Abstract
Despite regular advances in Blattodea systematics, several relationships remain controversial or untested in formal phylogenetic reconstructions. This common situation for understudied metazoan groups limits our power to answer questions about phenotypic evolution. In this study, we infer the evolutionary history of Blattodea using newly sampled taxa that improve phylogenetic resolution while also illuminating the evolutionary history of an unusual phenotype—the apically folded hind-wing. Taxa newly sequenced include those with a hind-wing apical fold (Anaplecta pulchella, A. pygmaea, A. sp. cf. malaysensis, Diplopterina parva, Prosoplecta semperi, Anaplectoidea klossi, and Oulopteryx illuminata sp. nov. that we describe herein, including its male genitalia) and other rare taxa (Dipteretrum hamstroemi, Duchailluia togoensis, Lauraesilpha mearetoi, Buboblatta vlasaki). The phylogenetic design utilizes 41 genes over 91 species in total, analyzed in a maximum likelihood and coalescent framework. To quantify the phylogenetic uncertainty of the analysis, support for various topologies is assessed. We find unambiguous support for the surprising position of Neotropical Oulopteryx (Oulopterygidae) as sister to New Caledonian/Australian Tryonicidae. This, and other phylogenetic findings, reveal that the apically folded hind-wing may have arisen nine times in Blattodea. Further investigations are needed, notably with an increased taxonomic sampling, to demonstrate stronger support for the placement of rogue taxa (e.g., Anaplecta) and to investigate the evolutionary correlates of wing evolution.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Cockroach systematic studies continue to further refine deep (e.g., Djernæs et al., 2020; Evangelista et al., 2020; Li, 2022) and recent (e.g., Beasley‐Hall et al., 2021; Velez-Bravo & Daza, 2021) relationships, but several of them remain highly disputed, poorly supported, or simply unstudied (for examples, see Evangelista et al., 2017; Legendre & Grandcolas, 2018). A number of morphologically distinct groups have not been phylogenetically sampled with adequate taxonomic breadth, or at all. While some taxa cannot be included yet because they remain to be discovered—estimates say only ¼ of cockroach species have been described (Legendre & Grandcolas pers. comm., Beccaloni & Eggleton, 2013)—others can be included, especially with the latest advances in sequencing (Card et al., 2021). Thus, increasing taxonomic sampling remains a central challenge of obtaining a backbone phylogeny of Blattodea (Heath et al., 2008; Zwickl & Hillis, 2002). The value of downstream phylogenetic analyses, like reconstructions of phenotype, is also highly dependent on thorough sampling (Garamszegi, 2014).
One phenotypic feature of interest is the apical folding membrane of the cockroach hind-wing (Fig. S1). This is polymorphic among winged cockroaches with most species lacking a membrane between the CuP and V[s] veins (sensu Li et al., 2018), while a few species have a membrane here. The membrane is generally devoid of veins except for V[1], which provides support towards the apical most point and lays along the folding line. The membrane can vary from a minute triangle (i.e., an “intercalated triangle”), to a large region rivaling the size of the rest of the wing (i.e., an “appendicular field”). Although the larger form is not often seen among cockroaches, it appears to be deeply conserved within a few extant clades (e.g., Anaplectidae Walker, 1868; Ectobiinae Brunner, 1865) and is present in a variety of other taxa, both contemporary (e.g., Anisyutkin, 2013) and fossilized (Anisyutkin & Gröhn, 2012). A recent review considered all forms (except Diploptera) of a hind-wing apical folding to be homologous and simply used the term “apical folding” to refer to such forms (Li et al., 2018). While similar structures are seen in other insects [e.g., beetles (Kukalova-Peck & Lawrence, 1993)], the precise homology of these structures is unexamined, and the authors are unaware of any examples of closely related insects (i.e., Mantodea, Alienoptera) with this hind-wing region.
The folded membrane allows for the hind-wing to extend further outwards during flight and have higher surface area. The cost of enlarged wings is mitigated by its ability to remain protected at rest through folding or rolling underneath the tegmina (Li et al., 2018). Although its function is unstudied, the added surface area from the apical wing region likely has some aerodynamic function (Wang et al., 2018). Previous phylogenetic studies have placed some important taxa with apically folded hind wings [e.g., Diploptera Saussure, 1864 (Evangelista et al., 2020; Legendre et al., 2017; Li, 2022), Ectobiinae (Djernæs et al., 2020; Evangelista et al., 2020; Li, 2022), Riatia Walker, 1868, Chorisoneura Brunner, 1865, Calhypnorna Saussure & Zehntner, 1893, Euhypnorna Hebard, 1921, Theganopteryx Brunner, 1865 (Evangelista et al., 2020), Anaplecta Burmeister, 1838 (Deng et al., 2023; Djernæs et al., 2020; Li, 2022)]. However, those relationships were not always strongly supported, and these taxa alone do not effectively represent the diversity of cockroaches with apically folded hind-wings.
Here, we investigate the placement of nine newly sequenced and taxonomically disparate taxa, five of which have an apically folded hind-wing: Anaplecta; Anaplectoidea Shelford, 1906; Oulopteryx Hebard, 1921; Diplopterina Princis, 1963; and Prosoplecta Saussure, 1864. Anaplecta (Anaplectidae) has been sampled previously in molecular (Bourguignon et al., 2018; Evangelista et al., 2018; Li, 2022; Wang et al., 2017), morphological (Klass & Meier, 2006), and combined data (Djernæs et al., 2015) studies, but its position has been highly controversial. On the other hand, Anaplectoidea has never been included in a phylogenetic study. This genus was placed in Anaplectidae before Roth (1996)’s morphological assessment placed it in Blaberoidea. Oulopteryx (Oulopterygidae Rehn, 1951) has also never been included in a phylogenetic study. Oulopterygidae was previously considered to be a clade of Corydioidea Saussure, 1864 (Roth, 2003), based on non-cladistic taxonomic treatment (Hebard, 1921). Recent work by Hinkelman et al. (2020) summarizes the taxonomic history of the group and concludes that its phylogenetic position remains uncertain, but the current best hypothesis is that it is a close relative of Blattidae. Finally, Diplopterina and Prosoplecta may be less controversial in their taxonomic affiliations but are morphologically distinct, lack phylogenetic study, and bear large folded apical regions on their hind-wings.
The four remaining newly sampled taxa we include—Buboblatta Hebard, 1920; Duchailluia Rehn, 1933; Lauraesilpha Grandcolas, 1997; and Dipteretrum Rehn, 1922—do not have apically expanded hind-wings but are important for phylogenetic sampling nonetheless. Buboblatta is currently considered a neotropical representative of Latindiinae Handlirsch 1925 (Evangelista et al., 2019a, b, c; Hebard, 1920; Princis, 1963). Latindiinae is a poorly sampled clade (Djernæs et al., 2015; Evangelista et al., 2018; Legendre et al., 2015) that may be among the closest relatives of Nocticolidae (Wang et al., 2017), but its monophyly has not been tested through robust geographic sampling in phylogenies. Duchailluia is perhaps the sister to all other Blattidae Latreille, 1810 (Djernæs et al., 2015; Djernæs & Murienne, 2022), and is thus a key taxon for both the internal blattid relationships and the placement of Blattidae in Blattoidea. Lauraesilpha is one of only two Tryonicidae McKittrick and Mackerras, 1965 genera (Murienne, 2009), and including it should also assist in better resolution of the blattodean backbone. Tryonicidae has only been represented by a single long-branch in recent phylogenomic studies (Evangelista et al., 2020, 2019a, b, c) and has been found in varying positions in studies relying on other data (e.g., Grandcolas, 1996; Klass & Meier, 2006; Murienne, 2009; Legendre et al., 2015). Finally, Dipteretrum is a poorly studied genus of African Blaberoidea, which is currently unplaced within that superfamily (Cockroach Species File, accessed 2021).
In attempting to resolve these phylogenetic issues and map the evolution of the hind-wing apical field, we present a phylogenomic study of 91 species using 41 genes. We explore phylogenetic support for alternative hypotheses and varied inference methods.
Methods
Genomic data collection
Blattodea and relevant outgroups were sampled widely from previously published studies (Evangelista et al., 2019a, b, c, 2020) with eleven targeted samples added through new sequencing (Table S1). The resulting taxon sampling comprised 86 blattodean species (cockroaches and termites) and 5 outgroups. Newly sampled taxa had DNA extracted with Qiagen DNEasy kit and target-enriched genome sequencing was done using Arbor Biosciences’ myReads/myBaits workflow. Probe sets used were from Evangelista et al. (2020). Paired-end 150 sequencing was done on NovaSeq S4 (Illumina Inc.).
Precleaned libraries were further cleaned in TrimGalore v. 0.4.5 (Krueger, 2017) to remove low-quality bases and trim adapters (Phred score cutoff: 20; max. trimming error rate: 0.1; min. required adapter overlap: 1 bp; min. required sequence length for both reads before a sequence pair gets removed: 20 bp). Newly sequenced libraries were assembled using Trinity RNA Seq V.2.11 (Grabherr et al., 2011) using the –no_bowtie option. All newly sequenced and previously published assembled libraries were homologized to single-copy orthologs using two methods: EggNOG (Huerta-Cepas et al., 2017) and OrthoGraph (Petersen et al., 2017). EggNOG (Huerta-Cepas et al., 2017) identifies orthologs and annotates libraries by identifying seed orthologs from pre-existing phylogenies and genomic databases. EggNOG Mapper V5.0 parameters were set to auto-adjust per query for taxonomic scope (Arthropods) and one-to-one ortholog restriction. OrthoGraph V0.7.1 (Petersen et al., 2017) identifies orthologs based on reciprocal hidden-markov searches among reference taxa. We utilized the following ODB V.10 (Kriventseva et al., 2019) reference genomes: Blattabacterium cuenoti, B. sp. Blaberus, B. sp. Blatta, B. sp. Blattella, B. sp. Cryptocercus, B. sp. Mastotermes, B. sp. Nauphoeta, Blattella germanica, Ladona fulva, Pediculus humanus, Tribolium castaneum, and Zootermopsis nevadensis. OrthoGraph options were set to default except for the following: minimum transcript length = 35, ORF extension on, ORF minimum overlap = 0.4, HMM search score threshold = 15, BLAST score threshold = 15, and MAFFT (Katoh & Toh, 2008)—any-symbol option enabled. Blattabacterium is a type of Flavobacteria (Kambhampati, 2010) with a specialized symbiosis with Blattodea (Patino-Navarrete et al., 2013). These were included as a way to identify sequences from non-target bacteria. Sequences with best hits to Blattabacterium spp. were removed.
Alignment and data preparation
We chose 15 loci from the EggNOG assemblies for which Oulopteryx sp. nov.—a species with an apical folded membrane and from a family never included in any phylogenetic analysis—had the greatest overlap with other species. We added 30 other loci from the OrthoGraph assemblies: 10 that had the greatest taxon coverage including Oulopteryx sp. nov. and the next 20 with the most overall taxon coverage. Orthologous loci were cross-checked for duplicates across orthology methods (there were four duplicates) and combined into individual files such that each taxon had a single contig of the longest possible length across each locus (some of which contained multiple non-overlapping domains). This resulted in 41 final loci.
Loci were aligned using MAFFT V7.475 (parameters localpair –maxiterate 1000 input) (Katoh & Toh, 2008) and manually fine-tuned in AliView (Larsson, 2014). Stop codons, either at the end of coding regions or erroneously placed mid-sequence, were removed, and alignments were reading-frame adjusted. Three modified sets of alignments were created (Table 1): (i) reduced, alignments reduced to only the positions with more than 40% completeness (12,508 nts); (ii) masked, alignments with apparent non-coding (intron and ribosomal DNA loops) and hard to align regions removed (37,149 nts); and (iii) reduced and masked, both of the above conditions combined (11,584 nts).
Tree reconstruction
An IQ-TREE (Nguyen et al., 2015) concatenation maximum likelihood inference was carried out on the above alignment sets. Partition blocks were assigned according to gene and codon position for protein-coding genes and gene only for rDNA sequences. IQ-TREE2 options were as follows: -m MFP -rcluster 25 -ninit 300 -ntop 100 -nbest 20 -allnni -B 1000 -bsam GENE -bnni -nstop 250. This yielded a ML tree, a consensus of all bootstrap pseudoreplicates, and all 1000 bootstrap trees.
Individual gene tree histories were also inferred using IQ-TREE (Nguyen et al., 2015) on the “Masked” alignments. The following options were used: -m MFP -rcluster 20 -allnni -ninit 300 -ntop 100 -nbest 20 -allnni -nstop 250. The ML gene trees were then used to infer a species tree using ASTRAL 5.7.7 (Zhang et al., 2018).
We put all concatenation trees and the consensus of all bootstrap trees into a single text file and computed the overall majority-rule consensus tree (CC-Tree). Finally, we computed the majority-rule consensus of the CC-Tree and the ASTRAL tree (Conservative Sp. Tree).
Topology testing and node support
We manually grafted the CC-Tree (Grafted sp. Tree) to reflect the assumption of monophyletic Corydioidea (not including Anaplecta) and monophyletic Ectobiinae (Ectobius, Mediastinia, and Ectoneura). We then subsequently modified the Grafted sp. Tree to create 13 additional trees. Each of the 13 unique topologies tested a previously suggested hypothesis for the placement of some taxa with apical folding: Anaplecta, Anaplectoidea, Oulopteryx, and Diploptera.
All trees were compared in an approximately unbiased test (Shimodaira, 2002) (-m MFP -rcluster 25 -p -zb 10000) under both the “Masked” and “Reduced and masked” alignments along with their original partitioning block definitions. Relationship-specific tests were replicated twice, using the “Masked tree” and “Grafted sp. Tree” to estimate parameters respectively.
The final tree chosen had node support values from all three bootstrap replicates mapped onto it as well as gene concordance factors (Minh et al., 2020) calculated from the optimized gene trees.
Ancestral state reconstruction
Morphological data (Supplementary data 1) was synthesized from literature sources, databased photographs (Cockroach Species File, accessed 2021; MNHN digital collections, accessed 2021), and preserved specimens. Apical field presence was defined as any visible membranous region between the anal and anterior wing sections. This included large appendicular fields whose as basal most angle exceeded 90°, small apical triangles that are not extended enough to be the most distal point on the wing (e.g., as commonly seen in Blattellinae), and all sizes in between. Apterous and brachypterous taxa were coded as missing data.
Morphological data were analyzed in R using PhyTools (Revell, 2012). The ancestral states of categorical characters were estimated under a Bayesian framework in SIMMAP (Bollback, 2006) using the symmetrical model of evolution. The first 1000 simulations were discarded as burn-in, and 500 additional simulations were done while sampling every 10 replicates. The result was then tested for sensitivity to the phylogenetic topology. To control for phylogenetic error, each of these analyses was repeated using the eight grafted trees discussed above.
Results
Five of the eight species trees evaluated were rejected due to low p-values in the AU tests (Table 2). Of the remaining three trees, we chose the “Masked tree” as the best tree since both it and the consensus tree of its bootstrap trees had high lnL and p-values (Table 2). Although the “Reduced and Masked” tree outperformed the rest in the AUTest using the “Reduced and Masked” alignment, this test was not definitive (i.e., none of the trees were rejected as implausible), and the “Masked tree” also performed fairly well in this analysis too. Also, the “Masked tree” was obtained in an analysis with significantly more data (Table 1).
The 15 additional AU-tests definitively rejected certain relationships (Table 3). There was no support for the placement of Anaplecta spp. in Blaberoidea or Tryonicidae. The test also rejected the placement of Anaplecta spp. in a monophyletic Corydioidea. Two relationships for Anaplecta spp. that were supported are sister to Kittrickea or sister to Buboblatta vlasaki Evangelista, Kotyková-Varadínová and Jůna, 2019 (which together are sister to Blattoidea). There was no discernable signal for Anaplectoidea in any placement other than as sister to all remaining Blaberoidea. Similarly, no other placement for Oulopteryx sp. nov. was supported, other than sister to Tryonicidae. The “masked tree” was grafted with respect to the above results (Fig. 1) and was taken as the final species tree for our study.
Phylogeny of Blattodea (a) and wing morphology illustration (b). a The phylogeny presented is the final species tree resulting from a number of topology tests and inference methods (IQ-TREE and ASTRAL). Node support values represent bootstrap frequency (3000 replicates from concatenation analyses of all three modified alignments; left) and gene concordance factors among the 41 loci (right). Taxa in bold have hind wings with a very large apical folding area (b – i, and b – ii). The apical region in Diploptera may not be homologous to those of other taxa so we use another symbol and did not count them as additional apical field gains. Symbols on branches show inferred points of apical field gain (pink with outline) and loss (purple without outline). Numbered tokens correspond to images of taxa in the corresponding clade. 1, Therea sp. (Corydiinae); 2, Melyroidea magnifica (Oulopterygidae); 3, Periplaneta australasiae; 4, Ectobius sylvestris; 5, Riatia orientis; 6, Prosoplecta semperi; 7, Megaloblatta longipennis; 8, Loboptera decipiens; 9, Episymploce asahinae; 10, Epilampra opaca; 11, Eublaberus distanti; 12, Diploptera punctata
This final topology (Fig. 1) shows a large degree of congruence with respect to the placement and monophyly of major clades in previous studies (Blaser et al., 2020; Evangelista et al., 2019a, b, c). Family-level relationships in Blaberoidea were identical to that in Evangelista et al. (2020). Diploptera minor (Brunner, 1865) and Diplopterina parva (Borg, 1902) were both placed as close relatives of Paraplecta minutissima (Shelford, 1908), although node support was low. Dipteretrum hamstroemi Princis (1963), was recovered as sister to Supella longipalpa (Fabricius, 1798). Prosoplecta semperi Shelford (1912), was recovered as closely related to Pachnepteryx sp. cf. signaticollis (Stål, 1877). Corydioidea was polyphyletic with respect to Buboblatta vlasaki, but the support values on this quartet are generally low. Lauraesilpha mearetoi Grandcolas (1997), was sister to Tryonicus parvus (Tepper, 1895). Duchailluia togoensis (Shelford, 1911) was strongly supported as sister to all other Blattidae.
Systematic entomology
Order
Blattodea Brunner (1882)
Unranked
Solumblattodea Evangelista and Wipfler (2019)
Super-family
Blattoidea Latreille (1810)
Family
Oulopterygidae Rehn (1951)
History
Princis (1965) placed Melyroidea in Oulopterygidae with Prosoplecta, Anareolaria Shelford (1909), Euhypnorna, and Oulopteryx. Shelford’s description of Anareolaria is not sufficient to determine its placement (Shelford, 1909), and it is currently considered unplaced in Blaberoidea (Roth, 2003). Phylogenetic analysis (Fig. 1) clearly demonstrates that Euhypnorna is a genus of Blattellinae and Prosoplecta belongs to Pseudophyllodromiinae, placements consistent with their morphologies (Anisyutkin, 2013; Evangelista et al., 2019a, b, c). Bonfils (1975) placed Oulopteryx in Anaplectinae without giving a morphological justification. Roth (2003) included Oulopteryx in Corydioidea and Melyroidea in Pseudophyllodromiinae. Aclavoidea socialis Vidlička and Vršanský (2020) was described as being closely related to Melyroidea and Oulopteryx. However, these three oulopterygid genera have no consistent morphological differences other than body dimensions and coloration.
Family diagnosis
Oulopterygidae (Oulopteryx, Aclavoidea, and Melyroidea) differs from Tryonicidae in the following characteristics: Neotropical distribution (as opposed to Australian), macropterous (as opposed to brachypterous), pronotum with raised anterior edge and other sculpting (as opposed to smooth and regular). Oulopterygidae differs from Anaplecta in the folded apical membrane of the hind-wing being rolled at rest (rather than folded). Oulopterygidae also has thicker sclerotization of body and forewings, giving it a blaberid-like appearance (as opposed to the more delicate ectobiid-like appearance of Anaplectidae).
Genus
Oulopteryx Hebard (1921)
Included species
Oulopteryx meliponarum Hebard (1921) (type species), O. dascilloides Hebard (1921), O. illuminata Evangelista and Legendre sp. nov.
Key to Oulopteryx species
-
1.
Relatively large (body length > 12 mm); pronotum elliptical; pronotum, supra-anal plate and elytra without setae; pattern of hind-wing cross venation between R and M roughly parallel; interocular space roughly similar to inter-antennal space; styli half as long as distance between styli, cerci short and stout … O. illuminata Evangelista & Legendre sp. nov.
-
2.
Relatively small (body length < 12 mm); pronotum nearly circular; pronotum, lateral margin of elytra and caudal margin of supra-anal plate setose; pattern of hind-wing cross venation between R and M tortuous for a few veins; interocular space distinctly smaller than inter-antennal space; styli length 1/3 of distance between styli; cerci relatively long and slender … O. meliponarum Hebard (1921)
-
3.
Relatively small (body length < 12 mm); pronotum and elytra without setae; supra-anal plate setose; pattern of hind-wing cross venation between R and M roughly parallel; interocular space slightly smaller than inter-antennal space; styli length 1/3 of distance between styli; cerci short and stout … O. dascilloides Hebard (1921)
Species
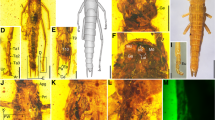
Oulopteryx illuminata Evangelista and Legendre sp. nov.
[Zoobank LSID: lsid:zoobank.org:act:0845FCB3-D4C6-440A-B0D3-D90A3E94E350]
(Fig. 2)
Oulopteryx illuminata sp. nov. is demonstrative of a typical species with an apical folded membrane, but having a rare method of concealment at rest (i.e., coiling, a feature shared among Oulopterygidae, Theganopteryx, Prosoplecta, and possibly a few others). Forewing (a, b) and hind-wing (c) morphology to scale with full body (f, g). Folding occurs alone dashed lines. Method of rolling the hind-wing apical field at rest shown (h). Genital morphology (d, e) of male holotype from posterior dorsal views (d) and dorsal view (e). Labels identify genital sclerites using the nomenclature of Klass (1997). See supplementary figures for more details. Wing venation (b, c) nomenclature based on Li et al. (2018) with modification. *Pcu not identified. See Schubnel et al. (2019) for a discussion of Pcu’s identity among Blattodea.
Type material
Holotype male: OUMNH-2005-065; Evergreen Forest, 1080 m alt.; @ M.V. light; coll. Mann, Hamel & Simmons.
Type locality
BOLIVIA: Dep. Santa Cruz; Bermejo, Refugio Los Volcanes, 18°06’S 63°36’W.
Description (male, female unknown)
General color reddish-chestnut. Head (Fig. 2g) large and round, yellow–brown, frons darker as well as four dark brown stripes on the vertex; clypeus unipartite but two-colored (tip hyaline); eyes black, bean-shaped; ocelli absent; maxillary palps five-segmented with the first two segments short, the third one the longest (ca. three times longer than the second one), the fourth segment half as long as the third, the fifth segment enlarged, oval, with fine setae mostly on the edges; interocular space slightly larger than inter-antennal space; antennae brown, darker at their tips, scape as long as the first two antennomeres. Pronotum (Fig. 2f) elliptical, rugose in its discal area, punctuated; anterior edge transverse, raised, lateral edges raised as well but to a lesser extent, posterior edge convex. Tegmina (Fig. 2a, b, f) fully developed, coriaceous, densely and regularly punctuated, apex sharply rounded; hind-wings (Fig. 2c, h) slightly infuscated, with a large apical field coiled at rest. Venation as in Fig. 2c, with no clear vein in the apical field. Anal plate symmetrical, concave medially. Subgenital plate small, convex, symmetrical, with two concave indentations where the small and elongate styli attached. Cerci short, stout, monomerous, flatten dorsad but with ventral side bulbous with several setae. Front femurs unarmed, only with a short genual spine, fifth tarsomere as long as the others combined, no pulvillus, claws symmetrical; Hindlegs with first tarsomere almost as long as the others combined, fifth tarsomere longer than the three previous ones, with pretarsal claws symmetrical and lacking pulvilli and arolia.
Genitalia as in Fig. 2d, e. Genitalia of congeneric species have never been illustrated or described.
Measurements
Body length (with tegmina): 16 mm; length of tegmina: 13 mm; largest width of tegmina: 4.5 mm; pronotum (width x length): 5.5 × 2.5 mm.
Diagnosis
Differs from M. magnifica Shelford (1912) and M. mimetica Shelford (1912) in coloration of pronotum and forewings (strongly reddish chestnut in O. illuminata) and the color of the hind-wing (mostly translucent in O. illuminata). Body coloration as in Aclavoidea socialis but more strongly reddened. Pronotum and body wide with stout legs. Strongly differs from A. socialis and M. magnifica in length of cerci (which are stout and roughly equal with the end of the male’s styli). Differs from A. socialis in presence of forewing PCu vein. Differs from Oulopteryx meliponarum Hebard (1921) in shape of pronotum (nearly circular in O. meliponarum), pattern of cross venation between radius and medial vein in hind-wing (roughly parallel in O. illuminata but not in O. meliponarum), non-setose pronotum, elytra and supra-anal plate (all setose in O. meliponarum), and body length larger.
Most similar to Oulopteryx dascilloides Hebard (1921) but with the following differences: body coloration reddish chestnut (as opposed to hazel-chestnut in O. dascilloides), styli half as long as distance between styli (as opposed to 1/3 the distance in O. dascilloides), cerci stouter with less than 1/3 extending past the supra-anal plate (as opposed to nearly half in of cerci extending past plate in O. dascilloides), supra-anal plate without setae (setose in O. dascilloides) and body length ~ 15 mm (~ 8 mm in O. dascilloides). Female unknown.
Etymology
The specific epithet is from the Latin illuminatus, meaning “lighting up or illuminating”. It has been chosen because this new species, belonging to the obscure Oulopterygidae family, has shed light on deep convergent evolution in cockroaches.
Discussion
Despite important advances in cockroach molecular systematics with increasing character and taxon sampling (e.g. > 50 genera in Legendre et al., 2015, 2017 and ca. 1 million nucleotides in Evangelista et al., 2019a, b, c), a vast majority of genera—and many more species—have never been included in any formal molecular phylogenetic analysis. We have shortened this gap with sampling of nine genera with unsettled phylogenetic affinities.
Arguably, the most outstanding result is the inference that Oulopteryx is sister to Tryonicidae (Fig. 1), since there was no previous finding of them sharing derived features. The biogeographical mismatch among this phylogenetic sister relationship is similarly surprising. This relationship may be between 65 MY [the youngest estimated age of crown Tryonicidae (Li, 2022)] and 200 MY old [the oldest estimated age of crown Blattidae + Tryonicidae (Li, 2022)], in which case it would be consistent with Gondwanan distribution and vicariance during the Jurassic, or a short dispersal during the Cretaceous.
Behaviorally, this relationship is of great interest, particularly with regard to diet and social behavior. In Blattodea, the prevailing hypothesis tightly links the evolution of eusociality with a xylophagous diet (Legendre & Grandcolas, 2018; Nalepa, 2015). Some Tryonicidae are wood-feeding and solitary (Grandcolas, 1997) while the diet of Oulopterygidae is unknown. Melyroidea has been found within excavated logs (Hinkelman et al., 2020), but their diet was not categorized. A recent study on Melyroidea in situ reported aggregation, group defensive behavior, and possibly parental care (Hinkelman et al., 2020). Further study is needed to reveal xylophagy or other aspects of social behavior in this and other lineages, exemplifying the desperate need of “natural history” data for numerous taxa (Greene, 2005). Other phylogenetic results are less intriguing but insightful nonetheless. While we did not obtain a robust placement for Anaplecta spp., we are able to narrow down possible placements to somewhere within, or sister to, Blattoidea. Recent analyses of mitogenomes (Bourguignon et al., 2018; Li, 2022) have recovered support for Anaplecta spp. as sister to Lamproblatta sp. Hebard, 1919, but studies have failed to reach congruence over the placement of the latter taxon (Bourguignon et al., 2018; Evangelista et al., 2019a, b, c; Li, 2022). On the other hand, we did find moderate support for Anaplectoidea sp. as sister to Blaberoidea s.s. This is consistent with Roth (1996)’s finding of genital synapomorphies with Blaberoidea. The genera Anaplectella Hanitsch, 1928, and Malaccina Hebard, 1929, should be included in future studies to ascertain potential monophyly with Anaplectoidea (Roth, 1996) and their position within, or sister to, Blaberoidea s.s.
The phylogenetic topology unambiguously suggests the hind-wing apically folded membrane was independently gained nine times and lost at least once (Fig. 1). While multiple acquisitions are strongly supported, the exact number of gains should be taken with caution, as it is influenced by a number of factors. First, the ancestral state reconstruction of the hind-wing apical field uses a very broad definition—any sized apical region of the membrane between the anterior (radial) and anal sectors of the wing. A more restricted definition—an apical expansion of the same membrane that is the apical most point on the wing—would remove the gains leading to Epilampra and Pycnoscelus. Although most Blattellinae fit the more liberal definition, a number of blattelline genera we included (Theganopteryx, Euhypnorna, Hemithrysocera) fit the more restricted one as well. Another potential source of error is an incomplete sampling of taxa with apically folded hind-wings. The Pseudophyllodromiidae genera Chorisoneura, Plectoptera (Rehn, 1951), and Calhypnorna (Saussure & Zehntner, 1893) also have apically expanded hind-wings. Phylogenetic studies have included Chorisoneura and Calhypnorna (Evangelista et al., 2014, 2020), and, while their findings are not entirely congruent, the evidence points to a close relationship with Macrophyllodromia—a large-bodied taxon lacking a hind-wing apical field. Other than Macrophyllodromia, these taxa were not included in our sample since their genetic data was non-overlapping with our 41 loci. If they were included, it may have resulted in an additional gain within Pseudophyllodromiidae or a deeper placement of the gain seen in Riatia sp. Another source of error in the ancestral reconstruction of total hind-wing apical field gains could be from the statistical inference method. The preference for two independent gains in Anaplectoidea and Ectobiinae as opposed to on the shorter stem of their ancestor is a feature of the Bayesian inference (Bollback, 2006). An equally parsimonious state reconstruction is a gain of an apical field at the ancestor of Anaplectoidea + Blaberoidea s.s. and a loss in the ancestor of Pseudophyllodromiidae + Orkrasomeria Evangelista et al. 2020. Finally, we chose to code Diploptera as not having apically expanded hind-wings. Although Diploptera’s hind-wings are long compared to the fore-wings, their expansion appears to be derived from the anterior section of the wing (Li & Wang, 2015). Coding their apical fields as “present” could have resulted in the inference of at least one additional gain of the apical field. Considering all of the above possible sources of error, the number of possible gains (7–12) and losses (1–3) could differ from our result of 9 and 1, respectively.
Regardless, there is extreme convergence in wing morphology among lineages that diverged deep in evolutionary time—some up to 230 MA (Li, 2022). The biological correlates of this remarkable convergent evolution are unexplored. Cockroach hind-wings are both flight organs (Nalepa et al., 2001) and used in mating rituals (Kotyk & Varadinova, 2017). In other animals, wing size relative to body size is known to affect flight efficacy (e.g., Le Roy et al., 2019; McCulloch & Waters, 2018), but such studies on taxa that are traditionally considered weak fliers are lacking. The interactions between relative wing size, body size (Grabow & Rüppell, 1995), wing membrane thickness, wing resilience (Dirks & Taylor, 2012), and flight capacity (Le Roy et al., 2019; Wootton, 1981) are largely unexplored in Blattodea. In addition to learning about aerodynamic evolution, comparing the folding mechanism of the wing to other organisms with similar apical wing folding, such as beetles or earwigs (Haas, 1999; Kukalova-Peck & Lawrence, 1993), could yield interesting outcomes as well. In fact, many of the cockroaches with expanded hind-wings are considered beetle mimics themselves (Shelford, 1912). Perhaps the morphological constraints of a beetle-like morphology somehow necessitate hind-wing folding and/or expansion. Exploring these intriguing patterns should be the subject of future studies on the evolutionary drivers of wing evolution.
Data availability
Newly sequenced genetic data are available on NCBI sequence read archive (PRJNA482916), and our main phylogenetic tree and alignments are available on Dryad digital repository (https://doi.org/10.5061/dryad.z34tmpgh0).
References
Anisyutkin, L. N. (2013). A description of a new species of the cockroach genus Prosoplecta Saussure, 1864 (Dictyoptera, Ectobiidae) from South Vietnam. Entomological Review, 93(2), 182–193. https://doi.org/10.1134/s0013873813020061
Anisyutkin, L. N., & Gröhn, C. (2012). New cockroaches (Dictyoptera: Blattina) from Baltic Amber, with the description of a new genus and species: Stegoblatta irmgardgroehni. Proceedings of the Zoological Institute RAS, 316(3), 193–202.
Beasley‐Hall, P. G., Rose, H. A., Walker, J., Kinjo, Y., Bourguignon, T., & Lo, N. (2021). Digging deep: A revised phylogeny of Australian burrowing cockroaches (Blaberidae: Panesthiinae, Geoscapheinae) confirms extensive nonmonophyly and provides insights into biogeography and evolution of burrowing. Systematic Entomology. https://doi.org/10.1111/syen.12487
Beccaloni, G., & Eggleton, P. (2013). Order: Blattodea. Zootaxa, 3703(1), 46. https://doi.org/10.11646/zootaxa.3703.1.10
Blaser, M., Misof, B., & Predel, R. (2020). The power of neuropeptide precursor sequences to reveal phylogenetic relationships in insects: A case study on Blattodea. Molecular Phylogenetics and Evolution, 143, 106686. https://doi.org/10.1016/j.ympev.2019.106686
Bollback, J. P. (2006). SIMMAP: Stochastic character mapping of discrete traits on phylogenies. BMC Bioinformatics, 7(88), 88. https://doi.org/10.1186/1471-2105-7-88
Bonfils, J. (1975). Blattoptera [Orthopteroidea] récoltés en Guyane Française par la mission du muséum national d’histoire naturelle. Annales de la Société Entomologique de France, 11(1), 29–62.
Bourguignon, T., Tang, Q., Ho, S. Y. W., Juna, F., Wang, Z., Arab, D. A., Cameron, S. L., Walker, J., Rentz, D., Evans, T. A., & Lo, N. (2018). Transoceanic dispersal and plate tectonics shaped global cockroach distributions: Evidence from mitochondrial phylogenomics. Molecular Biology and Evolution, 35(4), 1–14. https://doi.org/10.1093/molbev/msy013
Card, D. C., Shapiro, B., Giribet, G., Moritz, C., & Edwards, S. V. (2021). Museum genomics. Annual Review of Genetics, 55, 633–659. https://doi.org/10.1146/annurev-genet-071719-020506
Deng, W. L., Xinxing, Ho., & S. Y., Liao, S., Wang, Z., & Che, Y. (2023). Inclusion of rare taxa from Blattidae and Anaplectidae improves phylogenetic resolution in the cockroach superfamily Blattoidea. Systematic Entomology, 48, 23–39. https://doi.org/10.1111/syen.12560
Dirks, J. H., & Taylor, D. (2012). Veins improve fracture toughness of insect wings. PloS One, 7(8), e43411. https://doi.org/10.1371/journal.pone.0043411
Djernæs, M., Klass, K. D., & Eggleton, P. (2015). Identifying possible sister groups of Cryptocercidae+Isoptera: A combined molecular and morphological phylogeny of Dictyoptera. Molecular Phylogenetics and Evolution, 84, 284–303. https://doi.org/10.1016/j.ympev.2014.08.019
Djernæs, M., & Murienne, J. (2022). Phylogeny of Blattoidea (Dictyoptera: Blattodea) with a revised classification of Blattidae. Arthropod Systematics & Phylogeny, 80, 209–228. https://doi.org/10.3897/asp.80.e75819
Djernæs, M., Varadínová, Z. K., Eulitz, U., & Klass, K.-D. (2020). Phylogeny and life history evolution of Blaberoidea (Blattodea). Arthropod Systematics & Phylogeny, 78(1), 29–67. https://doi.org/10.26049/ASP78-1-2020-03
Evangelista, D., Simon, S., Wilson, M. M., Kawahara, A. Y., Kohli, M. K., Ware, J. L., Wipfler, B., Béthoux, O., Grandcolas, P., & Legendre, F. (2020). Assessing support for Blaberoidea phylogeny suggests optimal locus quality. Systematic Entomology, 46(1), 157–171. https://doi.org/10.1111/syen.12454
Evangelista, D., Thouzé, F., Kohli, M. K., Lopez, P., & Legendre, F. (2018). Topological support and data quality can only be assessed through multiple tests in reviewing Blattodea phylogeny. Molecular Phylogenetics and Evolution, 128, 112–122. https://doi.org/10.1016/j.ympev.2018.05.007
Evangelista, D. A., Bourne, G., & Ware, J. L. (2014). Species richness estimates of Blattodea s.s. (Insecta: Dictyoptera) from northern Guyana vary depending upon methods of species delimitation. Systematic Entomology, 39, 150–158. https://doi.org/10.1111/syen.12043
Evangelista, D. A., Djernæs, M., & Kohli, M. K. (2017). Fossil calibrations for the cockroach phylogeny (Insecta, Dictyoptera, Blattodea), comments on the use of wings for their identification, and a redescription of the oldest Blaberidae. Palaeontologia Electronica, 20.3(1FC), 1–23.
Evangelista, D. A., Kotyková-Varadínová, Z., Juna, F., Grandcolas, P., & Legendre, F. (2019a). New cockroaches (Dictyoptera: Blattodea) from French Guiana and a revised checklist for the region. Neotropical Entomology, 48(4), 645–659. https://doi.org/10.1007/s13744-019-00677-6
Evangelista, D. A., Wipfler, B., Béthoux, O., Donath, A., Fujita, M., Kohli, M. K., Legendre, F., Liu, S., Machida, R., Misof, B., Peters, R. S., Podsiadlowski, L., Rust, J., Schuette, K., Tollenaar, W., Ware, J. L., Wappler, T., Zhou, X., Meusemann, K., & Simon, S. (2019b). An integrative phylogenomic approach illuminates the evolutionary history of cockroaches and termites (Blattodea). Proceedings of the Royal Society B: Biological Sciences, 286(1895), 1–9. https://doi.org/10.1098/rspb.2018.2076
Evangelista, D. A., Wipfler, B., Béthoux, O., Donath, A., Fujita, M., Kohli, M. K., Legendre, F., Liu, S., Machida, R., Misof, B., Peters, R. S., Podsiadlowski, L., Rust, J., Schuette, K., Tollenaar, W., Ware, J. L., Zhou, X., Meusemann, K., & Simon, S. (2019c). An integrative phylogenomic approach illuminates the evolutionary history of cockroaches and termites (Blattodea). Proceedings of the Royal Society B: Biological Sciences, 286(1895), 1–9. https://doi.org/10.1098/rspb.2018.2076
Garamszegi, L. Z. (2014). Modern phylogenetic comparative methods and their application in evolutionary biology: Concepts and practice (1 ed.). Springer. https://doi.org/10.1007/978-3-662-43550-2
Grabherr, M., & BJ, H., Yassour, M., Levin, J. Z., Thompson, D. A., Amit, I., Adiconis, X., Fan, L., Raychowdhury, R., Zeng, Q., Chen, Z., Mauceli, E., Hacohen, N., Gnirke, A., Rhind, N., Palma, F. d., Birren, B. W., Nusbaum, C., Lindblad-Toh, K., … Regev, A. (2011). Trinity reconstructing a full-length transcriptome without a genome from RNA-Seq data. Nature Biotechnology, 29(7), 644–652.
Grabow, K., & Rüppell, G. (1995). Wing loading in relation to size and flight characteristics of European Odonata. Odonotologica, 12(2), 175–186.
Grandcolas, P. (1996). The phylogeny of cockroach families: A cladistic appraisal of morpho-anatomical data. Canadian Journal of Zoology, 74, 508–527.
Grandcolas, P. (1997). Systématique phylogénétique de la sous-famille des Tryonicinae (Dictyoptera, Blattaria, Blattidae). Mémoires Du Muséum National D’histoire Naturelle, 171, 91–124.
Greene, H. W. (2005). Organisms in nature as a central focus for biology. Trends in Ecology & Evolution, 20, 23–27.
Haas, F. (1999). Mechanische und evolutive Aspekte der Flugelfaltung bei Blattodea, Dermaptera und Coleoptera. Cour. Forsch.-Inst. Senckenberg, 215, 97–102.
Heath, T. A., Hedtke, S. M., & Hillis, D. M. (2008). Taxon sampling and the accuracy of phylogenetic analyses. Journal of Systematics and Evolution, 46(3), 239–257. https://doi.org/10.3724/SP.J.1002.2008.08016
Hebard, M. (1920). The Blattidae of Panama. American Entomological Society Memoirs, 4, 1–148.
Hebard, M. (1921). South American Blattidae from the Museum National d’Histoire Naturelle, Paris, France. Proceedings of the Academy of Natural Sciences of Philadelphia, 73(2), 193–304.
Hinkelman, J., Vrsansky, P., Garcia, T., Tejedor, A., Bertner, P., Sorokin, A., Gallice, G. R., Koubova, I., Nagy, S., & Vidlicka, L. (2020). Neotropical Melyroidea group cockroaches reveal various degrees of (eu)sociality. Naturwissenschaften, 107(5), 39. https://doi.org/10.1007/s00114-020-01694-x
Huerta-Cepas, J., Forslund, K., Coelho, L. P., Szklarczyk, D., Jensen, L. J., Mering, C. V., & Bork, P. (2017). Fast genome-wide functional annotation through orthology assignment by eggNOG-Mapper. Molecular Biology and Evolution, 34(8), 2115–2122.
Kambhampati, S. (2010). Family II. Blattabacteriaceae fam. nov. In S. J. Krieg NR, Brown DR, Hedlund BP, Paster BJ, Ward NL, Ludwig W, Whitman WB (Ed.), Bergey's Manual of Systematic Bacteriology (2 ed., Vol. 4, pp. 315). Springer.
Katoh, K., & Toh, H. (2008). Recent developments in the MAFFT multiple sequence alignment program. Briefings in Bioinformatics, 9(4), 286–298. https://doi.org/10.1093/bib/bbn013
Klass, K.-D. (1997). The external male genitalia and the phylogeny of Blattaria and Mantodea. Bonner Zoologische Monographien, 42, 1–340.
Klass, K.-D., & Meier, R. (2006). A phylogenetic analysis of Dictyoptera (Insecta) based on morphological characters. Entomologische Abhandlungen, 63(1–2), 3–50.
Kotyk, M., & Varadinova, Z. (2017). Wing reduction influences male mating success but not female fitness in cockroaches. Scientific Reports, 7(1), 2367. https://doi.org/10.1038/s41598-017-02647-7
Kriventseva, E. V., Kuznetsov, D., Tegenfeldt, F., Manni, M., Dias, R., Simão, F. A., & Zdobnov, E. M. (2019). OrthoDB v10: Sampling the diversity of animal, plant, fungal, protist, bacterial and viral genomes for evolutionary and functional annotations of orthologs. Nucleic Acids Research, 47(D1), D807–D811. https://doi.org/10.1093/nar/gky1053
Krueger, F. (2017). TrimGalore v. 0.4.5. Babraham Bioinformatics. Babraham Institute. https://www.bioinformatics.babraham.ac.uk/projects/trim_galore/
Kukalova-Peck, J., & Lawrence, J. F. (1993). Evolution of the hind wing in Coleoptera. The Canadian Entomologist, 125, 181–258.
Larsson, A. (2014). AliView: a fast and lightweight alignment viewer and editor for large data sets. Bioinformatics, 30(22), 3276–3278.
Le Roy, C., Cornette, R., Llaurens, V., & Debat, V. (2019). Effects of natural wing damage on flight performance in Morpho butterflies: What can it tell us about wing shape evolution? Journal of Experimental Biology, 222(Pt 16). https://doi.org/10.1242/jeb.204057
Legendre, F., & Grandcolas, P. (2018). The evolution of sociality in termites from cockroaches: A taxonomic and phylogenetic perspective. Journal of Experimental Zoology Part B: Molecular and Developmental Evolution, 330(5), 279–287. https://doi.org/10.1002/jez.b.22812
Legendre, F., Grandcolas, P., & Thouzé, F. (2017). Molecular phylogeny of Blaberidae (Dictyoptera, Blattodea) with implications for taxonomy and evolutionary studies. European Journal of Taxonomy, 291, 1–13. https://doi.org/10.5852/ejt.2017.291
Legendre, F., Nel, A., Svenson, G. J., Robillard, T., Pellens, R., & Grandcolas, P. (2015). Phylogeny of Dictyoptera: Dating the origin of cockroaches, praying mantises and termites with molecular data and controlled fossil evidence. PloS One, 10(7), e0130127. https://doi.org/10.1371/journal.pone.0130127
Li, X.-R. (2022). Phylogeny and age of cockroaches: A reanalysis of mitogenomes with selective fossil calibrations. Deutsche Entomologische Zeitschrift, 69(1), 1–18. https://doi.org/10.3897/dez.69.68373
Li, X., & Wang, Z. (2015). A taxonomic study of the beetle cockroaches (Diploptera Saussure) from China, with notes on the genus and species worldwide (Blattodea: Blaberidae: Diplopterinae). Zootaxa, 4018(1), 35–56. https://doi.org/10.11646/zootaxa.4018.1.2
Li, X.-R., Zheng, Y.-H., Wang, C.-C., & Wang, Z.-Q. (2018). Old method not old-fashioned: Parallelism between wing venation and wing-pad tracheation of cockroaches and a revision of terminology. Zoomorphology, 137(4), 519–533. https://doi.org/10.1007/s00435-018-0419-6
McCulloch, G. A., & Waters, J. M. (2018). Does wing reduction influence the relationship between altitude and insect body size? A case study using New Zealand’s diverse stonefly fauna. Ecology and Evolution, 8(2), 953–960. https://doi.org/10.1002/ece3.3713
Minh, B. Q., Hahn, M. W., & Lanfear, R. (2020). New methods to calculate concordance factors for phylogenomic datasets. Molecular Biology and Evolution, 37(9), 2727–2733. https://doi.org/10.1093/molbev/msaa106
Murienne, J. (2009). Molecular data confirm family status for the Tryonicus-Lauraesilpha group (Insecta: Blattodea: Tryonicidae). Organisms Diversity & Evolution, 9(1), 44–51. https://doi.org/10.1016/j.ode.2008.10.005
Nalepa, C. A. (2015). Origin of termite eusociality: Trophallaxis integrates the social, nutritional, and microbial environments. Ecological Entomology, 40(4), 323–335. https://doi.org/10.1111/een.12197
Nalepa, C. A., Miller, L. R., & Lenz, M. (2001). Flight characteristics of Mastotermes darwiniensis (Isoptera, Mastotermitidae). Insectes Sociaux, 48, 144–148.
Nguyen, L. T., Schmidt, H. A., von Haeseler, A., & Minh, B. Q. (2015). IQ-TREE: A fast and effective stochastic algorithm for estimating maximum-likelihood phylogenies. Molecular Biology and Evolution, 32(1), 268–274. https://doi.org/10.1093/molbev/msu300
Patino-Navarrete, R., Moya, A., Latorre, A., & Pereto, J. (2013). Comparative genomics of Blattabacterium cuenoti: The frozen legacy of an ancient endosymbiont genome. Genome Biology and Evolution, 5(2), 351–361. https://doi.org/10.1093/gbe/evt011
Petersen, M., Meusemann, K., Donath, A., Dowling, D., Liu, S., Peters, R. S., Podsiadlowski, L., Vasilikopoulos, A., Zhou, X., Misof, B., & Niehuis, O. (2017). Orthograph: A versatile tool for mapping coding nucleotide sequences to clusters of orthologous genes. BMC Bioinformatics, 18(1), 111. https://doi.org/10.1186/s12859-017-1529-8
Princis, K. (1963). Blattariae: Suborde Polyphagoidea: Fam. Homoeogamiidae, Euthyrrhaphidae, Latindiidae, Anacompsidae, Atticolidae, Attaphilidae; Subordo Blaberoidea: Fam. Blaberidae (Vol. Pars 4). W. Junk.
Princis, K. (1965). Blattariae: Subordo Blaberoidea: Fam. Oxyhaloidae, Panesthiidae, Cryptocercidae, Chorisoneuridae, Oulopterygidae, Diplopteridae, Anaplectidae, Archiblattidae, Nothoblattidae (Vol. Pars 7). W. Junk.
Rehn, J. W. H. (1951). Classification of the Blattaria as indicated by their wings (Orthoptera). Memoirs of the American Entomological Society, 14, 1–134.
Revell, L. J. (2012). phytools: An R package for phylogenetic comparative biology (and other things). Methods in Ecology and Evolution, 3, 217–223.
Roth, L. M. (1996). The cockroach genera Anaplecta, Anaplectella, Anaplectoidea, and Malaccina (Blattaria, Blattellidae: Anaplectinae and Blattellinae). Oriental Insects, 30(1), 301–372. https://doi.org/10.1080/00305316.1996.10434105
Roth, L. M. (2003). Systematics and phylogeny of cockroaches (Dictyoptera: Blattaria). Oriental Insects, 37, 1–186.
Saussure, H. D., & Zehntner, L. (1893). Insecta. Orthoptera. In F. D. Godman & O. Salvin (Eds.), Biologia Centrali-Americana (Vol. 1, pp. 1–285). R.H. Porter.
Schubnel, T., Desutter-Grandcolas, L., Legendre, F., Prokop, J., Mazurier, A., Garrouste, R., Grandcolas, P., & Nel, A. (2019). To be or not to be: Postcubital vein in insects revealed by microtomography. Systematic Entomology, 45(2), 327–336. https://doi.org/10.1111/syen.12399
Shelford, R. (1909). Descriptions of some new genera and species of Blattidae. Deutsche Entomologische Zeitschrift, 611–624.
Shelford, R. (1912). Mimicry amongst the Blattidae; with a revision of the genus Prosoplecta Sauss. and the description of a new genus. Proceedings of the Zoological Society of London, 82(2), 358–378.
Shimodaira, H. (2002). An approximately unbiased test of phylogenetic tree selection. Systematic Biology, 51(3), 492–508. https://doi.org/10.1080/10635150290069913
Velez-Bravo, A., & Daza, J. M. (2021). Molecular systematics and genital morphology of the Neotropical cockroaches from the genus Xestoblatta (Blattellidae). Zootaxa, 5057(3), 1–28. https://doi.org/10.11646/zootaxa.5057.3.1
Wang, C., Wang, C., Ning, Y., Chen, L., & Wang, X. (2018). Design and mechanical analysis of bionic foldable beetle wings. Applied Bionics and Biomechanics, 2018, 1308465. https://doi.org/10.1155/2018/1308465
Wang, Z., Shi, Y., Qiu, Z., Che, Y., & Lo, N. (2017). Reconstructing the phylogeny of Blattodea: Robust support for interfamilial relationships and major clades. Scientific Reports, 7(3903), 1–8. https://doi.org/10.1038/s41598-017-04243-1
Wootton, R. J. (1981). Support and deformability in insect wings. Journal of Zoology, London, 193, 447–468.
Zhang, C., Rabiee, M., Sayyari, E., & Mirarab, S. (2018). ASTRAL-III: Polynomial time species tree reconstruction from partially resolved gene trees. BMC Bioinformatics, 19(Suppl 6), 153. https://doi.org/10.1186/s12859-018-2129-y
Zwickl, D. J., & Hillis, D. M. (2002). Increased taxon sampling greatly reduces phylogenetic error. Systematic Biology, 51(4), 588–598. https://doi.org/10.1080/10635150290102339
Acknowledgements
The authors would like to thank Darren Mann for collecting and loaning specimens for this study, as well as motivation for the study of Oulopteryx. We also extend appreciation to Samuel Bennett for preliminary data collection, Elyane Barros de Pina for the concurrent study of cockroach wing morphology, and Olivier Béthoux for providing photographs of samples.
Funding
Funding for this study was provided by the US National Science Foundation award numbers 1608559 and 2209323.
Author information
Authors and Affiliations
Contributions
DAE and FL conceived of and designed the study. DAE carried out study from data collection to analysis, wrote the manuscript, and composed the figures. DN carried out bioinformatics. ZKV collected specimens, identified specimens, and provided contributions to the manuscript. FL collected specimens, provided systematic expertise, and revised the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Evangelista, D.A., Nelson, D., Varadínová, Z.K. et al. Phylogenomics and deep convergence in cockroach hind-wing morphology. Org Divers Evol 23, 929–940 (2023). https://doi.org/10.1007/s13127-023-00609-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13127-023-00609-8