Abstract
The diversification processes underlying why Amazonia hosts the most species-rich vertebrate fauna on earth remain poorly understood. We studied the spatio-temporal diversification of a tree frog clade distributed throughout Amazonia (Anura: Hylidae: Osteocephalus, Tepuihyla, and Dryaderces) and tested the hypothesis that Miocene mega wetlands located in western and central Amazonia impacted connectivity among major biogeographic areas during extensive periods. We assessed the group’s diversity through DNA-based (16S rRNA) species delimitation to identify Operational Taxonomic Units (OTUs) from 557 individuals. We then selected one terminal for each OTU (n = 50) and assembled a mitogenomic matrix (~14,100 bp; complete for 17 terminals) to reconstruct a Bayesian, time-calibrated phylogeny encompassing nearly all described species. Ancestral area reconstruction indicates that each genus was restricted to one of the major Amazonian biogeographic areas (western Amazonia, Guiana Shield and Brazilian Shield, respectively) between ~10 and 20 Mya, suggesting that they diverged and diversified in isolation during this period around the Pebas mega wetland. After 10 Mya and the transition to the modern configuration of the Amazon River watershed, most speciation within each genus continued to occur within each area. In Osteocephalus, only three species expanded widely across Amazonia (< 6 Mya), and all were pond-breeders. Species with other breeding modes remained mostly restricted to narrow ranges. The spectacular radiation of Osteocephalus was probably driven by climatic stability, habitat diversity and the acquisition of new reproductive modes along the Andean foothills and western Amazonia. Our findings add evidence to the importance of major hydrological changes during the Miocene on biotic diversification in Amazonia.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The Neotropics harbour the world’s highest species diversity for many animal and plant groups (Raven et al., 2020). The origin and distribution of this astonishing diversity have intrigued biologists for almost two centuries (e.g. Wallace, 1854), notably in Amazonia, which encompass > 6.5 million km2 of mostly rainforest vegetation. Our understanding of the diversification processes and assembling of biotas in the Neotropics has accelerated over the last decades, with Amazonia acting as the major biodiversity source for the entire continent (Antonelli et al., 2018). Nevertheless, the historical diversification of major lineages in the Neotropics has been mostly tackled at the continental scale, whereas diversification processes within Amazonia remain unresolved (Cracraft et al., 2020). This is at least partly because including sufficiently dense and taxonomically complete sampling across this broad region, together with rigorous temporal estimations of lineage diversification throughout Neogene (2.6–23 Mya), remains challenging.
Several diversification hypotheses have been formulated to explain the origin of Amazonian biodiversity. Hypotheses involving major landscape changes that lead to allopatric speciation have received the most attention (for a review, see Leite & Rogers, 2013). For example, the riverine barrier hypothesis (Wallace, 1854) postulates that populations of terrestrial organisms were isolated following the formation of the largest Amazonian rivers. Paleogeographic data suggests that this hypothesis should be largely limited to speciation since ~10 Mya, following the establishment of the modern Amazon River watershed (Albert et al., 2018; Hoorn et al., 2010). Another model of allopatric speciation is the marine incursions hypothesis (Webb, 1995), which postulates that during the middle Miocene, an extensive mega wetland with marine influence occupied western-central Amazonia (termed the ‘Pebas system’) and thus formed a barrier for rainforest organisms between ~10 and 23 Mya (Albert et al., 2018; Bicudo et al., 2019; Hoorn et al., 2010). However, it is important to keep in mind that large uncertainties surround Amazonia’s hydro-geological history, as illustrated by different models in Bicudo et al. (2019). The end of the Pebas system (draining north to the Caribbean Sea) was followed by the onset of the transcontinental Amazon River (draining east to the Atlantic Ocean) at 9–10 Mya (Hoorn et al., 2010, 2017). Nevertheless, the mega wetlands might have persisted in south-western Amazonia as the Acre system between 7 and 10 Mya (Hoorn et al., 2010). Therefore, over the last 10 Mya, western Amazonia underwent profound and progressive transformations, transitioning from a mega wetlands environment towards a fluvial and terra firme–dominated landscape (Albert et al., 2018).
Most comprehensive studies on speciation and biogeography of tetrapods in Amazonia have been conducted on birds (Ribas & Aleixo, 2019) and primates (e.g. Boubli et al., 2015; Byrne et al., 2018), primarily because their diversity and taxonomy are reasonably well known. A recent review suggests that most speciation in birds and primates within Amazonia occurred relatively recently (Pliocene and Pleistocene; 0.01–5.3 Mya) (Cracraft et al., 2020), and often implied successive dispersals across rivers from western to eastern Amazonia (Silva et al., 2019; Smith et al., 2014). In contrast, the diversity and biogeography of other groups, including Amazonian amphibians, are less well resolved. Frogs are a particularly valuable group to study due to their exceptional diversity in Amazonia and limited dispersal capabilities (Zeisset & Beebee, 2008).
Some frog groups originated out of Amazonia, notably in the Andes, and secondarily dispersed into Amazonia during the Miocene (5.3–23 Mya) (e.g. Centrolenidae, Castroviejo-Fisher et al., 2014; Dendrobatidae, Santos et al., 2009). Other groups are only found in the Amazonian lowlands and their diversification occurred in situ during the Miocene (e.g. Adenomera, Fouquet et al., 2014; Allobates, Réjaud et al., 2020; Amazophrynella, Moraes et al., 2022; Phyzelaphryninae, Fouquet et al., 2012; Synapturanus, Fouquet et al., 2021a; Pipa, Fouquet et al., 2022). In general, the diversification events in these ground-dwelling and aquatic frogs appear to reflect the presence of ancient mega wetlands (e.g. Pebas system) with separation between western and eastern Amazonian clades (> 10 Mya) and then establishment of the trans-continental Amazon River (~10 Mya) separating northern and southern Amazonian clades. However, there have been few biogeographic studies of the exceptionally diverse arboreal frog fauna in Amazonia, for which biogeographic patterns are expected to differ due to ecological differences with terrestrial species (Fouquet et al., 2015). For example, Fouquet et al. (2021b) found that diversification of tree frogs of the Boana albopunctata group is relatively recent (Pliocene and Pleistocene) and included dispersals widely across Amazonia. This diversification was thus similar to patterns observed in birds and primates but in contrast to those seen in ground and aquatic frogs.
The tree frog clade consisting of Osteocephalus (29 species), Tepuihyla (9 species) and Dryaderces (2 species) (Hylidae: Lophiohylinae; Frost, 2022; Ortiz & Ron, 2018) is endemic to Amazonia and the Pantepui region (Duellman et al., 2016; Jungfer et al., 2013). This clade is diverse and widespread and thus offers the potential for valuable insights into Amazonian biogeography. However, this clade also exemplifies the gap existing in our understanding of the extant Amazonian diversity because new species are frequently described (e.g. Chasiluisa et al., 2020; Duellman, 2019; Melo-Sampaio et al., 2021); many known populations are suspected to belong to undescribed species (Jungfer et al., 2013; Vacher et al., 2020), and some relationships among species and species groups remain unresolved (Blotto et al., 2020). Moreover, the spatio-temporal aspects of their diversification and biogeography remain virtually unexplored. Duellman et al. (2016) estimated the divergences among the three genera between ~25 and 28 Mya, the crown age of Osteocephalus ~18.2 Mya (15.5–20.9), and Salerno et al. (2012) estimated the crown age of Tepuihyla (including Tepuihyla exophthalma Smith & Noonan, 2001 as ‘Osteocephalus exophthalmus’ therein) to be 14.7–23.6 My old. However, these estimates are suspected to be overestimated based on recent genomic studies (Feng et al., 2017; Hime et al., 2021).
The spatial distribution of the three genera in this clade covers Amazonia. Dryaderces is endemic to the Brazilian Shield, Tepuihyla is restricted to the Pantepui region (Guiana Shield) and adjacent north-western Amazonia, and Osteocephalus has diversified extensively throughout Amazonia but is particularly diverse along the Andean foothills and western Amazonia (Duellman, 2019; Jungfer et al., 2013; Ron et al., 2016). Within Osteocephalus, five species groups are currently recognised based on their monophyly and reproductive modes (Jungfer et al., 2013). The O. leprieurii and O. taurinus groups are almost entirely pond-breeders, and the O. buckleyi group mostly breeds in streams. These three groups are widespread across Amazonia. The O. alboguttatus and O. planiceps groups are restricted to western Amazonia and reproduce in phytotelmata. The evolution of breeding ecology in Osteocephalus is of particular interest because it may have contributed to its extensive diversification, including enabling co-occurrence of species (Jungfer et al., 2013).
Based on the fact that the crown age of the most recent common ancestor between Osteocephalus and Trachycephalus (another genus in Lophiohylinae) has been estimated from genomics to be about 23 My old (Feng et al., 2017), the diversification of our focal clade must have taken place through the Neogene. Therefore, the origin of the three major clades (genera) distributed in the Amazonian areas described above (i.e. western Amazonia, Guiana Shield and Brazilian Shield) either predates 10 Mya and hence overlaps with the Pebas system as a potential barrier, or is more recent and occurred across major Amazonian rivers. Given the higher diversity of Osteocephalus in western Amazonia, we hypothesize that this region acted as a centre of diversification and that the different species groups dispersed throughout the Amazon River watershed after the demise of the Pebas and Acre systems.
To test these assumptions, we used spatially dense sampling of mitochondrial sequences (16S rRNA) from 557 individuals of the Amazonia-wide tree frog clade formed by Osteocephalus, Tepuihyla and Dryaderces to delimit Operational Taxonomic Units (OTUs). We subsequently built a mitogenomic matrix (complete for 17 OTUs) to reconstruct a time-calibrated phylogeny and estimate their ancestral geographic distribution, in order to imply diversification processes.
Materials and methods
DNA-based species delimitation
We analysed a 16S rRNA fragment to identify major mitochondrial DNA (mtDNA) lineages that may represent species (i.e. a general lineage species concept; de Queiroz, 1999, 2007). Thus, this study does not represent an integrative species delimitation. The 16S locus has been widely used in amphibian systematics (Vences et al., 2005) and, for our focal clade, sequences were available from previous studies encompassing most taxa and regions (e.g. Jungfer et al., 2013; Vacher et al., 2020). We generated new 16S sequences from recent field collections (n = 52) (laboratory protocols are in Electronic Supplement 1), which were aligned with available GenBank sequences (n = 505), thus resulting in 16S sequences from 557 individuals (Osteocephalus: n = 508; Tepuihyla: n = 34; and Dryaderces: n = 15). This dataset included sequences of 34 out of 40 described species (Frost, 2022; Ortiz & Ron, 2018) and encompassed the entire geographic range for the three genera. GenBank accession numbers and associated information to all analysed sequences, including sources, are available in Electronic Supplement 2. The entire dataset was aligned using the E-INS-i strategy in MAFFT 7 (Katoh & Standley, 2013). The alignment was manually reviewed and its extremes were trimmed in Aliview 1.26 (Larsson, 2014) to keep the majority of homologous nucleotides (583 bp) among individuals.
The species delimitation was undertaken using three single-locus methods (ABGD, mPTP and GMYC). The Automatic Barcode Gap Discovery (ABGD) is a distance-based method which infers a ‘gap’ putatively existing between intra- and interspecific genetic distances to partition the dataset (Puillandre et al., 2012). The analysis was conducted using a prior of intraspecific divergence (K80) of 0.1–10% (P = 0.001–0.1), X (relative gap width) = 1, and number of steps = 30 (Réjaud et al., 2020). The analysis was performed on the online web server (available at https://bioinfo.mnhn.fr/abi/public/abgd/abgdweb.html). We selected the 11th recursive partition (P = 0.0049) in the ranking of distances, which corresponds to the end of a plateau for a number of delimited groups ‘species’ and thus, it was considered stable (Puillandre et al., 2012). Partitions with ‘P’ values higher than this resulted in very few groups and the merging of many recognised species.
The multi-rate Poisson Tree Processes (mPTP) is a tree-based method that incorporates different levels of intraspecific divergence without input thresholds (Kapli et al., 2017; Zhang et al., 2013a). We first reconstructed a maximum likelihood tree in RAxML 8.2 (Stamatakis, 2014) using one partition, 1000 iterations, and the GTRCAT approximation. The outgroup was defined in the GMYC analysis (see below). After removing the outgroup, we performed mPTP using 50 million iterations of Markov chain Monte Carlo (MCMC), sampling every 100,000 iterations, and discarding an initial 10% as burn-in.
The General Mixed Yule Coalescent (GMYC) is also a tree-based method that classifies the branches as interspecific (using a Yule model of constant speciation rate and no extinction) or intraspecific (modelled using a neutral coalescent process) (Monaghan et al., 2009; Pons et al., 2006). We first reconstructed an ultrametric phylogeny in BEAST 2.6 (Bouckaert et al., 2019) to estimate the timing of diversification events using a birth–death model and the uncorrelated relaxed clock lognormal of rate variation among branches (Drummond et al., 2006). The substitution model for 16S (one partition) was defined using the Bayesian Information Criterion (BIC) as implemented in PartitionFinder 2.1 (Lanfear et al., 2016). We used Trachycephalus coriaceus Peters, 1867 as an outgroup to constrain the root age of the tree with a normal prior distribution. This secondary calibration corresponds to the time to the crown age of the most recent common ancestor (TMRCA) between Osteocephalus and Trachycephalus (23.52 Mya; 15.1–32.5), an estimation based on 95 nuclear genes (~88,000 bp) and relevant fossil calibration (Feng et al., 2017). We ran four independent chains on the CIPRES online cluster with 250 million MCMC steps, 10,000 generations of thinning, and discarding an initial 10% as burn-in. The sampling adequacy (ESS > 200) and convergence of parameters among independent runs were verified in Tracer 1.7 (Rambaut et al., 2018). We combined and re-sample the independent tree files (excluding the initial 10% as burn-in) in LogCombiner 2.6 (Bouckaert et al., 2019) resulting in a file with 9000 trees. From this file, we extracted the Maximum Clade Credibility (MCC) tree in TreeAnnotator 2.6 (Bouckaert et al., 2019) excluding an additional initial 10% as burn-in. After removing the outgroup, we performed GMYC on the ultrametric tree using ‘splits 1.0’ R-package (Ezard et al., 2009) within R 4.0.2 (R Core Team, 2020), an interval of 1–10 Mya, and the single-threshold method (Fujisawa & Barraclough, 2013).
Operational Taxonomic Units (OTUs) were defined by applying a majority rule consensus to the results of the three delimitation methods (i.e. a cluster of sequences corresponding to an independent OTU when recovered as such in at least two of the three methods). We then assigned the OTUs to known taxa based on a priori sequence identification, type locality and known geographic range (Frost, 2022). In addition, we calculated uncorrected p-distances (mean and range) for 16S among OTUs in MEGA X (Kumar et al., 2018).
Dated mitogenomic phylogeny
We selected 50 terminals to be represented in a phylogenetic reconstruction based on mitogenomic data. Forty-two of these terminals correspond to the defined OTUs from the 16S species delimitation (see Results). However, four of these OTUs clustered sequences of 2–4 recognised species each. Because these species display diagnosable external morphology and other evidence lines supporting their distinctiveness, we followed the current taxonomy and added six corresponding terminals to the mitogenomic matrix, totalling 48 terminals. Finally, terminals for two recognised taxa without 16S sequences and thus not included in the delimitation (Osteocephalus sangay Chasiluisa et al., 2020 and Tepuihyla shushupe Ron et al., 2016) were also added to the mitogenomic matrix (see justification on inclusion of additional terminals in Appendix). Our mitogenomic matrix included 36 out of 40 currently recognised species (90%), with the absence of Osteocephalus duellmani Jungfer, 2011; Osteocephalus germani Ron et al., 2012; Osteocephalus melanops Melo-Sampaio et al., 2021; and Tepuihyla luteolabris Ayarzagüena et al., 1993a.
We performed complete mitogenome sequencing for 16 of the 50 terminals, with these 16 terminals evenly distributed across the phylogeny. We used 200 ng of genomic DNA to generate high-quality mitogenomic assemblies through low-coverage shotgun sequencing. For library preparation, we used the Illumina TruSeq Nano DNA Sample Prep kit with supplier instructions at the Génopole facility (Toulouse, France). Genomic DNA was fragmented by sonication, then fragments were selected by size (50–400 bp), adenylated and ligated to indexed sequencing adapters. Eight cycles of polymerase chain reaction (PCR) were performed to amplify the libraries. We quantified and validated the libraries, which were then multiplexed and sequenced on one lane of an Illumina Hiseq3000 flow cell (Illumina Inc.). Finally, assembly of reads was conducted in ORGanelle ASseMbler (Boyer et al., 2016), following assembling and annotation steps described in Réjaud et al. (2020) and using as mitogenome reference Osteocephalus taurinus Steindachner, 1862 (GenBank accession: JX564881; Zhang et al., 2013b). For the remaining 34 terminals without complete mitogenomes, we gathered available GenBank accessions across multiple mitochondrial genes (12S, 16S, ND1, ND2, COI and CytB; see Electronic Supplement 2). Finally, we generated a complete mitogenome for the outgroup Trachycephalus coriaceus.
We extracted the non-coding (12S and 16S rRNAs) and all coding (CDS) regions considering the reading frame (therefore removing D-Loop and tRNAs) using Geneious v5.4 (https://www.geneious.com). We aligned each gene independently in MAFFT 7 using the strategies: E-INS-i for rRNA genes (multiple conserved domain and long gaps); and G-INS-i for protein-coding genes (global homology) (Katoh et al., 2019). Finally, we concatenated the independent alignments in Geneious v5.4. Our final alignment consisted of 51 terminals and 14,098 characters (50% of missing data): 17 terminals including the outgroup had complete mitogenome (~14,000 bp), and 34 terminals had on average 3555 bp (1176–5155). We predefined four partitions: one for combined 12S and 16S rRNAs, and one for each codon position of concatenated protein-coding genes. We estimated the best-fit substitution model for each partition using Bayesian Information Criteria (BIC) as implemented in PartitionFinder 2.1.
We reconstructed a time-calibrated phylogeny using BEAST 2.6. Initially, we used a similar tree model setting as described for GMYC delimitation method (see above); however, some inconsistencies were detected on the parameters’ posterior, probably caused because of the large CI of our calibration prior. Therefore, we used the Calibrated Yule tree model instead as this allows a proper sampling when a single calibration point is used (Heled & Drummond, 2012), as in our case. In absence of relevant fossils for calibration of our focal clade, we constrained the tree root prior to using the secondary calibration node estimated by Feng et al. (2017): TMRCA Osteocephalus-Trachycephalus (mean = 23.52 Mya, SD = 3.6 Mya, and normal distribution). Each partition was parametrized using the substitution model informed by PartitionFinder 2.1, but excluding the parameter of invariant sites proportion (Jia et al., 2014; Stamatakis, 2016). We used a linked, uncorrelated relaxed clock lognormal of rate variation among branches for all partitions (Drummond et al., 2006; Duchêne et al., 2020). MCMC parametrization in BEAST, runs assessment, and obtention of the MCC tree was similar as described for GMYC method (see above), except that each chain ran 100 million steps.
In addition, using the same file with 9000 trees (mitogenomic analysis) resulting from LogCombiner 2.6, we constructed a lineage through time (LTT) plot in ‘ape 5.5’ R-package (Paradis & Schliep, 2019) to explore changes in lineage accumulation over time. Finally, we conducted a maximum likelihood analysis on the mitogenomic matrix in RAxML 8.2 (Stamatakis, 2014), using 1000 iterations, and the GTRCAT approximation for the four partitions combined in a single tree.
Ancestral area reconstruction
We estimated the ancestral geographic range for the genera based on the mitogenomic phylogeny using ‘BioGeoBEARS 1.1.2’ R-package (Matzke, 2013). We coded the current distribution of our ingroup terminals using three areas: western Amazonia, the Guiana Shield and Brazilian Shield. These areas differ in geological history (Hoorn & Wesselingh, 2010), and are well-recognised biogeographically (Godinho & da Silva, 2018; Oliveira et al., 2017). The areas are approximately delimited by the rivers: Negro, Madeira and the lower course of the Amazon (see Electronic Supplement 3). The analysis compares the likelihood of several models of ancestral state reconstruction and their fit to the data, weighted by Akaike’s information criterion (AICc; Akaike, 1973). Six models were evaluated: Dispersal-Extinction Cladogenesis (DEC) model, a likelihood version of the Dispersal-Vicariance Analyses (DIVALIKE), and a Bayesian range-evolution model (BAYAREALIKE). The other three models correspond to modifications of the former ones by adding a parameter of founder-event speciation (+ J). We included these latter three models in the model selection in face of recent debate about using this parameter (Klaus & Matzke, 2020; Ree & Sanmartín, 2018).
Results
Species delimitation (16S)
There were considerable differences in the estimated number of groups ‘species’ found among the three methods: mPTP = 16, ABGD = 42 and GMYC = 159. The mPTP result (16 groups) was the most conservative, but 2–7 recognised, morphologically diagnosable species clustered together in some of these groups. This meant that the mPTP result failed to detect 19 described species. The ABGD result (42 groups) was more in agreement with current taxonomy, with 25 groups correctly assigned each to a described species, four groups contained 2–4 recognised species (thus failing to detect five described species and one paraphyletic lineage of ‘Osteocephalus cabrerai’ Cochran & Goin, 1970 from the Guiana Shield), and 13 groups corresponded to non-described species. Therefore, the number of recognised species that failed into being detected was approximately four times higher in mPTP than in ABGD. The GMYC method was deemed to over-split the dataset (159 groups). Nineteen described species were identified as single groups in GMYC result, but 15 species were formed by two or more groups. Therefore, GMYC method overall identified almost every clade in the tree as a different group. The majority rule consensus over the three methods was consistent with the intermediate result provided by ABGD, delimiting 42 OTUs (Fig. 1a). Results of each method and consensus are in Electronic Supplements 2 and 4.
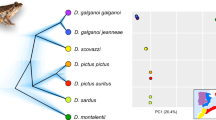
Panel a, time-calibrated phylogeny of Osteocephalus, Tepuihyla and Dryaderces in Amazonia, resulting from a mitogenomic analysis in BEAST and displaying the mean and 95% CI (blue bars) of divergences. Nodes are numbered in red (age details are in Electronic Supplement 1), and support values are depicted for Bayesian (following node symbol colours) and maximum likelihood analyses (following numbers below the nodes). Topological discrepancies between analyses for four low-supported nodes (21, 25, 34 and 43) are illustrated with grey dashed arrows. Terminals with complete mitogenome (~14,000 bp) are represented with asterisks, including the outgroup Trachycephalus coriaceus. Majority rule consensus results of 16S species delimitation are next to the terminals (each box corresponds to an Operational Taxonomic Unit ‘OTU’). Terminals are grouped according to genera or species groups (gr.) within Osteocephalus. Panel b, lineage through time plot for the ingroup depicting mean accumulation of lineages and 95% CI
A taxonomic account describing the correspondence between the 42 OTUs and known taxa is in the Appendix, including a detailed rationale for the addition of eight terminals to the mitogenomic matrix, thus leading to 50 putative species. Out of these, 14 OTUs could not be associated to any known taxon and were labelled as ‘aff.’ relative to their closest taxon in the mitogenomic phylogeny. The geographic distribution of all 50 putative species is illustrated in Electronic Supplement 3. The mean uncorrected p-distances for 16S among the 50 putative species, calculated within eight groups (the genera Dryaderces and Tepuihyla, and six Osteocephalus species groups; Electronic Supplement 1), was > 3% for the majority of the comparisons (57%). The rest of comparisons had distances between 1 and 3% (38%), or < 1% (5%).
Dated mitogenomic phylogeny
Thirty-nine out of 50 nodes in the Bayesian phylogeny were highly supported (~80%), displaying posterior probabilities (PP) \(\ge\) 0.95 (Fig. 1a). Nodes with PP < 0.9 were mostly bracketed by short branches and occurred between clades/terminals with and without complete mitogenome. The topology and bootstrap support (BS) of the maximum likelihood (ML) phylogeny was overall consistent with the Bayesian phylogeny, except for four low-supported nodes whose positions varied between analyses (Fig. 1a). One of these corresponds to the ‘O. mimeticus group’, which is the sister group of the O. buckleyi group (in Bayesian), but sister to the O. leprieurii group (in ML). A similar discrepancy in the position of this lineage (i.e. Osteocephalus mimeticus Melin, 1941) was reported by Blotto et al. (2020). We have considered the clade containing Osteocephalus mimeticus and O. aff. mimeticus as a new species group (‘O. mimeticus group’), and not as part of the O. buckleyi group as suggested by Jungfer et al. (2013), because it displays age of divergence similar to the other five traditional species groups (mean \(\ge\) 10 Mya).
Our phylogeny (Fig. 1a) is generally consistent with previously proposed topologies (Blotto et al., 2020; Duellman et al., 2016; Jungfer et al., 2013; Kok et al., 2015; Ron et al., 2016; Salerno et al., 2012). However, it differs on three major aspects that could be explained by the inclusion of more mtDNA data: (1) Osteocephalus is sister to Tepuihyla with high support (PP = 0.95; BS = 76), instead of Dryaderces according to previous studies; (2) most internal nodes within Tepuihyla are highly supported, and show that T. exophthalma diverged first in the genus and that Tepuihyla warreni Duellman & Hoogmoed, 1992 is sister to Tepuihyla tuberculosa Boulenger, 1882 + T. shushupe (similar to the ML topology of Blotto et al. (2020), although this inference was considered secondary therein and this relationship was not supported); and (3) internal nodes among Osteocephalus species groups are highly supported (except for the ‘O. mimeticus group’ position).
Dryaderces is estimated to have diverged from Tepuihyla + Osteocephalus 19.5 Mya (11.8–26.9); and Tepuihyla from Osteocephalus 17.4 Mya (10.4–24.2). Crown ages of the three genera are as follows: Osteocephalus (13.1 Mya (7.6–18.4)), Dryaderces (11.3 Mya (5.6–17.6)) and Tepuihyla (9.8 Mya (5.3–14.7)). Age information for all nodes depicted in Fig. 1a is in Electronic Supplement 1. Lineage through time plot shows a constant lineage accumulation (Fig. 1b).
Ancestral area reconstruction
Consistent results of ancestral areas were uncovered across models, with the few exceptions being around recent nodes (see Electronic Supplement 5). The best-fit model was DEC (Fig. 2) (model selection is in Electronic Supplement 1). However, other models supporting alternative states at particular nodes are mentioned when relevant. Without the inclusion of other Lophiohylinae genera, the ancestral area for the entire focal clade remains ambiguous within Amazonia (Fig. 2a).
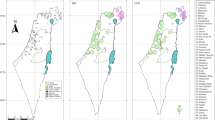
Panel a, ancestral area reconstruction for the diversification of Osteocephalus, Tepuihyla and Dryaderces in Amazonia, resulting from the best-fit model (DEC) in BioGeoBEARS and based on the mitogenomic phylogeny (Fig. 1a) and three areas (western Amazonia ‘W’, Guiana Shield ‘G’ and Brazilian Shield ‘B’). Species are grouped according to the three genera and six Osteocephalus species groups (gr.), with their generalised reproductive mode. Species displaying a reproductive mode being the exception within their groups are depicted with asterisks (O. vilarsi apparently displays both modes within its group; Ferrão et al., 2019). Two time slices show the hypothetic, ancestral distribution of the lineages (coloured polygons) at 16 Mya (panel b) and 9 Mya (panels c–e) according to DEC model and constrained by the extent of the Pebas and Acre mega wetlands. In panels b–e, the current distribution of the lineages is also plotted from individual 16S records; the species are combined in genera or species groups (see the distribution of each species in Electronic Supplement 2 and 3). An arrow indicates the flow direction of the Amazon River, and at the right bottom of the continent is the time span for that particular landscape configuration following Hoorn et al. (2010)
The ancestors of each genus diverged during the early-middle Miocene, each originating in a distinct Amazonian area (Osteocephalus: western Amazonia, Tepuihyla: the Guiana Shield and Dryaderces: Brazilian Shield; Fig. 2a, b). From these initial splits until ~10 Mya the diversification of each genus took place within each area, except for Tepuihyla which also included western Amazonia (Fig. 2a, c). However, other models indicate that the ancestral area of Tepuihyla crown age was predominantly the Guiana Shield (Electronic Supplement 5). In contrast, the inferred area for the ancestor of T. warreni (T. tuberculosa + T. shushupe) included consistently western Amazonia and the Guiana Shield in all models (Fig. 2a; Electronic Supplement 5), indicating that this node corresponds to the minimum age of colonization of Tepuihyla from the Guiana Shield to western Amazonia (mean 8 Mya).
From ~10 Mya to the present, the diversification of each genus continued mostly within their respective ancestral areas. In Osteocephalus, the inferred areas for all six species groups during ~5–10 Mya indicate that they co-occurred in western Amazonia (Fig. 2a, d–e). Posteriorly, the diversification for three out of the six species groups continued essentially within western Amazonia (O. alboguttatus, O. planiceps and ‘O. mimeticus’ groups). In contrast, the O. buckleyi, O. leprieurii and O. taurinus groups dispersed out of western Amazonia relatively recently (last ~6 My), and diversified in the rest of Amazonia (Fig. 2a). Each of these latter three groups currently contain species restricted to one of the three Amazonian areas, and a widespread, pond-breeding species occurring in all areas (Fig. 2a; Electronic Supplement 3). These dispersals out of western Amazonia are seemingly not synchronous among the three groups (Fig. 2a). The nodes associated to all dispersals display some ambiguity on the inferred ancestral areas in DEC model (Fig. 2a) and across models (see comments in Electronic Supplement 1). Combined, the models suggest that the dispersion sequence of Osteocephalus was as follows: O. taurinus group first, from western Amazonia to the Guiana Shield and subsequently to Brazilian Shield; O. buckleyi group second, from western Amazonia to Brazilian Shield and subsequently to Guiana Shield; and the terminal of Osteocephalus leprieurii Duméril & Bibron, 1841 third, from western Amazonia to the Guiana Shield or Brazilian Shield equally likely.
Discussion
Species diversity
Our final delimitation (n = 50 putative species), together with four missing taxa in our sampling (O. duellmani, O. germani, O. melanops and T. luteolabris), indicates that the three genera may harbour 54 species. The 14 unidentified OTUs (candidate species) represent a 35% increase in the current known diversity for the three genera combined (40 currently valid nominal species; Frost, 2022; Ortiz & Ron, 2018). The ratio of potential undocumented diversity in our focal clade (35%) is relatively low compared to what has been found in other widespread Neotropical frog groups using 16S (e.g. Fouquet et al., 2014, 2021a; Gehara et al., 2014; Vacher et al., 2020). The main reason for this lower ratio probably comes from better taxonomic knowledge because this clade has attracted more attention in recent years (Blotto et al., 2020; Chasiluisa et al., 2020; Jungfer et al., 2013; Moura et al., 2021; Ron et al., 2012, 2016).
Although our DNA-based delimitation is mostly based on a single and short fragment of 16S mtDNA, and thus is potentially prone to fail to indicate the existence or absence of species (e.g. Hickerson et al., 2006), most of the initially found 42 OTUs were readily confirmed by their correspondence with current taxonomy (Appendix). However, our results suggest that four of these OTUs included more than one nominal species since they were differentiated by their morphology and advertisement calls. This lack of accuracy is due to the combination of the small size of our locus and the recent divergence among some species (Fig. 1). Our results also corroborate that phenotypically recognised species with low 16S distances (< 3%) are frequent in this group (Jungfer et al., 2013).
Further investigations using more variable markers, including nuclear DNA (nDNA), and phenotypic data are needed to clarify the status of the 14 unidentified OTUs and that of widespread ‘species’ that may represent species complexes (i.e. Osteocephalus helenae Ruthven, 1919, O. leprieurii and O. taurinus; see Appendix; Jungfer et al. 2013). Finally, vast under-sampled regions remain in Amazonia and Pantepui regions, notably in Colombia and Venezuela (see Electronic Supplement 3), which may harbour undocumented species and extend the distribution of the known species. Therefore, although this study does not represent a thoroughly systematic review, it provides an exploratory assessment of diversity in this clade that will contribute to more detailed, integrative studies.
Diversification around the Pebas mega wetland
Our results suggest that the three genera diverged, and were isolated from each other, for the period of ~10–20 Mya (early–middle Miocene). This period coincides with the existence of the Pebas mega wetland system, and the ancestral area of each genus (as well as the bulk of their subsequent diversification) matches the putative distribution of suitable rainforest areas during this period (Fig. 2a, b). Therefore, this major hydrological barrier coincides with the deep spatio-temporal pattern of divergence in our clade. Additional support for the hypothesis that Miocene mega wetlands represented a major barrier preventing dispersals and diversification in western Amazonia comes from the fact that the three genera started to diversify during the last stages of the Pebas system (the period that preceded the onset of the transcontinental Amazon River, trigged by intensified Andean uplift). These early diversifications within genera (~10–13 Mya; Fig. 1a) remained mostly in situ, except for Tepuihyla which may have dispersed westwards around this period following the disappearance of the barrier once formed by the northward-drained Pebas (Fig. 2b, c). This timeframe was also relevant for the highest diversification period of rocket frogs (Allobates) in western Amazonia (10–14 Mya; Réjaud et al., 2020), suggesting shared patterns of isolation and diversification.
In Osteocephalus, the divergence of the ancestors of all six species groups (gr.) was probably driven by the availability of new and diverse rainforest environments along Andean slopes and nearby lowlands following the retraction of mega wetlands in western Amazonia (Hoorn et al., 2010). Also, the concomitant evolution of new reproductive modes (phytotelmata- and stream-breeding) from a probable pond-breeding ancestor (Jungfer et al., 2013), may at least partly explain their rapid diversification and current co-occurrence in western Amazonia (Fig. 2d–e). According to our phylogeny, the evolution of phytotelmata-breeding from pond-breeding happened in at least two instances in Osteocephalus (in the ancestor of O. alboguttatus gr. + O. planiceps gr. 10.4 Mya, and more recently in O. oophagus within the O. taurinus gr. < 2.6 Mya), and once in Tepuihyla (in the ancestor of T. tuberculosa + T. shushupe 3.8 Mya) (Fig. 2a). On the other hand, the evolution of stream-breeding from pond-breeding appeared once in Osteocephalus, in the ancestor of O. buckleyi gr. + ‘O. mimeticus gr.’ 9.8 Mya (Fig. 2a).
Other frog groups have similarly experienced transitions in reproductive mode during the Miocene in western Amazonia and Andes. Chiasmocleis antenori Walker, 1973 (Microhylidae), and presumably its closest relatives, transitioned from the generalised pond-breeding mode of Chiasmocleis to phytotelmata-breeding in bromeliads (de Sá et al., 2019; Peloso et al., 2014). The common ancestor of Dendrobatinae (Ranitomeya and relatives) transitioned from the generalised stream-breeding mode of dendrobatids to phytotelmata-breeding in bromeliads (Grant et al., 2006; Santos et al., 2009). Other Andean-derivate groups which diversified in the Miocene are strict stream-breeders (with some exceptions within Dendrobatidae) suggesting that their diversification was associated to Andean uplift and consequential availability of streams (e.g. Atelopus in Bufonidae (Lötters et al., 2011; Ramírez et al., 2020); Centrolenidae (Castroviejo‐Fisher et al., 2014); Dendrobatidae (Santos et al., 2009); Hyloscirtus in Hylidae (Coloma et al., 2012; Duellman et al., 2016)). Therefore, we hypothesize that intensified Andean uplift around 10 Mya (and subsequently) might have provided new reproductive habitats that led to ecological divergence in Osteocephalus along the Andean slopes and western Amazonia, to utilize varied stream habitats and water-filled cavities of bromeliads (which radiated extensively in the northern Andes; Givnish et al., 2014; Zizka et al., 2020).
Diversification and dispersion throughout the modern Amazon River drainage
The lineage accumulation for the three genera appears to have been constant (Fig. 1b), and most speciation events have occurred in situ with only a few trans-Amazonian expansions. The onset of the modern Amazon River resulted in the Pebas mega wetland being first reduced in north-western Amazonia, compared to the south-western region that remained vastly flooded as the Acre system (7–10 Mya; Hoorn et al., 2010). This large area probably remained unsuitable since it seems to have also prevented dispersals (see below). This suggests that the first connection for non-flying, rainforest organisms between the three major areas would have been established between western Amazonia and the Guiana Shield, while the Brazilian Shield remained more isolated from the rest. This is supported by Dryaderces that did not disperse out of the Brazilian Shield. Also, despite Dryaderces was the first genus to diverge, most of its speciation occurred relatively recently in the lowlands (last ~3 My), with the exception of the deep divergence of Dryaderces pearsoni Gaige, 1929 (~11.3 Mya) which is the only Dryaderces species known to occur close to the Andes (Electronic Supplement 3). The relatively low and recent diversification of Dryaderces in the Brazilian Shield may be related to the significant climate instability of the eastern Amazonia during the Pleistocene (e.g. Cheng et al., 2013; Silva et al., 2019; Wang et al., 2017).
Assessing diversification within Osteocephalus is particularly biogeographically interesting, given the diversity, distributional patterns and variation in breeding modes in this genus. Our results suggest that the six Osteocephalus species groups formed \(\ge\) 10 Mya in western Amazonia. Three of these dispersed and diversified eastwards relatively recently (since ~6 Mya; Fig. 2a). These dispersions occurred in one lineage in each of these groups and do not seem to have been synchronous (Fig. 2a). The O. taurinus gr. probably dispersed first towards the Guiana Shield, supporting the relatively early connection between western Amazonia and the Guiana Shield (as discussed above for Tepuihyla). On the other hand, expansion of terra firme rainforest after the Acre system disappeared (Albert et al., 2018) may explain the more recent dispersal of the O. buckleyi gr. from western Amazonia towards the Brazilian Shield (compared to that of the O. taurinus gr.). Nevertheless, the dispersion and diversification of the O. buckleyi gr. were not extensive in the latter area (Electronic Supplement 3).
The Osteocephalus groups that contain few and widespread species in the three major Amazonian areas display the ‘ancestral’ pond-breeding mode (O. leprieurii within O. leprieurii gr., O. taurinus within O. taurinus gr., and O. helenae within the O. buckleyi gr.; Fig. 2a) (O. helenae may represent a recent transition back to the pond-breeding state because it reproduces in ponds associated to black-water flooded forest (igapó) on the margin of rivers). In contrast, the groups that diversified in western Amazonia and the Andean foothills (O. alboguttatus gr., O. planiceps gr., ‘O. mimeticus gr.’ and most of O. buckleyi gr.) have high species diversity, species with relatively small distributions (Electronic Supplement 3), and ‘derived’ reproductive modes (i.e. phytotelmata- or stream-breeding; Fig. 2a). This pattern of distribution and reproductive mode suggests that the potential large availability of ponds across Amazonian lowlands might have facilitated range expansions in pond-breeding species, whereas phytotelmata- and stream-breeding habitat may have been relatively more restricted.
Resolving the details of dispersal and speciation events during the Pliocene and Pleistocene (last 5.3 My) is complicated because of spatio-temporal uncertainties over the formation of the large tributaries of the Amazon River (Albert et al., 2018), whether or not these acted as dispersal barriers for these frogs, and the possibility of river captures (Pupim et al., 2019; Ruokolainen et al., 2019). Nevertheless, the high diversification within Osteocephalus suggests that climatic stability in western Amazonia and steep environmental gradients along the Andean foothills (including their effects on stream habitats and bromeliad diversity and abundance) probably played a significant role in its diversification (Cheng et al., 2013; Ron et al., 2012).
Conclusions
We assessed the diversity and historical biogeography of a diverse, Amazonia-wide clade of tree frogs. The dense sampling of 16S sequences obtained from this and previous studies, combined with our mitogenomic reconstruction, led to a comprehensive phylogenetic and biogeographic assessment of Osteocephalus, Tepuihyla and Dryaderces. Our results suggest that the genera diverged and diversified in isolation around Miocene mega wetlands through most of the Neogene and Quaternary. Only Osteocephalus experienced a spectacular radiation along Andean foothills and western Amazonia including the appearance of new reproductive modes, as well as few relatively recent, trans-Amazonian dispersals in pond-breeding species. Nevertheless, the diversification within genera < 10 Mya needs to be further studied incorporating genomic and phenotypic data. These approaches will be particularly relevant to unveil cryptic diversity in widespread ‘species’. Overall, our study provides a spatio-temporal framework for future research on the evolution and systematics of these tree frogs, and adds evidence to the biogeographic understanding of arguably the world’s most biodiverse region.
Data availability
Information associated to all analysed DNA sequences is available in Electronic Supplement 2. Newly generated DNA sequences were deposited in GenBank under accessions: ON716130–65 (16S rRNA), and OP425862, OP572168–92 (mitogenomes). Final alignments are available in TreeBASE (http://purl.org/phylo/treebase/phylows/study/TB2:S29696).
Code availability
Not applicable.
References
Akaike, H. (1973). Information theory and an extension of the maximum likelihood principle. In B. N. Petrov & F. Csaki (Eds.), Proceedings of the 2nd international symposium on information theory (pp. 267–281). Budapest: Akademiai Kiado.
Albert, J. S., Val, P., & Hoorn, C. (2018). The changing course of the Amazon River in the Neogene: Center stage for Neotropical diversification. Neotropical Ichthyology, 16(3), e180033. https://doi.org/10.1590/1982-0224-20180033
Antonelli, A., Zizka, A., Carvalho, F. A., Scharn, R., Bacon, C. D., Silvestro, D., & Condamine, F. L. (2018). Amazonia is the primary source of Neotropical biodiversity. Proceedings of the National Academy of Sciences, 115(23), 6034–6039. https://doi.org/10.1073/pnas.1713819115
Ayarzagüena, J., Señaris, J. C., & Gorzula, S. J. (1993a). El grupo Osteocephalus rodriguezi de las tierras altas de la Guayana Venezolana: Descripción de cinco nuevas especies. Memoria De La Sociedad De Ciencias Naturales La Salle, 52, 113–142.
Ayarzagüena, J., Señaris, J. C., & Gorzula, S. J. (1993b). Un nuevo género para las especies del “grupo Osteocephalus rodriguezi” (Anura: Hylidae). Memoria De La Sociedad De Ciencias Naturales La Salle, 52, 213–221.
Bicudo, T. C., Sacek, V., de Almeida, R. P., Bates, J. M., & Ribas, C. C. (2019). Andean tectonics and mantle dynamics as a pervasive influence on Amazonian ecosystem. Scientific Reports, 9, 16879. https://doi.org/10.1038/s41598-019-53465-y
Blotto, B. L., Lyra, M. L., Cardoso, M. C. S., Rodrigues, M. T., Dias, I. R., Marciano-Jr, E., et al. (2020). The phylogeny of the casque-headed treefrogs (Hylidae: Hylinae: Lophyohylini). Cladistics, 37(1), 36–72. https://doi.org/10.1111/cla.12409
Boubli, J. P., Ribas, C., Lynch Alfaro, J. W., Alfaro, M. E., da Silva, M. N. F., Pinho, G. M., & Farias, I. P. (2015). Spatial and temporal patterns of diversification on the Amazon: A test of the riverine hypothesis for all diurnal primates of Rio Negro and Rio Branco in Brazil. Molecular Phylogenetics and Evolution, 82, 400–412. https://doi.org/10.1016/j.ympev.2014.09.005
Bouckaert, R., Vaughan, T. G., Barido-Sottani, J., Duchêne, S., Fourment, M., & Gavryushkina, A., et al. (2019). BEAST 2.5: An advanced software platform for Bayesian evolutionary analysis. PLOS Computational Biology, 15(4), e1006650. https://doi.org/10.1371/journal.pcbi.1006650
Boulenger, G. A. (1882). Catalogue of the Batrachia Salientia s. Ecaudata in the collection of the British museum (2nd ed.). Taylor and Francis.
Boyer, F., Coissac, E., & Viari, A. (2016). The ORGanelle ASseMbler. https://pypi.org/project/ORG.asm/
Byrne, H., Lynch Alfaro, J. W., Sampaio, I., Farias, I., Schneider, H., Hrbek, T., & Boubli, J. P. (2018). Titi monkey biogeography: Parallel Pleistocene spread by Plecturocebus and Cheracebus into a post-Pebas Western Amazon. Zoologica Scripta, 47(5), 499–517. https://doi.org/10.1111/zsc.12300
Castroviejo-Fisher, S., Guayasamin, J. M., Gonzalez-Voyer, A., & Vilà, C. (2014). Neotropical diversification seen through glassfrogs. Journal of Biogeography, 41(1), 66–80. https://doi.org/10.1111/jbi.12208
Chasiluisa, V. D., Caminer, M. A., Varela-Jaramillo, A., & Ron, S. R. (2020). Description and phylogenetic relationships of a new species of treefrog of the Osteocephalus buckleyi species group (Anura: Hylidae). Neotropical Biodiversity, 6(1), 21–36. https://doi.org/10.1080/23766808.2020.1729306
Cheng, H., Sinha, A., Cruz, F. W., Wang, X., Edwards, R. L., d’Horta, F. M., et al. (2013). Climate change patterns in Amazonia and biodiversity. Nature Communications, 4, 1411. https://www.nature.com/articles/ncomms2415
Cochran, D. M., & Goin, C. J. (1970). Frogs of Colombia. Bulletin of the United States National Museum, 288, 1–655.
Coloma, L. A., Carvajal-Endara, S., Dueñas, J. F., Paredes-Recalde, A., Morales-Mite, M., Almeida-Reinoso, D., et al. (2012). Molecular phylogenetics of stream treefrogs of the Hyloscirtus larinopygion group (Anura: Hylidae), and description of two new species from Ecuador. Zootaxa, 3364, 1–78.
Cope, E. D. (1874). On some Batrachia and Nematognathi brought from the upper Amazon by Prof. Orton. Proceedings of the Academy of Natural Sciences of Philadelphia, 26, 120–137.
Cracraft, J., Ribas, C. C., d’Horta, F. M., Bates, J., Almeida, R. P., Aleixo, A., et al. (2020). The origin and evolution of Amazonian species diversity. In V. Rull & A. C. Carnaval (Eds.), Neotropical Diversification: Patterns and Processes (pp. 225–244). Cham: Springer. https://doi.org/10.1007/978-3-030-31167-4_10
de Queiroz, K. (1999). The general lineage concept of species and the defining properties of the species category. In R. A. Wilson (Ed.), Species: New interdisciplinary essays (pp. 48–89). MIT Press.
de Queiroz, K. (2007). Species concepts and species delimitation. Systematic Biology, 56(6), 879–886. https://doi.org/10.1080/10635150701701083
de Sá, R. O., Tonini, J. F. R., van Huss, H., Long, A., Cuddy, T., Forlani, M. C., et al. (2019). Multiple connections between Amazonia and Atlantic Forest shaped the phylogenetic and morphological diversity of Chiasmocleis Mehely, 1904 (Anura: Microhylidae: Gastrophryninae). Molecular Phylogenetics and Evolution, 130, 198–210. https://doi.org/10.1016/j.ympev.2018.10.021
Dewynter, M., Marty, C., Courtois, E. A., Blanc, M., Gaucher, P., Martinez, Q., & Fouquet, A. (2016). L’identification des rainettes des genres Osteocephalus et Trachycephalus (Hylidae Lophyohylinae) en Guyane. Les Cahiers De La Fondation Biotope, 7, 1–16.
Drummond, A. J., Ho, S. Y. W., Phillips, M. J., & Rambaut, A. (2006). Relaxed phylogenetics and dating with confidence. PLOS Biology, 4(5), e88. https://doi.org/10.1371/journal.pbio.0040088
Duchêne, D. A., Tong, K. J., Foster, C. S. P., Duchêne, S., Lanfear, R., & Ho, S. Y. W. (2020). Linking branch lengths across sets of loci provides the highest statistical support for phylogenetic inference. Molecular Biology and Evolution, 37(4), 1202–1210. https://doi.org/10.1093/molbev/msz291
Duellman, W. E. (2019). The last one: A new species of Osteocephalus (Anura: Hylidae) from Colombia, with comments on the morphological and behavioral diversity within the genus. Phyllomedusa, 18(2), 141–157. https://doi.org/10.11606/issn.2316-9079.v18i2p141-157
Duellman, W. E., & Hoogmoed, M. S. (1992). Some hylid frogs from the Guiana highlands, northeastern South America: New species, distributional records, and a generic reallocation. Occasional Papers of the Museum of Natural History, University of Kansas, 147, 1–21.
Duellman, W. E., Marion, A. B., & Hedges, S. B. (2016). Phylogenetics, classification, and biogeography of the treefrogs (Amphibia: Anura: Arboranae). Zootaxa, 4104(1), 1–109. https://doi.org/10.11646/zootaxa.4104.1.1
Duméril, A. M. C., & Bibron, G. (1841). Erpétologie genérale ou histoire naturelle complète des reptiles (Vol. 8). Librarie Enclyclopedique de Roret.
Ezard, T., Fujisawa, T., & Barraclough, T. (2009). Splits: Species’ limits by threshold statistics. R package, Version 1.0.
Feng, Y. -J., Blackburn, D. C., Liang, D., Hillis, D. M., Wake, D. B., Cannatella, D. C., & Zhang, P. (2017). Phylogenomics reveals rapid, simultaneous diversification of three major clades of Gondwanan frogs at the Cretaceous-Paleogene boundary. Proceedings of the National Academy of Sciences, 114(29), E5864–E5870. https://doi.org/10.1073/pnas.1704632114
Ferrão, M., Moravec, J., Moraes, L. J. C. L., de Carvalho, V. T., Gordo, M., & Lima, A. P. (2019). Rediscovery of Osteocephalus vilarsi (Anura: Hylidae): An overlooked but widespread Amazonian spiny-backed treefrog. PeerJ, 7, e8160. https://doi.org/10.7717/peerj.8160
Fouquet, A., Loebmann, D., Castroviejo-Fisher, S., Padial, J. M., Orrico, V. G. D., Lyra, M. L., et al. (2012). From Amazonia to the Atlantic forest: Molecular phylogeny of Phyzelaphryninae frogs reveals unexpected diversity and a striking biogeographic pattern emphasizing conservation challenges. Molecular Phylogenetics and Evolution, 65(2), 547–561. https://doi.org/10.1016/j.ympev.2012.07.012
Fouquet, A., Cassini, C. S., Haddad, C. F. B., Pech, N., & Rodrigues, M. T. (2014). Species delimitation, patterns of diversification and historical biogeography of the Neotropical frog genus Adenomera (Anura, Leptodactylidae). Journal of Biogeography, 41(5), 855–870. https://doi.org/10.1111/jbi.12250
Fouquet, A., Courtois, E. A., Baudain, D., Lima, J. D., Souza, S. M., Noonan, B. P., & Rodrigues, M. T. (2015). The trans-riverine genetic structure of 28 Amazonian frog species is dependent on life history. Journal of Tropical Ecology, 31(4), 361–373. https://doi.org/10.1017/S0266467415000206
Fouquet, A., Leblanc, K., Framit, M., Réjaud, A., Rodrigues, M. T., Castroviejo-Fisher, S., et al. (2021a). Species diversity and biogeography of an ancient frog clade from the Guiana Shield (Anura: Microhylidae: Adelastes, Otophryne, Synapturanus) exhibiting spectacular phenotypic diversification. Biological Journal of the Linnean Society, 132(2), 233–256. https://doi.org/10.1093/biolinnean/blaa204
Fouquet, A., Marinho, P., Réjaud, A., Carvalho, T. R., Caminer, M. A., Jansen, M., et al. (2021b). Systematics and biogeography of the Boana albopunctata species group (Anura, Hylidae), with the description of two new species from Amazonia. Systematics and Biodiversity, 19(4), 375–399. https://doi.org/10.1080/14772000.2021.1873869
Fouquet, A., Cornuault, J., Rodrigues, M. T., Werneck, F. P., Hrbek, T., Acosta-Galvis, A. R., et al. (2022). Diversity, biogeography, and reproductive evolution in the genus Pipa (Amphibia: Anura: Pipidae). Molecular Phylogenetics and Evolution, 170, 107442. https://doi.org/10.1016/j.ympev.2022.107442
Frost, D. R. (2022). Amphibian species of the world: An online reference. Version 6.1. New York, USA: American Museum of Natural History. Electronic Database accessible at Retrieved date May 15, 2022, from https://amphibiansoftheworld.amnh.org/index.php
Fujisawa, T., & Barraclough, T. G. (2013). Delimiting species using single-locus data and the Generalized Mixed Yule Coalescent approach: A revised method and evaluation on simulated data sets. Systematic Biology, 62(5), 707–724. https://doi.org/10.1093/sysbio/syt033
Gaige, H. T. (1929). Three new tree-frogs from Panama and Bolivia. Occasional Papers of the Museum of Zoology, University of Michigan, 207, 1–6.
Gehara, M., Crawford, A. J., Orrico, V. G. D., Rodríguez, A., Lötters, S., Fouquet, A., et al. (2014). High levels of diversity uncovered in a widespread nominal taxon: Continental phylogeography of the Neotropical tree frog Dendropsophus minutus. PLoS ONE, 9(9), e103958. https://doi.org/10.1371/journal.pone.0103958
Givnish, T. J., Barfuss, M. H. J., Van Ee, B., Riina, R., Schulte, K., Horres, R., et al. (2014). Adaptive radiation, correlated and contingent evolution, and net species diversification in Bromeliaceae. Molecular Phylogenetics and Evolution, 71, 55–78. https://doi.org/10.1016/j.ympev.2013.10.010
Godinho, M. B. d. C., & da Silva, F. R. (2018). The influence of riverine barriers, climate, and topography on the biogeographic regionalization of Amazonian anurans. Scientific Reports, 8(1), 3427. https://doi.org/10.1038/s41598-018-21879-9
Gorzula, S., & Señaris, J. C. (1998). Contribution to the herpetofauna of the Venezuelan Guayana I A Data Base. Scientia Guaianae, 8, 1–270.
Grant, T., Frost, D. R., Caldwell, J. P., Gagliardo, R., Haddad, C. F. B., Kok, P. J. R., et al. (2006). Phylogenetic systematics of dart-poisson frogs and their relatives (Amphibia: Athesphatanura: Dendrobatidae). Bulletin of the American Museum of Natural History, 2006(299), 1–262. https://doi.org/10.1206/0003-0090(2006)299[1:PSODFA]2.0.CO;2
Heled, J., & Drummond, A. J. (2012). Calibrated tree priors for relaxed phylogenetics and divergence time estimation. Systematic Biology, 61(1), 138–149. https://doi.org/10.1093/sysbio/syr087
Hickerson, M. J., Meyer, C. P., & Moritz, C. (2006). DNA barcoding will often fail to discover new animal species over broad parameter space. Systematic Biology, 55(5), 729–739. https://doi.org/10.1080/10635150600969898
Hime, P. M., Lemmon, A. R., Lemmon, E. C. M., Prendini, E., Brown, J. M., Thomson, R. C., et al. (2021). Phylogenomics reveals ancient gene tree discordance in the amphibian tree of life. Systematic Biology, 70(1), 49–66. https://doi.org/10.1093/sysbio/syaa034
Hoorn, C., & Wesselingh, F. P. (Eds.). (2010). Amazonia: Landscape and species evolution. A look into the past. Wiley-Blackwell.
Hoorn, C., Wesselingh, F. P., ter Steege, H., Bermudez, M. A., Mora, A., Sevink, J., et al. (2010). Amazonia through time: Andean uplift, climate change, landscape evolution, and biodiversity. Science, 330(6006), 927–931. https://doi.org/10.1126/science.1194585
Hoorn, C., Bogotá-Angel, G. R., Romero-Baez, M., Lammertsma, E. I., Flantua, S. G. A., Dantas, E. L., et al. (2017). The Amazon at sea: Onset and stages of the Amazon River from a marine record, with special reference to Neogene plant turnover in the drainage basin. Global and Planetary Change, 153, 51–65. https://doi.org/10.1016/j.gloplacha.2017.02.005
Jia, F., Lo, N., & Ho, S. Y. W. (2014). The impact of modelling rate heterogeneity among sites on phylogenetic estimates of intraspecific evolutionary rates and timescales. PLoS ONE, 9(5), e95722. https://doi.org/10.1371/journal.pone.0095722
Jungfer, K. -H. (2011). A new tree frog of the genus Osteocephalus from high altitudes in the Cordillera del Cóndor, Ecuador (Amphibia: Anura: Hylidae). The Herpetological Journal, 21(4), 247–253.
Jungfer, K. -H., & Hödl, W. (2002). A new species of Osteocephalus from Ecuador and a redescription of O. leprieurii (Duméril & Bibron, 1841) (Anura: Hylidae). Amphibia-Reptilia, 23, 21–46.
Jungfer, K. -H., & Lehr, E. (2001). A new species of Osteocephalus with bicoloured iris from Pozuzo (Peru: Departamento de Pasco) (Amphibia: Anura: Hylidae). Zoologische Abhandlungen. Staatliches Museum Für Tierkunde in Dresden, 51, 321–329.
Jungfer, K. -H., & Schiesari, L. C. (1995). Description of a central Amazonian and Guianan tree frog, genus Osteocephalus (Anura, Hylidae), with oophagous tadpoles. Alytes, 13, 1–13.
Jungfer, K. -H., & Weygoldt, P. (1999). Biparental care in the tadpole-feeding Amazonian treefrog Osteocephalus oophagus. Amphibia-Reptilia, 20(3), 235–249.
Jungfer, K. -H., Faivovich, J., Padial, J. M., Castroviejo-Fisher, S., Lyra, M. M., Berneck, B. V. M., et al. (2013). Systematics of spiny-backed treefrogs (Hylidae: Osteocephalus): An Amazonian puzzle. Zoologica Scripta, 42(4), 351–380. https://doi.org/10.1111/zsc.12015
Jungfer, K. -H., Ron, S. R., Seipp, R., & Almendáriz, A. (2000). Two new species of hylid frogs, genus Osteocephalus, from Amazonian Ecuador. Amphibia-Reptilia, 21, 327–340.
Jungfer, K. -H., Verdade, V. K., Faivovich, J., & Rodrigues, M. T. (2016). A new species of spiny-backed treefrog (Osteocephalus) from Central Amazonian Brazil (Amphibia: Anura: Hylidae). Zootaxa, 4114(2), 171–181. https://doi.org/10.11646/zootaxa.4114.2.6
Kapli, P., Lutteropp, S., Zhang, J., Kobert, K., Pavlidis, P., Stamatakis, A., & Flouri, T. (2017). Multi-rate Poisson tree processes for single-locus species delimitation under maximum likelihood and Markov chain Monte Carlo. Bioinformatics, 33(11), 1630–1638. https://doi.org/10.1093/bioinformatics/btx025
Katoh, K., & Standley, D. M. (2013). MAFFT multiple sequence alignment software version 7: Improvements in performance and usability. Molecular Biology and Evolution, 30(4), 772–780. https://doi.org/10.1093/molbev/mst010
Katoh, K., Rozewicki, J., & Yamada, K. D. (2019). MAFFT online service: Multiple sequence alignment, interactive sequence choice and visualization. Briefings in Bioinformatics, 20(4), 1160–1166. https://doi.org/10.1093/bib/bbx108
Klaus, K. V., & Matzke, N. J. (2020). Statistical comparison of trait-dependent biogeographical models indicates that Podocarpaceae dispersal is influenced by both seed cone traits and geographical distance. Systematic Biology, 69(1), 61–75. https://doi.org/10.1093/sysbio/syz034
Kok, P. J. R., & Kalamandeen, M. (2008). Introduction to the taxonomy of the amphibians of Kaieteur National Park, Guyana. Abc Taxa: A series of manuals dedicated to capacity building in taxonomy and collection management (Vol. 5). Brussels: Belgian Development Corporation.
Kok, P. J. R., Ratz, S., Tegelaar, M., Aubret, F., & Means, D. B. (2015). Out of taxonomic limbo: A name for the species of Tepuihyla (Anura: Hylidae) from the Chimantá Massif, Pantepui region, northern South America. Salamandra, 51(4), 283–314.
Kumar, S., Stecher, G., Li, M., Knyaz, C., & Tamura, K. (2018). MEGA X: Molecular evolutionary genetics analysis across computing platforms. Molecular Biology and Evolution, 35(6), 1547–1549. https://doi.org/10.1093/molbev/msy096
Lanfear, R., Frandsen, P. B., Wright, A. M., Senfeld, T., & Calcott, B. (2016). PartitionFinder 2: New methods for selecting partitioned models of evolution for molecular and morphological phylogenetic analyses. Molecular Biology and Evolution, 34(3), 772–773. https://doi.org/10.1093/molbev/msw260
Larsson, A. (2014). AliView: A fast and lightweight alignment viewer and editor for large datasets. Bioinformatics, 30(22), 3276–3278. https://doi.org/10.1093/bioinformatics/btu531
Leite, R. N., & Rogers, D. S. (2013). Revisiting Amazonian phylogeography: Insights into diversification hypotheses and novel perspectives. Organisms Diversity & Evolution, 13(4), 639–664. https://doi.org/10.1007/s13127-013-0140-8
Lima, A. P., Magnusson, W. E., Menin, M., Erdtmann, L. K., Rodrigues, D. J., Keller, C., & Hödl, W. (2006). Guide to the frogs of Reserva Adolpho Ducke, Central Amazonia. Manaus: Áttema Design Editorial.
Lötters, S., van der Meijden, A., Coloma, L. A., Boistel, R., Cloetens, P., Ernst, R., et al. (2011). Assessing the molecular phylogeny of a near extinct group of vertebrates: The Neotropical harlequin frogs (Bufonidae; Atelopus). Systematics and Biodiversity, 9(1), 45–57. https://doi.org/10.1080/14772000.2011.557403
Lynch, J. D. (2002). A new species of the genus Osteocephalus (Hylidae: Anura) from the western Amazon. Revista De La Academia Colombiana De Ciencias Exactas, Físicas y Naturales, 26, 289–292.
Martins, M., & Cardoso, A. J. (1987). Novas espécies de hilídeos do Estado do Acre (Amphibia: Anura). Revista Brasileira De Biologia, 47, 549–558.
Matzke, N. J. (2013). BioGeoBEARS: Biogeography with Bayesian (and likelihood) evolutionary analysis in R scripts. R Package, Version, 1(1), 2.
Melin, D. E. (1941). Contributions to the knowledge of the Amphibia of South America. Göteborgs Kungl. Vetenskaps-och Vitterhets-samhälles. Handlingar. Serien b, Matematiska Och Naturvetenskapliga Skrifter, 1, 1–71.
Melo-Sampaio, P. R., Ferrão, M., & Moraes, L. J. C. L. (2021). A new species of Osteocephalus Steindachner, 1862 (Anura, Hylidae), from Brazilian Amazonia. Breviora, 572(1), 1–21. https://doi.org/10.3099/0006-9698-572.1.1
Monaghan, M. T., Wild, R., Elliot, M., Fujisawa, T., Balke, M., Inward, D. J. G., et al. (2009). Accelerated species inventory on Madagascar using coalescent-based models of species delineation. Systematic Biology, 58(3), 298–311. https://doi.org/10.1093/sysbio/syp027
Moraes, L. J. C. L., Werneck, F. P., Réjaud, A., Rodrigues, M. T., Prates, I., Glaw, F., et al. (2022). Diversification of tiny toads (Bufonidae: Amazophrynella) sheds light on ancient landscape dynamism in Amazonia. Biological Journal of the Linnean Society. https://doi.org/10.1093/biolinnean/blac006
Moravec, J., Aparicio, J., Guerrero-Reinhard, M., Calderón, G., Jungfer, K. -H., & Gvoždík, V. (2009). A new species of Osteocephalus (Anura: Hylidae) from Amazonian Bolivia: First evidence of tree frog breeding in fruit capsules of the Brazil nut tree. Zootaxa, 2215(1), 37–54.
Moura, P. H. A. G., Elias-Costa, A. J., Nunes, I., & Faivovich, J. (2021). Diversity and evolution of the extraordinary vocal sacs of casque-headed treefrogs (Anura: Hylidae). Biological Journal of the Linnean Society, 134, 423–442. https://doi.org/10.1093/biolinnean/blab083
Oliveira, U., Vasconcelos, M. F., & Santos, A. J. (2017). Biogeography of Amazon birds: Rivers limit species composition, but not areas of endemism. Scientific Reports, 7(1), 2992. https://doi.org/10.1038/s41598-017-03098-w
Ortiz, D. A., & Ron, S. R. (2018). Osteocephalus vilmae. In S. R. Ron, A. Merino-Viteri, & D. A. Ortiz (Eds.), Anfibios del Ecuador. Version 2021.0. Museo de Zoología, Pontificia Universidad Católica del Ecuador. Retrieved May 15, 2022, from https://bioweb.bio/faunaweb/amphibiaweb/FichaEspecie/Osteocephalus%20vilmae
Ortiz, D. A., & Ron, S. R. (2020). Osteocephalus duellmani. In S. R. Ron, A. Merino-Viteri, & D. A. Ortiz (Eds.), Anfibios del Ecuador. Version 2021.0. Museo de Zoología, Pontificia Universidad Católica del Ecuador. Retrieved May 15, 2022, from https://bioweb.bio/faunaweb/amphibiaweb/FichaEspecie/Osteocephalus%20duellmani
Ortiz, D. A., Moraes, L. J. C. L., Pavan, D., & Werneck, F. P. (2020). Molecular phylogenetic relationships and generic placement of Dryaderces inframaculata Boulenger, 1882 (Anura: Hylidae). Vertebrate Zoology, 7(3), 357–366. https://doi.org/10.26049/VZ70-3-2020-08
Paradis, E., & Schliep, K. (2019). ape 5.0: An environment for modern phylogenetics and evolutionary analyses in R. Bioinformatics, 35(3), 526–528. https://doi.org/10.1093/bioinformatics/bty633
Peloso, P. L. V., Sturaro, M. J., Forlani, M. C., Gaucher, P., Motta, A. P., & Wheeler, W. C. (2014). Phylogeny, taxonomic revision, and character evolution of the genera Chiasmocleis and Syncope (Anura, Microhylidae) in Amazonia, with descriptions of three new species. Bulletin of the American Museum of Natural History, 2014(386), 1–112. https://doi.org/10.1206/834.1
Peracca, M. G. (1904). Viaggio del Dr. Enrico Festa nell’ Ecuador e regioni vicine. Rettili ed anfibi. Bollettino dei Musei di Zoologia e Anatomia Comparata della R. Universita di Torino, 19(465), 1–41.
Peters, W. C. H. (1867). Über Fledertheire (Pteropus gouldii, Rhinolophus deckenii, Vespertilio lobipes, Vesperugo temminckii) und Amphibien (Hypsilurus godeffroyi, Lygosoma scutatum, Stenostoma narisostre, Onychocephalus unguirostris, Ahaetulla polylepis, Pseudechis scutellatus, Hoplobatrachus reinhardtii, Hyla coriacea). Monatsberichte Der Königlichen Preussische Akademie Des Wissenschaften Zu Berlin, 1867, 703–712.
Pons, J., Barraclough, T. G., Gomez-Zurita, J., Cardoso, A., Duran, D. P., Hazell, S., et al. (2006). Sequence-based species delimitation for the DNA taxonomy of undescribed insects. Systematic Biology, 55(4), 595–609. https://doi.org/10.1080/10635150600852011
Puillandre, N., Lambert, A., Brouillet, S., & Achaz, G. (2012). ABGD, automatic barcode gap discovery for primary species delimitation. Molecular Ecology, 21(8), 1864–1877. https://doi.org/10.1111/j.1365-294X.2011.05239.x
Pupim, F. N., Sawakuchi, A. O., Almeida, R. P., Ribas, C. C., Kern, A. K., Hartmann, G. A., et al. (2019). Chronology of terra firme formation in Amazonian lowlands reveals a dynamic Quaternary landscape. Quaternary Science Reviews, 210, 154–163. https://doi.org/10.1016/j.quascirev.2019.03.008
R Core Team. (2020). R: A language and environment for statistical computing. Vienna, Austria: R Foundation for Statistical Computing. https://www.r-project.org/
Rambaut, A., Drummond, A. J., Xie, D., Baele, G., & Suchard, M. A. (2018). Posterior summarization in Bayesian phylogenetics using Tracer 1.7. Systematic Biology, 67(5), 901–904. https://doi.org/10.1093/sysbio/syy032
Ramírez, J. P., Jaramillo, C. A., Lindquist, E. D., Crawford, A. J., & Ibáñez, R. (2020). Recent and rapid radiation of the highly endangered harlequin frogs (Atelopus) into Central America inferred from mitochondrial DNA sequences. Diversity, 12(9), 360. https://doi.org/10.3390/d12090360
Raven, P. H., Gereau, R. E., Phillipson, P. B., Chatelain, C., Jenkins, C. N., & Ulloa Ulloa, C. (2020). The distribution of biodiversity richness in the tropics. Science Advances, 6(37), eabc6228. https://doi.org/10.1126/sciadv.abc6228
Ree, R. H., & Sanmartín, I. (2018). Conceptual and statistical problems with the DEC+J model of founder-event speciation and its comparison with DEC via model selection. Journal of Biogeography, 45(4), 741–749. https://doi.org/10.1111/jbi.13173
Réjaud, A., Rodrigues, M. T., Crawford, A. J., Castroviejo-Fisher, S., Jaramillo, A. F., Chaparro, J. C., et al. (2020). Historical biogeography identifies a possible role of Miocene wetlands in the diversification of the Amazonian rocket frogs (Aromobatidae: Allobates). Journal of Biogeography, 47(11), 2472–2482. https://doi.org/10.1111/jbi.13937
Ribas, C. C., & Aleixo, A. (2019). Diversity and evolution of Amazonian birds: Implications for conservation and biogeography. Anais Da Academia Brasileira De Ciências, 91(3), e20190218. https://doi.org/10.1590/0001-3765201920190218
Rivero, J. A. (1968). A new species of Hyla (Amphibia, Salientia) from the Venezuelan Guayana. Breviora. Museum of Comparative Zoology, 307, 1–5.
Ron, S. R., & Pramuk, J. B. (1999). A new species of Osteocephalus (Anura: Hylidae) from Amazonian Ecuador and Peru. Herpetologica, 55(4), 433–446.
Ron, S. R., Venegas, P. J., Toral, E., Read, M., Ortiz, D. A., & Manzano, A. (2012). Systematics of the Osteocephalus buckleyi species complex (Anura, Hylidae) from Ecuador and Peru. ZooKeys, 229, 1–52. https://doi.org/10.3897/zookeys.229.3580
Ron, S. R., Venegas, P. J., Ortega-Andrade, H. M., Gagliardi-Urrutia, G., & Salerno, P. E. (2016). Systematics of Ecnomiohyla tuberculosa with the description of a new species and comments on the taxonomy of Trachycephalus typhonius (Anura, Hylidae). ZooKeys, 630, 115–154. https://doi.org/10.3897/zookeys.630.9298
Ruokolainen, K., Moulatlet, G. M., Zuquim, G., Hoorn, C., & Tuomisto, H. (2019). Geologically recent rearrangements in central Amazonian river network and their importance for the riverine barrier hypothesis. Frontiers of Biogeography, 11(3), e45046. https://doi.org/10.21425/F5FBG45046
Ruthven, A. G. (1919). The amphibians of the University of Michigan-Walker Expedition to British Guiana. Occasional Papers of the Museum of Zoology, University of Michigan, 69, 1–14.
Salerno, P. E., Ron, S. R., Señaris, J. C., Rojas-Runjaic, F. J. M., Noonan, B. P., & Cannatella, D. C. (2012). Ancient tepui summits harbor young rather than old lineages of endemic frogs. Evolution, 66(10), 3000–3013. https://doi.org/10.1111/j.1558-5646.2012.01666.x
Santos, J. C., Coloma, L. A., Summers, K., Caldwell, J. P., Ree, R., & Cannatella, D. C. (2009). Amazonian amphibian diversity is primarily derived from late Miocene Andean lineages. Plos Biology, 7(3), e1000056. https://doi.org/10.1371/journal.pbio.1000056
Silva, S. M., Peterson, A. T., Carneiro, L., Burlamaqui, T. C. T., Ribas, C. C., Sousa-Neves, T., et al. (2019). A dynamic continental moisture gradient drove Amazonian bird diversification. Science Advances, 5(7), eaat5752. https://doi.org/10.1126/sciadv.aat5752
Smith, B. T., McCormack, J. E., Cuervo, A. M., Hickerson, M. J., Aleixo, A., Cadena, C. D., et al. (2014). The drivers of tropical speciation. Nature, 515(7527), 406–409. https://doi.org/10.1038/nature13687
Smith, E. N., & Noonan, B. P. (2001). A new species of Osteocephalus (Anura: Hylidae) from Guyana. Revista De Biología Tropical, 49(1), 347–357.
Stamatakis, A. (2014). RAxML Version 8: A tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics, 30(9), 1312–1313. https://doi.org/10.1093/bioinformatics/btu033
Stamatakis, A. (2016). The RAxML v8.2.X manual. Heidelberg: Heidelberg Institute for Theoretical Studies.
Steindachner, F. (1862). Über zwei noch unbeschriebene Batrachier aus des Sammlung des K. K. zoologischn Museum zu Wien. Archivio per La Zoologia L”Anatomia E La Fisiologia, Genova, Fasciolo 1, 2, 77–82.
Vacher, J. -P., Chave, J., Ficetola, F. G., Sommeria-Klein, G., Tao, S., Thébaud, C., et al. (2020). Large-scale DNA-based survey of frogs in Amazonia suggests a vast underestimation of species richness and endemism. Journal of Biogeography, 47(8), 1781–1791. https://doi.org/10.1111/jbi.13847
Vences, M., Thomas, M., van der Meijden, A., Chiari, Y., & Vieites, D. R. (2005). Comparative performance of the 16S rRNA gene in DNA barcoding of amphibians. Frontiers in Zoology, 2(1), 5. https://doi.org/10.1186/1742-9994-2-5
Walker, C. F. (1973). A new genus and species of microhylid frog from Ecuador. Occasional Papers of the Museum of Natural History, University of Kansas, 20, 1–7.
Wallace, A. R. (1854). On the monkeys of the Amazon. Annals and Magazine of Natural History, 14(84), 451–454.
Wang, X., Edwards, R. L., Auler, A. S., Cheng, H., Kong, X., Wang, Y., et al. (2017). Hydroclimate changes across the Amazon lowlands over the past 45,000 years. Nature, 541(7636), 204–207. https://doi.org/10.1038/nature20787
Webb, S. D. (1995). Biological implications of the middle Miocene Amazon seaway. Science, 269(5222), 361–362. https://doi.org/10.1126/science.269.5222.361
Werner, F. (1901). Ueber Reptilien und Batrachier aus Ecuador und Neu Guinea. Verhandlungen Des Zoologisch-Botanischen Vereins in Wien, 51, 593–614.
Zeisset, I., & Beebee, T. J. C. (2008). Amphibian phylogeography: A model for understanding historical aspects of species distributions. Heredity, 101(2), 109–119. https://doi.org/10.1038/hdy.2008.30
Zhang, J., Kapli, P., Pavlidis, P., & Stamatakis, A. (2013a). A general species delimitation method with applications to phylogenetic placements. Bioinformatics, 29(22), 2869–2876. https://doi.org/10.1093/bioinformatics/btt499
Zhang, P., Liang, D., Mao, R. -L., Hillis, D. M., Wake, D. B., & Cannatella, D. C. (2013b). Efficient sequencing of anuran mtDNAs and a mitogenomic exploration of the phylogeny and evolution of frogs. Molecular Biology and Evolution, 30(8), 1899–1915. https://doi.org/10.1093/molbev/mst091
Zizka, A., Azevedo, J., Leme, E., Neves, B., da Costa, A. F., Caceres, D., & Zizka, G. (2020). Biogeography and conservation status of the pineapple family (Bromeliaceae). Diversity and Distributions, 26(2), 183–195. https://doi.org/10.1111/ddi.13004
Acknowledgements
This study is part of DAO’s PhD dissertation at James Cook University (JCU), Australia. DAO thanks JCU for his PhD scholarship. FPW thanks CNPq (Productivity Fellowship) and the L’Oréal-UNESCO For Women In Science Program. We thank researchers, curatorial staff and herpetological collections for providing tissue samples: M. Rodrigues (Universidade de São Paulo, Brazil), A. Lima, C. Ribas and M. Freitas (Instituto Nacional de Pesquisas da Amazônia, Brazil), R. Rojas (Universidad Nacional de la Amazonía Peruana, Peru), P. Kok (University of Lodz, Poland), J. C. Chaparro (Museo de Biodiversidad del Perú, Peru), B. Noonan (University of Mississippi, USA), I. Valente (Instituto de Desenvolvimento Sustentável Mamirauá, Brazil), D. Pavan (Ecosfera Consultoria e Pesquisa em Meio Ambiente Ltda., Brazil) and E. Courtois (Laboratoire Ecologie, Evolution, Interactions des Systèmes amazoniens, French Guiana). L. Moraes assisted with sample preparation for shipment; and D. Pazmiño with elaboration of Figs. 3, 4 and 5 in the Appendix. DAO thanks S. Ho for advice on BEAST analyses, and to organisers of the Taming the BEAST Down Under workshop 2019, University of Sydney, Australia.
Funding
Open Access funding enabled and organized by CAUL and its Member Institutions. This research was funded by the Center for the Study of Biodiversity in Amazonia-Labex CEBA (ref. ANR-10-LABX-25–01) under ‘SBOTA’ project granted to AF and DAO, and GUYAMAZON programme, an international French/Brazilian cooperation programme (IRD, CNRS, CTG, CIRAD and Brazilian Fundação de Amparo à Pesquisa do Estado do Amazonas-FAPEAM 062.00962/2018) under ‘COMBIA’ project co-coordinated by AF and FPW.
Author information
Authors and Affiliations
Contributions
DAO, CJH and AF designed the study. DAO, FPW and AF acquired samples and funding. DAO, SM and AF generated DNA sequences. DAO organized data and conducted analyses with contributions of CJH, AR and AF. DAO led the writing and preparation of figures with contributions from CJH, FPW, SRR and AF. All authors commented on the final draft.
Corresponding author
Ethics declarations
Ethics approval
Fieldwork and laboratory experiments have been conducted in accordance with relevant national legislation on the use of animals for research.
Consent to participate
Not applicable.
Consent for publication
Not applicable.
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Appendix
Appendix
Taxonomic account
Our mitogenomic phylogeny included 50 putative species as ingroup (Fig. 1a), and a distribution map based on our analysed sequences is illustrated in Electronic Supplement 3. This phylogeny included 36 out of 40 species currently recognised (Frost, 2022; Ortiz & Ron, 2018), with the absence of Osteocephalus duellmani Jungfer, 2011; Osteocephalus germani Ron et al., 2012 (see comments below on ‘Osteocephalus helenae’ species complex); Osteocephalus melanops Melo-Sampaio et al., 2021; and Tepuihyla luteolabris Ayarzagüena et al., 1993a. From the 40 recognised species, two of them (O. germani and Osteocephalus vilmae Ron et al., 2012) were considered synonyms of Osteocephalus helenae Ruthven, 1919 and Osteocephalus buckleyi Boulenger, 1882, respectively, by Jungfer et al. (2013) and currently are not listed in Frost (2022). However, we recognise these binomials as valid following discussion in this account (under O. buckleyi group) and justification in Ortiz and Ron (2018). Therefore, thirty-six out of the 50 putative species were associated to an available taxon name, while the remaining 14 (ten in Osteocephalus and four in Dryaderces) could not be associated to any described species and thus correspond to lineages (candidate species) pending of integrative study. They were labelled using ‘aff.’ relative to their closest related taxon in the mitogenomic phylogeny. Below we discuss our taxonomic conclusions under each ‘species group’ including the correspondence between described species and the 42 OTUs resulting from the 16S-based species delimitation, and a rationale for the addition of eight terminals to the mitogenomic matrix (thus n = 50). The species groups are the genera Dryaderces and Tepuihyla, and each clade within Osteocephalus that have diverged approximately \(\ge\) 10 million years ago ‘Mya’ to other clades (Fig. 1a).
Dryaderces Jungfer et al., 2013
The majority rule consensus delimited six OTUs within Dryaderces (Fig. 1a). Two of them unambiguously correspond to described species: Dryaderces inframaculata Boulenger, 1882; and D. pearsoni Gaige, 1929. One out of two sequences of D. inframaculata corresponds to its first molecular record (Ortiz et al., 2020). Both sequences of D. pearsoni are from nearby their type locality. The four undescribed OTUs are phenotypically similar to D. pearsoni, but genetically closer to D. inframaculata (thus labelled as D. aff. inframaculata 1–4 (‘morph pearsoni’)). Three of them (D. aff. inframaculata 1, 3 and 4) were previously considered as a single undescribed species, labelled as ‘D. pearsoni [Ca1] CCS’ by Jungfer et al. (2013).
Tepuihyla Ayarzagüena et al., 1993b
The majority rule consensus delimited four OTUs within Tepuihyla (Fig. 1a). Three of them unambiguously correspond to described species with sequences from their type locality or nearby sites, and/or from their known distribution: Tepuihyla exophthalma Smith & Noonan, 2001; T. tuberculosa Boulenger, 1882; and T. warreni Duellman & Hoogmoed, 1992. The fourth OTU is formed by sequences from specimens assigned to four recognised species (T. aecii Ayarzagüena et al., 1993a; T. edelcae Ayarzagüena et al., 1993a; T. obscura Kok et al., 2015; and T. rodriguezi Rivero, 1968). Given these species are phenotypically distinct (see ‘Definition and diagnosis’ section in Kok et al. (2015)), the low divergence among them in 16S led to not detect them as different in the majority rule consensus. Therefore, a terminal of each species was included in the mitogenomic matrix.
Finally, a terminal for T. shushupe Ron et al., 2016 (holotype) was also added to the mitogenomic matrix given it did not have 16S data and thus was not included in the delimitation analyses. Although T. shushupe morphologically resembles its sister species, T. tuberculosa, they differ in molecular data, iris colouration and advertisement call (Ron et al., 2016). Tepuihyla luteolabris Ayarzagüena et al., 1993a was absent in our mitogenomic phylogeny and no genetic sequences are known for this species to infer its relationships.
Osteocephalus Steindachner, 1862
The delimitation will be presented following the traditional five Osteocephalus species groups of Jungfer et al. (2013), plus an additional species group here labelled as ‘Osteocephalus mimeticus group’. The species in this latter group were previously considered as part of the O. buckleyi group by Jungfer et al. (2013), but we recognise it as an additional group because it displays an age of divergence similar to the other five species groups (mean \(\ge\) 10 Mya).
Osteocephalus alboguttatus group
The majority rule consensus delimited four OTUs within the Osteocephalus alboguttatus group (Fig. 1a). Three of them unambiguously correspond to described species with sequences from their type locality: Osteocephalus alboguttatus Boulenger, 1882 (Fig. 3a in the Appendix); O. heyeri Lynch, 2002; and O. subtilis Martins & Cardoso, 1987. The sequences of the fourth OTU (O. aff. subtilis) are from Serra do Divisor (border between Brazil and Peru), and form a highly divergent sister group, geographically close, to O. subtilis sensu stricto (Fig. 1a; Electronic Supplement 3). Osteocephalus melanops Melo-Sampaio et al., 2021 was recently described and thus it could not be included in our analyses. However, it is phenotypically distinct from the other species in this group and displays a sister relationship with O. alboguttatus (Melo-Sampaio et al., 2021).
Dorsolateral views of Osteocephalus. a O. alboguttatus sensu stricto, QCAZ15972, Puyo-Canelos Road, Pastaza, Ecuador. b O. mimeticus sensu stricto, AF4442 (sequenced), San José, San Martín, Peru. c O. buckleyi sensu stricto, Canelos, Pastaza, Ecuador. d O. vilmae, QCAZ51205, Pompeya-Iro Road km 80, Orellana, Ecuador. e O. mutabor, QCAZ56066, Lorocachi, Pastaza, Ecuador. f O. mutabor, QCAZ39588, Río Pucayacu, Pastaza, Ecuador. g O. taurinus sensu stricto, Reserva Ducke, Amazonas, Brazil. h O. oophagus sensu stricto, Reserva Ducke, Amazonas, Brazil. Photographs: a, d–f by Santiago Ron (BIOWEB, Licence: CC BY-NC-ND 4.0); b by Antoine Fouquet; c, g–h by Diego Ortiz
Osteocephalus buckleyi group
The majority rule consensus delimited 13 OTUs within the Osteocephalus buckleyi group (Fig. 1a). Ten of them unambiguously correspond to described species, with sequences from their type locality or nearby sites, and/or from their known distribution: Osteocephalus buckleyi Boulenger, 1882 (Fig. 3c in the Appendix); O. cabrerai Cochran & Goin, 1970 (Fig. 4a in the Appendix); O. camufatus Jungfer et al., 2016 (holotype); O. cannatellai Ron et al., 2012; O. carri Cochran & Goin, 1970; O. festae Peracca, 1904; O. helenae Ruthven, 1919 (Fig. 4b–d in the Appendix); O. mutabor Jungfer & Hödl, 2002 (Fig. 3e–f in the Appendix); O. verruciger Werner, 1901; and O. vilmae Ron et al., 2012 (Fig. 3d in the Appendix). The remaining three OTUs (O. aff. cabrerai, O. aff. helenae 1 and O. aff. verruciger) are discussed below.
Dorsolateral and ventral views of Osteocephalus. a O. cabrerai, CORBIDI120, Tarapoa, Sucumbíos, Ecuador. b O. helenae, AF2427 (sequenced), Nassau, Suriname. c O. helenae, Amapá, Brazil. d O. helenae, Voltaire, French Guiana. e–f O. aff. helenae 2 ‘morph cabrerai’, AF2357 (sequenced), Mapaou, French Guiana. g O. aff. helenae 2 ‘morph cabrerai’, AF3358 (sequenced), Bakhuis, Surinam. h O. aff. helenae 2 ‘morph cabrerai’, St. Georges, French Guiana. Photographs: a by Pablo Venegas (BIOWEB, Licence: CC BY-NC-ND 4.0); b–h by Antoine Fouquet
Dorsolateral views of Osteocephalus. a O. leprieurii sensu stricto, Kaw mountain, French Guiana. b O. leprieurii, AF1824, Mana, French Guiana. c O. yasuni sensu stricto, QCAZ69010, Estación Científica Yasuní, Orellana, Ecuador. d O. yasuni, QCAZ55994, Lorocachi, Pastaza, Ecuador. e O. deridens, QCAZ56031, Lorocachi, Pastaza, Ecuador. f O. fuscifacies, QCAZ59887, Parque Nacional Llanganates, Pastaza, Ecuador. g O. aff. leoniae 2, AF4513 (sequenced), San Martín, Peru. h O. planiceps, QCAZ55881, Lorocachi, Pastaza, Ecuador. Photographs: a–b, g by Antoine Fouquet; c–f, h by Santiago Ron (BIOWEB, Licence: CC BY-NC-ND 4.0)
Osteocephalus vilmae was synonymised under O. buckleyi by Jungfer et al. (2013). However, we associate the OTU containing three ‘O. buckleyi’ samples to the name O. vilmae in this study because these samples cluster with high support with other O. vilmae specimens (including the holotype) in previous phylogenies that include both mitochondrial (mtDNA) and nuclear (nDNA) data; and this clade is sister to O. buckleyi sensu stricto with high support (Chasiluisa et al., 2020; Jungfer et al., 2013). Further justification for the validity of O. vilmae is presented in Ortiz and Ron (2018).
The single sequence of O. aff. cabrerai (from Iça River, Amazonas, Brazil) is sister and geographically close to O. cabrerai sensu stricto (Fig. 1a; Electronic Supplement 3). The long branches separating them suggest that each represents a distinct species but morphological and bioacoustics data remain to be examined.
The single sequence of O. aff. helenae 1 is from Mataracú, Santa Cruz, Bolivia. It corresponds to the southern-most record of the O. buckleyi group (Electronic Supplement 3). Osteocephalus aff. helenae 1 is sister to O. helenae + O. aff. helenae 2 (Fig. 1a; see comments below on ‘O. helenae’ species complex).
The sequences of O. aff. verruciger are from the Caquetá Department, Colombia, lying geographically close to the type locality of a recently described species, previously considered part of O. verruciger (i.e. O. omega Duellman, 2019). Therefore, we tentatively labelled this OTU as O. cf. omega (Fig. 1a). However, is still necessary to determine whether they are conspecific.
Finally, a terminal for O. sangay Chasiluisa et al., 2020 (holotype) was added to the mitogenomic matrix because this species did not have 16S data and thus was not included in the delimitation analyses. Osteocephalus sangay is genetically and morphologically distinct from its sister species, O. cannatellai (Chasiluisa et al., 2020). Osteocephalus duellmani Jungfer, 2011 was absent in our mitogenomic phylogeny and no DNA sequences are known for this species to infer its relationships. It has been tentatively allocated within the O. buckleyi group given its morphological affinities (Jungfer, 2011). Nevertheless, O. duellmani may be a junior synonym of O. festae Peracca, 1904 (Ortiz & Ron, 2020).
Comments on the ‘Osteocephalus helenae’ complex (within O. buckleyi group)
The delimited OTU corresponding to ‘Osteocephalus helenae’ Ruthven, 1919 sensu Jungfer et al. (2013) is widespread across Amazonia (Electronic Supplement 3) and represents a species complex on its own. The sequences associated to the name ‘O. helenae’ consist of seven geographically structured clades which do not form a monophyletic group in our 16S tree given the positions of O. aff. helenae 1 and O. aff. helenae 2 (‘morph cabrerai’) (see below) (Electronic Supplement 4). Osteocephalus aff. helenae 1 was delimitated was an independent OTU, whereas O. aff. helenae 2 (‘morph cabrerai’) clustered in the same OTU with ‘O. helenae’ in the majority rule consensus (Fig. 1a).
In the Guiana Shield, this latter OTU is formed by sequences that belong to two co-occurring distinct species, thus the low divergence among them in 16S led to not detecting them as different. One of those (sp. 1) morphologically resembles O. buckleyi Boulenger, 1882 and has been associated to the name O. helenae Ruthven, 1919 even though the justification remains meagre (Jungfer et al., 2013). The holotype of ‘Hyla helenae’ (UMMZ52681) is a recently metamorphosed juvenile from Dunoon, Demerara River, Guyana. It apparently shows the typical characters of juveniles of most species of the O. buckleyi group according to Jungfer et al. (2013), who based on that, transferred the name ‘Hyla helenae’ to Osteocephalus under the O. buckleyi group. These authors assumed that there is only one species of the O. buckleyi group occurring in the Guiana Shield and therefore concluded to name all the samples within this clade under ‘O. helenae’, including previous reports of O. buckleyi from this region (e.g. Gorzula & Señaris, 1998; Kok & Kalamandeen, 2008) and other regions.
The other species from the Guiana Shield (sp. 2) was previously assigned to O. cabrerai Cochran & Goin, 1970 based on its morphological characteristics (Dewynter et al., 2016). However, the samples of ‘O. cabrerai’ from the Guiana Shield are not closely related to O. cabrerai sensu stricto from western Amazonia using mitochondrial DNA (mtDNA) (Electronic Supplement 4), thus representing another lineage closely related to ‘O. helenae’ instead. Therefore, we labelled sp. 2 as O. aff. helenae 2 (‘morph cabrerai’) and included an additional terminal for it in the mitogenomic matrix (Fig. 1a).
Both species, O. helenae sensu Jungfer et al. (2013) (Fig. 4b–d in the Appendix) and O. aff. helenae 2 ‘morph cabrerai’ (Fig. 4e–h in the Appendix) show a low-supported, sister relationship when analysing their complete mitogenomes (Fig. 1a), and differ in morphology and ecology. Osteocephalus aff. helenae 2 is larger than O. helenae; it has large and fleshy tubercles on all dorsal surfaces, including tarsus and lower jaw (tubercles smaller and more restricted to dorsum in O. helenae), and hidden surfaces are deep blue in O. aff. helenae 2 (tan or bluish grey in O. helenae) (Dewynter et al., 2016). In sympatric areas, both are allotopic species: O. aff. helenae 2 is found in rocky, fast, clear water streams, while O. helenae in ponds and flooded forests (igapó) around black water rivers; their calls also differ (A. Fouquet, pers. obs.). Finally, O. aff. helenae 2 may actually be circumscribed to the eastern Guiana Shield (Electronic Supplement 3) and its occurrence in northern Guyana still need to be confirmed but is unlikely. Moreover, the habitat found in Dunoon (flat, flooded riparian forest) matches the habitat of sp. 1 ‘O. helenae’ but not the one of sp. 2 ‘O. aff. helenae 2’ (rocky streams). Therefore, we consider that the assignation of Jungfer et al. (2013) for ‘O. helenae’ is correct. However, to completely solve this issue would require the sampling of adult specimens and the analysis of sequences from the type locality of O. helenae.
Finally, closely related samples from other Amazonian regions were also considered part of ‘O. helenae’ by Jungfer et al. (2013), including the synonymy of O. germani Ron et al., 2012. Osteocephalus germani was described from the Cusco Department, Andean slopes of Peru. Unfortunately, there were no 16S sequences available for the type material of O. germani to include it in our species delimitation and is absent in our mitogenomic phylogeny. However, a recent, dense-sampled phylogeny with a focus on the O. buckleyi group (Chasiluisa et al., 2020) showed that ‘O. germani’ individuals clustered in a clade that contain samples of O. aff. helenae 2 ‘morph cabrerai’, suggesting that both species may be closely related (in mtDNA) within the ‘O. helenae’ complex. Osteocephalus germani differs from O. aff. helenae 2 in morphology (Dewynter et al., 2016; Ron et al., 2012), and it is distributed in an opposite extreme of Amazonia (Andean slopes in south-western Amazonia) compared to O. helenae and O. aff. helenae 2 (eastern Guiana Shield) (Electronic Supplement 3), thus making unlikely that these three species are conspecific. Because ‘O. helenae’ is a species complex and this issue was unknown by Jungfer et al. (2013), we consider that their synonymy of O. germani under O. helenae was premature (see Ortiz & Ron, 2018). A thorough review of the ‘O. helenae’ complex is opportune including genomic nDNA and bioacoustics.
Osteocephalus leprieurii group
The majority rule consensus delimited one OTU for the entire Osteocephalus leprieurii group (Fig. 1a). Two species have been formally described in this group: Osteocephalus leprieurii Duméril & Bibron, 1841 (Fig. 5a–b in the Appendix) from the Guiana Shield; and O. yasuni Ron & Pramuk, 1999 (Fig. 5c–d in the Appendix) from western Amazonia. Given these two species are phenotypically distinct (see ‘Diagnosis’ section in Ron and Pramuk (1999)), the low divergence in 16S led to not detecting them as different in the majority rule consensus. Therefore, we included a terminal for each in the mitogenomic matrix from nearby sites to their type localities. Although with low support, three sequences clustering within the O. yasuni clade from Boa Vista, south bank of upper Negro River (MTR41294), and from the south bank of lower Japura River (MTR33656, 33683), Amazonas, Brazil, correspond to its first molecular records in Central Amazonia (Electronic Supplements 3 and 4). Two additional lineages phenotypically similar to O. leprieurii have been reported and labelled as ‘O. leprieurii [Ca1–2] UCS’ by Jungfer et al. (2013), but we did not include additional terminals in the mitogenomic matrix for them because of their current lack of morphological and bioacoustics information. We considered them here part of O. leprieurii; however, further systematic study and more variable markers are needed to delimit species and population structure within this widespread taxon.
‘Osteocephalus mimeticus group’
The majority rule consensus delimited two OTUs within the ‘Osteocephalus mimeticus group’ (Fig. 1a). One of them unambiguously corresponds to Osteocephalus mimeticus Melin, 1941 (Fig. 3b in the Appendix) containing a sequence from its type locality. The sequences of the other OTU (O. aff. mimeticus) are from the Cusco department, in Peru, and form a highly divergent sister group to O. mimeticus sensu stricto (Fig. 1a); they substitute each other latitudinally (Electronic Supplement 3).
Osteocephalus planiceps group
The majority rule consensus delimited ten OTUs within the Osteocephalus planiceps group (Fig. 1a). Six of them unambiguously correspond to described taxa with sequences from their type locality or nearby sites, and/or from their known distribution: Osteocephalus castaneicola Moravec et al., 2009 (holotype); O. deridens Jungfer et al., 2000 (Fig. 5e in the Appendix); O. fuscifacies Jungfer et al., 2000 (Fig. 5f in the Appendix); O. leoniae Jungfer & Lehr, 2001; O. planiceps Cope, 1874 (Fig. 5h in the Appendix); and O. vilarsi Melin, 1941. The sample of O. castaneicola (MTR18640) from the east bank of lower Purus River, Amazonas, Brazil, corresponds to its first molecular record in Central Amazonia (Electronic Supplement 3 and 4). The remaining four undescribed OTUs could not be assigned to any know taxa (O. aff. deridens, O. aff. leoniae 1, O. aff. leoniae 2 and O. aff. planiceps).
The sequences of O. aff. deridens are from nearby Iquitos, Loreto, Peru, and form a clade which is sister to Osteocephalus deridens sensu stricto (Fig. 1a).
Osteocephalus aff. leoniae 1 and O. aff. leoniae 2 (Fig. 5g in the Appendix) are sisters to each other, and both are sisters to O. leoniae sensu stricto (Fig. 1a). They are distributed allopatrically (Electronic Supplement 3).
The sequences of O. aff. planiceps are from Serra do Divisor (border between Brazil and Peru), and form a highly divergent sister group, geographically close, to Osteocephalus planiceps sensu stricto (Fig. 1a; Electronic Supplement 3).
Osteocephalus taurinus group
The majority rule consensus delimited two OTUs within the Osteocephalus taurinus group (Fig. 1a). The first OTU (labelled here as Osteocephalus aff. taurinus), containing two sequences from Amazonian Peru in western Amazonia, corresponds to an early diverging lineage within this group; it was previously labelled as O. taurinus [Ca1] CCS by Jungfer et al. (2013). The second OTU contains all the rest of sequences of O. taurinus Steindachner, 1862 (Fig. 3g in the Appendix) and O. oophagus Jungfer & Schiesari, 1995 (Fig. 3h in the Appendix) through Amazonia. Since both species differ in morphology and reproduction, this case represents another example of failure in detected them as different given the low divergence in 16S. Osteocephalus taurinus is considerably larger than O. oophagus, O. taurinus males have paired, lateral vocal sacs (single and subgular in O. oophagus males), dorsal tubercles with queratinized tips are abundant and prominent on dorsal surfaces in O. taurinus males and those may be also present but less noticeable in females (very few tubercles and barely discernible in O. oophagus males). Osteocephalus taurinus is an explosive pond-breeder with large clutch sizes of ~2000 eggs and parental care absent (O. oophagus has a small clutch of ~30 eggs which is laid within small water cavities of trees and leaf axils of terrestrial/epiphytic plants; water cavities are used by one pair at a time which also display parental care) (Jungfer & Weygoldt, 1999; Lima et al., 2006). These differences led us to include one terminal for each of these two species from their type locality in the mitogenomic matrix (Fig. 1a). The O. taurinus group contains several geographically structured mtDNA lineages (Electronic Supplement 4), and four of them have been suggested to correspond to candidate species (‘O. taurinus [Ca2–5] UCS’; Jungfer et al., 2013). However, we did not include additional terminals in the mitogenomic matrix for them because of their current lack of morphological and bioacoustics information. We considered them here part of O. taurinus; however, further systematic study and more variable markers are needed to delimit species and population structure within this widespread taxon.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Ortiz, D.A., Hoskin, C.J., Werneck, F.P. et al. Historical biogeography highlights the role of Miocene landscape changes on the diversification of a clade of Amazonian tree frogs. Org Divers Evol 23, 395–414 (2023). https://doi.org/10.1007/s13127-022-00588-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13127-022-00588-2