Abstract
The aims of the study were (i) to reveal the pattern of phylogeny of Pseudamnicola inhabiting the Aegean Islands, (ii) to describe and analyse the variation of the morphology in 17 populations of Pseudamnicola from the springs on the Aegean Islands not studied so far and considering also another seven populations studied earlier and (iii) to find out which model is more applicable to the island Pseudamnicola populations: either a model in which a relict fauna rich in endemics is differentiated in a way that mainly reflects the geological history of the area or a model in which a relatively young fauna is composed of more or less widely distributed taxa, with relatively high levels of gene flow among the springs they inhabit. To address the above issues, the morphology and the mitochondrial genes—cytochrome oxidase subunit I (COI) and ribosomal 16S—and nuclear genes—ribosomal 18S, 28S and histone 3 (H3)—were analysed. COI and COI+16S rRNA+18S datasets gave trees with identical topology in both ML and Bayesian inference. The 24 studied populations of Pseudamnicola form 16 clades, each of them generally having low levels of intrapopulation genetic differentiation. The generalised mixed Yule coalescent (GMYC) procedure and the Automatic Barcode Gap Discovery (ABGD) analysis for COI identified 16 Pseudamnicola entities coinciding with clades of the ML tree based on 44 haplotypes and 189 sequences. The present pattern of diversity, together with dating of divergence time, reflects a short story of colonisation/recolonisation, supported by the Late Pleistocene land bridges, rather than the consequences of earlier geological events. The principal component analysis (PCA) on the shells of the molecularly distinct clades showed differences, although variability ranges often overlap. Female reproductive organs showed no differences between the clades, and penile characters differed only in some cases.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
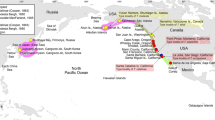
The history of the Mediterranean Basin is complex as a result of (i) tectonic events, e.g. the northward movement of the African tectonic plate and the eastward displacement of the Anatolian microplate; (ii) volcanic activity, e.g. the creation of new islands; and (iii) several climatic changes that caused sea level fluctuations, such as those that occurred during the glacial/interglacial periods of the Pleistocene. These events may significantly influence divergence in species richness and dispersion through the creation of favourable conditions for allopatric speciation by, for example, submergence and re-emergence of landmasses. It should be noted that the present geography of the Aegean is no more than 8–6 kya old (Kougioumoutzis et al. 2014 and references therein). Since the peak of the Wisconsin-Würm glaciation (24–10 kya), global sea level has risen 120–130 m and the Aegean Islands now lie where there was once dry land. In the Late Pleistocene, the eastern Aegean Islands were connected with Anatolia, and the strait between Europe and the Cyclade-mega-island, connecting the present Naxos, Paros, Tinos and Andros, was rather narrow (Kougioumoutzis et al. 2014).
Freshwater snails inhabiting springs on the islands—as ‘island on island’ habitats—are interesting organisms for the study of the relationships between geological events and phylogeography, but surprisingly, they are rarely used as subjects in this kind of study (e.g. Szarowska et al. 2014a, b). The genus Pseudamnicola Paulucci, 1878, contains species found in the Netherlands, Spain, France, Italy, the southern Balkans, Romania, Turkey and Cyprus, but the systematics of this group is poorly understood.
In Greece, six nominal species of Pseudamnicola have been distinguished based only on the shells (Schütt 1980; Szarowska et al. 2006). Four of those species are inhabiting the insular region [Pseudamnicola brachia (Westerlund, 1886), Pseudamnicola chia (Martens, 1889), Pseudamnicola negropontina (Clessin, 1878) and Pseudamnicola pieperi (Schütt 1980)]. Two are known from continental Greece [Pseudamnicola exilis (Frauenfeld, 1863) and Pseudamnicola macrostoma (Küster, 1853)]. The taxonomy of this genus is poorly understood, its shells are highly variable and its anatomy is simple (due to miniaturisation along with inevitably similar adaptations to internal fertilisation and development in freshwater) and variable even within a single population (Szarowska and Falniowski 2011). Therefore, morphology alone is insufficient to resolve the systematics of this group, hence the need for molecular data. For the Balkans Pseudamnicola, molecular data (cytochrome oxidase subunit I (COI), 18S) are available for P. macrostoma (Radea et al. 2013) from Attica, P. negropontina from Evvoia Island (Szarowska et al. 2006) and P. exilis (Szarowska and Falniowski 2011) from the southern Peloponnese and Kythira Island. However, it must be stressed that Schütt (1980) expanded the range of P. exilis for a large part of the Aegean, despite the fact that the species was described in northern Greece; thus, assignment of the Peloponnese and Kythira Island populations to P. exilis is doubtful and is certainly not representative of a single species (Szarowska and Falniowski 2011).
The aims of the present study were (i) to reveal, using molecular markers, the pattern of phylogeny of Pseudamnicola inhabiting the Aegean Islands; (ii) to describe and analyse the variation of the morphology of the shell and anatomical characters from 17 populations of Pseudamnicola from the Aegean Islands; and (iii) to find out which of the two models is more applicable to the island Pseudamnicola populations: either a model in which a relict fauna rich in endemics is differentiated in a way that mainly reflects the geological history of the area or a model in which a relatively young fauna is composed of more or less widely distributed taxa, with relatively high levels of gene flow among the springs they inhabit. To address the above issues, the morphology and the mitochondrial genes (COI and ribosomal 16S), as well as nuclear genes (ribosomal 18S, 28S and histone 3 (H3)), were analysed.
Materials and methods
Sample collection and fixation
The snails were collected from 17 freshwater spring populations across seven Aegean Islands (Table 1, Supplementary Fig. 1, Supplementary List 1) and analysed together with seven populations studied earlier (Table 1). Snails were fixed according to the procedures described in Szarowska et al. (2014a). The shells and soft parts (penis, female reproductive organs) were photographed with a CANON EOS 50D digital camera. For morphological comparisons, five males and five females from each locality were examined under a NIKON SMZ18 microscope with dark field (P-DF). The morphology of P. macrostoma and P. negropontina was described and illustrated in Szarowska et al. (2006) and in Szarowska and Falniowski (2011) for the populations from the Peloponnese and Kythira Island.
Morphometric techniques
We used a NIKON DS-5 digital camera measurement system to measure seven shell morphometric parameters: shell height, body whorl breadth, aperture height, spire height, aperture breadth, apex angle and angle between body whorl suture and horizontal surface (Szarowska 2006; Falniowski et al. 2012). For this analysis, ten randomly chosen specimens out of one population representing each of all nine clades, formed by sequences, were obtained in this study. They were as follows (clade/population): A1/A01, A2/A06, C/K03, CH/CH03, KAR/KAR01, N/N03, R1/R06, R2/R02 and TN/T02. The penes were measured in six dimensions (see figure caption) for each clade as well. The linear measurements were then logarithmically (log10) transformed; for angular measurements, the arcsine transformation was applied. We calculated Euclidean distances and computed minimum spanning tree (MST) using NTSYSpc (Rohlf 1998). The same program was used to compute principal component analysis (PCA), based on the matrix of correlation (Falniowski 2003). The original observations were projected into PC space, with a superimposed minimum spanning tree (for clarity, not presented in the figures) to detect local distortions in the data.
DNA extraction, PCR amplification and sequencing
DNA was extracted from foot tissue and hydrated in Tris-EDTA (TE) buffer (3 × 10 min); then, total genomic DNA was extracted with the SHERLOCK extracting kit (A&A Biotechnology), and the final product was resuspended in 20 μl TE buffer. The extracted DNA was stored at −80 °C at the Department of Malacology of Institute of Zoology of the Jagiellonian University in Kraków.
The following primers were used in the study: two mitochondrial gene fragments: (i) COI (with primers: LCOI490 5′-GGTCAACAAATCATAAAGATATTGG-3′; Folmer et al. 1994 and COR722b 5′-TAAACTTCAGGGTGACCAAAAAATYA-3′; Wilke and Davis 2000) and (ii) 16S (primers: LRJ12887 5′-CCGGTCTGAACTCAGATCACGT-3′ and LRN13398 5′-CGCCTGTTTAACAAAAACAT-3′; Kasper et al. 2004); two nuclear gene fragments: (iii) 18S (primers: SWAM18SF1 5′-GAATGGCTCATTAAATCAGTCGAGGTTCCTTAGATGATCCAAATC-3′ and SWAM 18SR1 5′-ATCCTCGTTAAAGGGTTTAAAGTGTACTCATTCCAATTACGGAGC-3′; Attwood et al. 2003) and (iv) 28S (primers: 28Sna1 5′-GACCC GTCTTGAAACACGGA-3′ and 28Sna2 5′-AGCCAATCCT TATCCCGAAG-3′, Kano et al. 2002); and (v) histone 3 (primers: H3F 5′-ATGGCTCGTACCAAGCAGACVGC-3′ and H3R 5′-ATATCCTTRGGCATRATRGTGAC-3′; Colgan et al. 1998).
The PCR conditions were as follows: for COI: initial denaturation step of 4 min at 94 °C, followed by 35 cycles of 1 min at 94 °C, 1 min at 55 °C, 2 min at 72 °C, and a final extension of 4 min at 72 °C; for 16S: initial denaturation step of 2 min 30 s at 90 °C, followed by 10 cycles of 50 s at 92 °C, 50 s at 43 °C, and 40 s at 72 °C, and 36 cycles of 30 s at 92 °C, 40 s at 44.4 °C, 40 s at 72 °C; for 18S: initial denaturation step of 4 min at 94 °C, followed by 40 cycles of 45 s at 94 °C, 45 s at 51 °C, 2 min at 72 °C; for 28S: initial denaturation step of 2 min at 94 °C, followed by 35 cycles of 30 s at 94 °C, 30 s at 50 °C, 90 s at 72 °C; and for H3: initial denaturation step of 2 min at 94 °C, followed by 35 cycles of 30 s at 94 °C, 30 s at 50 °C, 1 min at 72 °C, and after all cycles were completed, an additional elongation step of 4 min at 72 °C. Sequencing methods are described in Szarowska et al. (2014a).
Molecular analyses
For phylogenetic analysis, for COI, 16S and 18S, we used sequences of other available Pseudamnicola species from Greece: P. macrostoma from Attica (Radea et al. 2013; only COI), P. negropontina from Evvoia Island (Szarowska et al. 2006; only COI) and P. exilis from the southern Peloponnese and Kythira Island (Szarowska and Falniowski 2011) (Table 1). Adrioinsulana conovula (Frauenfeld, 1863) (Wilke et al. 2001) sequences were used as the outgroup.
Sequences were initially aligned in the MUSCLE (Edgar 2004) program in MEGA 6 (Tamura et al. 2013) and then checked in Bioedit 7.1.3.0 (Hall 1999). Haplotypes were distinguished with DnaSP 5.10 (Librado and Rozas 2009). The saturation test by Xia et al. (2003) performed with DAMBE (Xia 2013) revealed no saturation in any of the sequences studied.
Best-fit models of nucleotide evolution were selected according to the corrected Akaike Information Criteria (AICc) in jModelTest v. 2.1.4 (Guindon and Gascuel 2003; Darriba et al. 2012). The maximum likelihood approach was used to construct a phylogenetic tree with PhyML 3.0 (Guindon and Gascuel 2003). The Bayesian analyses were run with MrBayes ver. 3.2.3 (Ronquist et al. 2012) with default priors. Two simultaneous analyses were performed, each lasting 40,000,000 generations with one cold chain and three heated chains, starting from random trees and sampling trees every 1000 generations. The first 25 % trees were discarded as burn-in. The analyses were summarised on 50 % majority-rule tree. To infer haplotype networks of the markers used, a median-joining calculation was implemented in NETWORK 4.6.1.1 (Bandelt et al. 1999).
To apply the molecular clock, the data from COI were used. Two hydrobiids, Peringia ulvae Pennant, 1777 and Salenthydrobia ferreri (Wilke 2003; AF478401, AF478410), were used as outgroups (tree not shown). The divergence time between these two species was used to calibrate the molecular clock, with correction according to Falniowski et al. (2008). The likelihoods for trees with and without the molecular clock assumption for a likelihood ratio test (LRT) (Nei and Kumar 2000) were calculated with PAUP. The relative rate test (RRT) (Tajima 1993) was performed in MEGA. As Tajima’s RRTs and the LRT test rejected the equal evolutionary rate throughout the tree for Pseudamnicola, time estimates were calculated using a non-parametric rate smoothing (NPRS) analysis with the recommended Powell algorithm, in r8s v.1.7 for Linux (Sanderson 1997, 2003).
Although the aim of our study was not to delimit and/or describe species in the studied Pseudamnicola populations, some provisional distinction between local populations and putative species was useful. Thus, to provisionally delimit species from sequence data, without the prior definition of taxon and/or population, usually defined geographically, we used a procedure introduced by Pons et al. (2006), known as the generalised mixed Yule coalescent (GMYC) procedure. It is ‘general’ because it has relaxed some rather biologically unrealistic assumptions of the GMYC model, using a relaxed ultrametric tree and a single threshold approach. The multiple threshold approach gave similar results, with a few more species. Surely, there is no certain technique to delimit species with molecular data (e.g. Sauer and Hausdorf 2012); thus, we used this technique to gain some estimate only. For confirmation, we performed also the Automatic Barcode Gap Discovery (ABGD) analysis (Puillandre et al. 2012) in ‘abgd web’ (http://wwwabi.snv.jussieu.fr/cgi-bin/abgdWeb.cgi) selecting the K2P genetic distance and 100 steps (the remaining parameters were set to default). The ABGD estimates a maximum limit for intraspecific genetic divergence. After, this limit is used to separate sequences belonging to the same species (with smaller divergences) from sequences belonging to different species (with higher divergence level).
Results
Molecular phylogeny, preliminary species delimitation and time of divergence
In all the analyses, the topologies of the resulting phylograms were identical in both the maximum likelihood (ML) and Bayesian inference. The 18S sequences (401 bp, GenBank Accession numbers KT710543 - KT710575 ), as well as 28S sequences (512 bp, GenBank Accession numbers KT710576 - KT710594 ), show no differences between the studied populations (Table 1) and only slight differences with A. conovula (p-distance = 0.012) in 18S. In the H3 sequences (283 bp, GenBank Accession numbers KT710736 - KT710755 ), two haplotypes (p-distance = 0.007) were found. One of them was characteristic of populations from Rhodos and Karpathos, and the second was found in all the other populations.
We obtained 151 new sequences of COI (552 bp, GenBank Accession numbers KT710595 - KT710735), thus with the ones from GenBank (Table 1). In total, we analysed 189 sequences. The AICc selected the model GTR+I+G, with base frequencies of A = 0.257, T = 0.350, C = 0.215, G = 0.177, p-inv = 0.60 and G = 1.06. All analysed COI sequences were grouped into 44 haplotypes arranged into 16 main mitochondrial clades (Figs. 1 and 2, Table 1). p-distances between these clades are given in Table 2.
The maximum-likelihood phylogram (GTR+I+G, 10,000 bootstrap replicates) for the COI gene. The ML tree was rooted by using the Adrioinsulana conovula. Bootstrap support is shown only if above 70 %. Division into 16 clades (potential species) and connotations with earlier described Pseudamnicola species are also shown. Divergence times of main clades are also shown
The median-joining haplotype network of COI haplotypes. The colours indicate the specific clades, see Fig. 1
We also obtained 61 sequences of 16S ribosomal RNA (rRNA) (312 bp, GenBank Accession numbers KT710482 - KT710542 ), representing all clades discovered in COI analyses. 16S rRNA sequences showed no intrapopulation polymorphism and were unique for each population, with the exception of populations A03 and A04 from Andros that have the same haplotype. We analysed COI, 16S rRNA and 18S sequences together, since a partition homogeneity test indicated no conflicting phylogenetic signals between the datasets (p = 0.983). The AICc selected the model GTR+I+G, with base frequencies of A = 0.295, T = 0.336, C = 0.185, G = 0.183, p-inv = 0.790 and G = 0.016. The topology of this tree was identical with the COI tree but without two reference sequences (Fig. 3). For this reason, we performed the GMYC and ABGD analysis, as well as estimates of the time of divergence for the COI dataset.
The maximum-likelihood phylogram for the COI+16S rRNA+18S gene. The ML tree was rooted by using the Adrioinsulana conovula. Bootstrap support (>75 %) and Bayesian posterior probabilities (>0.95) were shown. Division into 16 clades (potential species) and connotations with earlier described Pseudamnicola species are also shown
The GMYC procedure, with a single threshold, gave the following results: likelihood for the null model = 290.7919, ML of the GMYC model = 295.343, LR = 9.102096, p = 0.02796, number of ML clusters = 13 (confidence interval = 10–15) and number of ML entities (including Adrioinsulana) = 17 (confidence interval = 13–22) (Supplementary Fig. 2). These data were confirmed by the ABGD method, which detected 17 stable candidate species (including Adrioinsulana), with estimated prior maximum divergence of intraspecific diversity (P) as large as 0.28 % (one-tail 95 % confidence interval). Sixteen Pseudamnicola entities (Fig. 4) coincide with the clades of the COI ML tree, with an exception of clade PN (P. negropontina), which belongs to clade TN in the ML tree. We will treat all these entities as molecular operational taxonomical unit (MOTUs) in the remaining part of the paper. Those MOTUs could be understood as ‘presumed species’, but all the data available are not sufficient for the exact delineation of the species in Pseudamnicola, and on the other hand, exact species delimitation was not one of our aims.
The two sequences formed divergent clades: PN and PM for P. negropontina and P. macrostoma, respectively (Fig. 1). Closest to PN is the TN clade, which consists of all the haplotypes from Tinos and one haplotype from Naxos. Two other haplotypes from Naxos form one separate clade, N. Haplotypes from Andros form two distinguished clades: A1 and A2. All haplotypes from Khios are grouped into one clade, CH, and the same applies to the case of Crete (clade C). The sequences from the Peloponnese form three distinct clades, whose names correspond to sample sites (Szarowska and Falniowski 2011): the most divergent are P1 (from the southernmost population) and P7 (from the western population). The third Peloponnese clade (P2) is closest to the Cretan clade (C). On Kythira, two separate clades (KIT4 and KIT3) were identified, representatives of each inhabiting one of the two studied populations. Kythira populations showed the highest divergence of all the islands studied. Similar to the 16S analysis, the haplotypes from Rhodes and Karpathos formed a distinct cluster, consisting of three clades. One of these (KAR) contains haplotypes from Karpathos and is close to one of two Rhodes clades (R2), which consists of sequences from the two southernmost Rhodes populations. The other Rhodes clade (R1) is more divergent from the northern population, both according to the COI and concatenated trees.
The results of the COI dating analysis are presented in Fig. 1. All of the populations from the northern and central Aegean area split less than 2 Mya. The two oldest clades, P1 and P7, emerged on the Peloponnese (4.52 and 3.27 Mya, respectively). The Rhodes and Karpathos eastern cluster separated 2.93 Mya. The Cretan clade separated 2.09 Mya, while the divergence time of the remaining western populations from the Peloponnese and Kythira was 2.4–2.55 Mya.
Morphological differentiation
Shell measurements are presented in Table 3. For clarity, PCA results are presented in one figure for Andros (A1, A2), Tinos, Naxos and Khios clades, and in another for Crete, Karpathos and Rhode (R1, R2) clades. For the first group, all the linear parameters had high character loading on PC1, and only spire height had smaller and the apex angle had the lowest loading for both dimensions. The highest loading for PC2 was the spire height and relatively high shell height and angle between the body whorl and horizontal surface (β). For the second group, all the linear dimensions had high and nearly identical loadings, the lowest being the apex angle. In PC2, the apex angle had the highest loading, the β angle had much lower and all the linear parameters had low loadings.
For the first group (Andros (A1, A2), Tinos, Naxos and Khios clades), the most ‘decisive’ characters were the spire height and β angle. Shells from the five populations from Andros Island (Fig. 5 A–R) represent two molecular clades (A1 and A2): shells from populations A01 (A–E) and A08 (Q–R) are low-spired (Table 3; clade A1). With one exception, all the A1 representatives are grouped together both by PCA (Fig. 8) and MST (Supplementary Fig. 3). Shells from populations A03 (F–I), A04 (J–L) and A06 (M–P) belonged to clade A2. The A2 clade markedly differs from clade A1 in PCA (Fig. 8). With the exception of three specimens mixed with clades CH and TN, all the representatives of A2 are joined by MST (Supplementary Fig. 3).
In populations from Tinos Island (Fig. 6A–F), all of the shells were small and low-spired (Table 3; clade TN). PCA (Fig. 8) as well as MST (Supplementary Fig. 3) mixes the clade TN with the clade CH from Khios Island representing P. chia (Fig. 6 N–U). The population N03 from Naxos Island (Fig. 6J–M) belongs to clade N (Table 3) whose distinctness confirms both PCA (Fig. 8) and MST (Supplementary Fig. 3).
For the second group (Crete, Karpathos and Rhode, (R1, R2) clades), they differ mostly in the apex angle. The shells of P. brachia from Crete Island (Fig. 7A–D) and the ones of Pseudamnicola from Rhode representing the clade R2 (population R02 (H–M) and R04 (N–Q), in PCA (Fig. 8)) were somewhat mixed, although MST (Supplementary Fig. 3) joins more representatives of the same group that could be seen in PCA. The shells of P. pieperi from Karpathos Island (Fig. 7 E–G) representing the clade KAR (Table 3), and the ones from populations R06 (R–U) from Rhode, representing the clade R1, with the exception of one outlier, were distinct both in PCA (Fig. 8) and MST (Supplementary Fig. 3).
Shell biometry (PCA) of Pseudamnicola for nine main clades, see COI phylogenetic tree. Measurements of shell were shown: a shell height, b body whorl breadth, c aperture height, d spire height, e aperture breadth, α apex angle and β angle between body whorl suture and horizontal surface. PC1 explains 58.73 % and PC2 explains 18.52 %, PC1 and PC2 cumulatively 77.25 % of the total variance for the Andros/Tinos/Khios/Naxos clades (above) and PC1 75.72 % and PC2 12.12 % cumulatively 87.84 % for the Rhodes/Karpathos/Crete clades (below)
The female reproductive organs of P. chia, as not yet described and illustrated in the literature, are presented in Fig. 9. In general, the female reproductive organs showed no differences between the presumed species. The penes of the studied Pseudamnicola (Fig. 10A–Z) were characterised by a simple form, with no prominent outgrowths and were highly variable in size. In PCA on the penes, the highest loading in PC1 was the breadth 0.25 of the length from the tip (5); in PC2, the highest loading was the length (1) and outline length (6). PCA with superimposed MST (Fig. 11) shows some distinctness of A2 TN or C clade, but in general, the variability widely overlaps.
Renal and pallial section of the female reproductive organs of Pseudamnicola chia from Khios Island, locality CH02: bc bursa copulatrix, cbc duct of bursa copulatrix, dgp gono-pericardial duct, ga albuminoid gland, gn nidamental gland, gp gonoporus, ov pallial oviduc, ovl loop of (renal) oviduct, rs receptaculum seminis, vc ventral canal
Discussion
The present research characterised 17 newly studied populations of Pseudamnicola from seven Aegean Islands and compared them with available data from five populations from the Peloponnese and Kythira Island and with two populations from Attica and Evvoia Island. This comparison allowed the detailed pattern of phylogeny of the Pseudamnicola in the Aegean area to be revealed.
Molecular phylogeny and putative species delimitation
The freshwater truncatelloideans in the Balkans appeared in the Palaeogene period, if not earlier (Kabat and Hershler 1993). Therefore, they are suitable for the evaluation of old (pre-Pleistocene) biogeographic relationships. Many of the present-day representatives of the freshwater Rissooidea are considered relicts, the distributions of which trace the Neogene drainage patterns fragmented by subsequent changes in climate and landscape (Hershler and Liu 2004). This hypothesis involves two conditions: (i) the springs inhabited by those obligatorily aquatic animals are stable habitats; and (ii) the gene flow among them, if at all present, is very low. In the abundant literature on the phylogeny and population genetic structure and gene flow of spring fauna, gastropods are well represented (e.g. Colgan and Ponder 1994; Ponder et al. 1995; Falniowski et al. 1998, 1999, 2009; Bohonak 1999; Hershler and Liu 2004; Brändle et al. 2005; Hershler et al. 2005; Falniowski and Szarowska 2011). Most studies indicate low levels of gene flow and high levels of endemism in spring snails (e.g. Colgan and Ponder 1994; Ponder et al. 1995; Finston and Johnson 2004), but other studies showed that some species were rather widespread, with high levels of gene flow among their populations (Falniowski et al. 1998, 1999; Hershler et al. 2005).
Phylogenetic analysis of Pseudamnicola indicates three main conclusions: (i) the studied populations of Pseudamnicola form 16 clades, according to the GMYC and ABGD results probably of species rank; (ii) intrapopulation differentiation level is generally low, slightly larger in populations occupying the mainland and islands close to the continent; and (iii) divergence times for most clades are less than 3 Mya. The short time of divergence, inferred from the low mitochondrial differentiation, suggesting thus close relationships between the studied Pseudamnicola populations, is confirmed by the low or no differentiation in the three studied nuclear loci.
According to the phylogenetic analysis and p-distance values for COI, the studied Pseudamnicola populations can be assigned to 16 clades, most of which are probably distinct species, according to the GMYC and ABGD analysis. The GMYC procedure has been applied in several previous studies (Barraclough et al. 2009; Papadopoulou et al. 2009a, b; Adolfsson et al. 2010; Pagès et al. 2010; Powell et al. 2011; Vuataz et al. 2011), including some on molluscs (Nekola et al. 2009; Lorion et al. 2010; Falniowski and Szarowska 2012; Jörger et al. 2012; Jörger and Schrödl 2013). Reliability of species delimitated by GMYC has also been discussed in the literature (e.g. Lohse 2009; Papadopoulou et al. 2009b; Jörger et al. 2012; Jörger and Schrödl 2013; Sauer and Hausdorf 2012), also with substantial criticism, and the method was further developed (Monaghan et al. 2009). However, in our work, the GMYC results were confirmed by ABGD analysis. Surely, with one short DNA fragment, species level distinction is always ambiguous; thus, we treat those clades as presumed species only, not attempting to distinguish and describe them formally—the latter should wait for more data available.
According to morphological criteria, six species of Pseudamnicola were distinguished in Greece (Schütt 1980; Szarowska et al. 2006). P. brachia was described from Crete (corresponding with clade C), P. chia from Khios (clade CH) and P. negropontina from Evvoia Island and may probably inhabit continental Greece as well (clade PN). P. pieperi is known from Karpathos (clade KAR). According to our results, P. pieperi may occur also on Rhodes Island; however, clades belonging to this cluster are more divergent, which rather suggests the existence of three rather than one species on these two islands. Recently, two molecularly distinct species from Rhodos were described (Radea et al. 2015). Similarly, P. negropontina may occupy the islands of Tinos, Naxos and probably Andros, but GMYC results indicate that there are four species in this area, with one case of sympatry (at locality A03). Two Pseudamnicola species come from the continent (Schütt 1980). P. macrostoma inhabits the northern Peloponnese and continental Greece. Unfortunately, we do not have well-fixed specimens of Pseudamnicola from this region, apart from the reference sequence (Radea et al. 2013). The second species, according to Schütt (1980), within the range of P. exilis (drawn of the map but not described and far from the type locality—see above) has been studied in the southern part of the Peloponnese and Kythira Island (Szarowska and Falniowski 2011). However, this presumed ‘P. exilis’ represented at least four distinct species, including one from Kythira Island, which was highly polymorphic (Fig. 29 in Szarowska and Falniowski 2011).
Morphological differentiation
Differences were noted in the shell habitus (height/width ratio, spire height, shell size) between the two presumed species of Pseudamnicola from the Peloponnese (population P01 vs P02 and P07) and two from the two localities in Kythira Island (Szarowska and Falniowski 2011: Figs. 2–14). No differences—apart from size—were found in the morphology of the penis and female reproductive organs between those populations (Szarowska and Falniowski 2011: Figs. 15–25 and 26); the lack of taxonomically useful characters and variable degrees of fixation-caused contraction may be an explanation for this.
On Andros Island, where there were three presumed species, shells were slightly variable and intrapopulation variability was sometimes higher than interpopulation variability. At locality A03, inhabited sympatrically by two presumed species, shell variation did not coincide with genotypic differences. Only the shells from localities A05 (not presented) and A06 were distinguishable, with their high penultimate whorls, from those of the other populations/species. However, PCA showed distinctness of the two Andros clades. The difference between the shells from the Naxos populations, N01 and N03, representing two presumed species, is clearly visible although it only concerns their size, and this is also reflected by their distinctness in PCA, mostly along the PC1 axis (TN versus N). Indeed, the shells from locality N03 do not differ from the ones from the two Tinos localities, representing the same presumed species. The shells of P. chia in PCA show distinctness from clade A1, but are more or less mixed with the shells of the other Pseudamnicola from the studied area.
In addition, there were no apparent differences distinguishing the shells of P. brachia (clade C) and P. pieperi (clade KAR), although PCA grouped them separately. On Rhodes Island, shells from localities R02 and R04 were characterised by their expanded apertures and differed in size and proportion from the shells in locality R06, representing another presumed species. PCA confirmed the distinctness of R01 and R02.
In the literature, the Greek Pseudamnicola species are distinguished through the application of only two criteria: shell morphology and geographic distribution (Schütt 1980). Our results certainly do not confirm those criteria. Morphometry shows some differences in the shell morphology of the particular taxa, but those differences concern no qualitative characters routinely used in species descriptions, like shape, general habitus, etc.
A similar lack of differences between the species in the characters of the female reproductive organs in Pseudamnicola was found by Szarowska and Falniowski (2011). Furthermore, the penes of the four presumed species from the islands of Andros, Tinos and Naxos are indistinguishable, as are those from the Peloponnese and Kythira Island (Szarowska and Falniowski 2011). In contrast, the penes of P. chia and P. brachia, although characteristic, cannot be used to distinguish these two species. Similar penes could also be found in the Pseudamnicola inhabiting Rhodes Island. Thus, in general, it is difficult to use penis characters to distinguish species within the Greek Pseudamnicola.
Female reproductive organ characteristics, such as receptacula, bursae copulatrix, etc., as well as penis morphology, have been found to be useful characteristics to infer phylogeny at higher levels, at least at the genus level, but certainly not at the species level (Szarowska 2006; Szarowska and Falniowski 2008). Pseudamnicola seems to be a further example of morphostatic radiation, as defined by Davis (1992), which is so common in truncatelloid snails (Wilke and Davis 2000; Wilke and Falniowski 2001; Falniowski and Szarowska 2011; Falniowski et al. 2012).
History of Pseudamnicola at the Aegean Islands
In fact, high genotypic diversity was noted on the islands of Kythira and Rhodes—two islands close to the mainland. Similarly high or even higher differences were noted between the Peloponnese populations. This may suggest that the continent offers conditions conducive to the longevity of metapopulations and may serve as a source of migrants, transported passively, most probably by birds. Two presumed species on Rhodes Island, which that are situated a few kilometres from each other, differ more from one another than each of them does from P. pieperi from the distant Karpathos Island. The relatively low genetic distances between the Cretan P. brachia and the other species despite the long distance and time of isolation of Crete are also notable. In contrast, relatively high diversity in the Andros/Tinos/Naxos complex is similar to that noted for plants by Kougioumoutzis et al. (2014). Another striking pattern is shown by the relatively close relationships between this group of islands and Khios, which could be explained by a nearly complete land bridge in the Late Pleistocene. The estimated times of divergence of Bythinella in continental Greece (Falniowski and Szarowska 2011) unexpectedly suggest that this genus has a rather short history in the studied area. We have found the same to be true in our study of Pseudamnicola.
According to geologic and climatic events, some key dates can be highlighted: the initiation of the mid-Aegean Trench split (around 12 Mya), its ending (around 9 Mya) and the Messinian crisis (around 5 Mya). The modern post-Alpine European topography in the Mediterranean emerged in the Late Tortonian (8 Mya) (Kostopoulos 2009). In the Upper Miocene (around 6 Mya), the Peloponnese and Evvoia were part of a continuous land mass with the continent (Popov et al. 2004). In the Middle-Late Pliocene (3.5–1.8 Mya), Evvoia was still a part of the continent, but the Peloponnese became separated from the continent by a broad strait (Creutzburg 1963; Dermitzakis and Papanikolaou 1981; Dermitzakis 1990; Popov et al. 2004). During the glacial maxima, the sea level was as much as 200 m lower than today (Beerli et al. 1996). During the glacial maxima in the Pleistocene, on the Greek island Andikythira (16 kya, and thus during the last glacial maximum: LGM) temperatures were some 5–8 °C below present levels, and climatic conditions were slightly drier (Digerfeldt et al. 2000; Gittenberger and Goodfriend 2006). For Bythinella from continental Greece, the estimated time of divergence did not exceed 4 Mya (Falniowski and Szarowska 2011). Schilthuizen et al. (2004) estimated a similarly short time for the Cretan Albinaria, at 3.5 Mya. In the Pliocene, some land connections disappeared due to a rise in sea level (Dermitzakis and Papanikolaou 1981; Dermitzakis 1990), but the present Peloponnese was isolated from the mainland, which coincides, in our study, with the dating of the divergence time between two of the three Peloponnese Pseudamnicola populations and the third one.
The Aegean Islands are divided into the following five groups: those associated with (i) Asia Minor (Khios and Rhodes in our study); (ii) the eastern shelf of Greece (Evvoia in our study); (iii) the Crete/Karpathos complex; (iv) the Cyclades that form a central, independent shelf (Andros, Tinos and Naxos in our study); and (v) Kythira. The central group (Cyclades) has been separated from Europe since the Middle Pleistocene (Foufopoulos and Ives 1999). Since the peak of the Wisconsin-Würm glaciation (24–10 kya), global sea level has risen 120–130 m and, in the present Aegean Islands, replaced dry land. Between 350 and 250 kya, the sea area was restricted again, and almost 50–60 % of the present Aegean Sea was land with extensive drainage systems and large lakes (Lykousis 2009). In the Late Pleistocene, the eastern Aegean Islands were connected with Anatolia, and the strait between Europe and the Cyclade-mega-island, connecting current-day Naxos, Paros, Tinos and Andros, was rather narrow (Kougioumoutzis et al. 2014). The present geography of the Aegean is no more than 8–6 kya old (Kougioumoutzis et al. 2014).
Levels of gene flow, as well as metapopulation structure in general, depend on several species-specific traits (Bohonak 1999; Bilton et al. 2001; Myers et al. 2001). The model of allopatric fragmentation and/or vicariant speciation in the Aegean Pseudamnicola does not agree with the mtCOI sequence divergence data, suggesting the relatively recent origin of the observed differentiation, which is attributable to geologic events only in some cases (see above). In the same manner, vicariance cannot explain the whole pattern in the land snail Albinaria (Douris et al. 1998, 2007) and other invertebrates and vertebrates in the Aegean (Dennis et al. 2000; Kasapidis et al. 2005). Just as in the central European Bythinella (Falniowski 1987; Falniowski et al. 1998, 1999; Szarowska 2000) and the Greek mainland Bythinella (Falniowski and Szarowska 2011), the present data did not confirm either the complete isolation of the Aegean Pseudamnicola island populations or their stability and longevity, thus confirming the second model in which a relatively young fauna is composed of more or less widely distributed taxa, with relatively high levels of gene flow among the springs they inhabit. For inhabitants of such miniature and changeable habitats, survival must depend on dynamic processes of colonisation and recolonisation events coupled with short- or (in some cases) long-distance dispersal. Pseudamnicola is not a typical oligostenothermic inhabitant of springs as it can also be found in streams, and it is somewhat amphibiotic as it can be found crawling and feeding above the water (personal observations), thus seeming capable of passive transportation by birds (for review of the literature on bird transportation, see Falniowski and Szarowska 2011).
References
Adolfsson, S., Michalakis, Y., Paczesniak, D., Bode, S. N. S., & Butlin, R. K. (2010). Evaluation of elevated ploidy and asexual reproduction as alternative explanations for geographic parthenogenesis in Eucypris virens ostracods. Evolution, 64, 986–997.
Attwood, S. W., Ambu, S., Meng, X.-H., Upatham, E. S., Xu, F.-S., & Southgate, V. R. (2003). The phylogenetics of Triculine snails (Rissooidea: Pomatiopsidae) from South-East Asia and southern China: historical biogeography and the transmission of human schistosomiasis. Journal of Molluscan Studies, 69, 263–271.
Bandelt, H. J., Forster, P., & Röhl, A. (1999). Median-joining networks for inferring intraspecific phylogenies. Molecular Biology and Evolution, 16, 37–48.
Barraclough, T. G., Hughes, M., Ashford-Hodges, N., & Fujisawa, T. (2009). Inferring evolutionarily significant units of bacterial diversity from broad environmental surveys of single-locus data. Biological Letters, 5, 425–428.
Beerli, P., Hotz, H., & Uzzell, T. (1996). Geologically dated sea barriers calibrate a protein clock for Aegean water frogs. Evolution, 50, 1676–1687.
Bilton, D. T., Freeland, J. R., & Okamura, B. (2001). Dispersal in freshwater invertebrates. Annual Reviews of Ecology and Systematics, 32, 159–181.
Bohonak, A. J. (1999). Dispersal, gene flow and population structure. The Quaternary Review of Biology, 74, 21–45.
Brändle, M., Westermann, I., & Brandl, R. (2005). Gene flow between populations of two invertebrates in springs. Freshwater Biology, 50, 1–9.
Colgan, D. J., & Ponder, W. F. (1994). The evolutionary consequences of restrictions on gene flow: examples from Hydrobiid snails. The Nautilus, 108, 25–43.
Colgan, D. J., McLauchlan, A., Wilson, G. D. F., Livingston, S., Edgecombe, G. D., Macaranas, J., Cassis, G., & Gray, M. R. (1998). Histone H3 and U2 snRNA sequences and arthropod molecular evolution. Australian Journal of Zoology, 46, 419–437.
Creutzburg, N. (1963). Palaeogeographic evolution of Crete from Miocene till our days. Cretean Annals, 15(16), 336–342.
Darriba, D., Taboada, G. L., Doallo, R., & Posada, D. (2012). jModelTest 2: more models, new heuristics and parallel computing. Nature Methods, 9, 772.
Davis, G. M. (1992). Evolution of prosobranch snails transmitting Asian Schistosoma: coevolution with Schistosoma: a review. Progress in Clinical Parasitology, 3, 145–204.
Dennis, R. L. H., Shreeve, T. G., Olivier, A., & Coutsis, J. G. (2000). Contemporary geography dominates butterfly diversity gradients within the Aegean archipelago (Lepidoptera: Papilionoidea, Hesperioidea). Journal of Biogeography, 27, 1365–1383.
Dermitzakis, M. D. (1990). Palaeogeography, geodynamic processes and event stratigraphy during the Late Cenozoic of the Aegean area. International Symposium on Biogeographical Aspects of Insularity, Roma 1987. Accademia Nazionale dei Lincei, 85, 263–288.
Dermitzakis, M. D., & Papanikolaou, D. J. (1981). Palaeogeography and geodynamics of the Aegean region during the Neogene. Annales Géologique des Pays Helléniques, 30, 245–289.
Digerfeldt, G., Olsson, S., & Sandgren, P. (2000). Reconstruction of lake-level changes in lake Xinias, central Greece, during the last 40 000 years. Palaeogeography, Palaeoclimatology, Palaeoecology, 158, 65–82.
Douris, V., Cameron, R. A. D., Rodakis, G. C., & Lecanidou, R. (1998). Mitochondrial phylogeography of the land snail Albinaria in Crete: long-term geological and short-term vicariance effects. Evolution, 52, 116–125.
Douris, V., Giokas, S., Thomaz, D., Lecanidou, R., & Rodakis, G. C. (2007). Inference of evolutionary patterns of the land snail Albinaria in the Aegean archipelago: is vicariance enough? Molecular Phylogenetics and Evolution, 44, 1224–1236.
Edgar, R. C. (2004). MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Research, 32, 1792–1797.
Falniowski, A. (1987). Hydrobioidea of Poland (Prosobranchia: Gastropoda). Folia Malacologica, 1, 1–122.
Falniowski, A. (2003). Metody numeryczne w taksonomii [Numerical techniques in taxonomy]. Kraków: Wydawnictwo Uniwersytetu Jagiellońskiego.
Falniowski, A., & Szarowska, M. (2011). Radiation and phylogeography in a spring snail Bythinella (Mollusca: Gastropoda: Rissooidea) in continental Greece. Annales Zoologici Fennici, 48, 67–90.
Falniowski, A., & Szarowska, M. (2012). Sequence-based species delimitation in the Balkan Bythinella Moquin-Tandon, 1856 (Gastropoda: Rissooidea) with general mixed Yule coalescent model. Folia Malacologica, 20, 111–120.
Falniowski, A., Szarowska, M., Fiałkowski, W., & Mazan, K. (1998). Unusual geographic pattern of interpopulation variation in a spring snail Bythinella (Gastropoda, Prosobranchia). Journal of Natural History, 32, 605–616.
Falniowski, A., Mazan, K., & Szarowska, M. (1999). Homozygote excess and gene flow in the spring snail Bythinella (Gastropoda: Prosobranchia). Journal of Zoological Systematics and Evolutionary Research, 37, 165–175.
Falniowski, A., Szarowska, M., Sirbu, I., Hillebrand, A., & Baciu, M. (2008). Heleobia dobrogica (Grossu & Negrea, 1989) (Gastropoda: Rissooidea: Cochliopidae) and the estimated time of its isolation in a continental analogue of hydrothermal vents. Molluscan Research, 28, 165–170.
Falniowski, A., Szarowska, M., & Sirbu, I. (2009). Bythinella Moquin-Tandon, 1856 (Gastropoda: Rissooidea: Bythinellidae) in Romania: species richness in a glacial refugium. Journal of Natural History, 43, 2955–2973.
Falniowski, A., Szarowska, M., Glöer, P., & Pešić, V. (2012). Molecules vs morphology in the taxonomy of the Radomaniola/Grossuana group of Balkan Rissooidea (Mollusca: Caenogastropoda). Journal of Conchology, 41, 19–36.
Finston, T. L., & Johnson, M. S. (2004). Geographic patterns of genetic diversity in subterranean amphipods of the Pilbara, Western Australia. Marine and Freshwater Research, 55, 619–628.
Folmer, O., Black, M., Hoeh, W., Lutz, R. A., & Vrijenhoek, R. C. (1994). DNA primers for amplification of mitochondrial cytochrome c oxidase subunit I from diverse metazoan invertebrates. Molecular Marine Biology and Biotechnology, 3, 294–299.
Foufopoulos, J., & Ives, A. R. (1999). Reptile extinctions on land-bridge islands: life-history attributes and vulnerability to extinction. The American Naturalist, 153, 1–25.
Gittenberger, E., & Goodfriend, G. A. (2006). Land snails from the last glacial maximum on Andikithira, southern Greece and their palaeoclimatic implications. Journal of Quaternary Science, 8, 109–116.
Guindon, S., & Gascuel, O. (2003). A simple, fast and accurate algorithm to estimate large phylogenies by maximum likelihood. Systematic Biology, 52, 696–704.
Hall, T. A. (1999). BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acids Symposium Series, 41, 95–98.
Hershler, R., & Liu, H.-P. (2004). A molecular phylogeny of aquatic gastropods provides a new perspective on biogeographic history of the Snake River region. Molecular Phylogenetics and Evolution, 32, 927–937.
Hershler, R., Mulvey, M., & Liu, H.-P. (2005). Genetic variation in the Desert Springsnail (Tryonia porrecta): implications for reproductive mode and dispersal. Molecular Ecology, 14, 1755–1765.
Jörger, K. M., & Schrödl, M. (2013). How to describe a cryptic species? Practical challenges of molecular taxonomy. Frontiers in Zoology, 10, 1–27.
Jörger, K. M., Norenburg, J. L., Wilson, N. G., & Schrödl, M. (2012). Barcoding against a paradox? Combined molecular species delineations reveal multiple cryptic lineages in elusive meiofaunal sea slugs. BMC Evolutionary Biology, 12(245), 1–18.
Kabat, A. R., & Hershler, R. (1993). The prosobranch snail family Hydrobiidae (Gastropoda: Rissooidea): review of classification and supraspecific taxa. Smithsonian Contributions to Zoology, 547, 1–94.
Kano, Y., Chiba, S., & Kase, T. (2002). Major adaptive radiation in neritopsine gastropods estimated from 28S rRNA sequences and fossil records. Proceedings of the Royal Society of London B, 269, 2457–2465.
Kasapidis, P., Magoulas, A., Mylonas, M., & Zouros, E. (2005). The phylogeography of the gecko Cyrtopodion kotschyi (Reptilia: Gekkonidae) in the Aegean archipelago. Molecular Phylogenetics and Evolution, 35, 612–623.
Kasper, M. L., Reeson, A. F., Cooper, S. J. B., Perry, K. D., & Austin, A. D. (2004). Assessment of prey overlap between a native (Polistes humilis) and an introduced (Vespula germanica) social wasp using morphology and phylogenetic analyses of 16S rDNA. Molecular Ecology, 13, 2037–2048.
Kostopoulos, D. S. (2009). The Pikermian Event: temporal and spatial resolution of the Turolian large mammal fauna in SE Europe. Palaeogeography, Palaeoclimatology, Palaeoecology, 274, 82–95.
Kougioumoutzis, K., Simaiakis, S. M. & Tiniakou, A. (2014). Network biogeographical analysis of the central Aegean archipelago. Journal Biogeography, 1–11. doi:10.1111/jbi.12342.
Librado, P., & Rozas, J. (2009). DnaSP v5 a software for comprehensive analysis of DNA polymorphism data. Bioinformatics, 25, 1451–1452.
Lohse, K. (2009). Can mtDNA barcodes be used to delimit species? A response to Pons et al. (2006). Systematic Biology, 58, 439–442.
Lorion, J., Buge, B., Cruaud, C., & Samadi, S. (2010). New insights into diversity and evolution of deep-sea Mytilidae (Mollusca: Bivalvia). Molecular Phylogenetics and Evolution, 57, 71–83.
Lykousis, V. (2009). Sea-level changes and shelf break prograding sequences during the last 400 ka in the Aegean margins: subsidence rates and palaeogeographic implications. Continental Shelf Research, 29, 2037–2044.
Myers, M. J., Sperling, F. A. H., & Resh, V. H. (2001). Dispersal of two species of Trichoptera from desert springs: Conservation implications for isolated vs connected populations. Journal of Insect Conservation, 5, 207–215.
Monaghan, M. T., Wild, R., Elliot, M., Fujisawa, T., Balke, M., Inward, D. J., Lees, D. C., Ranaivosolo, R., Eggleton, P., Barraclough, T. G., & Vogler, A. P. (2009). Accelerated species inventory on Madagascar using coalescent-based models of species delineation. Systematic Biology, 58, 298–311.
Nei, M., & Kumar, S. (2000). Molecular evolution and phylogenetics. Oxford: Oxford University Press.
Nekola, J. C., Coles, B. F., & Bergthorsson, U. (2009). Evolutionary pattern and process within the Vertigo gouldii (Mollusca: Pulmonata, Pupillidae) group of minute North American land snails. Molecular Phylogenetics and Evolution, 53, 1010–1024.
Pagès, M., Chaval, Y., Herbreteau, V., Waengsothorn, S., Cosson, J. F., Hugot, J.-P., Morand, S., & Michaux, J. (2010). Revisiting the taxonomy of the Rattini tribe: a phylogeny-based delimitation of species boundaries. BMC Evolutionary Biology, 10, 184.
Papadopoulou, A., Anastasiou, I., Keskin, B., & Vogler, A. P. (2009a). Comparative phylogeography of tenebrionid beetles in the Aegean archipelago: the effect of dispersal ability and habitat preference. Molecular Ecology, 18, 2503–2517.
Papadopoulou, A., Monaghan, M. T., Barraclough, T. G., & Vogler, A. P. (2009b). Sampling error does not invalidate the Yule-coalescent model for species delimitation. A response to Lohse (2009). Systematic Biology, 58, 442–444.
Ponder, W. F., Eggler, P., & Colgan, D. J. (1995). Genetic differentiation of aquatic snails (Gastropoda: Hydrobiidae) from artesian springs in arid Australia. Biological Journal of the Linnean Society, 56, 553–596.
Pons, J., Barraclough, T. G., Gomez-Zurita, J., Cardoso, A., Duran, D. P., Hazell, S., Kamoun, S., Sumlin, W. D., & Vogler, A. P. (2006). Sequence-based species delimitation for the DNA taxonomy of undescribed insects. Systematic Biology, 55, 595–609.
Popov, S. V., Rögl, F., Rozanov, A. Y., Steininger, F. F., Shcherba, I. G., & Kovac, M. (2004). Lithological-paleogeographic maps of the Paratethys. 10 maps Late Eocene to Pliocene. Courier Forschungsinstitut Senckenberg, 250, 1–46.
Powell, J. R., Monaghan, M. T., Opik, M., & Rillig, M. C. (2011). Evolutionary criteria outperform operational approaches in producing ecologically relevant fungal species inventories. Molecular Ecology, 20, 655–666.
Puillandre, N., Lambert, A., Brouillet, S., & Achaz, G. (2012). ABGD, Automatic Barcode Gap Discovery for primary species delimitation. Molecular Ecology, 21, 1864–1877.
Radea, C., Parmakelis, A., Papadogiannis, V., Charou, D., & Triantis, K. A. (2013). The hydrobioid freshwater gastropods (Caenogastropoda, Truncatelloidea) of Greece: new records, taxonomic reassessments using DNA sequence data and an update of the IUCN Red List Categories. ZooKeys, 350, 1–20.
Radea, C., Parmakelis, A., Velentzas, A. D., & Triantis, K. A. (2015). Systematics of Pseudamnicola (Gastropoda: Hydrobiidae): description of two new species from insular Greece and redescription of P. pieperi Schütt, 1980. Journal of Molluscan Studies, doi:10.1093/mollus/eyv031.
Rohlf, F. J. (1998). NTSYSpc, numerical taxonomy and multivariate analysis system. Version 2.0. [Computer software and manual] Exeter Software, Seatuket, NY.
Ronquist, F., Teslenko, M., van der Mark, P., Ayres, D. L., Darling, A., Höhna, S., Larget, B., Liu, L., Suchard, M. A., & Huelsenbeck, J. P. (2012). Efficient Bayesian phylogenetic inference and model choice across a large model space. Systematic Biology, 61, 539–542.
Sanderson, M. J. (1997). A nonparametric approach to estimating divergence times in the absence of rate constancy. Molecular Biology and Evolution, 14, 1218–1231.
Sanderson, M. J. (2003). R8s: inferring absolute rates of molecular evolution, divergence times in the absence of a molecular clock. Bioinformatics, 19, 301–302.
Sauer, J., & Hausdorf, B. (2012). A comparison of DNA-based methods for delimiting species in a Cretan land snail radiation reveals shortcomings of exclusively molecular taxonomy. Cladistics, 28, 300–316.
Schilthuizen, M., Gutteling, E., Van Moorsel, C. H. M., Welter-Schultes, F. W., Haase, M., & Gittenberger, E. (2004). Phylogeography of the land snail Albinaria hippolyti (Pulmonata: Clausiliidae) from Crete, inferred from ITS-1 sequences. Biological Journal of the Linnean Society, 83, 317–326.
Schütt, H. (1980). Zur Kenntnis griechischer Hydrobiiden. Archiv füer Molluskenkunde, 110, 115–149.
Szarowska, M. (2000). Environmental stress and stability of Bythinella populations in South Poland (Gastropoda: Prosobranchia: Hydrobioidea). Malakologische Abhandlungen, 20, 93–98.
Szarowska, M. (2006). Molecular phylogeny, systematics and morphological character evolution in the Balkan Rissooidea (Caenogastropoda). Folia Malacologica, 14, 99–168.
Szarowska, M. & Falniowski, A. (2008). There is no philosopher’s stone: coup de grace for the morphology-based systematics in the rissooidean gastropods? 5th Congress of the European Malacological Societies, Ponta Delgada: 28.
Szarowska, M., & Falniowski, A. (2011). Psudamnicola exilis (Frauenfeld, 1863) in southern Greece: a remnant of a flock of species (Rissooidea: Hydrobiidae)? Folia Malacologica, 19, 117–129.
Szarowska, M., Grzmil, P., & Falniowski, A. (2006). Pseudamnicola Paulucci, 1878 (Gastropoda: Hydrobiidae) in the Balkans. Folia Malacologica, 14, 179–190.
Szarowska, M., Hofman, S., Osikowski, A., & Falniowski, A. (2014a). Heleobia maltzani (Westerlund, 1886) (Caenogastropoda: Truncatelloidea: Cochliopidae) from Crete and species-level diversity of Heleobia Stimpson, 1865 in Europe. Journal of Natural History. doi:10.1080/00222933.2014.946109.
Szarowska, M., Hofman, S., Osikowski, A., & Falniowski, A. (2014b). Divergence preceding island formation among Aegean insular populations of the freshwater snail genus Pseudorientalia (Caenogastropoda: Truncatelloidea). Zoological Science, 31, 680–686.
Tajima, F. (1993). Simple methods for testing molecular clock hypothesis. Genetics, 135, 599–607.
Tamura, K., Stecher, G., Peterson, D., Filipski, A., & Kumar, S. (2013). MEGA6: molecular evolutionary genetics analysis version 6.0. Molecular Biology and Evolution, 30, 2725–2729.
Vuataz, L., Sartori, M., Wagner, A., & Monaghan, M. T. (2011). Toward a DNA taxonomy of alpine Rhithrogena (Ephemeroptera: Heptageniidae) using a mixed Yule-coalescent analysis of mitochondrial and nuclear DNA. PLoS ONE, 6(5), e19728. doi:10.1371/journal.pone.0019728.
Wilke, T. (2003). Salenthydrobia gen. nov. (Rissooidea: Hydrobiidae): a potential relict of the Messinian salinity crisis. Zoological Journal of the Linnean Society, 137, 319–336.
Wilke, T., & Davis, G. M. (2000). Infraspecific mitochondrial sequence diversity in Hydrobia ulvae and Hydrobia ventrosa (Hydrobiidae: Rissoacea: Gastropoda): do their different life histories affect biogeographic patterns and gene flow? Biological Journal of the Linnean Society, 70, 89–105.
Wilke, T., & Falniowski, A. (2001). The genus Adriohydrobia (Hydrobiidae: Gastropoda): polytypic species or polymorphic populations? Journal of Zoological Systematics and Evolutionary Research, 39, 227–234.
Wilke, T., Davis, G. M., Falniowski, A., Giusti, F., Bodon, M., & Szarowska, M. (2001). Molecular systematics of Hydrobiidae (Mollusca: Gastropoda: Rissooidea): testing monophyly and phylogenetic relationships. Proceedings of the Academy of Natural Sciences of Philadelphia, 151, 1–21.
Xia, X. (2013). DAMBE: A comprehensive software package for data analysis in molecular biology and evolution. Molecular Biology and Evolution, 30, 1720–1728.
Xia, X., Xie, Z., Salemi, M., Chen, L., & Wang, Y. (2003). An index of substitution saturation and its application. Molecular Phylogenetics and Evolution, 26, 1–7.
Acknowledgments
This study received funding from the National Science Centre [2011/01/B/NZ8/00041] as a grant awarded to Andrzej Falniowski. We wish to thank Dr Tim Barraclough from the Imperial College London, for providing the R scripts used for performing the GMYC analysis, and Dr Helen McCombie-Boundry from Brest University for improving the language of the paper.
Conflict of interest
The authors declare that they have no competing interests. The authors alone are responsible for the content and writing of the paper.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Supplementary figure 1
(DOCX 3972 kb)
Supplementary figure 2
(DOCX 1941 kb)
Supplementary figure 3
(DOCX 2479 kb)
Supplementary list 1
(DOCX 10 kb)
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Szarowska, M., Osikowski, A., Hofman, S. et al. Pseudamnicola Paulucci, 1878 (Caenogastropoda: Truncatelloidea) from the Aegean Islands: a long or short story?. Org Divers Evol 16, 121–139 (2016). https://doi.org/10.1007/s13127-015-0235-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13127-015-0235-5