.
Production of transgenic plants
Agrobacterium tumefaciens strain AGL0 containing the plasmid pCAMBIA3301 was used for transformation. Cotyledon explants of rapeseed cv. SLM046 were inoculated and co-cultivated with Agrobacterium inoculum on MS medium containing 1 mg/l 2,4-D and 4.5 mg/l BAP. After co-cultivation, cotyledonary explants were transferred to MS selection medium, containing 4.5 mg/l BAP and 4 mg/l phosphinothricine, 400 mg/l cefotaxime and 300 mg/l carbeniciline. The regenerated plants were analyzed by histochemical GUS assay according to the method reported by Jefferson et al. (1987).
The rooted transgenic plants were transferred into a mixture of peat and perlite (1:1, v/v), and they were grown in the greenhouse conditions. At five-leaf stage, the plants were incubated at 4 °C for 8 weeks to vernalize, and then they were moved to 25 °C for 16 h in light and 8 h in dark till maturity. The presence of transgene in T1 transgenic plants was confirmed by amplification of BnSHP sense, antisense cassette (F: 5′-AATACTAGTGGCGCGCCCCGTTAACCC TCCTACTG-3′, R: 5′-GCCTTAATTAAATTTAAATTTGAAGAGGAGGTTGGTC-3′, underlined part is a tail segment) and bar (F: 5′-ATCTCGGTGACGGGCAGGAC-3′, R: 5′-CGCAGGACCCGCAGGAGTG-3′) by PCR. A T2 transgenic line (cultured seeds from T1 line) was used for gene expression evaluation.
To study the expression of BnSHP alleles (SHP1, SHP2-a and SHP2-b), pod samples were taken from three transgenic plants of a T2 line and one non-transgenic plants (two replications from each plant) for RNA extraction. RT- and Real-time PCR was conducted with two specific primer pairs:
P14 F: 5′-TGAACTAGTCCATGGAGATCTTCTTCTCATGATCAGTCGCAGCATT-3′, P14 R: 5′-AGCTTAATTAAATTTAAATTTAAACAAGTTGAAGAGGAGGTTGG-3′ (producing a 151-bp fragment from three alleles, underlined part is a tail segment); P19 F: 5′-GAACAAGGCGCGAGATTGAATCC-3′ and P19 R: 5′-GATCATGAGAAGAAGACAGACCGG-3′ (producing a 94-bp fragment from SHP1). The GAPDH gene-specific primers: F: 5′-AGAGCCGCTTCCTTCAACATCATT-3′ and R: 5′-TGGGCACACGGAAGGACATACC-3′ (producing a 112-bp fragment) were used as a reference gene. The relative gene expression data were analyzed using the 2−∆∆CT method as described previously (Schmittgen and Livak 2008). The amount of BnSHP gene expression in transgenic and non-transgenic plants was calculated using the threshold data by 2−∆∆CT method (Pfaffl et al. 2002), and the data were statistically analyzed, and the graphs were drawn by Bio-Rad software package.
Results and discussion
Construction of RNAi cassette and transgenic plant production
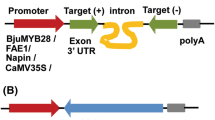
To construct the RNAi cassette, a 506-bp fragment (containing a 470-bp sequence downstream the MADS-box region of the BnSHP1 cDNA) was amplified and cloned in the sense and antisense direction in either side of a GUS intron in plasmid vector pGAS1252 (Fig. 1). The cassette was then sub-cloned in the Pst1 site in pCAMBIA3301, and the recombinant plasmid was designated as pCAMRNAi. The RNAi cassette was transferred into rapeseed, and putative transgenic plants were regenerated on medium containing phosphinotricin (Fig 2). Transformation efficiency of 3.7 % was obtained in SLM046 cultivar. Histochemical GUS assay in transgenic plants was done using Jefferson’s method (Jefferson et al. 1987), and blue leaf samples were observed in transgenic plants (Fig. 3). PCR analysis on T0 putative transgenic and wild-type plants showed the insertion of 501-bp antisense segment only in putative transgenic plants (Fig. 4). Seeds of putative T1 transgenic plants were sown in soil in pots, and growing plants at 5–6 leaf stage were subjected to 1.5 % basta herbicide. Three herbicide resistant lines were selected, and the insertion of 1,000-bp fragment of the GUS gene was confirmed by PCR (Fig. 5).
Schematic representation of the silencing cassette for the BnSHP gene. A 470-bp fragment of the BnSHP1 cDNA was cloned into pGSA1252 vector as sense and antisense segments, and the cassette was sub-cloned in pCAMBIA3301 transformation vector. CaMV35S: 35S promoter of a cauliflower mosaic virus, OCS3′: octopine synthase gene, GUS: β-glucuronidase gene, NOS A: terminator of the nopaline synthase gene
Development of transgenic B. napus. a Explants on shoot induction medium supplemented with 400 mg/l cefotaxime and 300 mg/l carbeniciline. b Non-transformed shoot on selection medium c, d Putative transgenic shoot on MS medium supplemented with 4.5 mg/l BAP, 4 mg/l phosphinothricine, 400 mg/l cefotaxime and 300 mg/l carbeniciline). e Flowering and fruiting (pod development) of transgenic plants in the greenhouse (Bar: 10 cm). f Ripening pods. Barsa, b, c, d and f: 1 cm
Detection of the antisense segment of BnSHPRNAi cassette in T0 putative transgenic B. napus plants. A 501-bp antisense fragment of the BnSHP gene was amplified using PCR. Lanes 1–4 and 6–7 produced clear bands of 501 bp for the transgene (higher band is related to the internal SHP gene); 5–8–9 blank; P positive control (plasmid vector of transformation); C control (non-transgenic plant); M 1 kb Plus DNA Ladder
Analysis of BnSHP1 gene in wild-type and transgenic plants
The effect of silencing cassette on gene expression was evaluated using T2 transgenic lines. The transgenic plants were grown in greenhouse conditions and showed normal vegetative and reproductive characteristics compared with control wild-type plants. The presence of the transgene was confirmed by amplification of a 400 bp of the bar gene in transgenic plants (Fig. 6).
Expression of the BnSHP gene was analyzed using RT-PCR. A sharp 100-bp band for BnSHP1 and BnSHP2 gene expression was observed in wild-type plants, while transgenic plants showed a weak band as was expected (Fig. 5). The primer P14 amplifies all three BnSHP alleles (SHP1, SHP2-a and SHP2-b), and P19 amplifies BnSHP1. A very slight difference in gene expression was observed in transgenic plants. Expression level for housekeeping GAPDH gene as control was the same in both transgenic and control plant, indicating that expression of BnSHP alleles was decreased in transgenic plants as judged with the GAPDH gene expression (Fig. 7).
Expression analysis of BnSHP gene in transgenic B. napus plants and wild type by RT-PCR: a amplification of a 105-bp band using the p14 specific primers, b amplification of a 94-bp band using the p19 specific primers and c amplification of a 100-bp band using the GAPDH specific primers. C non-transgenic control plant, T transgenic plant
Real-time PCR
To analyze the relative changes in gene expression, a quantitative Real-time PCR using two specific primers for BnSHP alleles and GAPDH gene was applied (Fig. 8). The efficiency of amplification using dilution series was 95 and 94 % for the GAPDH and the BnSHP genes. A higher CT value was observed for transgenic plants than non-transgenic plants, implying that the BnSHP gene expression has been decreased in transgenic plants. The data were statistically analyzed, and the related vertical graph showed the 97 % reduction in gene expression (Fig. 9).
Real-time analysis of the BnSHPgene and GAPDH expression in transgenic and non-transgenic B. napus plant using the P19 (a), P14 (b) BnSHP specific primers and GAPDH (c) primers. For BnSHP: delay in amplification in transgenic plant indicating the reduced level of the BnSHPRNAs and for GAPDH: no difference in amplification curves, indicating the same amount of RNA in the samples. High PCR specificity is shown by melting curve analysis
The genetic variation in shatter resistance has been reported in Brassica species, including B. rapa L., B. juncea L., B. hirta L. and within wild relatives of Brassica (Kadkol et al. 1985; Wang et al. 2007, Bagheri et al. 2012). However, to avoid windrowing of crops on a routine basis, the level of resistance is insufficient (Raman et al. 2011). Random impact test (RIT) on 229 accessions of B. napus for shatter resistance detected only two shatter resistant lines (Wen et al. 2008). Using introgression method, shatter resistance could be improved in B. juncea (Kadkol 2009; Raman et al. 2011). However, unwanted traits could be present too as reported by Summers et al. (2003). The DK142 line (resynthesized B. napus using B. oleracea alboglabra and B. rapa chinensis) showed higher shatter resistance in all locations, but significantly lower seed yield than commercial variety (Summers et al. 2003).
Indehiscent and harder siliques were obtained by constitutive MADSB expression in winter rape lines compared with wild-type winter rape plants, and precocious seed dispersal was prevented (Chandler et al. 2005). However, for unclear reason, the transgenic summer rape lines did not show a non-opening silique phenotype. This indicated that the tender rape cultivars differences could affect the co-ordination of signaling events involved in fruit dehiscence. A number of genes have been shown to involve in fruit dehiscence and seed shattering. Among the reported genes, SHP, IND and ALC genes have been suggested as ideal candidates for manipulation in breeding shatter resistant lines (Liljegren et al. 2000; Tan et al. 2009). In B. napus, three SHP alleles have been reported, and due to more probable redundancy of these alleles, resistance to shattering may need control of all these alleles simultaneously (Tan et al. 2009).
In Arabidopsis, double mutant of both SHATTERPROOF1 (SHP1) and SHATTERPROOF2 (SHP2) produced indehiscent pods (Liljegren et al. 2000). Pod shatter resistance was also observed in mutants of the INDEHISCENT gene (Liljegren et al. 2004; Wu et al. 2006), and the ALCATRAZ gene in Arabidopsis (Rajani and Sundaresan 2001). Sorefan et al. (2009) showed that the INDEHISCENT (IND) mutant produced indehiscent fruits by preventing differentiation of tissue in dehiscence zone into an abscission layer.
Over-expression of FRUITFUL (FUL), a repressor of SHP and IND, produced indehiscent siliques in Arabidopsis (Ferrandiz et al. 2000). Ectopic expression of FUL gene in B. juncea produced indehiscent fruits, but they could not be threshed in a combine harvester without seed damage (Ostergaard et al. 2006). In the present study, introduction of the BnSHP gene-silencing cassette into B. napus showed a drastic reduction in BnSHP expression. Morphologically, transgenic plants were normal and set seeds at greenhouse conditions. At ripening stage, pods failed to shatter. Further phenotypic evaluation of shatter resistance such as the cantilever test, pendulum test (Kadkol et al. 1984) and the RIT are needed to be done in produced transgenic plants.
References
Bagheri H, El-Soda M, Oorschot I, Hanhart C, Bonnema G, Jansen-van den Bosch T, Mank R, Keurentjes J, Meng L, Wu J, Koornneef M, Aarts M (2012) Genetic analysis of morphological traits in a new, versatile, rapid-cycling Brassica rapa recombinant inbred line population. Front Plant Sci 3:183. doi:10.3389/fpls.2012.00183
Basalma D (2008) The correlation and Path analysis of yield and yield components of different winter rapeseed (Brassica napus ssp. oleifera L.) cultivars. Res J Agric Biol Sci 4:120–125
Chandler J, Corbesier L, Spielmann P, Dettendorfer J, Stahl D, Apel K, Melzer S (2005) Modulating flowering time and prevention of pod shatter in oilseed rape. Mol Breed 15:87–94
Child RD, Evans DE (1989) Improvement of recoverable yields in oilseed rape (Brassica napus) with growth retardants. Asp Appl Biol 23:135–143
Daminato M, Guzzo F, Casadoro G (2013) A SHATTERPROOF-like gene controls ripening in non-climacteric strawberries, and auxin and abscissic acid antagonistically affect its expression. J Exp Bot. doi:10.1093/jxb/ert214
Dellaporta SL, Wood J, Hickes JB (1983) A plant DNA minipreparation: version II. Plant Mol Biol Rep 1:19–21
Ferrándiz C, Fourquin C (2014) Role of the FUL–SHP network in the evolution of fruit morphology and function. J Exp Bio. doi:10.1093/jxb/ert479
Ferrandiz C, Liljegran SJ, Yanofsky MF (2000) Negative regulation of the SHATTERPROOF genes by FRUITFULL during Arabidopsis development. Science 280:436–438
Flanagan CA, Hu Y, Ma H (1996) Specific expression of the AGL1 MADS-box gene suggests regulatory functions in Arabidopsis gynoecium and ovule development. J Plant 10:343–353
Jefferson RA, Kavanagh TA, Bevan MW (1987) GUS fusion. B-glucuronidase as a sensitive and versatile gene fusion marker in higher plants. J EMBO 6:3901–3907
Kadkol GP (2009) Brassica shatter-resistance research update. In: Proceedings of the 16th Australian research assembly on brassicas conference, Ballarat Victoria, pp 104–109
Kadkol GP, MacMillan RH, Burrow RP, Halloran GM (1984) Evaluation of Brassica genotypes for resistance to shatter. I. Development of a laboratory test. Euphytica 33:63–73
Kadkol GP, Halloran GM, MacMillan RH (1985) Evaluation of Brassica genotypes for resistance to shatter. II. Variation in siliqua strength within and between accessions. Euphytica 34:915–924
Liljegren SJ, Ditta GS, Eshed Y, Savidge B, Bowman JL, Yanofsky MF (2000) SHATTERPROOF MADS-box genes control seed dispersal in Arabidopsis. Nature 404:766–770
Liljegren SJ, Roeder AH, Kempin S, Gremski K, Ostergaard L, Guimil S, Yanofsky MF (2004) Control of Fruit Patterning in Arabidopsis by INDEHISCENT. Cell 116:843–853
Macleod J (1981) Harvesting in oilseed rape. Cambridge Agricultural Publishing, Cambridge, pp 107–120
Meakin PJ, Roberts JA (1990) Dehiscence of fruit in oilseed rape (Brassica napus L.), II. The role of cell wall degrading enzymes and ethylene. J Exp Bot 41:1003–1011
Ostergaard L, Kempin SA, Bies D, Klee HJ, Yanofsky MF (2006) Pod shatter-resistant Brassica fruit produced by ectopic expression of the FRUITFULL gene. J Plant Biotechnol 3:1–7
Petersen M, Sander L, Child R, Onckelen H, Ulvskov P, Boekhardt B (1996) Isolation and characterization of a pod dehiscence zone-specific polyglactronase from Brassica napus. Plant Mol Biol 31:517–527
Pfaffl WM, Horgan GW, Dempfle L (2002) Relative expression software tool for group-wise comparison and statistical analysis of relative expression results in Real-time PCR. Nucleic Acids Res 9:1–10
Prakash S, Chopra VL (1990) Reconstruction of allopolyploid Brassicas through non-homologous recombination introgression of resistance to pod shatter in Brassica napus. Genet Res 26:1–2
Price JS, Neale MA, Hobson RN, Bruce DM (1996) Seed losses in commercial harvesting of Oilseed Rape. Eng J Agric Res 80:343–350
Rajani S, Sundaresan V (2001) The Arabidopsis myc/bHLH gene ALCATRAZ enables cell separation in fruit dehiscence. Curr Biol 11:1914–1922
Raman R, Raman H, Kadkol GP, Coombes N, Taylor B, Luckett D (2011) Genome-wide association analyses of loci for shatter resistance in Brassicas. In: Proceedings of the 11th Australian research assembly on brassicas (ARAB) conference, WaggaWagga, NSW, pp 36–41
Savidge B, Rounsley SD, Yanofsky MF (1995) Temporal relationship between the transcription of two Arabidopsis MADS box genes and the floral organ identity genes. Plant Cell 7:721–733
Schmittgen TD, Livak KJ (2008) Analyzing Real-time PCR data by the comparative CT method. Nat Protoc 3:1101–1108
Sorefan K, Girin T, Liljegren SJ, Ljung K, Robles P, Galvan-Ampudia CS, Offringa R, Friml J, Yanofsky MF, Ostergaard L (2009) A regulated auxinminimumis required for seed dispersal in Arabidopsis. Nature 459:583–587
Summers JE, Bruce DM, Vancanneyt G, Redig P, Werner CP, Morgan C, Child RD (2003) Pod shatter resistance in the resynthesised Brassica napus line DK142. J Agric Sci 140:43–52
Tan X, Xia Z, Zhang L, Zhang Z, Guo Z, Qi C (2009) Cloning and sequence analysis of oilseed rape (Brassica napus) SHP2 gene. Bot Stud 50:403–412
Wang R, Ripley VL, Rakow G (2007) Pod shatter resistance evaluation in cultivars and breeding lines of B. napus, B. juncea and Sinapis alba. Plant Breed 126:588–595
Wen YC, Fu TD, Tu JX, Ma CZ, Shen JX, Zhang SF (2008) Screening and analysis of resistance to siliquae shattering in rape (Brassica napus L.). Acta Agronomic Sinica 34:163–166
Wu H, Mori A, Jiang X, Wang Y, Yang M (2006) The INDEHISCENT protein regulates unequal cell divisions in Arabidopsis fruit. Planta 224:971–979
Acknowledgments
This research was supported by Agricultural Research, Education and Extension Organization (AREEO) of Iran.
Conflict of interest
The authors certify that there is no conflict of interest with any financial organization regarding the material discussed in our manuscript.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
This article is published under license to BioMed Central Ltd.Open Access This article is distributed under the terms of the Creative Commons Attribution License which permits any use, distribution, and reproduction in any medium, provided the original author(s) and the source are credited.
About this article
Cite this article
Kord, H., Shakib, A.M., Daneshvar, M.H. et al. RNAi-mediated down-regulation of SHATTERPROOF gene in transgenic oilseed rape. 3 Biotech 5, 271–277 (2015). https://doi.org/10.1007/s13205-014-0226-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13205-014-0226-9