Abstract
MicroRNAs (miRNAs) are small single-stranded non-coding RNA molecules that regulate gene expression at the post-transcriptional level. A cross-kingdom regulatory function has been unveiled for plant miRNAs (xenomiRs), which could shape inter-species interactions of plants with other organisms (bacteria and humans) and thus, be key functional molecules of plant-based food in mammals. However, discrepancies regarding the stability and bioavailability of dietary plant miRNAs on the host cast in doubt whether these molecules could have a significant impact on human physiology. The aim of the present study was to identify miRNAs in edible plants and determine their bioavailability on humans after an acute intake of plant-based products. It was found that plant food, including fruits, vegetables and greens, nuts, legumes, and cereals, contains a wide range of miRNAs. XenomiRs miR156e, miR159 and miR162 were detected in great abundance in edible plants and were present among many plant foods, and thus, they were selected as candidates to analyse their bioavailability in humans. These plant miRNAs resisted cooking processes (heat-treatments) and their relative presence increased in faeces after and acute intake of plant-based foods, although they were not detected in serum. Bioinformatic analysis revealed that these miRNAs could potentially target human and bacterial genes involved in processes such as cell signalling and metabolism. In conclusion, edible plants contain miRNAs, such as miR156e, miR159 and miR162, that could resist degradation during cooking and digestion and reach the distal segments of the gastrointestinal tract. Nevertheless, strategies should be developed to improve their absorption to potentially reach host tissues and organs and modulate human physiology.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
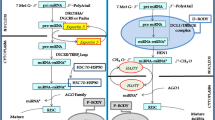
MicroRNAs (miRNAs) are non-coding RNAs of ~ 22 nucleotides that are master regulators of gene expression at the post-transcriptional level [8]. In plants, miRNAs play part in biological functions, such as growth and development, stress responses and metabolism [22, 67]. Remarkably, a key role in cross-kingdom communication have been unveiled for plant miRNAs, which could modulate the interactions of plants with microorganisms and animals [35]. Plant miRNAs can be up-taken by pathogens inhibiting their virulence, and by intestinal bacteria thus having an impact on gut microbiota composition, localization, and metabolite production [58, 70]. Plant miRNAs can also interact with mammalian cells and exert several biological functions, including anti-viral, anti-tumour, immuno-modulatory and anti-inflammatory as well as anti-apoptotic effects [35]. In fact, it has been suggested that the use of plant miRNAs as therapeutic agents could be an effective strategy to treat a wide variety of diseases, such as cancer, COVID-19 and chronic-inflammatory diseases [11, 42, 55, 58, 59]. The effect of plants miRNAs in human gene expression and biological functions has been studied in vitro. For instance, plant miRNAs could exert anti-inflammatory effects through the modulation of CLEC7A, NFAM1 genes or by direct binding to TLR3 receptor in immune system cells [11, 20]. In addition, plant miRNAs promote anti-tumoral responses by enhancing apoptosis and suppressing proliferation of human tumour cells [42, 47]. Plant miRNAs could also regulate the expression of metabolic genes, such as LDLRAP1 (intestinal cells), QKI and MK2 (hepatocytes) and decrease lipid accumulation in hepatocytes [19, 69].
Diet would be the main source of plant miRNAs in mammals. However, it is still unclear whether dietary miRNAs could reach host cells and display a significant biological impact, by which plant miRNA could act as cross-kingdom gene expression regulators. Plant miRNAs have been found in urine, faeces, serum and tissues of mammals such as mice, pigs and humans [12, 37, 39, 40, 62, 65, 66, 69, 72]. In addition, gene expression and physiological changes in animals, such the improvement of metabolic parameters or the decrease of inflammation, have been detected upon the intake of exogenous plant miRNAs (plant xenomiRs) [4, 59, 69]. However, these results are not supported by other authors, who reported negligible levels of xenomiRs on animals, suggesting that the delivery of dietary plant miRNAs on the host would be ineffective [18, 46, 63].
It is well known that plant foods possess beneficial properties that could be used in the management of human diseases, such as obesity, non-alcoholic fatty liver disease, respiratory diseases, cancer, diabetes, anxiety and depression [7, 16, 27, 32, 48, 60]. We hypothesize that the beneficial properties of plants in human physiology could partially rely on cross-kingdom communication through specific plant miRNAs. For this reason, it is important to determine (1) if miRNAs could be relevant effector molecules of the therapeutic effects of plants in humans, and (2) if dietary intake of plants could provide sufficient bioavailability of miRNAs on the host, to be considered as bioactive ingredients in humans. According to this, in the present study, we aimed to identify miRNAs from edible plants and to determine their presence in the human gut and circulatory system upon an acute intake of different plant foods. Moreover, we selected some of the most abundant and conserved plant miRNAs and performed a bioinformatic analysis of their putative biological effects on human cells and bacterial cells.
Materials and Methods
RNA isolation from edible plants, human serum and faecal samples
Total RNA was extracted from cereals (rice), vegetables and greens (green beans, green peppers, lettuces, and spinaches), fruits (apples, olives, oranges, pears, and tomatoes), legumes (chickpeas and lentils), and nuts (walnuts), using miRNeasy Serum/Plasma Kit (Qiagen, Hilden, Germany), according to manufacturer’s instructions. Briefly, raw plants products were used, except for green beans, legumes, and cereals: green beans, lentils and chickpeas were boiled for 4.5, 15 and 30 min, respectively, in a pressure cooker (chickpeas and lentils were soaked previously during 12 h and 2 h, respectively). Rice was boiled for 20 min using a casserole. 0.2 g of cereals, 0.1 g of vegetables and greens, fruits and legumes, and 0.05 g of nuts, were grounded in 1 ml of QIAzol Lysis Reagent with Ultra-Turrax T25 Basic (Basic IKA- Werke, Staufen, Germany) for 1 min (vegetables and greens, fruits, and nuts) or 2 min (cereals and legumes), on ice and at the highest speed. Samples were centrifuged at 12,000 g at 4 ºC for 15 min (olive, vegetables and greens) or 5 min (fruits except olive, cereals, legumes, and nuts). Olive samples formed a fat layer, which was removed, and samples were centrifuged at 12,000 g at 4 ºC for 5 min. The supernatants from plant samples were used for RNA isolation. Prior to the RNA extraction of plant samples used for quantitative real-time PCR (qPCR) assays, 1 µl of spike-in controls (RNA Spike-In Kit, For RT. Qiagen).
Plant miRNA expression analyses in humans were performed as a proof-of-concept study from anonymous healthy volunteers (> 18 yr.). They had a two-day wash out period with poor vegetable intake and then increased their plant intake for three-days. Specifically, volunteers followed an enriched plant-based diet which included several groups of plant foods, such as nuts, legumes, fruits, vegetables and cereals, in the proportion and variety they preferred. Faecal and serum samples were collected before and after the time course; no personal data were obtained from these subjects. Faeces were collected in stool nucleic acid collection and preservation tubes (Norgen Biotek Corp., ON, Canada), and 250 µl of sample was used to isolate total RNA with RNeasy PowerMicrobiome Kit (Qiagen). Venous blood was collected in BD Vacutainer® SST™ II Advance tubes (BD Vacutainer Systems, Plymouth, United Kingdom), sera was separated by centrifugation at 16,000 g at 4 ºC for 10 min, and 200 µl of sample was used to isolate total RNA with miRNeasy Serum/Plasma Advanced Kit (Qiagen). Prior to the extraction, 1 µl of spike-in controls was added to each serum and faecal sample according to manufacturer’s instructions.
Identification of miRNAs in plant samples by next-generation sequencing
Next generation sequencing (NGS) analyses were conducted with total RNA from plants samples, including vegetables (spinach), nuts (walnut) and fruits (apple, olive, pear, orange, and tomato), which were selected after performing quality controls. Library preparation and amplification was carried out using NEBNext® Small RNA Library Prep Set for Illumina® kit and NEBNext® Multiplex Oligos for Illumina (Index Primers Set 1‐4) (New England Biolabs, Ipswich, MA, USA), following manufacturer’s instructions. Removal of big fragment was achieved by bead-based purification as follows: samples were incubated twice with AgenCourt AMPure XP beads (Beckman Coulter, Brea, CA, USA), firstly at a ratio of 1.3x, and secondly at a ratio of 3.7x. Size distribution of the libraries was estimated using Agilent Bioanalyzer (Agilent Technologies, Santa Clara, CA, USA). Quantification of final libraries was carried out by qPCR using the KAPA Library Quantification Kit (KAPA Biosystems Inc., Wilmington, MA, USA), prior to the amplification with Illumina's cBot. Libraries were sequenced 1 × 50 + 8 bp on Illumina's HiSeq2500. Output raw data were processed with skewer [29] to remove the adapter, and reads (from 15 to 30 bp) were aligned with a plant reference genome (Prunus persica, NCBIv2 and annotation NCBIv2.52 restricted to miRNAs) (https://www.ncbi.nlm.nih.gov/assembly/GCF_000346465.2/#/def_asm_Primary_Assembly), with maximum 2 mismatches in the seed area, using miRNA annotation from miRbase (version 22) and ShortStack [5]. Htseq-count [3] was used to count mapped tags, considering the strand information. Raw reads were normalized with DESeq2. Sequencing data were deposited in NCBI's Gene Expression Omnibus [23], and are accessible through GEO Series accession number GSE234786 (https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE234786).
Plant-derived miR156e, miR159 and miR162 expression analysis
The expression of the xenomiRs miR156e, miR159 and miR162 (identified by NGS) was analysed by qPCR in RNA from: cereals (cooked rice), vegetables and greens (green peppers, lettuces, spinaches, raw and cooked green beans), fruits (apples, olives, oranges, pears, tomatoes), cooked legumes (chickpeas and lentils), nuts (walnuts), and human samples (serum and faeces). The expression of the exogenous spike-in UniSp4 was used as a positive control in plant and human samples to determine the efficiency of the experimental procedure. In addition, in human samples, the expression of the endogenous hsa-miR-141-3p and hsa-miR-103a-3p was analysed to check the sample quality.
Reverse-transcription was performed with 4 µl of total RNA with miRCURY LNA RT Kit (Qiagen), in a final volume reaction of 10 µl. Reactions were conducted at 42 °C for 60 min and 95 ºC for 5 min, in a MyCycler Thermal Cycler (Bio-Rad, Hercules, CA, USA). cDNA from plant samples was centrifuged at 4 ºC for 1 min at the highest speed to collect the supernatant for qPCR. cDNA (diluted 1/10) was amplified with miRCURY LNA miRNA PCR Assays (Table 1) and miRCURY LNA SYBR Green PCR Kit (Qiagen), in a CFX384 Touch Real-Time PCR detection System (Bio-Rad). Control samples (non-template) were added to each reaction. The cycling conditions were 95 ºC for 2 min, 40 cycles at 95 ºC for 10 s and 56 ºC for 1 min. miRNA expression levels (Cq values) in serum and faecal samples were normalized with the spike-in UniSp4 (ΔCq) and the following formula was applied to compare the number of copies of plant miRNAs before and after the acute intake of plant foods: 2 (ΔCq before plant acute intake – ΔCq after plant acute intake).
Bioinformatic analyses to predict potential human and bacterial targets of plant miRNAs
The small RNA target analysis servers TAPIR (https://www.zhaolab.org/psRNATarget/) [9] and psRNATarget (scoring schemas V1 and V2) (https://bioinformatics.psb.ugent.be/webtools/tapir/) [15] were used to identify putative human target genes of miR156e (5'-UGACAGAAGAGAGUGAGCAC-3'), miR159 (5'-UUUGGAUUGAAGGGAGCUCUA-3') and miR162 (5'-UCGAUAAACCUCUGCAUCCAG-3'). Plant miRNA sequences were aligned to the cDNA library “Homo sapiens (human), transcript, Human genomic sequencing project” (available in the psRNATarget server), applying the default parameters. The online tools Genecodis (https://genecodis.genyo.es/) [10] and PANTHER (https://pantherdb.org/) [45] were used to conduct Gene Ontology (GO) enrichment analysis and biological process classification. In addition, pathway analyses were performed with Genecodis, by which annotations from different sources were used (KEGG Pathways, Panther Pathways and WikiPathways). These analyses were carried out independently for each miRNA, with the putative target genes identified with TAPIR and psRNATarget. A cut-off threshold for the expectation value, which represents the penalty for the mismatches between miRNA and target sequence, was applied to filtered psRNATarget scoring schema V2 putative targets: the cut-off value was set between 3.5–4.5 in order to select up to 20 top target genes. For PANTHER analyses (access on 26 May 2023) “Homo sapiens” was selected as reference organism.
Prediction analyses of bacterial targets of plants miRNAs were conducted with TargetRNA3 (https://cs.wellesley.edu/~btjaden/TargetRNA3/), applying the default parameters [61]. Ten genomes from eight different bacteria species were selected Escherichia coli (Escherichia coli ATCC 25922 (GCF_017357505.1); Escherichia coli str. K-12 substr. MG1655 (GCF_000005845.2); Escherichia coli O157:H7 str. Sakai Sakai substr. RIMD 0509952 (GCF_000008865.2)), Enterococcus faecalis (Enterococcus faecalis OG1RF (GCF_000172575.2)), Bifidobacterium longum (Bifidobacterium longum subsp. longum JCM 1217 (GCF_000196555.1)), Lactobacillus acidophilus (Lactobacillus acidophilus La-14 (GCF_000389675.2)), Levilactobacillus brevis (Levilactobacillus brevis NPS-QW-145 (GCF_001676805.1)), Limosilactobacillus fermentum (Limosilactobacillus fermentum SCB0035 (GCF_022819245.1)), Ligilactobacillus salivarius (Ligilactobacillus salivarius LPM01 (GCF_900094615.1)), Lactobacillus jensenii (Lactobacillus jensenii SNUV360 (GCF_001936235.1)).
Statistical analysis
Wilcoxon matched-pairs signed rank test was applied to determine differences in human samples before and after the acute intake of plant food products. Differences were considered statistically significant at p-value p < 0.05.
Results
miRNAs diversity and abundance in edible plants
The miRNA profile was analysed by NGS in the following edible plants: fruits (apple, olive, orange pear and tomato), vegetables (spinach), and nuts (walnut): They were selected because of their very good results in the quality controls (bioanalyzer determinations). The results revealed that 176 miRNAs were present in at least one of the plants used for this study. miR156e (gene:ENSRNA049996234), miR159 (gene:ENSRNA049996936), and miR162 (gene:ENSRNA049996910) were selected as candidate miRNAs for subsequent analyses due to their broad presence among the selected plant species and high abundance (number of reads) (Table S1).
The expression of miR156e, miR159 and miR162 in edible plants was validated by qPCR in an extended selection of raw and cooked food matrices: nuts (walnuts), fruits (apple, orange, olive, tomato, and pear), vegetables and greens (raw and cooked green beans, lettuce, spinach and green pepper), cooked legumes (lentils and chickpeas) and cooked cereals (rice). The three plant miRNAs were present in all the plant foods analysed (Fig. 1). Notably, the three miRNAs were also present in boiled green beans, lentils, chickpeas, and rice, which is the process that they usually undergo before being consumed. Since the objective was to identify miRNAs that were present in the plants, rather than quantifying miRNAs and comparing their abundance between different samples, data were not normalized. Nonetheless, spike-in UniSp4 was added prior to the RNA isolation and whose detection was used as a positive (reference) control to determine the reliability of the results.
Detection of miR156e, miR159 and miR162 in raw and cooked edible plant products by quantitative PCR. The results are presented as the Cq values of (a) miR156e (5'-UGACAGAAGAGAGUGAGCAC-3'), (b) miR159 (5'-UUUGGAUUGAAGGGAGCUCUA-3'), and (c) miR162 (5'-UCGAUAAACCUCUGCAUCCAG-3') in fruits (apple, orange, olive, tomato, and pear), vegetables and greens (raw and cooked green beans, lettuce, spinach and green pepper), nuts (walnuts), cooked legumes (lentil and chickpea) and cooked cereals (rice). Results are expressed as the mean ± standard error of the mean (SEM) (n = 2)
Plant xenomiRs miR156e, miR159 and miR162 expression is increased in human faecal samples after an acute intake of plant-origin foods
To determine the presence of plant miRNAs in human samples, seven healthy volunteers were subjected to a three days-acute intake of a wide variety of plant products (see MM). The expression profile of miR156e, miR159 and miR162 was analysed in faecal and serum samples before and after the intervention. The expression levels of two endogenous human miRNAs, hsa-miR-141-3p and hsa-miR-103a-3p, were also evaluated in faecal and serum samples, respectively to determine sample quality and data reliability. Providing that no universal endogenous miRNA normalizer (housekeeping) has been standardized yet, exogenous spike-in UniSp4 expression was used to eliminate variability of RNA extraction, reverse-transcription, and qPCR processes.
Plant miR156e, miR159 and miR162 were present in all the faecal samples and their expression levels increased 5.34 ± 1.72 (p < 0.05), 1.68 ± 0.51 (ns), and 2.21 ± 0.28 (p < 0.05) times, respectively, after the high intake of plant foods as compared to their respective levels at baseline (Fig. 2). Undetectable levels of plant miRNAs were reported in serum samples of the same set of individuals, suggesting that the selected plant miRNAs were not absorbed. However, the endogenous control hsa-miR-103a-3p was amplified (Cq values between 24–28). This demonstrates that human-origin miRNAs are present in serum samples while (exogenous) plant xenomiRs are not present or are available in such low levels that are far beyond qPCR detection limit.
Relative quantification of plant miRNAs miR156e, miR159 and miR162 in human faeces after a three-day time course acute intake of plant products. Plant miRNAs miR156e, miR159 and miR162 expression was analysed by qPCR in human faecal samples from 7 volunteers before and after a dietary intervention consisting of a high consumption of plant products for three days. Cq values were normalized (ΔCq) with the spike-in UniSp4 and expression level differences before and after the interventions were expressed as 2 (ΔCq before plant acute intake – ΔCq after plant acute intake). Results are presented as the mean ± standard error of the mean (SEM) (n = 7 volunteers). p-value: * p < 0.05 as compared vs. before the acute intake
Plant miRNAs miR156e, miR159 and miR162 could potentially target human and bacterial genes and modulate host biological functions and pathways
To evaluate if plant miR156e, miR159 and miR162 could exert biological effects on human cells, bioinformatic analyses were carried out to potentially verify plant miRNA impact on shaping host gene expression. miR156e, miR159 and miR162 mature sequences were aligned with the human transcriptome to predict putative human target genes using psRNATarget (scoring schemas V1 and V2) and TAPIR algorithms. miR156e could potentially modulate the expression of 166 different transcripts (152 different human genes) (Table S2). Of note, the F11R gene (transcript NM_016946) appeared in the three prediction algorithms (Table S2). In addition, the three algorithms predicted that a total of 132 different transcripts (120 different genes) could be potentially regulated by miR159, of which the transcript NM_005444 (RQCD1 gene) was a common output (Table S3). psRNATarget scoring schema V2 was the only algorithm that reported putative targets (40 different transcripts, 37 different genes) for miR162 (Table S4).
To identify the human biological functions and pathways that could be modulated by the predicted targets of miR156e, miR159 and miR162, we conducted Gene Ontology (GO) and KEGG, Panther and WikiPathways analyses:
-
For miR156e putative targes, 19 genes were analysed (HIF3A, F11R, SUCLG2, ALG2, SV2B, C12orf74, KCTD18, HORMAD2, EHMT1, FHL1, ANKRD13A, LPGAT1, VSX1, ARF3, ARL4C, POLR3H, SEC23IP, CCDC88C, ZFP62). In PANTHER software, one gene was not detected (C12orf74) and ten genes were unclassified (ALG2, HORMAD2, FHL1, EHMT1, ZFP62, SV2B, LPGAT1, ANKRD13A, SEC23IP, KCTD18). In KEGG Pathways, Panther Pathways and WikiPathways, the genes that were reported as unannotated inputs were: ANKRD13A, KCTD18, SEC23IP, VSX1, ZFP62, C12orf74 (KEGG, Panther and Wiki Pathways), ARL4C, CCDC88C, HIF3A, HORMAD2 (KEGG and Panther Pathways), SV2B (Panther and Wiki Pathways), F11R, SUCLG2, ALG2, EHMT1, FHL1, LPGAT1, POLR3H (Panther Pathways), ARF3 (Wiki Pathways).
-
For miR159, 14 genes were analysed (AMOT, RQCD1, TRIM14, RBAK, PPM1E, RGAG4, AK1, LPP, IRS1, ENKUR, PIM3, RIC3, SEH1L, STON1). In PANTHER, one gene was not detected (RGAG4), two were unclassified (ENKUR, TRIM14). In GO analysis with Genecodis, RGAG4 was also an unannotated input. In KEGG Pathways, Panther Pathways and WikiPathways, the genes that were reported as unannotated inputs were: ENKUR, PIM3, PPM1E, RIC3, RGAG4, STON1, TRIM14 (KEGG, Panther and Wiki Pathways), LPP (KEGG and Panther Pathways), RBAK, SHE1L (Panther and Wiki Pathways), AMOT, RQCD1 (Panther Pathways), AK1 (Wiki Pathways).
-
For miR162, seven genes were introduced (MAPK14, CISD3, RAB3D, PLG, PRR5L, DNM2, RBAK). In PANTHER analyses, two were unclassified (DNM2, CISD3). In KEGG Pathways, Panther Pathways and WikiPathways, the genes that were reported unannotated as inputs were: CISD3 (KEGG, Panther and Wiki Pathways), PRR5L (KEGG and Panther Pathways), RAB3D, RBAK (Panther and Wiki Pathways), DNM2 (Panther Pathways).
In terms of biological process, 7, 10 and 6 groups were identified by PANTHER for miR156e, miR159 and miR162 predicted target genes, respectively, of which “cellular process” was the top group (Fig. 3). Predicted targets were also clustered in other groups, including “cell signalling” (miR159 and miR162 putative targets), “metabolic process” and “response to stimulus” (all three). GO results performed with Genecodis showed that miR156e putative targets are enriched in biological processes that include stress-activated protein kinase signalling cascade (CCDC88C), memory T cell extravasation, and regulation of membrane permeability and bicellular tight junction assembly (F11R), (Table 2), and the Huntington disease pathway (ARL4C and ARF3) (Table 5). Predicted target genes of plant miR159 were enriched in processes such as negative regulation of insulin secretion involved in cellular response to glucose stimulus (PIM3), positive regulation of fatty acid beta-oxidation and glucose metabolism process (IRS1) (Table 3), and pathways such as insulin signalling and type II diabetes mellitus (IRS1) (Table 5). The biological functions of miR162 target genes are gathered in Table 4, and include micropinocytosis, positive regulation of endocytosis (DNM2), stress-induced premature senescence, regulation of cytokine production involved in inflammatory response, fatty acid oxidation and response to dietary excess (MAPK14), while no biological pathways were identified with a significant adjusted p value (Table 5).
Gene ontology analysis of predicted target genes of plant miRNAs, miR156e, miR159 and miR162, performed by PANTHER. Putative target identified with TAPIR and psRNATarget scoring schemas V1 and V2 were classified based on biological process annotation. Results are presented in pie charts, filtering genes with no PANTHER category assigned
To determine if plant miR156e, miR159 and miR162 could potentially have an impact on (gut-present) bacteria, eight bacteria species that have been found in the human gut were selected (Escherichia coli, Enterococcus faecalis and several species of Lactobacillus) [17, 26, 41, 44, 52, 54, 57, 73]. Bioinformatic analyses were conducted with TargetRNA3 algorithm to predict targets in prokaryotes (Table 6). miR156e could potentially modulate the expression of two different targets in Escherichia Coli: fadK (a short chain acyl-CoA synthetase) and the hypothetical protein ECs_5262. Moreover, MukB (a chromosome partitioning protein) and the hypothetical protein ECs_5262 from Escherichia Coli, and pknB (a Stk1 family PASTA domain-containing Ser/Thr kinase) from Ligilactobacillus salivarius, were predicted as targets of miR159. Finally, several putative targets of miR162 were identified: cysI (a sulfite reductase subunit) of Escherichia coli, OG1RF_RS11400 (ribonuclease J) of Enterococcus faecalis, BLLJ_RS06465 (ABC transporter ATP-binding protein) of Bifidobacterium longum, and ppx (exopolyphosphatase) of Levilactobacillus brevis.
Discussion
This study contributed to the identification of the miRNA expression profile of edible plants. Small-RNA sequencing results revealed that plant-based diets could be a major source of plant xenomiRs. miR156e, miR159 and miR162 were selected as candidates for downstream analysis due to their abundance and high conservation degree across the selection of analysed plant foods. However, this selection does not exclude the possibility that other identified miRNAs could be interesting research elements for new investigations. Notably, these miRNAs were present in plant food products (legumes, cereals, and green beans) after boiling, suggesting that extreme processes do not jeopardise the stability (and presumably functionality) of plant miRNAs. These results are in agreement with other studies reporting that miR166, miR167 and miR168 from soybean and rice resisted storage, processing and cooking conditions [51], and miR156a, miR166a and miR168a were also stable in cooked rice [69]. Indeed, results from the present study might suggest that miR156e, miR159 and miR162 could be good candidates to generally estimate the bioavailability of plant miRNAs in humans, since they were present in great amount in all groups of plant foods tested (legumes, nuts, fruits, cereals, vegetables and greens) and they resisted physical treatments (soaking and heat-treatments). Of note, no universal plant miRNA housekeeping has been identified to potentially quantify the relative expression of target xenomiRs. We evaluated miRNA levels as a proof-of-concept (validation) of their presence in different food matrices (qualitative value). Therefore, we consider that normalizing the relative miRNA expression is not necessary for the specific aim in the framework of the present study (we did not want to quantify, just qualitatively demonstrate their abundance: present/not present).
Plant miR156e, miR159 and miR162 were detected in faeces of healthy volunteers, and the relative abundance of plant miR156e and miR162 increased after a three-day acute intake of plant products. These results revealed that plant miRNAs could reach the gastrointestinal tract resisting the digestive processes. In addition, it was unveiled that dietary modifications such as increasing plant intake enhance exogenous miRNA bioavailability. Importantly, efforts should not be focused exclusively on the evaluation of plant miRNA effects in distal tissues and organs and on developing strategies to eventually promote their absorption, since their biological impact could be exerted and limited to at local-gut level. Several published studies support this hypothesis. For instance, Teng et al. reported that ginger EVs can be taken by gut microorganisms and their miRNA cargo modulates bacteria gene expression, shaping immune system responses and improving gut barrier function [58]. In this study, we demonstrate that plant miR156e, miR159 and miR162 reach the human gastrointestinal tract in agreement with previous studies, which showed that miR156 could reach mice gut and regulate enterocyte growth [36]. In concordance with the studies that unveiled a crosstalk between plant miRNAs mdo-miR7267-3p and bol-miR159 and bacteria which shaped metabolite production, composition, localization, and growth of gut microbes [58, 64], we conducted bioinformatic analyses to eventually identify non-human (prokaryote) targets of plant miRNAs. Our results revealed a potential interaction of plant miR156e, miR159 and miR162 and bacterial targets. Thus, miR156e, miR159, and miR162 could potentially modulate the expression of targets involved in bacterial metabolism (i.e., fadK and cysI), bacteria growth (i.e., mukB), and stress responses (i.e., ppx), and thus, could contribute to gut eubiosis and subsequent impact in host physiology [6, 24, 33, 49]. In addition, we performed bioinformatic predictions, GO and pathways analysis to predict putative human target genes of plant miR156e, miR159 and miR162 and biological functions. The data presented here suggest that miR156 could potentially modulate the expression of human genes involved in intestinal homeostasis, such as the establishment of the endothelial intestinal barrier, in agreement with the results of Li et al. [36]. Importantly, Li et al. also showed that the administration of synthetic miR156 could modulate mammalian enterocyte growth by targeting WNT10B in vitro and in mice [36]. Our in silico analyses show that this effect in intestinal homeostasis could potentially be achieved through the modulation of F11R gene, which appeared in all the prediction programs used (psRNATarget scoring schema V1 and V2 and TAPIR). Knockdown of F11R gene (which encodes for JAM-A protein) has been previously associated with the leaky gut and gut barrier disruption [14, 53]. By contrast, positive effects have been achieved though F11R downregulation, which comprise counteraction of cancer progression [34, 68]. However, the potential biological impact of bioinformatics predictions in this article is still to be fully determined.
Nonetheless, plant miRNAs were undetectable in serum samples of the same volunteers. Our data suggest that plant miRNA absorption and distribution to peripheral organs and tissues cannot be achieved by implementing dietary modifications (increase of plant food intake) or under our experimental conditions. These results do not support the observation made by other authors, which found that plant miRNAs could be detected in the circulatory system of a diverse type of animals, such as mice, pigs and humans, and be distributed to tissues and organs [37, 38, 40, 65, 66, 69]. Interestingly, Chen et al. analysed blood samples from NGS datasets and detected differences in the expression profile of plant miRNAs in animals with different dietary regimes, determining that plant miRNA abundance in blood was higher in herbivores, followed by omnivorous and they were barely detected in carnivores [12]. However, absorption of plant miRNAs in animals has been questioned by several reports, which claimed that detection of plant miRNAs might be artifacts of human sequence contaminations during NGS [25, 30, 62, 71]. In this context, the results presented in this article are consistent with the observations made by other studies where absorption of plant miRNAs could be undetected or even inexistent. Increasing plant intake, such pollen, corn, rice or extra virgin olive oil or fruits, in a wide range of animals, including honey bees, mice and humans, reported no measurable levels of plants miRNAs in blood or tissues [18, 28, 43, 46, 56]. The discrepancies concerning the bioavailability of plant miRNAs in animals may be explained by different factors: (1) plant miRNA administration methods (i.e., oral intake, gavage, administration of isolated RNA or plant-based products), dosage/amount administered and exposure time; (2) sensibility of the miRNA detection techniques; (3) gut permeability; and (4) miRNA stability (each plant miRNA could have different availability depending on its physical–chemical stability, and eventually protective factors like encapsulation in extracellular vesicles (EVs) [21, 65, 66]. As we report here, plant miRNAs are not detected in blood despite increasing plant intake.
Notably, our bioinformatic analyses suggest that plant miRNAs could not only have a potential impact at the local-gut level, but also at peripheral tissues and organs. miR159 could potentially modulate the expression of genes related with metabolic pathways (i.e., glucose metabolism and insulin signalling pathways) and immune system. Moreover, RQCD1 could be a putative target gene of miR159; it appeared in all the prediction algorithms and its upregulation has been linked to breast cancer progression [1]. Notably, functional validation of miR159 interactions with mammalian genes, that were not identified in this work using TAPIR and psRNATarget has been already unveiled [4, 13]. Aquilano et al. [4] reported that mimics for plant miR159 could target Tnfrsf1a gene in obese mice, supressing inflammation and improving the metabolic profile. Chin et al. demonstrated that miR159 targets TCF7 gene and its administration counteracted xenograft breast tumour growth in mice [13]. miR156e could also have an impact beyond the gut level. For instance, GO and pathways analyses unveiled and association between its predicted human targets and neuron differentiation and Huntington disease. In this context, in silico analyses have already revealed the therapeutic potential of plants miRNAs to treat neurological disorders, such as Alzheimer [50]. However, the results of the present work suggest that plant miRNAs might not surpass the gut barrier, since they were not detected in serum samples, at least in the conditions of our study. Despite these results, we hypothesize that other (dietary) strategies could be addressed to potentially promote the absorption of miRNAs and enhance their potential therapeutic effects in vivo. In this context, miRNA stability and bioavailability could be improved by encapsulation in EVs. Notably, plant EVs have been postulated as promising therapeutic delivery systems, which confer protection to their cargo, can be internalized by mammalian cells, are stable in several conditions, such as storage and digestion, and their internal content is highly bioavailable [2, 31]. Therefore, the administration of nanocarriers designed to contain cocktails of plant miRNAs with therapeutic benefits could be worthwhile to promote their absorption since the evidence presented here unveil that they were not present in the circulatory system, but they could potentially exert a biological impact beyond the gut.
In conclusion, the present article has contributed to the identification of miRNA expression profile in plant foods, revealing that edible plant-based products contain large number of miRNAs, which can resist cooking processes. In addition, it has been unveiled that a short plant-based dietary intervention can increase the abundance of exogenous miRNAs in the gut. Therefore, significant effects of plant miRNAs on human physiology could potentially be displayed thought the interaction with gut cells (i.e. enterocytes) and bacteria. Nevertheless, the negligible levels of plant miRNAs detected in serum highlight the need to develop novel strategies to improve the bioavailability of plant miRNAs beyond the gut. For instance, the administration of nanocarriers designed to contain cocktails of plant miRNAs with therapeutic benefits could be worthwhile. In any case, our data suggest that miR156e, miR159 and miR162 have the potential to affect the expression of human and bacterial genes and modulate important biological pathways.
Data availability
The data supporting the conclusions of this work are available by the authors on request. Next generation sequencing data are publicly available at https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE234786.
Code availability
Not applicable.
References
Ajiro M, Katagiri T, Ueda K, Nakagawa H, Fukukawa C, Lin M-L, Park J-H, Nishidate T, Daigo Y, Nakamura Y (2009) Involvement of RQCD1 overexpression, a novel cancer-testis antigen, in the Akt pathway in breast cancer cells. Int J Oncol 35:673–681
Alzahrani FA, Khan MI, Kameli N, Alsahafi E, Riza YM (2023) Plant-Derived Extracellular Vesicles and Their Exciting Potential as the Future of Next-Generation Drug Delivery. Biomolecules 13:839. https://doi.org/10.3390/biom13050839
Anders S, Pyl PT, Huber W (2015) HTSeq–a Python framework to work with high-throughput sequencing data. Bioinformatics 31:166–169. https://doi.org/10.1093/bioinformatics/btu638
Aquilano K, Ceci V, Gismondi A, De Stefano S, Iacovelli F, Faraonio R, Di Marco G, Poerio N, Minutolo A, Minopoli G, Marcone A, Fraziano M, Tortolici F, Sennato S, Casciardi S, Potestà M, Bernardini R, Mattei M, Falconi M, Montesano C, Rufini S, Canini A, Lettieri-Barbato D (2019) Adipocyte metabolism is improved by TNF receptor-targeting small RNAs identified from dried nuts. Commun Biol 2:317. https://doi.org/10.1038/s42003-019-0563-7
Axtell MJ (2013) ShortStack: comprehensive annotation and quantification of small RNA genes. RNA 19:740–751. https://doi.org/10.1261/rna.035279.112
Baena C, Blasco A, Zúñiga Cabrera M, Monedero V (2013) Accumulation of Polyphosphate in Lactobacillus spp. and Its Involvement in Stress Resistance. Appl Environ Microbiol 80. https://doi.org/10.1128/AEM.03997-13
Bagherniya M, Nobili V, Blesso CN, Sahebkar A (2018) Medicinal plants and bioactive natural compounds in the treatment of non-alcoholic fatty liver disease: A clinical review. Pharmacol Res 130:213–240. https://doi.org/10.1016/j.phrs.2017.12.020
Bartel DP (2018) Metazoan MicroRNAs. Cell 173:20–51. https://doi.org/10.1016/j.cell.2018.03.006
Bonnet E, He Y, Billiau K, Van de Peer Y (2010) TAPIR, a web server for the prediction of plant microRNA targets, including target mimics. Bioinformatics 26:1566–1568. https://doi.org/10.1093/bioinformatics/btq233
Carmona-Saez P, Chagoyen M, Tirado F, Carazo JM, Pascual-Montano A (2007) GENECODIS: a web-based tool for finding significant concurrent annotations in gene lists. Genome Biol 8:R3. https://doi.org/10.1186/gb-2007-8-1-r3
Cavalieri D, Rizzetto L, Tocci N, Rivero D, Asquini E, Si-Ammour A, Bonechi E, Ballerini C, Viola R (2016) Plant microRNAs as novel immunomodulatory agents. Sci Rep 6:25761. https://doi.org/10.1038/srep25761
Chen X, Liu L, Chu Q, Sun S, Wu Y, Tong Z, Fang W, Timko MP, Fan L (2021) Large-scale identification of extracellular plant miRNAs in mammals implicates their dietary intake. PLoS ONE 16:e0257878. https://doi.org/10.1371/journal.pone.0257878
Chin AR, Fong MY, Somlo G, Wu J, Swiderski P, Wu X, Wang SE (2016) Cross-kingdom inhibition of breast cancer growth by plant miR159. Cell Res 26:217–228. https://doi.org/10.1038/cr.2016.13
Chopyk DM, Kumar P, Raeman R, Liu Y, Smith T, Anania FA (2017) Dysregulation of junctional adhesion molecule-A contributes to ethanol-induced barrier disruption in intestinal epithelial cell monolayers. Physiol Rep 5:e13541. https://doi.org/10.14814/phy2.13541
Dai X, Zhao PX (2011) psRNATarget: a plant small RNA target analysis server. Nucleic Acids Res 39:W155–W159. https://doi.org/10.1093/nar/gkr319
Datta S, Luthra R, Bharadvaja N (2022) Medicinal Plants for Glioblastoma Treatment. Anticancer Agents Med Chem 22:2367–2384. https://doi.org/10.2174/1871520622666211221144739
Díaz R, Torres-Miranda A, Orellana G, Garrido D (2021) Comparative Genomic Analysis of Novel Bifidobacterium longum subsp. longum Strains Reveals Functional Divergence in the Human Gut Microbiota. Microorganisms 9:1906. https://doi.org/10.3390/microorganisms9091906
Dickinson B, Zhang Y, Petrick JS, Heck G, Ivashuta S, Marshall WS (2013) Lack of detectable oral bioavailability of plant microRNAs after feeding in mice. Nat Biotechnol 31:965–967. https://doi.org/10.1038/nbt.2737
Díez-Sainz E, Aranaz P, Amri E-Z, Riezu-Boj JI, Lorente-Cebrián S, Milagro FI (2024) Plant miR8126-3p and miR8126-5p Decrease Lipid Accumulation through Modulation of Metabolic Genes in a Human Hepatocyte Model That Mimics Steatosis. Int J Mol Sci 25:1721. https://doi.org/10.3390/ijms2503172
Díez-Sainz E, Lorente-Cebrián S, Aranaz P, Amri E-Z, Riezu-Boj JI, Milagro FI (2023) miR482f and miR482c-5p from edible plant-derived foods inhibit the expression of pro-inflammatory genes in human THP-1 macrophages. Front Nutr 10:1287312. https://doi.org/10.3389/fnut.2023.1287312
Díez-Sainz E, Lorente-Cebrián S, Aranaz P, Riezu-Boj JI, Martínez JA, Milagro FI (2021) Potential Mechanisms Linking Food-Derived MicroRNAs, Gut Microbiota and Intestinal Barrier Functions in the Context of Nutrition and Human Health. Front Nutr 8:586564. https://doi.org/10.3389/fnut.2021.586564
Dong Q, Hu B, Zhang C (2022) microRNAs and Their Roles in Plant Development. Front Plant Sci 13:824240. https://doi.org/10.3389/fpls.2022.824240
Edgar R, Domrachev M, Lash AE (2002) Gene Expression Omnibus: NCBI gene expression and hybridization array data repository. Nucleic Acids Res 30:207–210. https://doi.org/10.1093/nar/30.1.207
Edwards AL, Sangurdekar DP, Jeong KS, Khodursky AB, Rybenkov VV (2013) Transient growth arrest in Escherichia coli induced by chromosome condensation. PLoS ONE 8:e84027. https://doi.org/10.1371/journal.pone.0084027
Fromm B, Kang W, Rovira C, Cayota A, Witwer K, Friedländer MR, Tosar JP (2019) Plant microRNAs in human sera are likely contaminants. J Nutr Biochem 65:139–140. https://doi.org/10.1016/j.jnutbio.2018.07.019
Gao H, Li X, Chen X, Hai D, Wei C, Zhang L, Li P (2022) The Functional Roles of Lactobacillus acidophilus in Different Physiological and Pathological Processes. J Microbiol Biotechnol 32:1226–1233. https://doi.org/10.4014/jmb.2205.05041
Garima S, Ajit Kumar P, Marcy DM, Sakthivel R, Bhim Pratap S, Nachimuthu Senthil K (2020) Ethnobotanical survey of medicinal plants used in the management of cancer and diabetes. J Tradit Chinese Med = Chung i tsa chih ying wen pan 40:1007–1017. https://doi.org/10.19852/j.cnki.jtcm.2020.06.012
Huang H, Davis CD, Wang TTY (2018) Extensive Degradation and Low Bioavailability of Orally Consumed Corn miRNAs in Mice. Nutrients 10:215. https://doi.org/10.3390/nu10020215
Jiang H, Lei R, Ding S-W, Zhu S (2014) Skewer: a fast and accurate adapter trimmer for next-generation sequencing paired-end reads. BMC Bioinform 15:182. https://doi.org/10.1186/1471-2105-15-182
Kang W, Bang-Berthelsen CH, Holm A, Houben AJS, Müller AH, Thymann T, Pociot F, Estivill X, Friedländer MR (2017) Survey of 800+ data sets from human tissue and body fluid reveals xenomiRs are likely artifacts. RNA 23:433–445. https://doi.org/10.1261/rna.059725.116
Karamanidou T, Tsouknidas A (2021) Plant-Derived Extracellular Vesicles as Therapeutic Nanocarriers. Int J Mol Sci 23:191. https://doi.org/10.3390/ijms23010191
Karri S, Sharma S, Hatware K, Patil K (2019) Natural anti-obesity agents and their therapeutic role in management of obesity: A future trend perspective. Biomed Pharmacother 110:224–238. https://doi.org/10.1016/j.biopha.2018.11.076
Kobayashi K (2019) Inactivation of cysL Inhibits Biofilm Formation by Activating the Disulfide Stress Regulator Spx in Bacillus subtilis. J Bacteriol 201:e00712-e718. https://doi.org/10.1128/JB.00712-18
Lauko A, Mu Z, Gutmann DH, Naik UP, Lathia JD (2020) Junctional Adhesion Molecules in Cancer: A Paradigm for the Diverse Functions of Cell-Cell Interactions in Tumor Progression. Cancer Res 80:4878–4885. https://doi.org/10.1158/0008-5472.CAN-20-1829
Li D, Yang J, Yang Y, Liu J, Li H, Li R, Cao C, Shi L, Wu W, He K (2021) A Timely Review of Cross-Kingdom Regulation of Plant-Derived MicroRNAs. Front Genet 12:613197. https://doi.org/10.3389/fgene.2021.613197
Li M, Chen T, Wang R, Luo J-Y, He J-J, Ye R-S, Xie M-Y, Xi Q-Y, Jiang Q-Y, Sun J-J, Zhang Y-L (2019) Plant MIR156 regulates intestinal growth in mammals by targeting the Wnt/β-catenin pathway. Am J Physiol Cell Physiol 317:C434–C448. https://doi.org/10.1152/ajpcell.00030.2019
Liang G, Zhu Y, Sun B, Shao Y, Jing A, Wang J, Xiao Z (2014) Assessing the survival of exogenous plant microRNA in mice. Food Sci Nutr 2:380–388. https://doi.org/10.1002/fsn3.113
Liang H, Zhang S, Fu Z, Wang Y, Wang N, Liu Y, Zhao C, Wu J, Hu Y, Zhang J, Chen X, Zen K, Zhang C-Y (2015) Effective detection and quantification of dietetically absorbed plant microRNAs in human plasma. J Nutr Biochem 26:505–512. https://doi.org/10.1016/j.jnutbio.2014.12.002
Liu Y-C, Chen WL, Kung W-H, Huang H-D (2017) Plant miRNAs found in human circulating system provide evidences of cross kingdom RNAi. BMC Genom 18:112. https://doi.org/10.1186/s12864-017-3502-3
Luo Y, Wang P, Wang X, Wang Y, Mu Z, Li Q, Fu Y, Xiao J, Li G, Ma Y, Gu Y, Jin L, Ma J, Tang Q, Jiang A, Li X, Li M (2017) Detection of dietetically absorbed maize-derived microRNAs in pigs. Sci Rep 7:645. https://doi.org/10.1038/s41598-017-00488-y
Martinson JN V, Walk ST (2020) Escherichia coli Residency in the Gut of Healthy Human Adults. EcoSal Plus 9. https://doi.org/10.1128/ecosalplus.ESP-0003-2020
Marzano F, Caratozzolo MF, Consiglio A, Licciulli F, Liuni S, Sbisà E, D’Elia D, Tullo A, Catalano D (2020) Plant miRNAs Reduce Cancer Cell Proliferation by Targeting MALAT1 and NEAT1: A Beneficial Cross-Kingdom Interaction. Front Genet 11:552490. https://doi.org/10.3389/fgene.2020.552490
Masood M, Everett CP, Chan SY, Snow JW (2016) Negligible uptake and transfer of diet-derived pollen microRNAs in adult honey bees. RNA Biol 13:109–118. https://doi.org/10.1080/15476286.2015.1128063
Messaoudi S, Manai M, Kergourlay G, Prévost H, Connil N, Chobert J-M, Dousset X (2013) Lactobacillus salivarius: Bacteriocin and probiotic activity. Food Microbiol 36:296–304. https://doi.org/10.1016/j.fm.2013.05.010
Mi H, Muruganujan A, Casagrande JT, Thomas PD (2013) Large-scale gene function analysis with the PANTHER classification system. Nat Protoc 8:1551–1566. https://doi.org/10.1038/nprot.2013.092
Micó V, Martín R, Lasunción MA, Ordovás JM, Daimiel L (2016) Unsuccessful Detection of Plant MicroRNAs in Beer, Extra Virgin Olive Oil and Human Plasma After an Acute Ingestion of Extra Virgin Olive Oil. Plant Foods Hum Nutr 71:102–108. https://doi.org/10.1007/s11130-016-0534-9
Minutolo A, Potestà M, Gismondi A, Pirrò S, Cirilli M, Gattabria F, Galgani A, Sessa L, Mattei M, Canini A, Muleo R, Colizzi V, Montesano C (2018) Olea europaea small RNA with functional homology to human miR34a in cross-kingdom interaction of anti-tumoral response. Sci Rep 8:12413. https://doi.org/10.1038/s41598-018-30718-w
Moragrega I, Ríos JL (2021) Medicinal Plants in the Treatment of Depression: Evidence from Preclinical Studies. Planta Med 87:656–685. https://doi.org/10.1055/a-1338-1011
Morgan-Kiss RM, Cronan JE (2004) The Escherichia coli fadK (ydiD) gene encodes an anerobically regulated short chain acyl-CoA synthetase. J Biol Chem 279:37324–37333. https://doi.org/10.1074/jbc.M405233200
Patel M, Mangukia N, Jha N, Gadhavi H, Shah K, Patel S, Mankad A, Pandya H, Rawal R (2019) Computational identification of miRNA and their cross kingdom targets from expressed sequence tags of Ocimum basilicum. Mol Biol Rep 46:2979–2995. https://doi.org/10.1007/s11033-019-04759-x
Philip A, Ferro VA, Tate RJ (2015) Determination of the potential bioavailability of plant microRNAs using a simulated human digestion process. Mol Nutr Food Res 59:1962–1972. https://doi.org/10.1002/mnfr.201500137
Pramanick R, Gazara R, Ahmad R (2022) Gut Microbiome Prediction: From Current Human Evidence to Future Possibilities. Biorxiv. https://doi.org/10.1101/2022.11.16.516694
Rahman K, Desai C, Iyer SS, Thorn NE, Kumar P, Liu Y, Smith T, Neish AS, Li H, Tan S, Wu P, Liu X, Yu Y, Farris AB, Nusrat A, Parkos CA, Anania FA (2016) Loss of Junctional Adhesion Molecule A Promotes Severe Steatohepatitis in Mice on a Diet High in Saturated Fat, Fructose, and Cholesterol. Gastroenterology 151:733-746.e12. https://doi.org/10.1053/j.gastro.2016.06.022
Repoila F, Le Bohec F, Guérin C, Lacoux C, Tiwari S, Jaiswal AK, Santana MP, Kennedy SP, Quinquis B, Rainteau D, Juillard V, Furlan S, Bouloc P, Nicolas P, Miyoshi A, Azevedo V, Serror P (2022) Adaptation of the gut pathobiont Enterococcus faecalis to deoxycholate and taurocholate bile acids. Sci Rep 12:8485. https://doi.org/10.1038/s41598-022-12552-3
Saiyed AN, Vasavada AR, Johar SRK (2022) Recent trends in miRNA therapeutics and the application of plant miRNA for prevention and treatment of human diseases. Futur J Pharm Sci 8:24. https://doi.org/10.1186/s43094-022-00413-9
Snow JW, Hale AE, Isaacs SK, Baggish AL, Chan SY (2013) Ineffective delivery of diet-derived microRNAs to recipient animal organisms. RNA Biol 10:1107–1116. https://doi.org/10.4161/rna.24909
Sun J, Qi C, Zhu H, Zhou Q, Xiao H, Le G, Chen D, Yu R (2019) IgA-Targeted Lactobacillus jensenii Modulated Gut Barrier and Microbiota in High-Fat Diet-Fed Mice. Front Microbiol 10:1179. https://doi.org/10.3389/fmicb.2019.01179
Teng Y, Ren Y, Sayed M, Hu X, Lei C, Kumar A, Hutchins E, Mu J, Deng Z, Luo C, Sundaram K, Sriwastva MK, Zhang L, Hsieh M, Reiman R, Haribabu B, Yan J, Jala VR, Miller DM, Van Keuren-Jensen K, Merchant ML, McClain CJ, Park JW, Egilmez NK, Zhang HG (2018) Plant-Derived Exosomal MicroRNAs Shape the Gut Microbiota. Cell Host Microbe 24:637-652.e8. https://doi.org/10.1016/j.chom.2018.10.001
Teng Y, Xu F, Zhang X, Mu J, Sayed M, Hu X, Lei C, Sriwastva M, Kumar A, Sundaram K, Zhang L, Park JW, Chen S, Zhang S, Yan J, Merchant ML, Zhang X, McClain CJ, Wolfe JK, Adcock RS, Chung D, Palmer KE, Zhang H-G (2021) Plant-derived exosomal microRNAs inhibit lung inflammation induced by exosomes SARS-CoV-2 Nsp12. Mol Ther 29:2424–2440. https://doi.org/10.1016/j.ymthe.2021.05.005
Timalsina D, Pokhrel KP, Bhusal D (2021) Pharmacologic Activities of Plant-Derived Natural Products on Respiratory Diseases and Inflammations. Biomed Res Int 2021:1636816. https://doi.org/10.1155/2021/1636816
Tjaden B (2023) TargetRNA3: predicting prokaryotic RNA regulatory targets with machine learning. Genome Biol 24:276. https://doi.org/10.1186/s13059-023-03117-2
Witwer KW (2018) Alternative miRNAs? Human sequences misidentified as plant miRNAs in plant studies and in human plasma. F1000Research 7:244. https://doi.org/10.12688/f1000research.14060.1
Witwer KW, McAlexander MA, Queen SE, Adams RJ (2013) Real-time quantitative PCR and droplet digital PCR for plant miRNAs in mammalian blood provide little evidence for general uptake of dietary miRNAs. RNA Biol 10:1080–1086. https://doi.org/10.4161/rna.25246
Xu Q, Qin X, Zhang Y, Xu K, Li Y, Li Y, Qi B, Li Y, Yang X, Wang X (2023) Plant miRNA bol-miR159 Regulates Gut Microbiota Composition in Mice. In Vivo Evidence of the Crosstalk between Plant miRNAs and Intestinal Microbes. J Agric Food Chem 71:16160–16173. https://doi.org/10.1021/acs.jafc.3c06104
Yang J, Farmer LM, Agyekum AAA, Elbaz-Younes I, Hirschi KD (2015) Detection of an Abundant Plant-Based Small RNA in Healthy Consumers. PLoS ONE 10:e0137516. https://doi.org/10.1371/journal.pone.0137516
Yang J, Farmer LM, Agyekum AAA, Hirschi KD (2015) Detection of dietary plant-based small RNAs in animals. Cell Res 25:517–520. https://doi.org/10.1038/cr.2015.26
Zhang F, Yang J, Zhang N, Wu J, Si H (2022) Roles of microRNAs in abiotic stress response and characteristics regulation of plant. Front Plant Sci 13:919243. https://doi.org/10.3389/fpls.2022.919243
Zhang H, Zhang R, Yao J, Hu X, Pu Y, He S, Yu J, Zhu H, Mu B, Zhao C (2022) Effect of F11R Gene Knockdown on Malignant Biological Behaviors of Pancreatic Cancer Cells. J Oncol 2022:3379027. https://doi.org/10.1155/2022/3379027
Zhang L, Hou D, Chen X, Li D, Zhu L, Zhang Y, Li J, Bian Z, Liang X, Cai X, Yin Y, Wang C, Zhang T, Zhu D, Zhang D, Xu J, Chen Q, Ba Y, Liu J, Wang Q, Chen J, Wang J, Wang M, Zhang Q, Zhang J, Zen K, Zhang C-Y (2012) Exogenous plant MIR168a specifically targets mammalian LDLRAP1: evidence of cross-kingdom regulation by microRNA. Cell Res 22:107–126. https://doi.org/10.1038/cr.2011.158
Zhang T, Zhao Y-L, Zhao J-H, Wang S, Jin Y, Chen Z-Q, Fang Y-Y, Hua C-L, Ding S-W, Guo H-S (2016) Cotton plants export microRNAs to inhibit virulence gene expression in a fungal pathogen. Nat Plants 2:16153. https://doi.org/10.1038/nplants.2016.153
Zhang Y, Wiggins BE, Lawrence C, Petrick J, Ivashuta S, Heck G (2012) Analysis of plant-derived miRNAs in animal small RNA datasets. BMC Genom 13:381. https://doi.org/10.1186/1471-2164-13-381
Zhao Q, Liu Y, Zhang N, Hu M, Zhang H, Joshi T, Xu D (2018) Evidence for plant-derived xenomiRs based on a large-scale analysis of public small RNA sequencing data from human samples. PLoS ONE 13:e0187519. https://doi.org/10.1371/journal.pone.0187519
Zhao Y, Yu L, Tian F, Zhao J, Zhang H, Chen W, Xue Y, Zhai Q (2022) Environment-Related Genes Analysis of Limosilactobacillus fermentum Isolated from Food and Human Gut: Genetic Diversity and Adaption Evolution. Foods 11:3135. https://doi.org/10.3390/foods11193135
Acknowledgements
The authors acknowledge the Centro for Genomic Regulation (CRG) for conducting the next-generation sequencing analysis and bioinformatic analysis to identify miRNAs of plant food samples.
Funding
Open Access funding provided thanks to the CRUE-CSIC agreement with Springer Nature. This work was supported by CIBERobn, (grant number CB12/03/30002), Ministerio de Economía y Competitividad (MINECO; RTI2018-102205-B-I00, BFU2015-65937-R, PID2022-141766OB-I00, and PID2022-141313OB-I00 projects), a Center for Nutrition Research (University of Navarra) pre-doctoral fellowship, and an EMBO scientific exchange grant awarded to E.D.
Author information
Authors and Affiliations
Contributions
Conceptualization: S.L., F.M. and E.D.; Methodology: E.D. and P.A.; Writing-original draft preparation: E.D.; Writing-review and editing: S.L., F.M., E.D., P.A., and J.R.; Funding acquisition: F.M., J.R. and S.L; Supervision: F.M. and S.L.; Project administration: F.M. J.R. and S.L. All authors have read and approved the final manuscript. The authors declare that all data were generated in-house and that no paper mill was used.
Corresponding author
Ethics declarations
Ethics approval
Not applicable.
Consent to participate
Not applicable.
Consent for publication
Not applicable.
Conflict of Interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Keypoints
1. Food-derived plants (fruits, vegetables and greens, nuts, legumes, and cereals) contain a wide variety of microRNAs.
2. Plant miR156e, miR159 and miR162 are widely present among edible plants and can resist degradation during cooking and digestion, reaching the gut, although they are not detected in blood.
3. Plant miR156e, miR159 and miR162 could potentially influence human physiology through the modulation of the expression of genes involved in many functions, such as metabolism and cell signalling.
4. New approaches such as the encapsulation of plant miRNAs in extracellular vesicles should be taken into consideration to improve bioavailability of miRNAs.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Díez-Sainz, E., Milagro, F.I., Aranaz, P. et al. MicroRNAs from edible plants reach the human gastrointestinal tract and may act as potential regulators of gene expression. J Physiol Biochem (2024). https://doi.org/10.1007/s13105-024-01023-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s13105-024-01023-0