Abstract
Acacia longifolia is an aggressive invader in Mediterranean-type ecosystems severely impacting biodiversity and ecosystem functions. The species’ invasiveness has been linked to its ability to thrive in nutrient poor soils, high seed production, and quick establishment after fire. In this study, we identify and compare the bacterial endophytes of A. longifolia seeds collected from populations in the species’ native (Australia) and invasive (Portugal) ranges. For this, we characterised the morphology (length, width, and weight) of seeds from two sites in each range and isolated and cultivated bacteria from seeds. DNA fingerprinting and cluster analyses revealed slightly higher, and distinct, bacterial diversity associated with seeds collected from native range populations in comparison to those collected from invasive populations. Sequencing of the 16S rDNA gene identified 119 bacterial isolates from 15 genera, with Curtobacterium strains being common in both ranges. Several differences in bacterial genera were found among ranges and sites: Dermacoccus, Frigoribacterium, Kocuria, Pantoea and Phyllobacterium taxa were each unique to seeds from the native populations, while Brevundimonas, Microbacterium, Rhizobium and Sphingomonas taxa were only found in the invasive seeds. The genus Paraburkholderia occurred in all invasive-range seeds but was not isolated from the native-range. Bacillus and Paenibacillus co-occurred in seeds collected from all invaded sites, but the simultaneous presence of both taxa was not found in native-range seeds. We propose that the bacterial endophytes present in invasive-range seeds may be important players for the invasiveness of A. longifolia, due to their role as plant growth promoters, providing extra capabilities helping acacia expansion.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
The global movement of humans and goods has led to the introduction of many alien species beyond their native ranges, where some have become invasive (Jeschke et al. 2014). Invasive species are one of the main drivers of global change and impact the composition and functioning of ecosystems worldwide (Van der Putten et al. 2007; Essl et al. 2020). The ecology of invasive species has been the subject of many studies, yet invasiveness – the inherent ability of an alien species to colonise and spread in certain environments – are still not fully understood (but see Gioria et al. 2023). Knowing these factors is important to better design and implement management strategies that can be applied on a global scale (Pyšek et al. 2020).
Some of the most problematic woody plant invaders are from the genus Acacia (family Fabaceae), commonly known as wattles. The genus contains 1082 species, most of which are found in Australia (Binggeli 1996; Richardson et al. 2023) and many of which have been introduced outside Australia. Numerous Acacia species are considered as some of the world’s worst invasive species, especially in Mediterranean and temperate climate regions (Lorenzo et al. 2010; Richardson et al. 2011). For example, in Portugal approximately 20% of the total area covered by invasive species is invaded by wattles (Marchante et al. 2003). One of the most widespread wattles in the country is A. longifolia (Andrews) Willd., which was introduced at the end of the nineteenth century for coastal dune stabilisation, forestry, ornamental use, and to prevent sand erosion (Marchante 2011). This species threatens native plant communities and reduces native species cover and diversity (Marchante et al. 2003). Acacia longifolia is also a major invader in other parts of the world, notably in South Africa, and it is considered a “transformer” species (Richardson et al. 2000) that negatively impacts ecosystem services (Le Maitre et al. 2011), soil physicochemical properties and functions (Marchante et al. 2008), and that has high water and resource consumption (Morris et al. 2011). These changes can be partially evaluated considering carbon and nitrogen cycles and isotopic analyses can provide insights into several ecological processes ascribed to plant growth and physiology within seed formation, especially in relation to climatic conditions (Kelly et al. 2005; Gonzalvez et al. 2009; McCue et al. 2020).
The success of invasive wattles has partly been attributed to their ability to establish mutualisms with nitrogen-fixing bacteria, known as rhizobia (Novoa et al. 2023). Rhizobia are free-living soil bacteria capable of forming specialised structures, called root nodules, on most legumes. These bacteria fix atmospheric nitrogen in root nodules into forms that legumes can utilise, in turn receiving carbon-rich photosynthates from their host plants (Spaink 2000). Forming symbioses with compatible rhizobia can be an important factor underlying the invasiveness of legumes (Parker 2001; Parker et al. 2006). Acacia species are generally able to associate with a wide range of rhizobia, primarily from the genus Bradyrhizobium (Le Roux and Wandrag 2023). Despite this promiscuity, many invasive wattles have been co-introduced with their Bradyrhizobium strains from Australia into new environments (Rodríguez-Echeverría 2010; Crisóstomo et al. 2013; Ndlovu et al. 2013; Warrington et al. 2019; Le Roux and Wandrag 2023). Strains in other rhizobia genera, such as Rhizobium, have also been isolated from invasive wattles (Rodríguez-Echeverría et al. 2011; Birnbaum et al. 2016; Keet et al. 2017; Jesus et al. 2020).
Multiple and independent acacia-rhizobium co-introductions, coupled with the known symbiotic promiscuity of acacias, raises the question about the role of other seed microbiome taxa in the invasiveness of these plants. Seed microbiota is diverse, mostly comprising epiphytic or endophytic bacteria (Malfanova et al. 2013; Truyens et al. 2015) and fungi (Rodriguez et al. 2009). Epiphytes are microorganisms found on the seed’s surface and can be internalised within seed tissues and transferred vertically or horizontally to seedlings (Nelson 2018). Seed endophytes are microorganisms that are present in the inner seed tissues and are transferred vertically (Truyens et al. 2015; Nelson 2018). Vertical transmission allows the retention of certain microbiota over successive generations, including mutualists (Ewald 1987; Truyens et al. 2015). The direct and indirect benefits of seed endophytes for plant growth are well known. For example, direct effects are related to nutrient acquisition, including nitrogen, phosphorus, and iron, and phytohormone production (e.g. Khan et al. 2014; Goswami et al. 2016; Brígido et al. 2019). Indirect effects may include protection from pathogens through the production of antibiotics and lytic enzymes by seed endophytes (Ma et al. 2016; Santoyo et al. 2016; Liu et al. 2017).
Among the studies considering bacterial seed endophytes in a wide range of plant taxa, four different phyla seem to always be represented, with Pseudomonadota (formerly Proteobacteria) being dominant, followed by Actinomycetota (formerly Actinobacteria), Bacillota (formerly Firmicutes), and Bacteroidota (formerly Bacteroidetes). Among these, some genera are commonly represented, including Bacillus, Methylobacterium, Pseudomonas, Paenibacillus, Pantoea, Rhizobium, Sphingomonas, while others are less common, e.g., Acinetobacter, Micrococcus and Staphylococcus (Truyens et al. 2015; Nelson 2018; Simonin et al. 2022). Plant associated bacterial endophytic communities are thought to be more deterministic while fungal endophytes appear more randomly associated with plants (Powell et al. 2015). This may be due to their response to different edaphoclimatic conditions, since fungal endophytes are more often described to be associated with plant resistance to biotic and abiotic stresses compared to bacterial endophytes (Yan et al. 2019). The most common phylum of plant fungal endophytes is Ascomycota, including the genera Alternaria, Aspergillus, Cladosporium, Fusarium and Penicillium, but also Mucor from the phylum Mucoromycota (Shahzad et al. 2018).
Considering that seeds are the natural propagules responsible for dissemination and invasion, in the present study, we wanted to compare A. longifolia seeds from the native and invasive ranges. Specifically, we compared seed bacterial endophyte communities given their key physiological roles allowing a more successful early plant establishment and survival. We hypothesised that (1) invasive range seeds would harbour unique endophytic bacterial communities, characterised by taxa involved in nitrogen fixation and plant growth promotion and (2) bacterial endophytes would be more diverse in the native than in the invasive range, due to loss of endophyte diversity associated with the introduction events of the species into Portugal. Furthermore, seeds from the species’ native and invasive ranges were compared morphologically and isotopically, to check for differences between ranges.
2 Materials and Methods
2.1 Seeds collection and characterisation
Seeds of Acacia longifolia were collected at each of two sites from the species’ native (Australia) and invaded (Portugal) ranges. Australian seeds were acquired from the Indigo Nursery in Sydney, New South Wales, and were collected from Long Reef (LR) in November 2007 and Belrose (BE) in November 2010 (GPS coordinates not available). Portuguese seeds were collected in July 2020 from Vagos (VA; 40.52451, -8.67253) and Vila Nova de Milfontes (VN; 37.682060, -8.762254). Collected seeds were stored in paper bags at room temperature until further use. Fifty seeds from each site were randomly selected and morphologically characterised (length, width, and weight). Seed volumes were calculated using an oblate spheroid as a proxy for seed shape, using the following equation:
where a is the semi-major axis (i.e., seed length) and b is the semi-minor (i.e., seed width), being the spheroid made by the semi rotation of the shortest axis (i.e., seed width is equal to seed height).
2.2 Isotopic analysis
Isotopic analysis was conducted for Carbon (C) and Nitrogen (N) fractionation for A. longifolia seeds from the species’ native and invasive (Portugal) ranges. For these analyses, seeds were dried for 48 h at 60 °C, combined into composite samples (5 composites per site with 10 seeds each), and ground using a ball mill (Retsch, Haan, Germany). Stable isotope ratio analysis was performed at the Stable Isotopes Analysis Facility (SIIAF) at the Faculdade de Ciências from the University of Lisbon (Portugal). Carbon and nitrogen isotopic fractionation signatures were determined by continuous flow isotope mass spectrometry (CF-IRMS) (Preston and Owens 1983) on a Sercon Hydra 20–22 (Sercon, UK) stable isotope ratio mass spectrometer, coupled to a EuroEA (EuroVector, Italy) elemental analyser for online sample preparation by Dumas-combustion. The \(\delta\) N and \(\delta\) C values were determined as \(\delta N/C= \left[\frac{\left({R}_{sample}-{R}_{standard}\right)}{{R}_{standard}}\right].1000\), where R is the ratio between the heavier and lighter isotope of N or C. δ15NAir values are referred to Air and δ13CVPDB values are referred to PDB (Pee Dee Belemnite). The reference materials used were IAEA N1, IAEA N2, IAEA-C3 and IAEA-CH7 (Coleman and Meier-Augenstein 2014) and the laboratory standard used was Rice Flour. Uncertainty of the isotope ratio analysis, calculated using values from 6 to 9 replicates of laboratory standard interspersed among samples in every batch of analysis, was ≤ 0. 1‰. The major mass signals of C and N were used to calculate total C and N abundances, using Wheat Flour Standard OAS (Elemental Microanalysis, UK, with 1.47%N, 39.53%C) as elemental composition reference materials.
2.3 Seed surface-disinfection and pre-germination
To isolate endophytic bacteria, the seeds were surface-sterilised using a modified protocol from Vincent (1970). The seeds were immersed in 70% ethanol for 1 min, followed by immersion in commercial bleach for 6 min, followed by five thorough rinses with sterile distilled water at room temperature. After this, the seeds were soaked for 5 min in sterile distilled water at approximately 100 °C to induce germination. The efficiency of the protocol was tested by surface printing the seeds on Tryptic Soy Agar (TSA) plates, incubated at 28 °C for four days. Seeds that did not exhibit visible microbial growth (i.e., contamination) were germinated with a photoperiod of 16 h at 23 ± 2 °C until root emergence.
2.4 Isolation of seed endophytic bacteria
We tested for seed viability and only used germinated seeds for endophyte isolations, leading to the inclusion of 40 seeds (10 per collection site). In each pool, two seeds were macerated in 500 μL 0.85% sodium chloride. Serial dilutions (100—10–5) were used to inoculate Yeast Mannitol Agar (YMA) and TSA plates supplemented with 0.01% cycloheximide to avoid fungal growth. The plates were incubated for 5 – 10 days at 28 °C and checked daily for visible bacterial growth. Colonies with different morphologies were selected and sub-cultured until purification, which was determined by morphological and microscopic observations. Five isolation pools (of two seeds/collection site) were plated out as described above until no new morphologically distinct bacterial colonies were observed (assessed through macro- and microscopic observation).
2.5 Phenotypic and biochemical characterisation
To evaluate bacterial growth patterns we characterised colony size, pigmentation, form, margin, and elevation as described by Cappuccino and Sherman (1998). Gram-staining allowed us to distinguish between Gram-positive and Gram-negative rods and cocci. Other routine tests were performed such as potassium hydroxide (3% KOH, or string test, where Gram-negative bacteria cell walls are broken down releasing stringy or viscous material), catalase (bubble formation when a colony loop is mixed with 3% hydrogen peroxide), and oxidase tests (checks for cytochrome oxidase; in its presence the reduced colourless reagent becomes an oxidized coloured product). All tests were performed after 24—48 h of visible colony growth following streaking onto new growth media. Phenotypical and biochemically different isolates were used for further clustering analyses, together with the molecular fingerprinting.
2.6 DNA extraction and fingerprinting
Bacterial DNA extraction was performed using a guanidium thiocyanate, ethylenediamine tetraacetic acid (EDTA) and Sarkosyl (GES) protocol with some modifications (Pitcher et al. 1989). A loopful of one pure colony from each isolate was used and DNA was resuspended in 50 μL of 1 × Tris–EDTA (TE) buffer.
Genetic discrimination between the different isolates was performed using repetitive sequence-based polymerase chain reaction (rep-PCR) fingerprinting. DNA fragments were amplified in two separate PCR reactions using the primers csM13 (5’ – GAGGGTGGCGGTTCT – 3’) (Meyer et al. 1993) and (GTG)5 (5’- GTGGTGGTGGTGGTG -3’) (Versalovic et al. 1994). Each PCR reaction was performed in a final reaction volume of 12.5 μL, containing 1 × NZYTaq II Green Master Mix (NZYTech, Portugal), 0.5 µM of each primer and 1 µL (10 ng) of DNA template. All PCR reactions were performed in a BioRad T100 thermal cycler (BioRad, USA) with the following thermal cycle: initial denaturation at 95 °C, followed by 40 cycles of 95 °C for 1 min, 50 °C for 2 min, 72 °C for 2 min, and a final extension at 72 °C for 5 min. PCR products were separated by electrophoresis on a 1% (w/v) agarose gel using 0.5 × TBE Buffer, ran at 85 V for 5 h. The gel was stained with 0.5 ng. μL−1 ethidium bromide solution for 10 min and washed in water to remove excess staining. Gel images were obtained using an Alliance 4.7 UV Transilluminator (Uvitec, Cambridge) and processed with the associated software, Alliance version 15.15 (Uvitec, Cambridge).
2.7 Taxonomic identification of bacterial endophytes via 16S rRNA sequencing
For 16S rRNA gene sequencing, isolates representative of all clusters obtained by DNA fingerprinting were included. For these samples, the 16S rRNA gene was amplified using the primers PA(8f) (5’–AGAGTTTGATCCTGGCTCAG–3’) (Massol-Deya et al. 1995) and 907r (5-CCGTCAATTCMTTTRAGTTT-3) (Lane et al. 1985). The final PCR volume for each reaction was 25μL, containing 2μL of template DNA, 1U of NZYTaq II DNA polymerase (NZYTech, Portugal), 1 μM of each primer, 3 mM MgCl2 (NZYTech, Portugal), 0.2 mM dNTPs and 1 × PCR buffer (NZYTech, Portugal). The thermocycle for amplification was as follows: initial denaturation at 95 °C, followed by 40 cycles of 95 °C for 1 min, 55 °C for 2 min, 72 °C for 1 min, and a final extension at 72 °C for 5 min. Aliquots of 5 μL of each PCR product were ran on a 1% (w/v) agarose gel to confirm amplification success. All PCR products were purified and sequenced using the reverse primer (STAB VIDA, Portugal).
DNA sequences were edited and analysed using the software Geneious (v5.3 2010). All the sequences were compared with data available in GenBank using BLAST (16S ribosomal RNA database). Analysis of the dendrogram obtained with the fingerprinting clusters allowed us to assign identification based on the sequences isolates that fell in same the clustering. All DNA sequences were submitted to GenBank (accession numbers OR039116 to OR039161; Supplementary Table S1).
2.8 Statistical analyses
Seed morphological and ecophysiological data (i.e., seed length, width, weight, N and C isotopic signatures), were analysed using Kruskal–Wallis tests (α = 0.05). The link between seed volume and seed weight was evaluated through a Spearman correlation for each site separately (Fig. S1). Data analyses were performed using the package stats and rstatix in R (v.4.2.2) (R Core Team 2022).
The software BioNumerics (Applied Maths, Belgium) was used to construct the dendrograms, which compares and clusters DNA fingerprints. For better resolution and discrimination between isolates, fingerprinting profiles obtained from the amplification with both csM13 and (GTG)5 primers were used in the analysis. For clustering we used the unweighted pair-group method with arithmetic mean (UPGMA) with Pearson correlation coefficients. A cut-off level for each dendrogram was established based on the analysis of the reproducibility, in which a random sample of 10% of the total isolates was used. Diversity and evenness between the isolates at each location were assessed and compared as the Shannon-Weiner (H’), Simpson (D), and Pielou (J’) indices (Krebs 1989).
3 Results
3.1 Seed characterisation
Acacia longifolia seed origin significantly affected the width of the seed, considering the two sites in each range \(\left(\uprho <0.05\right)\) (Table 1). Native seeds displayed the largest differences in width, from 2.46 to 3.04 mm, while the seeds from the invaded range were more uniform (2.7–2.81 mm). For seed shape, seeds from Belrose (BE) were more elongated and narrower, than Long Reef (LR) seeds, which were shorter and wider. Seeds weight also revealed differences, with BE presenting the lighter seeds (16.8 mg) followed by VN (20.2 mg) (ρ < 0.05). LR and Vagos (VA) seeds were heavier (25.3–23.7 mg) however differences between these two sites were not statistically significant. Similar to seed width, the highest differences in seed weight were in the native range (16.8–25.3 mg vs. 20.2–23.7 for invasive range). For seed length, significant differences \(\left(\uprho <0.05\right)\) were found in seeds between sites in each range (native BE vs LR and invaded VA vs Vila Nova de Mil Fontes (VN). Seed volume was similar between sites in the invaded region (146 and 145 mm3 for VA and VN, respectively). LR seeds had the highest volume (172 mm3) while BE had the lowest volume (124 mm3). Seed volume significantly correlated with seed weight \(\left(\uprho <0.001\right)\) in all sites (Fig. S1), with higher values in the invaded region \((\uprho =0.88,\uprho <2.2\times {10}^{-16} {\text{ and }}\uprho =0.82,\uprho =2\times {10}^{-13})\) for VA and VN, respectively). Once again, the lowest correlation was found for BE followed by LR (\(\uprho =0.35,\uprho =0.013 {\text{ and }}\uprho =0.45,\uprho =0.00093\), respectively).
Seed viability was similar among seeds from different origins (native and invasive) and sites (results not shown).
3.2 Isotopic analysis
The analysis performed on A. longifolia seeds showed significant differences when comparing sites for most parameters of isotopic composition (\(\uprho <0.05\)), but these were unrelated to range (Table 2). Regarding δ13C, differences were statistically significant between all the sites. The seeds from native populations, LR and BE showed average values of -25.9 and -28.9 ‰, respectively, while in invaded seeds means were more uniform (-27.1 in VN to -28.2 ‰ in VA). Values of δ15N were close to 0 ‰ in all locations, but seeds from the invaded range had the highest variation (-0.21 and + 1.42 ‰ for VA and VN respectively, (\(\uprho <0.05\)). Australian seeds had mean values of -0.44 and + 0.80 ‰ for BE and LR, respectively (\(\uprho <0.05\)). Nitrogen values (%N) were different (\(\uprho <0.05\)) between sites within Australia (BE vs LR) and sites within Portugal (VA vs VN). BE seeds had the highest nitrogen content (%N; 4.64), followed by similar values for LR (4.35) and VA (4.31). Seeds from VN had the lowest %N (3.95). The carbon content (%C) did not differ between native and invaded ranges, with a low interval of values (40.5–41.6%), but seeds from invaded range had statistically significant differences. C/N ratios did not show high variability, ranging from 8.78 to 10.5. Differences were significant between sites (\(\uprho <0.05\)), except for the comparison between LR and VA.
3.3 Bacterial isolation and fingerprinting
We observed no microbial growth after imprinting seed coats of sterilised seeds onto growth media. In total, 119 bacterial isolates were obtained from our culture-dependent method. Sixty-five isolates were obtained from native A. longifolia seeds: 30 from BE and 35 from LR and 54 isolates were obtained from the invaded range: 28 from VA and 26 from VN (Table 3). Our UPGMA clustering analysis (Fig. 1) separated BE13AUS (later identified as Kocuria) from all the other accessions (cluster I). A second cluster contained eight accessions, all from the native range (LR34AUS—LR14AUS, mostly Curtobacterium and Dermacoccus), followed by another group consisting of the native range isolates BE15AUS (Kocuria) and BE16AUS (Frigobacterium) (cluster III). In another hierarchical level, two main clusters, IV-A and IV-B, are evident, one including most of the bacteria isolated from the invasive range and the other containing most of the isolates from the native range, respectively. However, both these clusters contained bacteria isolated from both ranges (Fig. 1). This is evident for the subclusters containing Curtobacterium and Bacillus (including Paenibacillus) isolates from both the native and invasive ranges. Some bacteria show identical profiles for both primers.
Dendrogram based on cluster analysis of fingerprinting PCR profiles using universal primers M13 and (GTG)5, using UPGMA clustering algorithm with Pearson correlation coefficient, of all the bacterial isolates obtained from A. longifolia seeds in native (Australia) and invasive (Portugal) range. Native: BE- Belrose; LR- Long Reef. Invaded: VA- Vagos, VN-Vila Nova de Milfontes are the sites from where bacteria were isolated from the seeds. The cut-off level determined was 84.23%, whereby isolates below this level were considered different. On the left side the labels with genus identification of several isolates are included. The main clusters are represented, from I to IV (A, B)
Cluster analysis was also performed for accessions obtained from each site separately (Fig. S2). In Belrose (BE) a major cluster (IV-B) contained the Bacillus group with some Kocuria accessions; in Long Reef (LR) two different clusters of Curtobacterium were present; in Vagos (VA) Rhizobium, Curtobacterium, Bacillus and Paenibacillus fell into four clusters, while in Vila Nova de Milfontes (VN) Curtobacterium, Bacillus and Paenibacillus clustered separately.
3.4 Bacterial identification based on 16S rRNA gene sequence analysis
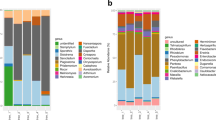
BLAST results on partially sequenced 16S rRNA genes identified 41 isolates to genus level with up to 98.9% pairwise identity (Table S1). For 14 isolates, identification was only possible to family level. These isolates (referred as Unclassified) corresponded to Australian seeds, 10 from Belrose (with a combined relative abundance of 33.33%) and four isolates from Long Reef (with a combined relative abundance of 11.43%) (Table 3, Table S1). Based on our BLAST results, it was possible to group bacterial genera as ‘core taxa’ (i.e., present at all seed collection sites) or ‘flexible taxa’ (only present in some seed collection sites; Simonin et al. 2022). Curtobacterium was the only core taxon while all other genera, including Bacillus, Brevundimonas, Dermacoccus, Frigoribacterium Kocuria, Microbacterium, Niallia, Paenibacillus, Pantoea, Paraburkholderia, Phyllobacterium, Rhizobium, Sphingomonas and Staphylococcus, were considered flexible (Fig. 2). Native and invasive seeds harboured the same richness of endophytic bacterial genera (10 in each) that differed compositionally (Table 3). Native and invasive range seeds had five genera in common, Bacillus, Curtobacterium, Niallia, Paenibacillus, and Staphylococcus. The genus Bacillus was dominant in native range seeds and Paenibacillus in invasive range seeds. The genera Dermacoccus, Frigoribacterium, Kocuria, Pantoea, and Phyllobacterium were only present in native range seeds, while Brevundimonas, Microbacterium, Paraburkholderia, Rhizobium and Sphingomonas were only present in invasive range seeds.
In native A. longifolia seeds, Bacillus was considered the most represented genus in Belrose with 12 isolates (corresponding to 40% of all sequenced isolates), while Curtobacterium was the most represented genus in Long Reef with 14 isolates (corresponding to 40% of all sequenced isolates). In Portuguese seeds, Rhizobium was the most represented genus in Vagos (corresponding to 28.57% of all sequenced isolates) and Paenibacillus was the most represented genus in Vila Nova de Milfontes (corresponding to 50% of all sequenced isolates) (Table 3, Fig. 3). An interesting result was the presence of Rhizobium only in seeds from Vagos (Portugal) and the absence of Bradyrhizobium strains in all locations. Isolates from Australia and Portugal were from three different phyla: Bacillota (formerly Firmicutes), Actinomycetota (formerly Actinobacteria) and Pseudomonadota (formerly Proteobacteria), with different representation at each site (Table 3). While the limited number of sites we sampled in each region precluded us from doing statistical comparisons of diversity between regions we do note a higher bacterial diversity in seeds from the native range and no dominance by a specific genus in both seed origins (Australia vs. Portugal; Table 4). The Shannon-Weiner diversity of native seeds was higher in BE (H’ = 1.21) in comparison to LR (H’ = 1.13), while non-native seeds from VA had higher diversity than VN (respectively H’ = 0.96 and H’ = 0.93). The Simpson diversity index was highest in the native-range seeds from LR (D’ = 0.98), while in invasive seeds, VN had the highest diversity (D’ = 0.92). Simpson diversity indexes were very similar both within native and invasive ranges. The Pielou evenness index was similar (J’ = 0.92—0.94) in all locations, from native and invasive range, meaning that the isolates found were equally distributed and that no dominant isolate was found.
Relative abundance (%) of the different bacterial genera found in the different seed origins [BE (n = 30);LR (n = 35);VA (n = 28);VN (n = 26)]. Unclassified: sequences which could not be classified to the genus level. (BE – Belrose, AUS; LR – Long Reef, AUS; VA – Vagos, PT; VN – Vila Nova de Milfontes, PT) AUS, Australia; PT, Portugal
4 Discussion
Understanding plant invasiveness is an area of immense research and for several invasive species differences in traits between native and invaded ranges have been identified. Yet, few studies to date have compared specific seed traits between ranges and how these may facilitate invasion, either considering morphological characteristics or the endophytic communities they harbour. Here, we compared seeds from native and invasive sites of the globally important invader, Acacia longifolia, based on morphological, isotopic, and bacterial endophyte community data.
The link between seed size and range (invasive vs. native) has been receiving a lot of research attention, however, no clear picture has emerged. For example, contradictory results have been found in a study on Cytisus scoparius (Scotch broom) and Ulex europaeus (European gorse), with C. scoparius (Scotch broom) producing significantly heavier seeds in its invasive range compared to in the native range, while U. europaeus produced seeds of similar size in both ranges (Buckley et al. 2003). Our study illustrates that, although seeds of A. longifolia do not differ morphologically between native and invasive populations, significant differences were found among sites within the same range (e.g. seed length within the native and invasive ranges) and between different ranges and sites (e.g. for seed width).
Isotopic analyses can provide information on the physiological responses of plants to local environmental conditions such as their water use efficiency (WUE) (Farquhar et al. 1982, 1989; Brugnoli and Farquhar 2000). For example, higher δ13C values (less negative) are generally linked to drier habitats (Cernusak et al. 2013). In the present study, one can infer that native LR seeds (δ13C = -25.9) collected from plants close to the sea, are more influenced by salt spray than plants from BE (δ13C = -28.9), from more inland areas. Similarly, in the invaded range VN seeds (δ13C = -27.1) originated from trees closer to the Atlantic Ocean than VA seeds (δ13C = -28.2).
The isotopic analysis of the δ15N revealed values close to 0 ‰ in seeds from all sites, suggesting that most of the nitrogen present in the seeds was derived from the atmosphere and not soil (i.e. soil nitrate or ammonium). While δ15N of seeds collected from the native and invasive range of A. longifolia were generally similar, we note differences among sites that suggest different soil conditions between BE (δ15N = -0.44) and LR (δ15N = 0.8), as well as between VA (δ15N = -0.21) and VN (δ15N = 1.42). Our results also suggest that for VN and LR, some NO3− and NH4+ were utilised by plants and translocated to seeds. The total % N obtained in our study are within the range of values reported previously (2.8–5.1) for other Acacia seedlings (Konate et al. 2016). Overall, in accordance with other seed studies (e.g. Alegria et al. 2020), our isotope results suggest a site-dependent response due to local climatic and geological characteristics. Therefore, isotope analyses have been used to trace the geographical origins of seeds from diverse species such as coffee (Rodrigues et al. 2011), Quercus species (Alegria et al. 2020) and argan (Elgadi et al. 2021). In conclusion, the seeds isotopic composition of A. longifolia reflects site characteristics and no distinctive pattern was found that could differentiate native from invasive seeds.
Our study also provided clear evidence that A. longifolia seeds harbour a large variety of endophytic bacteria. The 15 cultivable genera that we identified in these communities are likely a gross underestimate of the true endosymbiont diversity associated with the seeds of this species. Our finding that most cultivable endophytic bacteria belong to the phyla Pseudomonadota (formerly Proteobacteria) and Actinomycetota (formerly Actinobacteria), and some to Bacillota (formerly Firmicutes), is in accordance with a recent meta-analysis on the diversity of seed microbiomes (Simonin et al. 2022). This meta-analysis also identified the phylum Bacteroidota (formerly Bacteroidetes) as being commonly associated with seed microbiomes, which we did not find, either because in the present study we only characterised cultivable bacteria or because they are not present in A. longifolia seeds. Different studies have shown that seed vigour can be influenced by the microorganisms living inside and on the surface of seeds (e.g. Nelson 2018), by protecting seedlings from pathogens or harsh environmental conditions (e.g., Sahu et al. 2019; Xia et al. 2022). As outlined below, we propose that the seed endophytic bacteria identified in our study may be associated with the establishment and growth of young plants (see below), facilitating the invasiveness of A. longifolia.
Seed endophytes have been found to colonise the ovules of plants (Sahu et al. 2019), to facilitate vertical transfer from the mother plant through developmental stages (embryogenesis). Another possible pathway of transmission is horizontal transfer from various sources such as soil, water, air, and insects, which may explain why the diversity of bacterial seed endophytes in our study was site-specific (Afzal et al. 2019). These different pathways for transmission can explain the core (transmitted across multiple generations) and flexible (depending on habitat) bacteriomes (Shahzad et al. 2018; Simonin et al. 2022) that we found in our study. The dominance of Curtobacterium strains in all seeds (Figs. 2 and 3) suggests the existence of a possible vertical transmission pathway for this taxon and warrants further research. Curtobacterium is an Actinomycetota (formerly Actinobacteria) with a cosmopolitan distribution, predominantly found in soils. Although traditionally described as pathogens, the genus plays various ecological roles such as the decomposition of organic matter (Chase et al. 2016), attributed to its genomic potential for carbohydrates degradation, specifically structural polysaccharides (Chase et al. 2016). In the case of A. longifolia, we can hypothesise that the presence of Curtobacterium as a core endophytic phytopathogen may benefit its establishment, since it may negatively affect the native flora.
Considering the Bacillota phylum, Bacillus was a prominent genus isolated from A. longifolia seeds collected from most sites (except LR), while Paenibacillus was present in seeds from most sites (except BE). These two genera co-occurred in the seeds collected from invasive populations. Previous research has described these genera as including plant growth promoting (PGP) bacteria, capable of phytohormones production, but also involved in atmospheric nitrogen fixation and phosphate solubilisation (e.g., Goswami et al. 2016; Grady et al. 2016). These functions may contribute to the successful establishment of invasive plants, in terms of growth, by granting them a competitive advantage over native plants (Rout et al. 2013). Some Bacillus and Paenibacillus species also produce several metabolites that have antifungal activities and play an important role in pathogen suppression (Hu et al. 2017). These genera are also considered as drought resistant (Köberl et al 2013), which recently have been associated with the drought tolerance capacity of plants. Drought affects plant growth, and some studies have correlated this abiotic stress with the presence of drought-resistant bacteria in endophyte communities that are capable of producing PGP substances, as well as compounds that cope oxidative stress (Ullah et al. 2019; Verma et al. 2021; Ashry et al. 2022). In the context of ongoing climate change, these bacterial endophytes may be important to their host plants and may increase the performance strength of invasive species. Furthermore, these bacteria may also be involved in auxin and cytokinin production (de Melo Pereira et al. 2012; Xia et al. 2022). Apart from all the roles played by these phytohormones in plant development, higher auxin levels have also been directly linked to higher nodulation in legumes (Pii et al. 2007). The presence of Bacillus and Paenibacillus in A. longifolia seeds in both VA and VN sites suggests that plants in the invaded range benefit from these endosymbionts, the extent of which remains to be determined.
Kocuria and Rhizobium species have also been identified as capable of auxin production (Haidar et al. 2018). Pantoea is one of the most common bacterial genera associated with plant seeds (Nelson 2018) and has been associated with biocontrol activity against a wide range of phytopathogens (Walterson and Stavrinides 2015). The presence of Pantoea species in the seeds of A. longifolia suggest similar functions may be present and deserves further research. The role played by these bacteria in the seed is difficult to prove unequivocally, and the ecological characterisation of microorganisms is difficult, since they display dynamic alterations of life cycles depending on the environmental conditions they live in (Chase et al. 2016).
In this study, we also wanted to find out whether rhizobia are common endophytes of the seeds of A. longifolia, as has been shown for other legume seeds (Chimwamurombe et al. 2016; Aguilar et al. 2018; Laranjeira et al. 2022). Rhizobium was the only representative genus found in the seeds of A. longifolia, from one site in the species’ invasive range. Rhizobium strains fix atmospheric nitrogen and solubilise phosphorus, which are important attributes to legume performance. Bradyrhizobium, commonly associated with Australian acacias (Lafay and Burdon 2001; Rodríguez-Echeverría et al. 2007; Birnbaum et al. 2016), was not found in our study in either native or invasive seeds. It has been identified in A. longifolia nodules as the predominant bacterium from both invasive and native plants (Rodríguez-Echeverría et al. 2012; Crisóstomo et al. 2013; Birnbaum et al. 2016; Jesus et al. 2020). Previous phylogenetic studies have shown that Bradyrhizobium species have been co-introduced from Australia (native range) with various Acacia species into Brazil, California, New Zealand, Portugal and South Africa (Crisóstomo et al. 2013; Warrington et al. 2019; Le Roux and Wandrag 2023). Assessing whether Bradyrhizobium strains can act as Acacia seed endosymbionts provides exiting future research opportunities.
5 Conclusion
Although several studies have reported on the seed endophytic communities of plants, as far as we know, our study is the first to report on the bacterial endophyte communities of Acacia longifolia seeds. We show that seed endophytic communities from the species’ native and invasive ranges share common taxa, such as Curtobacterium while some ranges harboured unique associations (e.g., the presence of Paraburkholderia or the co-occurrence of Bacillus and Paenibacillus in seeds from the invasive range). We propose that the seed endophytic bacteria of A. longifolia might be playing an important role underlying its invasiveness. The bacterial taxa we identified are known to aid plant establishment and early development, especially in stressful environments. Future studies should include an evaluation of the ecological roles of seed endophytic bacteria and their contributions to plant performance.
Data availability
Further data and details are provided in the Supplementary Information file. The full dataset used in this study is available upon request to the corresponding Author.
References
Afzal I, Shinwari ZK, Sikandar S, Shahzad S (2019) Plant beneficial endophytic bacteria: Mechanisms, diversity, host range and genetic determinants. Microbiol Res 221:36–49. https://doi.org/10.1016/j.micres.2019.02.001
Aguilar A, Mora Y, Dávalos A, Girard L, Mora J, Peralta H (2018) Analysis of genome sequence and symbiotic ability of rhizobial strains isolated from seeds of common bean (Phaseolus vulgaris). BMC Genom 19:645. https://doi.org/10.1186/s12864-018-5023-0
Alegria C, Antunes C, Giovanetti M, Abreu M, Máguas C (2020) Acorn Isotopic Composition: A New Promising Tool for Authenticity Maps of Montado’s High-Value Food Products. Molecules 25(7):1535. https://doi.org/10.3390/molecules25071535
Ashry NM, Alaidaroos BA, Mohamed SA, Badr OAM, El-Saadony MT, Esmael A (2022) Utilization of drought-tolerant bacterial strains isolated from harsh soils as a plant growth-promoting rhizobacteria (PGPR). Saudi J Biol Sci 29(3):1760–1769. https://doi.org/10.1016/j.sjbs.2021.10.054
Binggeli P (1996) A taxonomic, biogeographical and ecological overview of invasive woody plants. J Veg Sci 7:121–124. https://doi.org/10.2307/3236424
Birnbaum C, Bissett A, Thrall PH, Leishman MR (2016) Nitrogen-fixing bacterial communities in invasive legume nodules and associated soils are similar across introduced and native range populations in Australia. J Biogeogr 43(8):1631–1644. https://doi.org/10.1111/jbi.12752
Brígido C, Menéndez E, Paço A, Glick BR, Belo A, Félix MR, Oliveira S, Carvalho M (2019) Mediterranean native leguminous plants: A reservoir of endophytic bacteria with potential to enhance chickpea growth under stress conditions. Microorganisms 7(10):392. https://doi.org/10.3390/microorganisms7100392
Brugnoli E, Farquhar GD (2000) Photosynthetic fractionation of carbon isotopes. In: Leegood RC, Sharkey TD, von Caemmerer S (eds) Photosynthesis: Physiology and Metabolism – Advances in Photosynthesis, vol 9. Kluwer Academic Publishers, The Netherlands, pp 399–434
Buckley YM, Downey P, Fowler SV, Hill R, Memmot J, Norambuena H, Pitcairn M, Shaw R, Sheppard AW, Winks C, Wittenberg R, Rees M (2003) Are Invasives Bigger? A Global Study of Seed Size Variation in Two Invasive Shrubs. Ecol 84:1434–1440. https://doi.org/10.1890/0012-9658(2003)084[1434:AIBAGS]2.0.CO;2
Cappuccino J, Sherman N (1998) Experiment 2 - Techniques for Isolation of Pure Cultures. In: Cappuccino J, Sherman N (eds) Microbiology - A Laboratory Manual, 5th edn. Benjamin-Cummings Pub Co, San Francisco, CA, pp 13–16
Cernusak LA, Ubierna N, Winter K, Holtum JAM, Marshall JD, Farquhar GD (2013) Environmental and physiological determinants of carbon isotope discrimination in terrestrial plants. New Phytol 200:950–965. https://doi.org/10.1111/nph.12423
Chase AB, Arevalo P, Polz MF, Berlemont R, Martiny JBH (2016) Evidence for Ecological Flexibility in the Cosmopolitan Genus Curtobacterium. Front Microbiol 7:1874. https://doi.org/10.3389/fmicb.2016.01874
Chimwamurombe PM, Grönemeyer JL, Reinhold-Hurek B (2016) Isolation and characterization of culturable seed-associated bacterial endophytes from gnotobiotically grown Marama bean seedlings. FEMS Microbiol Ecol 92(6):fiw083. https://doi.org/10.1093/femsec/fiw083
Coleman M, Meier-Augenstein W (2014) Ignoring IUPAC guidelines for measurement and reporting of stable isotope abundance values affects us all. Rapid Commun Mass Spectrom 28:1953–1955. https://doi.org/10.1002/rcm.6971
Crisóstomo JA, Rodríguez-Echeverría S, Freitas H (2013) Co-introduction of exotic rhizobia to the rhizosphere of the invasive legume Acacia saligna, an intercontinental study. Appl Soil Ecol 64:118–126. https://doi.org/10.1016/j.apsoil.2012.10.005
de Melo Pereira GV, Magalhães KT, Lorenzetii ER, Souza TP, Schwan R (2012) A Multiphasic Approach for the Identification of Endophytic Bacterial in Strawberry Fruit and their Potential for Plant Growth Promotion. Microb Ecol 63:405–417. https://doi.org/10.1007/s00248-011-9919-3
Elgadi S, Ouhammou A, Taous F, Zine H, Papazoglou EG, Elghali T, Amenzou N, El Allali H, Aitlhaj A, El Antari A (2021) Combination of Stable Isotopes and Fatty Acid Composition for Geographical Origin Discrimination of One Argan Oil Vintage. Foods 10:1274. https://doi.org/10.3390/foods10061274
Essl F, Lenzner B, Bacher S, Bailey S, Capinha C, Daehler C, Dullinger S, Genovesi P, Hui C, Hulme PE, Jeschke JM, Katsanevakis S, Kühn I, Leung B, Liebhold A, Liu C, MacIsaac HJ, Meyerson LA, Nuñez MA, Pauchard A, Pyšek P, Rabitsch W, Richardson D, Roy H, Ruiz GM, Russell JC, Sanders N, Sax D, Scalera R, Seebens H, Springborn M, Turbelin A, van Kleunen M, von Holle B, Winter M, Zenni R, Mattsson B, Roura-Pascual N (2020) Drivers of future alien species impacts: An expert-based assessment. Glob Change Biol 26:4880–4893. https://doi.org/10.1111/gcb.15199
Ewald PW (1987) Transmission Modes and Evolution of the Parasitism-Mutualism Continuum. Ann N Y Acad Sci 503(1):295–306. https://doi.org/10.1111/j.1749-6632.1987.tb40616.x
Farquhar GD, O’Leary MH, Berry JA (1982) On the relationship between carbon isotope discrimination and the intercellular carbon dioxide concentration in leaves. Aust J Plant Physiol 9:121–137. https://doi.org/10.1071/PP9820121
Farquhar GD, Ehleringer JR, Hubick KT (1989) Carbon isotope discrimination and photosynthesis. Annu Rev Plant Physiol Plant Mol Biol 40:503–537. https://doi.org/10.1146/annurev.pp.40.060189.002443
Geneious v5.3 (2010) Available online: http://www.geneious.com. Accessed Apr 2023
Gioria M, Hulme PE, Richardson DM, Pyšek P (2023) Why Are Invasive Plants Successful? Annu Rev Plant Biol 74:635–670. https://doi.org/10.1146/annurev-arplant-070522-071021
Gonzalvez A, Armenta S, De La Guardia M (2009) Trace-element composition and stable isotope ratio for discrimination of foods with Protected Designation of Origin. Trends Anal Chem 28:1295–1311. https://doi.org/10.1016/j.trac.2009.08.001
Goswami D, Thakker JN, Dhandhukia PC (2016) Portraying mechanics of plant growth promoting rhizobacteria (PGPR): A review. Cogent Food Agric 2:1. https://doi.org/10.1080/23311932.2015.1127500
Grady EN, MacDonald J, Liu L, Richman A, Yuan Z-C (2016) Current knowledge and perspectives of Paenibacillus: a review. Microb Cell Fact 15:203. https://doi.org/10.1186/s12934-016-0603-7
Haidar B, Ferdous M, Fatema B, Ferdous AS, Islam MR, Khan H (2018) Population diversity of bacterial endophytes from jute (Corchorus olitorius) and evaluation of their potential role as bioinoculants. Microbiol Res 208:43–53. https://doi.org/10.1016/j.micres.2018.01.008
Hu H-J, Chen Y-L, Wang Y-F, Tang Y-Y, Chen S-L, Yan S-Z (2017) Endophytic Bacillus cereus Effectively Controls Meloidogyne incognita on Tomato Plants Through Rapid Rhizosphere Occupation and Repellent Action. Plant Dis 101:448–455. https://doi.org/10.1094/PDIS-06-16-0871-RE
Jeschke JM, Bacher S, Blackburn TM, Dick JTA, Essl F, Evans T, Gaertner M, Hulme PE, Kühn I, Mrugała A, Pergl J, Pyšek P, Rabitsch W, Ricciardi A, Richardson DM, Sendek A, Vilà M, Winter M, Kumschick S (2014) Defining the impact of non-native species. Conserv Biol 28(5):1188–1194. https://doi.org/10.1111/cobi.12299
Jesus JG, Tenreiro R, Máguas C, Trindade H (2020) Acacia longifolia: A host of many guests even after fire. Diversity 12(6):250. https://doi.org/10.3390/D12060250
Keet JH, Ellis AG, Hui C, Le Roux JJ (2017) Legume-rhizobium symbiotic promiscuity and effectiveness do not affect plant invasiveness. Ann Bot 119(8):1319–1331. https://doi.org/10.1093/aob/mcx028
Kelly S, Heaton K, Hoogewerff J (2005) Tracing the geographical origin of food: the application of multi-element and multi-isotope analysis. Trends Food Sci Technol 16:555–567. https://doi.org/10.1016/j.tifs.2005.08.008
Khan AL, Waqas M, Kang SM, Al-Harrasi A, Hussain J, Al-Rawahi A, Al-Khiziri S, Ullah I, Ali L, Jung H, Lee IJ (2014) Bacterial endophyte Sphingomonas sp. LK11 produces gibberellins and IAA and promotes tomato plant growth. J Microbiol 52:689–695. https://doi.org/10.1007/s12275-014-4002-7
Köberl M, Schmidt R, Ramadan EM, Bauer R, Berg G (2013) The microbiome of medicinal plants: diversity and importance for plant growth, quality and health. Front Microbiol 4:400. https://doi.org/10.3389/fmicb.2013.00400
Konate NM, Dreyer E, Epron D (2016) Differences in carbon isotope discrimination and whole-plant transpiration efficiency among nine Australian and Sahelian Acacia species. Ann for Sci 73:995–1003. https://doi.org/10.1007/s13595-016-0589-7
Krebs CJ (1989) Chapter 10, Species diversity measures. In: Ecological methodology. Harper Collins, New York, pp 328–368
Lafay B, Burdon JJ (2001) Small-Subunit rRNA Genotyping of Rhizobia Nodulating Australian Acacia spp. Appl Environ Microbiol 67(1):396–402. https://doi.org/10.1128/AEM.67.1.396-402.2001
Lane DJ, Pace B, Olsen GJ, Stahl DA, Sogin ML, Pace NR (1985) Rapid determination of 16S ribosomal RNA sequences for phylogenetic analyses. Proc Natl Acad Sci 82(20):6955–6959. https://doi.org/10.1073/pnas.82.20.6955
Laranjeira SS, Alves IG, Marques G (2022) Chickpea (Cicer arietinum L.) Seeds as a Reservoir of Endophytic Plant Growth-Promoting Bacteria. Curr Microbiol 79:277. https://doi.org/10.1007/s00284-022-02942-1
Le Maitre DC, Gaertner M, Marchante E, Ens EJ, Holmes PM, Pauchard A, O’Farrell PJ, Rogers AM, Blanchard R, Blignaut J, Richardson DM (2011) Impacts of invasive Australian acacias: Implications for management and restoration. Divers Distrib 17(5):1015–1029. https://doi.org/10.1111/j.1472-4642.2011.00816.x
Le Roux JJ, Wandrag EM (2023) Co-invasion by Australian Acacia species and rhizobium mutualists. In: Richardson DM, Le Roux JJ, Marchante E (eds) Wattles: Aust Acacia species around world; CABI, UK, Invasive Species Series, pp 284–299. https://doi.org/10.1079/9781800622197.0018
Liu H, Carvalhais LC, Crawford M, Singh E, Dennis PG, Pieterse CMJ, Schenk PM (2017) Inner plant values: Diversity, colonization and benefits from endophytic bacteria. Front Microbiol 8(12):1–17. https://doi.org/10.3389/fmicb.2017.02552
Lorenzo P, González L, Reigosa MJ (2010) The genus Acacia as invader: the characteristic case of Acacia dealbata Link in Europe. Ann for Sci 67(1):101. https://doi.org/10.1051/forest/2009082
Ma Y, Rajkumar M, Zhang C, Freitas H (2016) Beneficial role of bacterial endophytes in heavy metal phytoremediation. J Environ Manage 174:14–25. https://doi.org/10.1016/j.jenvman.2016.02.047
Malfanova N, Lugtenberg BJJ, Berg G (2013) Bacterial endophytes: who and where, and what are they doing there? In: De Brujin FJ (ed) Molecular microbial ecology of the rhizosphere. John Wiley & Sons, Inc., Hoboken, New Jersey, pp 391–403. https://doi.org/10.1002/9781118297674.ch36
Marchante H, Marchante E, Freitas H (2003) Invasion of the Portuguese dune ecosystems by the exotic species Acacia longifolia (Andrews) Willd.: effects at the community level. In: Child L, Brock JH, Brundu G, Prach K, Pyšek K, Wade PM, Williamson M (eds) Plant Invasion: Ecological Threats and Management Solutions. Backhuys Publishers, Leiden, Netherlands, pp 75–85
Marchante E, Kjøller A, Struwe S, Freitas H (2008) Short- and long-term impacts of Acacia longifolia invasion on the belowground processes of a Mediterranean coastal dune ecosystem. Appl Soil Ecol 40(2):210–217. https://doi.org/10.1016/j.apsoil.2008.04.004
Marchante H (2011) Invasion of Portuguese dunes by Acacia longifolia: present status and perspectives for the future. Doctoral dissertation, Faculty of Sciences and Technology, University of Coimbra
Massol-Deya AA, Odelson DA, Hickey RF, Tiedje JM (1995) Bacterial community fingerprinting of amplified 16S and 16–23S ribosomal DNA gene sequences and restriction endonuclease analysis (ARDRA). In: Akkermans ADL, Van Elsas JD, De Brujin FJ (eds) Molecular Microbial Ecology Manual. Springer, Netherlands, pp 289–296
McCue M, Javal M, Clusella-Trullas S, Le Roux JJ, Jackson MC, Ellis AG, Richardson DM, Valentine AJ, Terblanche JS (2020) Using stable isotope analysis to answer fundamental questions in invasion ecology: Progress and prospects. Methods Ecol Evol 11:196–214. https://doi.org/10.1111/2041-210X.13327
Meyer W, Mitchell TG, Freedman EZ, Vilgalys R (1993) Hybridization probes for conventional DNA fingerprinting used as single primers in the polymerase chain reaction to distinguish strains of Cryptococcus neoformans. J Clin Microbiol 31(9):2274–2280. https://doi.org/10.1128/jcm.31.9.2274-2280.1993
Morris TL, Esler KJ, Barger NN, Jacobs SM, Cramer MD (2011) Ecophysiological traits associated with the competitive ability of invasive Australian acacias. Divers Distrib 17(5):898–910. https://doi.org/10.1111/j.1472-4642.2011.00802.x
Ndlovu J, Richardson DM, Wilson JRU, Le Roux JJ (2013) Co-invasion of South African ecosystems by an Australian legume and its rhizobial symbionts. J Biogeogr 40(7):1240–1251. https://doi.org/10.1111/jbi.12091
Nelson EB (2018) The seed microbiome: Origins, interactions, and impacts. Plant Soil 422(1–2):7–34. https://doi.org/10.1007/s11104-017-3289-7
Novoa A., Wilson JRU, Le Roux JJ, Gioria M, Pyšek P, Richardson DM (2023) The ‘WATTLES’ invasion syndrome. In: Richardson DM, Le Roux JJ, Marchante E (eds) Wattles: Aust Acacia species around world; CABI, UK, Invasive Species Series, pp 514–525. https://doi.org/10.1079/9781800622197.0031
Oren A, Garrity GM (2021) Valid publication of the names of forty-two phyla of prokaryotes. Int J Syst Evol Microbiol 71:005056. https://doi.org/10.1099/ijsem.0.005056
Parker MA (2001) Mutualism as a constraint on invasion success for legumes and rhizobia. Divers Distrib 7(3):125–136. https://doi.org/10.1046/j.1472-4642.2001.00103.x
Parker MA, Malek W, Parker IM (2006) Growth of an invasive legume is symbiont limited in newly occupied habitats. Divers Distrib 12(5):563–571. https://doi.org/10.1111/j.1366-9516.2006.00255.x
Pii Y, Crimi M, Cremonese G, Spena A, Pandolfini T (2007) Auxin and nitric oxide control indeterminate nodule formation. BMC Plant Biol 7:21. https://doi.org/10.1186/1471-2229-7-21
Pitcher DG, Saunders NA, Owen RJ (1989) Rapid extraction of bacterial genomic DNA with guanidium thiocyanate. Lett Appl Microbiol 8(4):151–156. https://doi.org/10.1111/j.1472-765X.1989.tb00262.x
Powell JR, Karunaratne S, Campbell CD, Yao H, Robinson L, Singh BK (2015) Deterministic processes vary during community assembly for ecologically dissimilar taxa. Nat Commun 6:8444. https://doi.org/10.1038/ncomms9444
Preston T, Owens NJP (1983) Interfacing an automatic elemental analyser with an isotope ratio mass spectrometer: the potential for fully automated total nitrogen and nitrogen-15 analysis. Analyst 108:971–977. https://doi.org/10.1039/AN9830800971
Pyšek P, Hulme PE, Simberloff D, Bacher S, Blackburn TM, Carlton JT, Dawson W, Essl F, Foxcroft LC, Genovesi P, Jeschke JM, Kühn I, Liebhold AM, Mandrak NE, Meyerson LA, Pauchard A, Pergl J, Roy HE, van Kleunen M, Wingfield MJ, Richardson DM (2020) Scientists’ warning on invasive alien species. Biol Rev 95(6):1511–1534. https://doi.org/10.1111/brv.12627
R Core Team (2022) R: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria. http://www.r-project.org/index.html. Accessed Apr 2023
Richardson DM, Pyšek P, Rejmánek M, Barbour MG, Dane Panetta F, West CJ (2000) Naturalization and invasion of alien plants: Concepts and definitions. Divers Distrib 6(2):93–107. https://doi.org/10.1046/j.1472-4642.2000.00083.x
Richardson DM, Carruthers J, Hui C, Impson FAC, Miller JT, Robertson MP, Rouget M, Le Roux JJ, Wilson JRU (2011) Human-mediated introductions of Australian acacias - a global experiment in biogeography. Divers Distrib 17(5):771–787. https://doi.org/10.1111/j.1472-4642.2011.00824.x
Richardson DM, Marchante E, Le Roux JJ (2023) Australian acacia species around the world: historical, social, evolutionary and ecological insights into one of the planet’s most widespread plant genera. In: Richardson DM, Le Roux JJ, Marchante E (eds) Wattles: Australian Acacia species around world; CABI, UK, Invasive Species Series, pp 1–26. https://doi.org/10.1079/9781800622197.0001
Rodrigues C, Brunner M, Steiman S, Bowen GJ, Nogueira JMF, Gautz L, Prohaska T, Máguas C (2011) Isotopes as Tracers of the Hawaiian Coffee-Producing Regions. J Agric Food Chem 59(18):10239–10246. https://doi.org/10.1021/jf200788p
Rodriguez RJ, White JF, Arnold AE, Redman RS (2009) Fungal endophytes: Diversity and functional roles. New Phytol 182(2):314–330. https://doi.org/10.1111/j.1469-8137.2009.02773.x
Rodríguez-Echeverría S (2010) Rhizobial hitchhikers from Down Under: Invasional meltdown in a plant-bacteria mutualism? J Biogeogr 37(8):1611–1622. https://doi.org/10.1111/j.1365-2699.2010.02284.x
Rodríguez-Echeverría S, Crisóstomo JA, Freitas H (2007) Genetic Diversity of Rhizobia Associated with Acacia longifolia in Two Stages of Invasion of Coastal Sand Dunes. Appl Environ Microbiol 73:5066–5070. https://doi.org/10.1128/AEM.00613-07
Rodríguez-Echeverría S, Le Roux JJ, Crisóstomo JA, Ndlovu J (2011) Jack-of-all-trades and master of many? How does associated rhizobial diversity influence the colonization success of Australian Acacia species? Divers Distrib 17(5):946–957. https://doi.org/10.1111/j.1472-4642.2011.00787.x
Rodríguez-Echeverría S, Fajardo S, Ruiz-Díez B, Mercedes F-P (2012) Differential effectiveness of novel and old legume–rhizobia mutualisms: Implications for invasion by exotic legumes. Oecologia 170:253–261. https://doi.org/10.1007/s00442-012-2299-7
Rout ME, Chrzanowski TH, Westlie TK, DeLuca TH, Callaway RM, Holben WE (2013) Bacterial endophytes enhance competition by invasive plants. Am J Bot 100:1726–1737. https://doi.org/10.3732/ajb.1200577
Sahu PK, Singh S, Gupta A, Singh UB, Brahmaprakash G, Saxena AK (2019) Antagonistic potential of bacterial endophytes and induction of systemic resistance against collar rot pathogen Sclerotium rolfsii in tomato. Biol Control 137:104014. https://doi.org/10.1016/j.biocontrol.2019.104014
Santoyo G, Moreno-Hagelsieb G, del Carmen O-M, Glick BR (2016) Plant growth-promoting bacterial endophytes. Microbiol Res 183:92–99. https://doi.org/10.1016/j.micres.2015.11.008
Shahzad R, Khan AL, Bilal S, Asaf S, Lee I-J (2018) What is there in seeds? Vertically transmitted endophytic resources for sustainable improvement in plant growth. Front Plant Sci 9:24. https://doi.org/10.3389/fpls.2018.00024
Simonin M, Briand M, Chesneau G, Rochefort A, Marais C, Sarniguet A, Barret M (2022) Seed microbiota revealed by a large-scale meta-analysis including 50 plant species. New Phytol 234(4):1448–1463. https://doi.org/10.1111/nph.18037
Spaink HP (2000) Root nodulation and infection factors produced by rhizobial bacteria. Annu Rev Microbiol 54:257–288. https://doi.org/10.1146/annurev.micro.54.1.257
Truyens S, Weyens N, Cuypers A, Vangronsveld J (2015) Bacterial seed endophytes: Genera, vertical transmission and interaction with plants. Environ Microbiol Rep 7(1):40–50. https://doi.org/10.1111/1758-2229.12181
Ullah A, Nisar M, Ali H, Hazrat A, Hayat K, Keerio AA, Ihsan M, Laiq M, Ullah S, Fahad S, Khan A, Khan AH, Akbar A, Yang X (2019) Drought tolerance improvement in plants: an endophytic bacterial approach. Appl Microbiol Biotechnol 103:7385–7397. https://doi.org/10.1007/s00253-019-10045-4
Van der Putten WH, Klironomos JN, Wardle DA (2007) Microbial ecology of biological invasions. ISME J 1:28–37. https://doi.org/10.1038/ismej.2007.9
Verma H, Kumar D, Kumar V, Kumari M, Singh SK, Sharma VJ, Droby S, Santoyo G, White JF, Kumar A (2021) The Potential Application of Endophytes in Management of Stress from Drought and Salinity in Crop Plants. Microorganisms 9:1729. https://doi.org/10.3390/microorganisms9081729
Versalovic J, Schneider M, De Brujin FJ, Lupski JR (1994) Genomic fingerprinting of bacteria using repetitive sequence-based PCR (rep-PCR). Methods Mol Cell Biol 5:25–40
Vincent JM (1970) A Manual for the Practical Study of Root-nodule Bacteria. Blackwell Scientific Publishers, Oxford, p 164
Walterson AM, Stavrinides J (2015) Pantoea: insights into a highly versatile and diverse genus within the Enterobacteriaceae. FEMS Microbiol Rev 39(6):968–984. https://doi.org/10.1093/femsre/fuv027
Warrington S, Ellis S, Novoa A, Wandrag EM, Hulme PE, Duncan RP, Valentine A, Le Roux JJ (2019) Cointroductions of Australian acacias and their rhizobial mutualists in the Southern Hemisphere. J Biogeogr 46:1519–1531. https://doi.org/10.1111/jbi.13602
Xia Y, Liu J, Chen C, Mo X, Tan Q, He Y, Wang Z, Yin J, Zhou G (2022) The Multifunctions and Future Prospects of Endophytes and Their Metabolites in Plant Disease Management. Microorganisms 10:1072. https://doi.org/10.3390/microorganisms10051072
Yan L, Zhu J, Zhao X, Shi J, Jiang C, Shao D (2019) Beneficial effects of endophytic fungi colonization on plants. Appl Microbiol Biotechnol 103:3327–3340. https://doi.org/10.1007/s00253-019-09713-2
Acknowledgements
This research was funded by Fundação para a Ciência e a Tecnologia (FCT, Portugal), FCT/MCTES through financial support to cE3c, Research Unit grant number UIDB/00329/2020 (https://doi.org/10.54499/UIDB/00329/2020). We thank Rodrigo Maia from SIIAF for the technical support on isotopic analysis.
Funding
Open access funding provided by FCT|FCCN (b-on).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflicts of interest
On behalf of all the authors, the corresponding author states that there are no conflicts of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Condessa, M., Jesus, J.G., Máguas, C. et al. The seeds of invasion: a comparison of endophytic seed bacteria of Acacia longifolia between its native and invasive ranges. Symbiosis 93, 29–42 (2024). https://doi.org/10.1007/s13199-024-00987-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13199-024-00987-3