Abstract
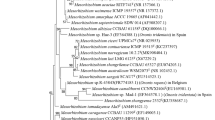
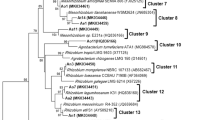
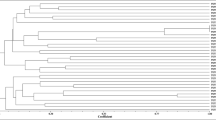
The genetic diversity of bacterial populations’ nodulating Retama sphaerocarpa grown in the soils of Maamora cork forest (Morocco) was examined. ERIC-PCR fingerprinting of 30 strains distributed them in 2 groups, of which a representative strain from each group was studied by multilocus sequence analysis of the 16S rRNA, atpD, and gyrB genes. The two representative strains RSM25 and RSM32, grouped with “Microvirga tunisiensis”. This is the first description of Retama nodule bacteria as members of the genus Microvirga. A nodC-based phylogeny confirmed that the two representative strains RSM25 and RSM32 are affiliated with symbiovar mediterranense. The 2 strains were capable of nodulating not only R. sphaerocarpa but also R. monosperma, R. dasycarpa and L. luteus, and unable to nodulate Phaseolus vulgaris, Vachellia gummifera, Cicer arietinum, Vigna unguiculata and Glycine max. The inoculation of R. sphaerocarpa with RSM25 or RSM32 produced a 1.22-, and 1.36-fold increase in the dry weight of the plants compared to those grown in the presence of 0.05% KNO3. The 2 strains used monosaccharides and disaccharides as the sole C source, but fructose, glucose, galactose, arabinose and starch were not used. They were unable to grow on glycine as a N source. Phosphate solubilization and siderophore production was not detected, but IAA or IAA-related compounds were produced. The alkaline pH of the sampling site soil in the Maamora forest where Retama grows could explain why Microvirga was the dominant species in the root nodules of the plants.



Similar content being viewed by others
References
Aafi A, Achhal EL Kadmiri A, Benabid A, Rouchdi M (2005) Richesse et diversité floristique de la subéraie de la Mamora (Maroc). Acta Bot Mal 30:127–138. https://doi.org/10.24310/abm.v30i0.7187
Ahnia H, Bourebaba Y, Durán D, Boulila F, Palacios JM, Rey L, Ruiz-Argüeso T, Boulila A, Imperial J (2018) Bradyrhizobium algeriense sp. nov., a novel species isolated from effective nodules of Retama sphaerocarpa from northeastern Algeria. Syst Appl Microbiol 41(4):333–339. https://doi.org/10.1016/j.syapm.2018.03.004
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ (1990) Basic local alignment search tool. J Mol Biol 215:403–410
Ardley JK, Parker MA, De Meyer SE, Trengove RD, O'Hara GW, Reeve WG, Yates RJ, Dilworth MJ, Willems A, Howieson JG (2012) Microvirga lupini sp. nov., Microvirga lotononidis sp. nov. and Microvirga zambiensis sp. nov. are alphaproteobacterial root nodule bacteria that specifically nodulate and fix nitrogen with geographically and taxonomically separate legume hosts. Int J Syst Evol Microbiol 62:2579–2588. https://doi.org/10.1099/ijs.0.035097-0
Armas C, Rodríguez-Echeverría S, Pugnaire FI (2011) A field test of the stress-gradient hypothesis along an aridity gradient. J Veg Sci 22:818–827
Belghazi B, Mounir F (2016) Analyse de la vulnérabilité au changement climatique du couvert forestier. Forêt de la Maâmora (Maroc). Edit. FAO, rapport technique, 124 p
Beligala DH, Michaels HJ, Devries M, Phuntumart V (2017) Multilocus sequence analysis of root nodule Bacteria associated with Lupinus spp. and Glycine max. Adv Microbiol 7(11):790–812. https://doi.org/10.4236/aim.2017.711063
Benabid A, Fennane M (1994) Connaissances sur la végétation du Maroc. Phytogéographie, phytosociologie et séries de végétation. Lazaroa 14:21–97
Boulila F, Depret G, Boulila A, Belhadi D, Benallaoua S, Laguerre G (2009) Retama species growing in different ecological-climatic areas of northeastern Algeria have a narrow range of rhizobia that form a novel phylogenetic clade within the Bradyrhizobium genus. Syst Appl Microbiol 32:245–255. https://doi.org/10.1016/j.syapm.2009.01.005
Brenner DJ, Mcwhorter AC, Leete Knutson JK, Steigerwalt AG (1982) Escherichia vulneris a new species of Enterobacteriaceae associated with human wounds. J Clin Microbiol 15:1133–1140
de Lajudie P, Andrews M, Ardley J, Eardly B, Jumas-Bilak E, Kuzmanović N, Lassalle F, Lindström K, Mhamdi R, Martínez-Romero E, Moulin L, Abdollah Mousavi S, Nesme X, Peix A, Puławska J, Steenkamp E, Stępkowski T, Tian CF, Vinuesa P, Wei G, Willems A, Zilli J, Young P (2019) Minimal standards for the description of new genera and species of rhizobia and agrobacteria. Int J Syst Evol Microbiol 69:1852–1863. https://doi.org/10.1099/ijsem.0.003426
El Boukhari EM, Brhadda N, Gmira N (2016) Contribution à l’étude de la régénération artificielle du chêne liège (Quercus suber L.) vis-à-vis du contenu minéral des feuilles et des paramètres physicochimiques des sols de la Maâmora (Maroc). Nature Technol 14:26–23
Estrada-de los Santos P, Palmer M, Steenkamp ET, Maluk M, Beukes C, Hirsch AM, James EK, Venter SN (2018) Trinickia dabaoshanensis sp. nov., a new name for a lost species. Arch Microbiol 201(9):1313–1316
Fan K, Weisenhorn P, Gilbert JA, Shi Y, Bai Y, Chu H (2018) Soil pH correlates with the co-occurrence and assemblage process of diazotrophic communities in rhizosphere and bulk soils of wheat fields. Soil Biol Biochem 121:185–192. https://doi.org/10.1016/j.soilbio.2018.03.017
Gaddas RR (1976) Aménagement et Amélioration des Parcours, Royaume du Maroc. Les Sols de la Mamora et du Rif Occidental, (zones caractéristiques d'intervention). Document de Travail
Graham PH, Parker CA (1964) Diagnostic features in the characterization of the root nodule bacteria of legumes. Plant Soil 20:338–396
Guerrouj K, Ruíz-Díez B, Chahboune R, Ramírez-Bahena MH, Abdelmoumen H, Quiñones M, Missbah El Idrissi M, Velázquez E, Fernández-Pascual M, Bedmar EJ, Peix A (2013) Definition of a novel symbiovar (sv. retamae) within Bradyrhizobium retamae sp. nov., nodulating Retama sphaerocarpa and Retama monosperma. Syst Appl Microbiol 36:218–223. https://doi.org/10.1016/j.syapm.2013.03.001
Han LL, Wang ET, Han TX, Liu J, Sui XH, Chen WF, Chen WX (2009) Unique community structure and biogeography of soybean rhizobia in the saline-alkaline soils of Xinjiang, China. Plant Soil 324:291–305. https://doi.org/10.1007/s11104-009-9956-6
Hannane FZ, Kacem M, Kaid-Harche M (2014) Retama-rhizobia symbiosis studies in some countries of the Mediterranean Basin. Ecol Medit 40:5–18
Kimura M (1980) A simple method for estimating evolutionary rates of base substitutions through comparative studies of nucleotide sequences. J Mol Evol 16(2):111–120. https://doi.org/10.1007/BF01731581
Kumar S, Stecher G, Tamura K (2016) MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol Biol Evol 33(7):1870–1874. https://doi.org/10.1093/molbev/msw054
Laguerre G, Nour SM, Macheret V, Sanjuan J, Drouin P, Amarger N (2001) Classification of rhizobia based on nodC and nifH gene analysis reveals a close phylogenetic relationship among Phaseolus vulgaris symbionts. Microbiol 147:981–993
Lakshmanan V, Shantharaj D, Li G, Seyfferth AM, Sherrier DJ, Bais HP (2015) A natural rice rhizospheric bacterium abates arsenic accumulation in rice (Oryza sativa L.). Planta 242:1037–1050. https://doi.org/10.1007/s00425-015-2340-2
Lamin H, Alami S, Bouhnik O, ElFaik S, Abdelmoumen H, Bedmar EJ, Missbah-El Idrissi M (2019) Nodulation of Retama monosperma by Ensifer aridi in an Abandonned Lead mine soils in eastern Morocco. Front Microbiol 10:1456. https://doi.org/10.3389/fmicb.2019.01456
Laouina A, Nafaa R, Wtfeh A (1997) Occupation des sols et dégradation des terres, le cas de la Mamora et de ses bordures. Méditerranée, vol. 86, n° 1–2 ("Impact anthropique en milieu méditerranéen"). p. 45–51. https://doi.org/10.3406/medit.1997.2989
León-González AJ, Navarro I, Acero N, Muñoz Mingarro D, Martín-Cordero C (2018) Genus Retama: a review on traditional uses, phytochemistry, and pharmacological activities. Phytochem Rev 17:701–731. https://doi.org/10.1007/s11101-018-9555-3
Lindström K, Lehtomäki S (1988) Metabolic properties, maximum growth temperature and phage sensitivity of Rhizobium sp. (Galega) compared with other fast-growing rhizobia. FEMS Microbiol Let 50:277–287
Martens M, Delaere M, Coopman R, De Vos P, Gillis M, Willems A (2007) Multilocus sequence analysis of Ensifer and related taxa. Int J Syst Evol Microbiol 57:489–503
Menna P, Barcellos FG, Hungria M (2009) Phylogeny and taxonomy of a diverse collection of Bradyrhizobium strains based on multilocus sequence analysis of the 16S rRNA gene, ITS region and glnII, recA, atpD and dnaK genes. Int J Syst Evol Microbiol 59:2934–2950
Missbah El-Idrissi M, Aujjar N, Belabed A, Dessaux Y, Filali-Maltouf A (1996) Characterization of rhizobia isolated from carob tree (Ceratonia siliqua). J Appl Bacteriol 80(2):165–173. https://doi.org/10.1111/j.1365-2672.1996.tb03205.x
Missbah El-Idrissi M, Lamin H, ElFaik S, Tortosa G, Peix A, Bedmar EJ, Abdelmoumen H (2019) Microvirga sp. symbiovar mediterranense nodulates Lupinus cosentinii grown wild in Morocco. J Appl Microbiol 128(4):1109–1118. https://doi.org/10.1111/jam.14526
Msaddak A, Rejili M, Durán D, Rey L, Imperial J, Palacios J, Ruiz-Argüeso T, Mars M (2017a) Members of Microvirga and Bradyrhizobium genera are native endosymbiotic bacteria nodulating Lupinus luteus in northern Tunisian soils. FEMS Microbiol Ecol 93. https://doi.org/10.1093/femsec/fix068
Msaddak A, Durán D, Rejili M, Mars M, Ruiz-Argüeso T, Imperial J, Palacios JM, Rey L (2017b) Diverse Bacteria affiliated with the genera Microvirga, Phyllobacterium, and Bradyrhizobium Nodulate Lupinus micranthus growing in soils of northern Tunisia. Appl Environ Microbiol 83(6). https://doi.org/10.1128/AEM.02820-16
Msaddak A, Rejili M, Durán D, Rey L, Palacios JM, Imperial J, Ruiz-Argüeso T, Mars M (2018) Definition of two new symbiovars, sv. lupini and sv. mediterranense, within the genera Bradyrhizobium and Phyllobacterium efficiently nodulating Lupinus micranthus in Tunisia. Syst Appl Microbiol 41(5):487–493
Msaddak A, Rejili M, Durán D, Mars M, Palacios JM, Ruiz-Argüeso T, Imperial J (2019) Microvirga tunisiensis sp. nov., a root nodule symbiotic bacterium isolated from Lupinus micranthus and L. luteus grown in northern Tunisia. Syst Appl Microbiol 42(6):126015. https://doi.org/10.1016/j.syapm.2019.126015
Patil A, Kale A, Ajane G, Sheikh R, Patil S (2017) Plant growth-promoting rhizobium: mechanisms and biotechnological prospective. In: Hansen A, Choudhary D, Agrawal P, Varma A (eds) Rhizobium Biology and Biotechnology. Soil Biology, vol 50. Springer, Cham. https://doi.org/10.1007/978-3-319-64982-5_7
Pikovskaya RI (1948) Mobilization of phosphorus in soil in connection with the vital activity of some microbial species. Mikrobiologiya 17:362–370
Prieto I, Kikvidze Z, Pugnaire FI (2010) Hydraulic lift: soil processes and transpiration in the Mediterranean leguminous shrub Retama sphaerocarpa (L.) Boiss. Plant Soil 329:447–456. https://doi.org/10.1007/s11104-009-0170-3
Rathi S, Tak N, Bissa G, Chouhan B, Ojha A, Adhikari D, Barik SK, Satyawada RR, Sprent JI, James EK, Gehlot HS (2018) Selection of Bradyrhizobium or Ensifer symbionts by the native Indian caesalpinioid legume Chamaecrista pumila depends on soil pH and other edaphic and climatic factors. FEMS Microbiol Ecol 94(11):fiy180. https://doi.org/10.1093/femsec/fiy180
Rejili M, Msaddak A, Filali I, Benabderrahim MA, Mars M, Marín M (2019) New chromosomal lineages within Microvirga and Bradyrhizobium genera nodulate Lupinus angustifolius growing on different Tunisian soils. FEMS Microbial Ecol 95(9):fiz118. https://doi.org/10.1093/femsec/fiz118
Rodríguez-Echeverría S, Pérez-Fernández MA (2003) Soil fertility and herb facilitation mediated by Retama sphaerocarpa. J Veg Sci 14(6):807–814. https://doi.org/10.1111/j.1654-1103.2003.tb02213.x
Rodríguez-Echeverría S, Pérez-Fernández MA (2005) Potential use of Iberian shrubby legumes and rhizobia inoculation in revegetation projects under acidic soil conditions. Appl Soil Ecol 29(2):203–208. https://doi.org/10.1016/j.apsoil.2004.11.004
Rogel MA, Ormeño-Orrillo E, Martinez Romero E (2011) Symbiovars in rhizobia reflect bacterial adaptation to legumes. Syst Appl Microbiol 34:96–104
Roughley RJ (1984) Effect of Soil Environmental Factor on Rhizobia. In R. Shiblies (ed.) Proceedings of World Soybean Research Conference III. August 12-17 1984. Westview Press London. 903-910
Ruiz-Díez B, Fajardo S, Puertas-Mejía MA, del Rosario de Felipe M, Fernández-Pascual M (2009) Stress tolerance, genetic analysis and symbiotic properties of root-nodulating bacteria isolated from Mediterranean leguminous shrubs in Central Spain. Arch Microbiol 191(1):35–46. https://doi.org/10.1007/s00203-008-0426-y
Ruiz-Díez B, Quiñones MA, Fajardo S, López MA, Higueras P, Fernández-Pascual M (2012) Mercury-resistant rhizobial bacteria isolated from nodules of leguminous plants growing in high Hg-contaminated soils. Appl Microbiol Biotechnol 96:543–554
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Vargas LK, Volpiano CG, Lisboa BB, Giongo A, Beneduzi A, Passaglia LMP (2017) Potential of Rhizobia as Plant Growth-Promoting Rhizobacteria. In A. Zaidi et al. (eds.), Microbes for Legume Improvement, Springer International Publishing AG 2017. https://doi.org/10.1007/978-3-319-59174-2_7
Versalovic J, Koeuth T, Lupski R (1991) Distribution of repetitive DNA sequences in eubacteria and application to fingerprinting of bacterial genomes. Nucleic Acids Res 19:6823–6831
Versalovic J, Schneider M, de Bruijn FJ, Lupski JR (1994) Genomic fingerprinting of bacteria using repetitive sequence-based Polymerase Chain Reaction. Methods Mol Cell Biol 5:25–40
Vincent JM (1970) A manual for the practical study of the root-nodule Bacteria. Blackwell Scientific Publications, Oxford
Vinuesa P, Silva C, Werner D, Martínez-Romero E (2005) Population genetics and phylogenetic inference in bacterial molecular systematics: the roles of migration and recombination in Bradyrhizobium species cohesion and delineation. Mol Phylogenet Evol 34:29–54. https://doi.org/10.1016/j.ympev.2004.08.020
Wang C, Zhou X, Guo D, Zhao JH, Yan L, Feng GZ, Gao Q, Yu H, Zhao LP (2019) Soil pH is the primary factor driving the distribution and function of microorganisms in farmland soils in northeastern China. Ann Microbiol 69:1461–1473. https://doi.org/10.1007/s13213-019-01529-9
Weisburg WG, Barns SM, Pelletier DA, Lane DJ (1991) 16S ribosomal DNA amplification for phylogenetic study. J Bacteriol 173(2):697–703
Yoon S-H, Ha S-M, Kwon S, Lim J, Kim Y, Seo H, Chun J (2017) Introducing EzBioCloud: a taxonomically united database of 16S rRNA gene sequences and whole-genome assemblies. Int J Syst Evol Microbiol 67(5):1613–1617
Zaim S, Bekkar AA, Belabid L (2017) Rhizobium as a Crop Enhancer and Biofertilizer for Increased Non-legume Production. A.P. Hansen et al. (eds.), Rhizobium Biology and Biotechnology, Soil Biology 50, Springer International Publishing AG 2017. ISBN978-3-319-64981-8. https://doi.org/10.1007/978-3-319-64982-5_4
Zerhari K, Aurag J, Khbaya B, Kharchaf D, Filali-Maltouf A (2000) Phenotypic characteristics of rhizobia isolates nodulating Acacia species in the arid and Saharan regions of Morocco. Lett Appl Microbiol 30(5):351–357. https://doi.org/10.1046/j.1472-765x.2000.00730.x
Zohary M (1959) A revision of the genus Retama ( Boiss). Bull Res Counc Isr 7(D):1–2
Acknowledgements
The authors want to thank the Hassan II Academy of Sciences for funding this research in the frame of the BIOMICS project. Financial support was also obtained from the ERDF-cofinanced project AGL2017–85676R from Ministerio de Economía, Industria y Competitividad (Spain).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare there is no conflict of interest.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
ESM 1
(DOCX 184 kb)
Rights and permissions
About this article
Cite this article
Mouad, L., Hanane, L., Omar, B. et al. Nodulation of Retama species by members of the genus Microvirga in Morocco. Symbiosis 82, 249–258 (2020). https://doi.org/10.1007/s13199-020-00725-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13199-020-00725-5