Abstract
Relevance
The proteasome is a crucial mechanism that regulates protein fate and eliminates misfolded proteins, playing a significant role in cellular processes. In the context of lung cancer, the proteasome’s regulatory function is closely associated with the disease’s pathophysiology, revealing multiple connections within the cell. Therefore, studying proteasome inhibitors as a means to identify potential pathways in carcinogenesis and metastatic progression is crucial in in-depth insight into its molecular mechanism and discovery of new therapeutic target to improve its therapy, and establishing effective biomarkers for patient stratification, predictive diagnosis, prognostic assessment, and personalized treatment for lung squamous carcinoma in the framework of predictive, preventive, and personalized medicine (PPPM; 3P medicine).
Methods
This study identified differentially expressed proteasome genes (DEPGs) in lung squamous carcinoma (LUSC) and developed a gene signature validated through Kaplan–Meier analysis and ROC curves. The study used WGCNA analysis to identify proteasome co-expression gene modules and their interactions with the immune system. NMF analysis delineated distinct LUSC subtypes based on proteasome gene expression patterns, while ssGSEA analysis quantified immune gene-set abundance and classified immune subtypes within LUSC samples. Furthermore, the study examined correlations between clinicopathological attributes, immune checkpoints, immune scores, immune cell composition, and mutation status across different risk score groups, NMF clusters, and immunity clusters.
Results
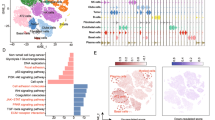
This study utilized DEPGs to develop an eleven-proteasome gene-signature prognostic model for LUSC, which divided samples into high-risk and low-risk groups with significant overall survival differences. NMF analysis identified six distinct LUSC clusters associated with overall survival. Additionally, ssGSEA analysis classified LUSC samples into four immune subtypes based on the abundance of immune cell infiltration with clinical relevance. A total of 145 DEGs were identified between high-risk and low-risk score groups, which had significant biological effects. Moreover, PSMD11 was found to promote LUSC progression by depending on the ubiquitin–proteasome system for degradation.
Conclusions
Ubiquitinated proteasome genes were effective in developing a prognostic model for LUSC patients. The study emphasized the critical role of proteasomes in LUSC processes, such as drug sensitivity, immune microenvironment, and mutation status. These data will contribute to the clinically relevant stratification of LUSC patients for personalized 3P medical approach. Further, we also recommend the application of the ubiquitinated proteasome system in multi-level diagnostics including multi-omics, liquid biopsy, prediction and targeted prevention of chronic inflammation and metastatic disease, and mitochondrial health-related biomarkers, for LUSC 3PM practice.









Similar content being viewed by others
Data availability
All the data used in this study were collected in this article and supplemental materials.
Code availability
All protein and gene accession codes can be available in the Swiss-Prot and Genbank databases.
Abbreviations
- ADRM1:
-
ADRM1 26S proteasome ubiquitin receptor
- AIC:
-
Akaike Information Criterion
- AUC:
-
Area under the curve
- BP:
-
Biological process
- CAR-T:
-
Chimeric antigen receptor T cell
- CC:
-
Cellular component
- CD274:
-
Programmed cell death 1 ligand 1
- CD276:
-
CD276 antigen
- CD80:
-
T-lymphocyte activation antigen CD80
- CD86:
-
T-lymphocyte activation antigen CD86
- CHST7:
-
Carbohydrate sulfotransferase 7
- CMTM6:
-
CKLF-like MARVEL transmembrane domain-containing protein 6
- CSN5:
-
COP9 signalosome complex subunit 5
- CTLA4:
-
Cytotoxic T-lymphocyte protein 4
- DEGs:
-
Differentially expressed genes
- DTP:
-
Developmental Therapeutics Program
- GAP43:
-
Gap junction alpha-1 protein
- GO:
-
Gene ontology
- GSEA:
-
Gene Set Enrichment Analysis
- IL:
-
Interleukin
- ITGA8:
-
Integrin alpha-8
- KEGG:
-
Kyoto Encyclopedia of Genes and Genomes
- LUSC:
-
Lung squamous carcinomas
- MAD:
-
Median absolute deviation
- MF:
-
Molecular function
- MHC:
-
Major Histocompatibility Complex
- NCBP1:
-
Nuclear cap-binding protein subunit 1
- NCI:
-
National Cancer Institute
- NMF:
-
Nonnegative matrix factorization method
- OPA1:
-
Dynamin-like 120 kDa protein
- PD-1:
-
Programmed cell death protein 1
- PDCD1:
-
Programmed cell death protein 1
- PDCD1LG2:
-
Programmed cell death 1 ligand 2
- PD-L1:
-
Programmed cell death 1 ligand 1
- PPI:
-
Protein-protein interaction
- PSMA1:
-
Proteasome subunit alpha type-1
- PSMA2:
-
Proteasome subunit alpha type-2
- PSMA3:
-
Proteasome subunit alpha type-3
- PSMA4:
-
Proteasome subunit alpha type-4
- PSMA5:
-
Proteasome subunit alpha type-5
- PSMA6:
-
Proteasome subunit alpha type-6
- PSMA7:
-
Proteasome subunit alpha type-7
- PSMB1:
-
Proteasome subunit beta type-1
- PSMB10:
-
Proteasome subunit beta type-10
- PSMB11:
-
Proteasome subunit beta type-11
- PSMB2:
-
Proteasome subunit beta type-2
- PSMB3:
-
Proteasome subunit beta type-3
- PSMB4:
-
Proteasome subunit beta type-4
- PSMB5:
-
Proteasome subunit beta type-5
- PSMB6:
-
Proteasome subunit beta type-6
- PSMB7:
-
Proteasome subunit beta type-7
- PSMB8:
-
Proteasome subunit beta type-8
- PSMB9:
-
Proteasome subunit beta type-9
- PSMC1:
-
26S proteasome regulatory subunit 4
- PSMC2:
-
26S proteasome regulatory subunit 7
- PSMC3:
-
26S proteasome regulatory subunit 6A
- PSMC4:
-
26S proteasome regulatory subunit 6B
- PSMC5:
-
26S proteasome regulatory subunit 8
- PSMC6:
-
26S proteasome regulatory subunit 10B
- PSMD1:
-
26S proteasome non-ATPase regulatory subunit 1
- PSMD11:
-
26S proteasome non-ATPase regulatory subunit 11
- PSMD12:
-
26S proteasome non-ATPase regulatory subunit 12
- PSMD13:
-
26S proteasome non-ATPase regulatory subunit 13
- PSMD14:
-
26S proteasome non-ATPase regulatory subunit 14
- PSMD2:
-
26S proteasome non-ATPase regulatory subunit 2
- PSMD3:
-
26S proteasome non-ATPase regulatory subunit 3
- PSMD4:
-
26S proteasome non-ATPase regulatory subunit 4
- PSMD6:
-
26S proteasome non-ATPase regulatory subunit 6
- PSMD7:
-
26S proteasome non-ATPase regulatory subunit 7
- PSMD8:
-
26S proteasome non-ATPase regulatory subunit 8
- PSMD9:
-
26S proteasome non-ATPase regulatory subunit 9
- PSME1:
-
Proteasome activator complex subunit 1
- PSME2:
-
Proteasome activator complex subunit 2
- PSME3:
-
Proteasome activator complex subunit 3
- PSME4:
-
Proteasome activator complex subunit 4
- ROC:
-
Receiver operating characteristic
- Rpn10:
-
26S proteasome non-ATPase regulatory subunit 4
- Rpn13:
-
26S proteasome regulatory subunit RPN13
- SEM1:
-
26S proteasome complex subunit SEM1
- ssGSEA:
-
Single-sample Gene Set Enrichment Analysis
- TCA:
-
Tricarboxylic acid
- TCGA:
-
The Cancer Genome Atlas
- TMB:
-
Tumor mutation burden
- TME:
-
Tumor microenvironment
- TNF:
-
Tumor necrosis factor
- UPS:
-
Ubiquitin-proteasome system
- VTCN1:
-
V-set domain-containing T cell activation inhibitor 1
- WGCNA:
-
Weighted gene co-expression network analysis
References
Socinski MA, Obasaju C, Gandara D, Hirsch FR, Bonomi P, Bunn PA, et al. Current and emergent therapy options for advanced squamous cell lung cancer. J Thorac Oncol. 2018;13(2):165–83. https://doi.org/10.1016/j.jtho.2017.11.111.
Comprehensive genomic characterization of squamous cell lung cancers. Nature. 2012;489(7417):519–25. https://doi.org/10.1038/nature11404.
Felip E, Altorki N, Zhou C, Csőszi T, Vynnychenko I, Goloborodko O, et al. Adjuvant atezolizumab after adjuvant chemotherapy in resected stage IB-IIIA non-small-cell lung cancer (IMpower010): a randomised, multicentre, open-label, phase 3 trial. Lancet (London, England). 2021;398(10308):1344–57. https://doi.org/10.1016/S0140-6736(21)02098-5.
Rousseau A, Bertolotti A. Regulation of proteasome assembly and activity in health and disease. Nat Rev Mol Cell Biol. 2018;19(11):697–712. https://doi.org/10.1038/s41580-018-0040-z.
Wang S, Wang T, Yang Q, Cheng S, Liu F, Yang G, et al. Proteasomal deubiquitylase activity enhances cell surface recycling of the epidermal growth factor receptor in non-small cell lung cancer. Cell Oncol (Dordr). 2022;45(5):951–65. https://doi.org/10.1007/s13402-022-00699-0.
Tong L, Shen S, Huang Q, Fu J, Wang T, Pan L, et al. Proteasome-dependent degradation of Smad7 is critical for lung cancer metastasis. Cell Death Differ. 2020;27(6):1795–806. https://doi.org/10.1038/s41418-019-0459-6.
Liu J, Guan D, Dong M, Yang J, Wei H, Liang Q, et al. UFMylation maintains tumour suppressor p53 stability by antagonizing its ubiquitination. Nat Cell Biol. 2020;22(9):1056–63. https://doi.org/10.1038/s41556-020-0559-z.
Collins GA, Goldberg AL. The logic of the 26S proteasome. Cell. 2017;169(5):792–806. https://doi.org/10.1016/j.cell.2017.04.023.
Kimura Y, Tanaka K. Regulatory mechanisms involved in the control of ubiquitin homeostasis. J Biochem. 2010;147(6):793–8. https://doi.org/10.1093/jb/mvq044.
Deng L, Meng T, Chen L, Wei W, Wang P. The role of ubiquitination in tumorigenesis and targeted drug discovery. Signal Transduct Target Ther. 2020;5(1):11. https://doi.org/10.1038/s41392-020-0107-0.
Lu M, Chen W, Zhuang W, Zhan X. Label-free quantitative identification of abnormally ubiquitinated proteins as useful biomarkers for human lung squamous cell carcinomas. EPMA J. 2020;11(1):73–94. https://doi.org/10.1007/s13167-019-00197-8.
Bhat SA, Vasi Z, Adhikari R, Gudur A, Ali A, Jiang L, et al. Ubiquitin proteasome system in immune regulation and therapeutics. Curr Opin Pharmacol. 2022;67:102310. https://doi.org/10.1016/j.coph.2022.102310.
Çetin G, Klafack S, Studencka-Turski M, Krüger E, Ebstein F. The ubiquitin-proteasome system in immune cells. Biomolecules. 2021;11(1). https://doi.org/10.3390/biom11010060.
Kammerl IE, Meiners S. Proteasome function shapes innate and adaptive immune responses. Am J Physiol Lung Cell Mol Physiol. 2016;311(2):L328–36. https://doi.org/10.1152/ajplung.00156.2016.
Wang P, Chen Y, Wang C. Beyond tumor mutation burden: tumor neoantigen burden as a biomarker for immunotherapy and other types of therapy. Front Oncol. 2021;11:672677. https://doi.org/10.3389/fonc.2021.672677.
Xuan DTM, Wu C-C, Kao T-J, Ta HDK, Anuraga G, Andriani V, et al. Prognostic and immune infiltration signatures of proteasome 26S subunit, non-ATPase (PSMD) family genes in breast cancer patients. Aging. 2021;13(22):24882–913. https://doi.org/10.18632/aging.203722.
Xu J, Brosseau J-P, Shi H. Targeted degradation of immune checkpoint proteins: emerging strategies for cancer immunotherapy. Oncogene. 2020;39(48):7106–13. https://doi.org/10.1038/s41388-020-01491-w.
Zengin T, Önal-Süzek T. Comprehensive profiling of genomic and transcriptomic differences between risk groups of lung adenocarcinoma and lung squamous cell carcinoma. J Personalized Med. 2021;11(2). https://doi.org/10.3390/jpm11020154.
Liu Z, Wang W, Zhou Y, Li L, Zhou W. PSMA1, a poor prognostic factor, promotes tumor growth in lung squamous cell carcinoma. Dis Markers. 2023;2023:5386635. https://doi.org/10.1155/2023/5386635.
Zhan X, Lu M, Yang L, Yang J, Zhan X, Zheng S, Guo Y, Li B, Wen S, Li J, Li N. Ubiquitination-mediated molecular pathway alterations in human lung squamous cell carcinomas identified by quantitative ubiquitinomics. Front Endocrinol (Lausanne). 2022;13:970843. https://doi.org/10.3389/fendo.2022.970843.
Amit S, Ben-Neriah Y. NF-kappaB activation in cancer: a challenge for ubiquitination- and proteasome-based therapeutic approach. Semin Cancer Biol. 2003;13(1):15–28.
Sun Y, Wang Y, Zhao J, Gu M, Giscombe R, Lefvert AK, et al. B7–H3 and B7–H4 expression in non-small-cell lung cancer. Lung Cancer. 2006;53(2):143–51.
Song X, Zhou Z, Li H, Xue Y, Lu X, Bahar I, et al. Pharmacologic suppression of B7–H4 glycosylation restores antitumor immunity in immune-cold breast cancers. Cancer Discov. 2020;10(12):1872–93. https://doi.org/10.1158/2159-8290.CD-20-0402.
Baravalle G, Park H, McSweeney M, Ohmura-Hoshino M, Matsuki Y, Ishido S, et al. Ubiquitination of CD86 is a key mechanism in regulating antigen presentation by dendritic cells. J Immunol. 2011;187(6):2966–73. https://doi.org/10.4049/jimmunol.1101643.
Dyck L, Mills KHG. Immune checkpoints and their inhibition in cancer and infectious diseases. Eur J Immunol. 2017;47(5):765–79. https://doi.org/10.1002/eji.201646875.
Corcoran K, Jabbour M, Bhagwandin C, Deymier MJ, Theisen DL, Lybarger L. Ubiquitin-mediated regulation of CD86 protein expression by the ubiquitin ligase membrane-associated RING-CH-1 (MARCH1). J Biol Chem. 2011;286(43):37168–80. https://doi.org/10.1074/jbc.M110.204040.
Mezzadra R, Sun C, Jae LT, Gomez-Eerland R, de Vries E, Wu W, et al. Identification of CMTM6 and CMTM4 as PD-L1 protein regulators. Nature. 2017;549(7670):106–10. https://doi.org/10.1038/nature23669.
Burr ML, Sparbier CE, Chan Y-C, Williamson JC, Woods K, Beavis PA, et al. CMTM6 maintains the expression of PD-L1 and regulates anti-tumour immunity. Nature. 2017;549(7670):101–5. https://doi.org/10.1038/nature23643.
Lim S-O, Li C-W, Xia W, Cha J-H, Chan L-C, Wu Y, et al. Deubiquitination and Stabilization of PD-L1 by CSN5. Cancer Cell. 2016;30(6):925–39. https://doi.org/10.1016/j.ccell.2016.10.010.
Ding L, Chen X, Zhang W, Dai X, Guo H, Pan X, et al. Canagliflozin primes antitumor immunity by triggering PD-L1 degradation in endocytic recycling. J Clin Investigation. 2023;133(1). https://doi.org/10.1172/JCI154754.
Debeljak Ž, Dundović S, Badovinac S, Mandić S, Samaržija M, Dmitrović B, et al. Serum carbohydrate sulfotransferase 7 in lung cancer and non-malignant pulmonary inflammations. Clin Chem Lab Med. 2018;56(8):1328–35. https://doi.org/10.1515/cclm-2017-1157.
Hung C-C, Lin C-H, Chang H, Wang C-Y, Lin S-H, Hsu P-C, et al. Astrocytic GAP43 induced by the TLR4/NF-κB/STAT3 axis attenuates astrogliosis-mediated microglial activation and neurotoxicity. J Neurosci. 2016;36(6):2027–43. https://doi.org/10.1523/JNEUROSCI.3457-15.2016.
Zhang F, Ying L, Jin J, Feng J, Chen K, Huang M, et al. GAP43, a novel metastasis promoter in non-small cell lung cancer. J Transl Med. 2018;16(1):310. https://doi.org/10.1186/s12967-018-1682-5.
Gebhardt A, Habjan M, Benda C, Meiler A, Haas DA, Hein MY, et al. mRNA export through an additional cap-binding complex consisting of NCBP1 and NCBP3. Nat Commun. 2015;6:8192. https://doi.org/10.1038/ncomms9192.
Zhang H, Wang A, Tan Y, Wang S, Ma Q, Chen X, et al. NCBP1 promotes the development of lung adenocarcinoma through up-regulation of CUL4B. J Cell Mol Med. 2019;23(10):6965–77. https://doi.org/10.1111/jcmm.14581.
Li X, Zhu G, Li Y, Huang H, Chen C, Wu D, et al. LINC01798/miR-17-5p axis regulates ITGA8 and causes changes in tumor microenvironment and stemness in lung adenocarcinoma. Front Immunol. 2023;14:1096818. https://doi.org/10.3389/fimmu.2023.1096818.
Wang Y, Li Y, Jiang X, Gu Y, Zheng H, Wang X, et al. OPA1 supports mitochondrial dynamics and immune evasion to CD8+ T cell in lung adenocarcinoma. Peer J. 2022;10:e14543. https://doi.org/10.7717/peerj.14543.
Abaza Y, Kantarjian HM, Faderl S, Jabbour E, Jain N, Thomas D, et al. (2018) Hyper-CVAD plus nelarabine in newly diagnosed adult T-cell acute lymphoblastic leukemia and T-lymphoblastic lymphoma. Am J Hematol. 2018;93(1):91–9. https://doi.org/10.1002/ajh.24947.
Oo ZY, Proctor M, Stevenson AJ, Nazareth D, Fernando M, Daignault SM, et al. Combined use of subclinical hydroxyurea and CHK1 inhibitor effectively controls melanoma and lung cancer progression, with reduced normal tissue toxicity compared to gemcitabine. Mol Oncol. 2019;13(7):1503–18. https://doi.org/10.1002/1878-0261.12497.
Tacconi EM, Badie S, De Gregoriis G, Reisländer T, Lai X, Porru M, et al. Chlorambucil targets BRCA1/2-deficient tumours and counteracts PARP inhibitor resistance. EMBO Mol Med. 2019;11(7):e9982. https://doi.org/10.15252/emmm.201809982.
Heng WS, Cheah S-C. Chelerythrine chloride downregulates β-catenin and inhibits stem cell properties of non-small cell lung carcinoma. Molecules. 2020; 25(1). https://doi.org/10.3390/molecules25010224.
Chaikovsky AC, Li C, Jeng EE, Loebell S, Lee MC, Murray CW, et al. The AMBRA1 E3 ligase adaptor regulates the stability of cyclin D. Nature. 2021;592(7856):794–8. https://doi.org/10.1038/s41586-021-03474-7.
Yang Y-C, Zhao C-J, Jin Z-F, Zheng J, Ma L-T. Targeted therapy based on ubiquitin-specific proteases, signalling pathways and E3 ligases in non-small-cell lung cancer. Front Oncol. 2023;13:1120828. https://doi.org/10.3389/fonc.2023.1120828.
Sun Q, Zhang J, Li X, Yang G, Cheng S, Guo D, et al. The ubiquitin-specific protease 8 antagonizes melatonin-induced endocytic degradation of MT1 receptor to promote lung adenocarcinoma growth. J Adv Res. 2022; 41. https://doi.org/10.1016/j.jare.2022.01.015.
Zhan Z, Xie X, Cao H, Zhou X, Zhang XD, Fan H, et al. Autophagy facilitates TLR4- and TLR3-triggered migration and invasion of lung cancer cells through the promotion of TRAF6 ubiquitination. Autophagy. 2014;10(2):257–68. https://doi.org/10.4161/auto.27162.
Kikuchi N, Soga T, Nomura M, Sato T, Sakamoto Y, Tanaka R, et al. Comparison of the ischemic and non-ischemic lung cancer metabolome reveals hyper activity of the TCA cycle and autophagy. Biochem Biophys Res Commun. 2020;530(1):285–91. https://doi.org/10.1016/j.bbrc.2020.07.082.
Lieu EL, Nguyen T, Rhyne S, Kim J. Amino acids in cancer. Exp Mol Med. 2020;52(1):15–30. https://doi.org/10.1038/s12276-020-0375-3.
Xu Y, Pan J, Lin Y, Wu Y, Chen Y, Li H. Ceramide synthase 1 inhibits brain metastasis of non-small cell lung cancer by interacting with USP14 and downregulating the PI3K/AKT/mTOR signaling pathway. Cancers. 2023;15(7). https://doi.org/10.3390/cancers15071994.
Blijlevens M, Komor MA, Sciarrillo R, Smit EF, Fijneman RJA, van Beusechem VW. Silencing core spliceosome SM gene expression induces a cytotoxic splicing switch in the proteasome subunit beta 3 mRNA in non-small cell lung cancer cells. Int J Mol Sci. 2020;21(12). https://doi.org/10.3390/ijms21124192.
Rouette A, Trofimov A, Haberl D, Boucher G, Lavallée V-P, D’Angelo G, et al. Expression of immunoproteasome genes is regulated by cell-intrinsic and -extrinsic factors in human cancers. Sci Rep. 2016;6:34019. https://doi.org/10.1038/srep34019.
Lu Z, Song Q, Yang J, Zhao X, Zhang X, Yang P, et al. Comparative proteomic analysis of anti-cancer mechanism by periplocin treatment in lung cancer cells. Cell Physiol Biochem. 2014;33(3):859–68. https://doi.org/10.1159/000358658.
Yim J-H, Yun HS, Lee S-J, Baek J-H, Lee C-W, Song J-Y, et al. Radiosensitizing effect of PSMC5, a 19S proteasome ATPase, in H460 lung cancer cells. Biochem Biophys Res Commun. 2016; 469(1). https://doi.org/10.1016/j.bbrc.2015.11.077.
Tanimoto A, Matsumoto S, Takeuchi S, Arai S, Fukuda K, Nishiyama A, et al. Proteasome inhibition overcomes ALK-TKI resistance in ALK-rearranged/TP53-mutant NSCLC via noxa expression. Clin Cancer Res. 2021;27(5):1410–20. https://doi.org/10.1158/1078-0432.CCR-20-2853.
Crigna AT, Samec M, Koklesova L, Liskova A, Giordano FA, Kubatka P, et al. Cell-free nucleic acid patterns in disease prediction and monitoring-hype or hope? EPMA J. 2020;11(4):603–27. https://doi.org/10.1007/s13167-020-00226-x.
Hu X, Huang W, Sun Z, Ye H, Man K, Wang Q, et al. Predictive factors, preventive implications, and personalized surgical strategies for bone metastasis from lung cancer: population-based approach with a comprehensive cancer center-based study. EPMA J. 2022;13(1):57–75. https://doi.org/10.1007/s13167-022-00270-9.
Qian S, Golubnitschaja O, Zhan X. Chronic inflammation: key player and biomarker-set to predict and prevent cancer development and progression based on individualized patient profiles. EPMA J. 2019;10(4):365–81. https://doi.org/10.1007/s13167-019-00194-x.
Zhang G, Wang Z, Song P, Zhan X. DNA and histone modifications as potent diagnostic and therapeutic targets to advance non-small cell lung cancer management from the perspective of 3P medicine. EPMA J. 2022;13(4):649–69. https://doi.org/10.1007/s13167-022-00300-6.
Koklesova L, Mazurakova A, Samec M, Kudela E, Biringer K, Kubatka P, et al. Mitochondrial health quality control: measurements and interpretation in the framework of predictive, preventive, and personalized medicine. EPMA J. 2022;13(2):177–93. https://doi.org/10.1007/s13167-022-00281-6.
Golubnitschaja O. What is the routine mitochondrial health check-up good for? A holistic approach in the framework of 3P medicine. In book: Predictive, preventive, and personalised medicine: from bench to bedside. Springer 2023. https://doi.org/10.1007/978-3-031-34884-6_3.
Sharma R, Rakshit B. Global burden of cancers attributable to tobacco smoking, 1990–2019: an ecological study. EPMA J. 2023;14(1):167–82. https://doi.org/10.1007/s13167-022-00308-y.
Acknowledgements
The authors acknowledge The Cancer Genome Atlas (TCGA) project organizers as well as all study participants to provide the publicly available TCGA RNA-seq data and clinical data.
Funding
This work was supported by the Shandong Provincial Taishan Scholar Engineering Project Special Funds (NO.tstp20221143 to X.Z.), the Shandong Provincial Natural Science Foundation (ZR2021MH156 to X.Z.; ZR2022QH112 to N.L.), the Shandong First Medical University Talent Introduction Funds (to X.Z.), and China National Nature Scientific Funds (82203592 to N.L.).
Author information
Authors and Affiliations
Contributions
J.Y. analyzed the data and wrote the manuscript. S.Y.O. edited and critically revised the manuscript. J.W., Z.L., X.F. and Z.Y. participated in partial data analysis. S.Z. carried out partial experiments. X.Z. and N.L. conceived the concept, designed the manuscript, coordinated and critically revised the manuscript, and was responsible for its financial support and the corresponding works. All authors approved the final manuscript.
Corresponding authors
Ethics declarations
Ethical approval
Not applicable.
Consent to participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Yang, J., Ouedraogo, S.Y., Wang, J. et al. Clinically relevant stratification of lung squamous carcinoma patients based on ubiquitinated proteasome genes for 3P medical approach. EPMA Journal 15, 67–97 (2024). https://doi.org/10.1007/s13167-024-00352-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13167-024-00352-w
Keywords
- Lung squamous carcinoma
- Ubiquitin–proteasome system
- Proteasome complex
- Lung squamous carcinomas
- Novel prognostic model
- Mutation
- Immune microenvironment
- Drug sensitivity
- Pathway and network alterations
- Therapeutic target
- Biomarkers
- Patient stratification
- Predictive diagnosis
- Prognostic assessment
- Personalized treatment
- Predictive, preventive, and personalized medicine (PPPM / 3PM)
- Multi-level diagnostics
- Multi-omics
- Liquid biopsy
- Prediction and targeted prevention of chronic inflammation and metastatic disease
- Mitochondrial health-related biomarkers