Abstract
Allergic asthma is the most common type of asthma. It is characterized by TH2 cell–driven inflammation in which interleukin-13 (IL-13) plays a pivotal role. Cytoplasmic RNAs (Y-RNAs), a variety of non-coding RNAs that are dysregulated in many cancer types, are also differentially expressed in patients with allergic asthma. Their function in the development of the disease is still unknown. We investigated the potential role of RNY3 RNA (hY3) in the TH2 cell inflammatory response using the Jurkat cell line as a model. hY3 expression levels were modulated to mimic the upregulation effect in allergic disease. We evaluated the effect of hY3 over cell stimulation and the expression of the TH2 cytokine IL13. Total RNA was isolated and retrotranscribed, and RNA levels were assessed by qPCR. In Jurkat cells, hY3 levels increased upon stimulation with phorbol 12-myristate 13-acetate (PMA) and ionomycin. When transfecting with high levels of hY3 mimic molecules, cell proliferation rate decreased while IL13 mRNA levels increased upon stimulation compared to stimulated control cells. Our results show the effect of increased hY3 levels on cell proliferation and the levels of IL13 mRNA in Jurkat cells. Also, we showed that hY3 could act over other cells via exosomes. This study opens up new ways to study the potential regulatory function of hY3 over IL-13 production and its implications for asthma development.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Allergic diseases have become a global public health issue in the last decades, affecting both developed and developing countries. Allergies are caused by hypersensitivity of the immune system to substances termed allergens that are perceived as a threat and otherwise would be considered harmless. They produce the characteristic type 2 immune response, with a subsequent inflammatory process, that, when affecting lower airways, produces allergic asthma, the most common type of asthma.
Allergic asthma is a chronic inflammatory disorder of the lung airways causing airflow obstruction, which can be triggered by airborne allergens such as house dust mites, pollen from grass, weeds, trees, animal dander, or fungal spores. Classically, this form of asthma is associated with an increase in serum immunoglobulin E (IgE) antibodies and eosinophilia, as well as the presence of interleukin 4 (IL-4)–, IL-5-, and IL-13-producing cells in bronchoalveolar lavage (BAL) [18]. Among them, IL-13 has been shown to play a central role in asthma, mediating many of its key physiologic and pathological features [4]. Its actions in asthma include switching plasma cell antibody synthesis from IgM to IgE production, promoting migration of eosinophils into the lung, increasing permeability and sloughing of airway epithelial cells, increasing mucus production, production of inducible nitric oxide synthase by airway epithelial cells, the transformation of airway fibroblasts into myofibroblasts, proliferation of airway smooth muscle, and stimulation of airway hyperresponsiveness, among others [4]. Despite the signaling cascades that lead to IL13 transcription being well known [35], little is known about the post-transcriptional regulation of IL13. Although initially it was suggested that HuR—an RNA stabilizing protein—regulated post-transcriptionally IL13, stabilizing IL13 mRNA upon cell activation in Jurkat cells [3], and that HuR regulated mRNA stability through the 3′ UTR of different cytokine transcripts [29], Karginov et al. showed that HuR indeed exerted its control through upstream factors during T cell transcription activation [13].
Like other allergic diseases, allergic asthma represents a clear example of a multifactorial disorder, in which predisposing genetic variants cannot explain its rapid prevalence increase. Immunologic development and different environmental cues, like changes in lifestyle, hygiene, or access to antibiotics, have been proposed to be essential factors in allergic asthma development [26]. In recent years, epigenetics has emerged as the underlying mechanism linking genetics, development, and environmental triggers for the onset and progression of allergic diseases and asthma [10]. Thus, several mechanisms have been described to fine-tune different processes, such as gene expression or protein translation. In fact, small non-coding RNAs (sncRNAs) are involved in the epigenetic regulation of several allergic diseases, and many micro-RNAs—a type of sncRNAs—have been associated with allergic asthma by a large number of studies [30].
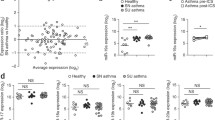
Notwithstanding, investigating other non-coding RNAs (ncRNAs) in allergy is still at an early stage. In a previous transcriptomic study performed with allergic asthmatic patients, we found differential transcription patterns between allergic asthmatic patients and controls, with most of the top 50 differentially expressed transcripts being a kind of ncRNAs called Y-RNA [23]. Recently, we have validated the overexpression of these Y-RNAs in a broader population of 208 allergic patients compared with 96 non-allergic controls and performed an in silico analysis of their potential RNA targets where several genes associated with immune response pathways were identified [11]. This family of sncRNAs was named cytoplasmic RNAs (Y-RNAs) as opposed to the nuclear RNA (U-RNAs) family, both first described as components of the ribonucleoprotein complexes targeted by autoantibodies in patients with systemic erythematous lupus [19].
Y-RNAs have a size that varies between 75 and 115 nucleotides, and although they differ slightly in their primary and secondary structures, they have a typical structure consisting of a double helix stem with paired bases, formed by the 5ʹ and 3ʹ ends of the RNA. Near the 5ʹ end, an unpaired cytosine forms a bulge in the structure necessary for its recognition by Ro60 [28]. Four different Y-RNAs have been described in humans: RNY1, RNY3, RNY4, and RNY5. Also, over 1000 pseudogenes exist that are the product of relatively recent reverse transposition events, with around 90% identity with some of the four Y-RNAs [24].
Even though little is known about its biology, two functions of these RNAs have been described. The best known is to form part of a ribonucleoprotein complex with Ro60 and other proteins. Functional studies of this complex suggest a role in the quality control of non-coding RNAs that require unique folding, i.e., rRNAs or tRNA, in mRNA processing and also the binding to misfolded 3ʹ ends of free RNA. Another function of the Y-RNAs described to date is related to DNA replication and does not rely on the association with the Ro60 ribonucleoprotein complex. [6]. In addition to the functions described, their presence in extracellular fluids, from saliva to blood serum, either as part of ribonucleoprotein complexes or associated with extracellular vesicles, may indicate a role in signal transmission, amplification, or modulation of different responses in their cellular targets both at the local and systemic level [6]. While the number of studies on the effects of extracellular Y-RNA remains limited, they have been described as involved in regulating immune responses and associated with both pro- and anti-inflammatory effects [6, 9].
The role of Y-RNAs in immune system regulation is, therefore, beginning to be understood. Here we aimed to explore the function of these small non-coding RNAs in the modulation of the allergic response and the subsequent inflammatory processes using the Jurkat E6-1 cell line as a model for T cells, an essential cell type for the allergic response.
Materials and methods
Cell culture
Human Jurkat T cell line (E6-1 clone from American Type Culture Collection) was cultured in RPMI medium supplemented with l-glutamine, charcoal-stripped fetal bovine serum (FBS) at a final concentration of 10%, and penicillin/streptomycin at a final concentration of 1%. Cells were cultured at 37 °C in an air atmosphere containing 5% CO2. When required for colorimetric assays, RPMI medium without phenol red was used. Cells were maintained at a concentration between 1 × 105 and 1 × 106 viable cells per mL, and fresh medium was added every 2 to 3 days. Experiments were performed in triplicate.
Cell stimulation
Jurkat cells were simultaneously stimulated with phorbol 12-myristate 13-acetate (PMA) and ionomycin for 24 h [1]. Briefly, cells were incubated in the presence of PMA 10 ng/mL and ionomycin 1 µM. As a control, DMSO was added to untreated cells at the same amount used in PMA and ionomycin. After stimulation, cells were harvested, and media were recovered and stored at − 20 °C for analysis.
Cell proliferation assay
The Amersham Cell Proliferation BioTrak ELISA System kit was used (GE Healthcare, Chicago, IL, USA) to assess cell proliferation according to the manufacturer’s guidelines. It consists of a bromodeoxyuridine (BrdU) labeling. Media were removed from cells and incubated overnight in the presence of BrdU before fixation. Then, cells were blocked, incubated with peroxidase-labeled anti-BrdU, and washed before reacting with TMB (3,3ʹ,5,5ʹ-tetramethylbenzidine) substrate. The reaction was stopped with 1 M sulfuric acid when color development was sufficient for optical density measurement. Plates were read within 10 min after stopping the reaction in a Multiskan Ascent Plate Reader (Thermo Fisher Scientific, Waltham, MA, USA).
Transient transfection
Jurkat cells were transfected with a synthetic RNY3 RNA (hY3), consistent in 102-mer RNA with an identical sequence to human RNY3 RNA (NCBI Reference Sequence: NR_004392.1; Fig. 1b) provided by Integrated DNA Technologies (Coralville, IA, USA). According to the manufacturer’s instructions, the transfection was performed with Xfect™ RNA Transfection Reagent (TaKaRa Bio, Shiga, Japan). Briefly, 5 × 105 cells per mL were plated in RPMI medium without FBS, and 50 pmol of RNA per mL was used for optimal transfection. RNA reaction buffer, RNA, and Xfect™ RNA Transfection polymer complexes were incubated for 10 min before adding to cultures. Cells were incubated for 4 h at 37 °C in a CO2 incubator and then centrifuged to remove nanoparticle complex–containing media. Fresh medium with charcoal-stripped FBS was added to the cells, and then, they were incubated for 48 h before the gene and protein expression assays. Control cells were transfected in parallel with IDTE buffer alone (Integrated DNA Technologies, Coralville, IA, USA). Experiments were performed in triplicate. Average and standard deviation were used for analysis and interpretation.
RNA isolation
Total RNA was isolated using the RNeasy Plus Mini kit (Qiagen, Hilden, Germany) and DNase treated (Qiagen, Hilden, Germany). Reverse transcription (RT) was performed on 1000 ng of total RNA using Superscript III First-Strand Synthesis System for RT-PCR (Invitrogen, Carlsbad, CA, USA).
Gene expression analysis
Relative qPCR was performed as previously described [21, 34]. Briefly, the LightCycler480 Instrument was used with SYBR Green I Master (Roche, Basel, Switzerland). Fold induction was calculated by the comparative Ct method using the 2−(ΔΔCt) or 2− (ΔCt) formulas [21]. GAPDH was used as the reference gene. The experiments were replicated three times, and every sample was performed in triplicate, with non-template controls and a calibrator. All procedures were performed following MIQE guidelines [2]. Table 1 lists the primers used in this study.
Cytokine quantification
Cell supernatants were harvested, centrifuged (400 g, 10 min), and stored at − 80 °C. According to the manufacturer’s guidelines, cytokine levels were determined using Human Cytokine Uncoated ELISA kits provided by Invitrogen (Carlsbad, CA, USA). After incubation, plates were read in a Multiskan Ascent Plate Reader (Thermo Fisher Scientific, Waltham, MA, USA).
Exosome purification
According to the manufacturer’s indications, exosomes delivered to culture media were purified with ExoQuick TC® ULTRA EV Isolation Kit for Tissue Culture Media (System Biosciences, Palo Alto, USA), and a fraction was used to assess purification by Western blot using the exosome markers CD81, CD9, and CD63 (EXO AB kit, System Biosciences) (Supp. Fig. 1). Western blot was performed as described previously [32]. Cells were harvested after the indicated incubation times and centrifuged (400 g) for 5 min. Cell pellets were stored for later processing, and the cell culture media supernatant was collected and stored at − 80 °C until processing. Five milliliters of media were used to purify exosomes. Exosome purification was assessed by Western blot using exosome antibodies provided by System Biosciences (Palo Alto, USA). As a control, media containing charcoal-stripped FBS were purified in parallel.
Statistical analysis
Experiments were performed in triplicate. The average and standard deviation (SD) of the different measurements were calculated. Levene’s test was used to assess sample homoscedasticity, and statistical differences were calculated by t-test for independent samples. Statistical significance for the tests was established at 0.05. Statistical analysis was performed using SPSS software (version 23) (IBM, Armonk, NY, USA).
Results
Jurkat cell line expresses RNY3
A previous transcriptomic analysis by our group found Y-RNAs as the most abundant group of differentially expressed non-coding RNAs in allergic patients [23], and three of them were analyzed in an independent population of allergic patients and healthy controls [11]. These results prompted us to investigate the role of Y-RNAs in the immunologic response observed in allergic patients. To explore the physiological effects of Y-RNA modulation, we decided to use the Jurkat E6-1 cell line as an in vitro model for T cells, as this cell type plays an essential role in the allergic response. Jurkat is an immortalized human T lymphocyte cell that has been widely utilized as a cell model to understand the immune responses associated with T cell signaling.
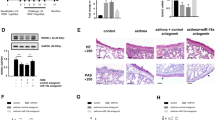
We could confirm by qPCR that the three Y-RNAs described in our previous publications were expressed at a similar magnitude range when normalized with the reference gene GAPGH (Fig. 1a). As these three Y-RNAs shared high homology with the RNY3 consensus sequence (Fig. 1b), one of the four types of Y-RNAs described in humans, the possibility of finding interference effects among them led us to deepen the study of RNY3 (hY3) general mechanisms in our model.
hY3 levels increase in stimulated cells
Our first approach to study the possible role of hY3 in Jurkat physiology was to analyze hY3 levels upon stimulation. We used PMA and the calcium ionophore ionomycin to stimulate Jurkat cells, a well-established model for studying T cell activation [1]. When cells were stimulated, cell proliferation increased significantly as per BrdU incorporation (p = 0.005) (Fig. 2a). Under these conditions, both IL2 mRNA levels (p = 0.016) and IL-2 protein levels (p = 0.003) were strongly increased after 24 h (Fig. 2b), supporting the notion that the T cells were correctly stimulated under our experimental conditions [1]. The analysis of hY3 levels in those cells showed a significant increase (p = 0.01) upon stimulation (Fig. 2c).
RNY3 is overexpressed under stimulation of Jurkat cells. a Proliferation index in cells stimulated with PMA and ionomycin compared with untreated cells. b IL2 mRNA levels were measured by RT-qPCR (left), and IL-2 protein secreted to the media by ELISA (right) in stimulated and unstimulated cells. c hY3 RNA levels were measured by RT-qPCR in unstimulated and stimulated cells. Proliferation was assessed by BrdU incorporation (AU, arbitrary units; *p < 0.05; **p < 0.01)
Effect of increased amounts of hY3 on cell proliferation
A previous publication from our group showed an increased expression of three Y-RNAs in peripheral blood from allergic asthmatic patients compared to healthy individuals [11]. To investigate the functional effects of increased Y-RNA levels on Jurkat cells, we transfected them with synthetic hY3. After 48 h, a significant increase in the intracellular gene expression levels of hY3 (p = 0.048) could be detected by qPCR (Fig. 3a). Evaluating cell proliferation rates (Fig. 3b), we found reduced but non-significant levels (p = 0.094) in cell proliferation of hY3-transfected cells when comparing control non-transfected vs. control hY3-transfected cells.
hY3 transfection affects cell proliferation. a hY3 RNA levels were measured by RT-qPCR after transfection. b Comparison of proliferation rates of cells transfected with hY3 and the control upon stimulation with PMA and ionomycin for 24 h and in normal conditions. c Proliferation fold change was calculated as the proliferation of cells stimulated with PMA and ionomycin for 24 h divided by the proliferation of unstimulated cells. Proliferation was assessed by BrdU incorporation. (AU, arbitrary units; *p < 0.05; **p < 0.01)
No significant differences in cell proliferation rates were found upon stimulation between non-transfected and hY3-transfected cells (Fig. 3b). However, the fold change (control vs. stimulated) was significantly higher (p = 0.008) in hY3-transfected cells (Fig. 3c).
hY3 levels affect IL13 mRNA levels upon stimulation
Since allergic asthma is associated with the presence of IL-13-producing cells in bronchoalveolar lavage (BAL) [18] and plays a relevant role in the development of the disease, we decided to study the IL13 gene expression in our model by using qPCR (Fig. 4).
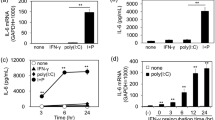
hY3 transfection effect over IL13 and IL2 mRNA levels. a IL2 mRNA levels were measured in cells transfected with hY3 compared with cells not transfected (ns). b IL2 mRNA levels were measured in cells stimulated with PMA and ionomycin for 24 h compared to cells not stimulated (ns). c IL13 mRNA levels were measured in cells transfected with hY3 compared with cells not transfected (ns). d IL13 mRNA levels were measured in cells stimulated with PMA and ionomycin for 24 h compared to cells not stimulated. mRNA levels were assessed by RT-qPCR (*p < 0.05; **p < 0.01; ***p < 0.001)
As mentioned above, we analyzed IL2 mRNA levels as a control to monitor that the Jurkat cells were correctly stimulated. Although both IL2 and IL13 could be detected at low levels in the non-stimulated cells (Fig. 4a, c) their levels substantially increase upon stimulation (Fig. 4b, d). Moreover, IL13 RNA significantly increased in those cells transfected with hY3 compared to the non-transfected (p = 0.041) (Fig. 4d).
We also measured the cytokine levels secreted to the culture media by ELISA. After 24 h of stimulation, IL-13 levels were below the detection limit, while IL-2 levels were around 600 pg/mL. The differences observed in the levels of the secreted cytokines can be due to different cues for their production in Jurkat cells, as has been discussed for other cytokines [20]. The low IL-13 production might depend on the intrinsic characteristics of this cell line, although changes in IL13 mRNA levels have been extensively described [3, 5].
To determine whether the increase of IL13 transcripts observed in hY3-transfected cells upon stimulation responded to an increase in transcription, we studied the expression of the transcription factor GATA3 in our experimental conditions. GATA3 is a master regulator of TH2 cells that is upregulated during TH2 differentiation; it regulates IL13 expression by binding to its promoter, and their levels substantially increase upon stimulation [25]. Interestingly, we did not detect any significant change in GATA3 expression when comparing hY3-transfected cells with non-transfected cells upon stimulation (Fig. 5). This result might suggest a role of hY3 in IL13 mRNA stabilization rather than in IL13 transcription.
hY3 is present in Jurkat-derived exosomes
The presence of Y-RNAs in extracellular fluids, such as saliva or blood serum, seems to be associated with ribonucleoprotein complexes or extracellular vesicles, suggesting a role in signal transmission, amplification, or modulation of different responses in their target cells [6]. Considering this, a set of experiments was designed to detect RNY3 RNA in exosomes from Jurkat cell culture medium.
We purified exosomes from the medium after 72 h of incubation. We detected hY3 at high abundance when normalized with GAPDH (Fig. 6a). In fact, the exosome hY3 level was higher than that of other Y-RNAs, such as ENSG00000207499 (Fig. 6a). Next, we investigated whether hY3 transfection would lead to an increase in exosomes, purifying them from the medium 48 h post-transfection. Transfected cells showed a 12.8-fold increase in hY3 expression compared to the mock control. Interestingly, we also found an exponential enrichment in hY3 in the purified exosomes (Fig. 6b).
hY3 presence in Jurkat-derived exosomes. a Comparison of hY3 and ENSG00000207499 in Jurkat-derived exosomes. b hY3 RNA levels in Jurkat-derived exosomes after transfection with hY3. c Proliferation index in cells incubated for 24 h with the supernatants of control cells and cells transfected with hY3. d Proliferation index in cells incubated for 24 h with exosomes purified from supernatants of control cells and cells transfected with hY3. RNA levels were assessed by RT-qPCR. Proliferation was assessed by BrdU incorporation. (AU, arbitrary units; *p < 0.05)
As mentioned above, exosomes may be a vehicle to transfer hY3 signaling to target cells. To check this hypothesis, we incubated Jurkat cells for 24 h with conditioned media from both control cells and hY3-transfected cells and also with exosomes isolated from those media, and found a reduction in the proliferation index of the cells incubated with hY3-enriched exosomes and the media containing them (Fig. 6c, d). However, the differences in proliferation did not reach significance (p = 0.112). Furthermore, stimulation of Jurkat cells incubated with hY3-enriched exosomes caused no significant effect on proliferation (Supp. Fig. 2).
Discussion
Allergic diseases, particularly allergic asthma, have increased over the last decades, becoming a public health burden. Interest and need to understand the mechanisms underlying these diseases have become increasingly urgent. As in other multifactorial diseases, the development of allergic diseases not only resides on heredity or genetic predisposition but also on other factors, i.e., environmental conditions and epigenetic modulation, that have become crucial components to understanding the onset and evolution of the disease. In this regard, understanding the role of non-coding RNAs in asthma is currently a hot topic. Previous works by our group found significant overexpression of three Y-RNAs (ENSG00000207499, ENSG00000201228, and ENSG00000201555) in allergic asthmatic patients [11, 23]. Unlike miRNAs, little is known about the function of these small non-coding RNAs in these patients. Their sequence homology to the highly conserved Y-RNA RNY3 led us to evaluate the effect of increasing RNY3 RNA levels using the Jurkat cell line as an in vitro model for lymphocyte T stimulation. When cells were stimulated with PMA and ionomycin, hY3 levels increased significantly. Using this stimulation strategy, we could observe a significant increase in IL13 mRNA levels in cells transfected with synthetic RNY3 RNA compared to cells with hY3 basal levels. Furthermore, we detected increased cell proliferation upon stimulation in hY3-transfected cells. In addition, we could confirm that Jurkat-derived exosomes contained hY3 RNA molecules, that hY3 transfection produces an increase of hY3 in exosomes, and finally, according to the trend in the diminution of the proliferation observed, that these exosomes seem to have a minor effect over cell proliferation.
The study of T cell signaling in Jurkat cells is well established and has been extensively reported [1]. It has also been used for studying T cell activation in allergy. Here, we used this model to disentangle the role of hY3 upon cell stimulation, finding an increase in hY3 levels upon stimulation with phorbol 12-myristate 13-acetate and ionomycin (Fig. 2). This result aligns with our previous studies, where we observed an overexpression of three Y-RNAs in allergic patients [11, 23].
Numerous studies have shown different functions of Y-RNAs. Several models have shown that Y-RNAs act both in favor of and against cell proliferation since they are involved in DNA replication and cell cycle progression [17]. Although we could not detect any significant differences in cell proliferation when hY3 was overexpressed, the significant increase in cell proliferation upon stimulation (Fig. 3c) might suggest a possible role in the cellular response to stimulation.
Accordingly, IL2 and IL13 transcripts significantly increased upon stimulation, as previously described [15]. Interestingly, a significant increase of IL13 transcripts upon stimulation was found when hY3 levels were increased upon transfection (Fig. 4d). Notwithstanding, no changes were observed in the expression of the IL13 transcription factor GATA3, suggesting that IL13 transcription is not affected by hY3 (Fig. 5). It is crucial to consider several circumstances to establish how hY3 can modify IL13 transcript levels. First, hY3 is part of ribonucleoprotein complexes and binds to proteins involved in mRNA processing and splicing, such as HuR (ELAVL1) [16]. Interestingly, HuR is involved in T cell activation response and differentiation [13, 31]. Also, Karginov et al. showed that HuR produced a broad control through upstream factors during T cell activation and that its depletion produced NFAT1 upregulation and NFAT2 downregulation [13]. Both NFAT2 and NFAT1 are critical regulators of early gene transcription in response to T cell activation and can bind to IL13 promoter in Jurkat cells [22]. Finally, hY3 has been proposed as a molecular sponge for HuD (ELAVL4) activity, an RNA-binding protein that plays a fundamental role during neuronal differentiation controlling neuronal cell fate [12]. hY3 competes with HuD target mRNAs and limits HuD access to the polysomal compartment [27]. Considering this, we could speculate that hY3 could act as a molecular sponge of HuR, impairing its interactions with other transcripts. Moreover, although HuR does not impact NFAT2 expression through its 3′ UTR [13], an association of NFAT5 mRNA with HuR has been found [7]; thus, we cannot rule a similar relation for NFAT1.
In addition to the role of Y-RNAs inside the cell, extracellular vesicles containing Y-RNAs have been found in different biological fluids and related to several diseases, including different types of cancer and cardiovascular diseases [6, 8]. In fact, Driedonks et al. recently showed that most full-length Y-RNAs are present in extracellular vesicles [5]. These vesicles seem to be a vehicle for mediating intercellular RNA transfer that could alter gene expression in target cells [33].
We have detected hY3 transcripts in exosomes derived from Jurkat cells, confirming what was previously reported elsewhere [14]. Moreover, exosomes derived from cells transfected with hY3 contained high levels of this transcript (Fig. 6b). Despite not being statistically significant, cell proliferation seemed impaired when growing in a medium supplemented with exosomes derived from hY3-overexpressing Jurkat cells (Fig. 6d). Nevertheless, no effect could be detected upon stimulation. This observation suggests that Y-RNAs can be transported in exosomes, but whether these exosomes have a function other than stimulating proliferation remains to be further explored and opens a new avenue in T cell signaling research.
Our study presents some limitations. First, we used the immortalized cell line Jurkat E6-1 as a model of T cell, which might not be as good a proxy to TH2 cell response as a primary T cell culture. Additionally, concerning IL13, only mRNA and not secreted protein could be detected at confident levels. Nevertheless, we could see increased IL13 and GATA3 expression levels upon cell activation, characteristic of the TH2 response. Finally, our observations over exosomes and hY3 were made to study the mere effect of Y-RNA content on T cell proliferation.
In conclusion, we have shown the effect of hY3 modulation in our T cell model and observed its influence on the cell proliferation and IL13 mRNA levels upon stimulation. Because IL-13 is a pivotal cytokine in T2-type asthma, this study opens up new ways to study the potential regulatory function of hY3 over IL-13 production and its implications for asthma development.
References
Abraham RT, Weiss A (2004) Jurkat T cells and development of the T-cell receptor signalling paradigm. Nat Rev Immunol 4:301–308. https://doi.org/10.1038/nri1330
Bustin SA, Benes V, Garson JA, Hellemans J, Huggett J, Kubista M, Mueller R, Nolan T, Pfaffl MW, Shipley GL, Vandesompele J, Wittwer CT (2009) The MIQE Guidelines: minimum information for publication of quantitative real-time PCR experiments. Clin Chem 55:611–622. https://doi.org/10.1373/clinchem.2008.112797
Casolaro V, Fang X, Tancowny B, Fan J, Wu F, Srikantan S, Asaki SY, De Fanis U, Huang S-K, Gorospe M, Atasoy UX, Stellato C (2008) Posttranscriptional regulation of IL-13 in T cells: role of the RNA-binding protein HuR. J Allergy Clin Immunol 121:853-859.e4. https://doi.org/10.1016/j.jaci.2007.12.1166
Corren J (2013) Role of interleukin-13 in asthma. Curr Allergy Asthma Rep 13:415–420. https://doi.org/10.1007/s11882-013-0373-9
Driedonks TAP, Mol S, Bruin S, Peters A, Zhang X, Lindenbergh MFS, Beuger BM, Stalborch AD, Spaan T, Jong EC, Vries E, Margadant C, Bruggen R, Vlaar APJ, Groot Kormelink T, Nolte-‘T Hoen ENM (2020) Y-RNA subtype ratios in plasma extracellular vesicles are cell type- specific and are candidate biomarkers for inflammatory diseases. J Extracell Vesicles 9:1764213. https://doi.org/10.1080/20013078.2020.1764213
Driedonks TAP, Nolte-‘T Hoen ENM (2018) Circulating Y-RNAs in extracellular vesicles and ribonucleoprotein complexes; implications for the immune system. Front Immunol 9:3164. https://doi.org/10.3389/fimmu.2018.03164
DuMond JF, Ramkissoon K, Zhang X, Izumi Y, Wang X, Eguchi K, Gao S, Mukoyama M, Burg MB, Ferraris JD (2016) Peptide affinity analysis of proteins that bind to an unstructured NH 2 -terminal region of the osmoprotective transcription factor NFAT5. Physiol Genomics 48:290–305. https://doi.org/10.1152/physiolgenomics.00110.2015
Guglas K, Kolenda T, Stasiak M, Kopczyńska M, Teresiak A, Ibbs M, Bliźniak R, Lamperska K (2020) YRNAs: new insights and potential novel approach in head and neck squamous cell carcinoma. Cells 9:1281. https://doi.org/10.3390/cells9051281
Hizir Z, Bottini S, Grandjean V, Trabucchi M, Repetto E (2018) RNY (YRNA)-derived small RNAs regulate cell death and inflammation in monocytes/macrophages. Cell Death Dis 8:e2530–e2530. https://doi.org/10.1038/cddis.2016.429
Isidoro-García M, Dávila I, Pascual de Pedro M, Sanz C, Lorente F (2007) Interactions between genes and the environment. Epigenetics in allergy. Allergol Immunopathol (Madr) 35:254–258. https://doi.org/10.1157/13112992
Isidoro-García M, García-Sánchez A, Sanz C, Estravís M, Marcos-Vadillo E, Pascual M, Roa S, Marqués-García F, Triviño JC, Dávila I (2019) YRNAs overexpression and potential implications in allergy. World Allergy Organ J 12:100047. https://doi.org/10.1016/j.waojou.2019.100047
Jung M, Lee EK (2021) RNA–binding protein HuD as a versatile factor in neuronal and non–neuronal systems. Biology (Basel) 10:361. https://doi.org/10.3390/biology10050361
Karginov FV (2019) HuR controls apoptosis and activation response without effects on cytokine 3’ UTRs. RNA Biol 16:686–695. https://doi.org/10.1080/15476286.2019.1582954
Kaur S, Elkahloun AG, Arakelyan A, Young L, Myers TG, Otaizo-Carrasquero F, Wu W, Margolis L, Roberts DD (2018) CD63, MHC class 1, and CD47 identify subsets of extracellular vesicles containing distinct populations of noncoding RNAs. Sci Rep 8:2577. https://doi.org/10.1038/s41598-018-20936-7
Keen JC, Cianferoni A, Florio G, Guo J, Chen R, Roman J, Wills-Karp M, Casolaro V, Georas SN (2006) Characterization of a novel PMA-inducible pathway of interleukin-13 gene expression in T cells. Immunology 117:29–37. https://doi.org/10.1111/j.1365-2567.2005.02260.x
Köhn M, Ihling C, Sinz A, Krohn K, Hüttelmaier S (2015) The Y3** ncRNA promotes the 3′ end processing of histone mRNAs. Genes Dev 29:1998–2003. https://doi.org/10.1101/gad.266486.115
Kowalski MP, Krude T (2015) Functional roles of non-coding Y RNAs. Int J Biochem Cell Biol 66:20–29. https://doi.org/10.1016/j.biocel.2015.07.003
Lambrecht BN, Hammad H (2015) The immunology of asthma. Nat Immunol 16:45–56. https://doi.org/10.1038/ni.3049
Lerner MR, Boyle JA, Hardin JA, Steitz JA (1981) Two novel classes of small ribonucleoproteins detected by antibodies associated with lupus erythematosus. Science 211:400–402. https://doi.org/10.1126/science.6164096
Liu M, Yasmeen R, Fukagawa N, Yu L, Kim Y, Wang T (2017) Dose-dependent responses of I3C and DIM on T-cell activation in the human T lymphocyte Jurkat cell line. Int J Mol Sci 18:1409. https://doi.org/10.3390/ijms18071409
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2-ΔΔCT method. Methods 25:402–408. https://doi.org/10.1006/meth.2001.1262
MacLean Scott E, Solomon LA, Davidson C, Storie J, Palikhe NS, Cameron L (2018) Activation of Th2 cells downregulates CRTh2 through an NFAT1 mediated mechanism. PLoS ONE 13:e0199156. https://doi.org/10.1371/journal.pone.0199156
Pascual M, Roa S, García-Sánchez A, Sanz C, Hernandez-Hernandez L, Greally JM, Lorente F, Dávila I, Isidoro-García M (2014) Genome-wide expression profiling of B lymphocytes reveals IL4R increase in allergic asthma. J Allergy Clin Immunol 134:972–975. https://doi.org/10.1016/j.jaci.2014.05.015
Perreault J, Noël J-F, Brière F, Cousineau B, Lucier J-F, Perreault J-P, Boire G (2005) Retropseudogenes derived from the human Ro/SS-A autoantigen-associated hY RNAs. Nucleic Acids Res 33:2032–2041. https://doi.org/10.1093/nar/gki504
Rayees S, Malik F, Bukhari SI, Singh G (2014) Linking GATA-3 and interleukin-13: implications in asthma. Inflamm Res 63:255–265. https://doi.org/10.1007/s00011-013-0700-6
Renz H, Autenrieth IB, Brandtzæg P, Cookson WO, Holgate S, von Mutius E, Valenta R, Haller D (2011) Gene-environment interaction in chronic disease: a European Science Foundation Forward Look. J Allergy Clin Immunol 128:S27–S49. https://doi.org/10.1016/j.jaci.2011.09.039
Tebaldi T, Zuccotti P, Peroni D, Köhn M, Gasperini L, Potrich V, Bonazza V, Dudnakova T, Rossi A, Sanguinetti G, Conti L, Macchi P, D’Agostino V, Viero G, Tollervey D, Hüttelmaier S, Quattrone A (2018) HuD is a neural translation enhancer acting on mTORC1-responsive genes and counteracted by the Y3 small non-coding RNA. Mol Cell 71:256-270.e10. https://doi.org/10.1016/j.molcel.2018.06.032
Teunissen SW, Kruithof MJ, Farris AD, Harley JB, Venrooij WJ, Pruijn GJ (2000) Conserved features of Y RNAs: a comparison of experimentally derived secondary structures. Nucleic Acids Res 28:610–619. https://doi.org/10.1093/nar/28.2.610
Wang JG, Collinge M, Ramgolam V, Ayalon O, Fan XC, Pardi R, Bender JR (2006) LFA-1-dependent HuR nuclear export and cytokine mRNA stabilization in T cell activation. J Immunol 176:2105–2113. https://doi.org/10.4049/jimmunol.176.4.2105
Weidner J, Bartel S, Kılıç A, Zissler UM, Renz H, Schwarze J, Schmidt-Weber CB, Maes T, Rebane A, Krauss-Etschmann S, Rådinger M (2021) Spotlight on microRNAs in allergy and asthma. Allergy 76:1661–1678. https://doi.org/10.1111/all.14646
Yu S, Tripod M, Atasoy U, Chen J (2021) HuR plays a positive role to strengthen the signaling pathways of CD4+ T cell activation and Th17 cell differentiation. J Immunol Res 2021:1–13. https://doi.org/10.1155/2021/9937243
Zhai K, Duan H, Khan GJ, Xu H, Han F, Cao W, Gao G, Shan L, Wei Z-J (2018) Salicin from Alangium chinense ameliorates rheumatoid arthritis by modulating the Nrf2-HO-1-ROS pathways. J Agric Food Chem 66:6073–6082. https://doi.org/10.1021/acs.jafc.8b02241
Zhai K, Duan H, Wang W, Zhao S, Khan GJ, Wang M, Zhang Y, Thakur K, Fang X, Wu C, Xiao J, Wei Z (2021) Ginsenoside Rg1 ameliorates blood–brain barrier disruption and traumatic brain injury via attenuating macrophages derived exosomes miR-21 release. Acta Pharm Sin B 11:3493–3507. https://doi.org/10.1016/j.apsb.2021.03.032
Zhai K, Zheng J, Tang Y, Li F, Lv Y, Zhang Y, Gao Z, Qi J, Yu B, Kou J (2017) The saponin D39 blocks dissociation of non-muscular myosin heavy chain IIA from TNF receptor 2, suppressing tissue factor expression and venous thrombosis. Br J Pharmacol 174:2818–2831. https://doi.org/10.1111/bph.13885
Zhu J (2015) T helper 2 (Th2) cell differentiation, type 2 innate lymphoid cell (ILC2) development and regulation of interleukin-4 (IL-4) and IL-13 production. Cytokine 75:14–24. https://doi.org/10.1016/j.cyto.2015.05.010
Acknowledgements
We thank Alicia Landeira-Viñuela and Clara Sanabria-Luis for contributing to this work as technicians. The authors are very grateful to María Begoña García Cenador for technical support.
Funding
Open Access funding provided thanks to the CRUE-CSIC agreement with Springer Nature. This research was funded by the Instituto de Salud Carlos III and the European Regional Development Fund “A way to make Europe,” grant numbers PI17/00818, PI20/00268, and IMP/00009, a grant from Junta de Castilla y León cofinanced by the European Regional Development Fund “A way to make Europe” (IES161P20), and a grant from the Gerencia Regional de Salud de la Junta de Castilla y León (GRS2156/A/20); Instituto de Salud Carlos III’s Thematic Network of Cooperative Research in Health—RETICS (Red temática de investigación en salud Asma, Reacciones Adversas y Alérgicas, ARADyAL), grant number RD16/0006/0019; and Instituto de Salud Carlos III’s Health Outcomes-Oriented Cooperative Research Networks (RICORS) Inflammatory Diseases (RD21/0002/0037), funded by the European Union – NextGenerationEU, Mecanismo para la Recuperación y la Resiliencia (MRR). M.J.M. and M.E. salaries were fully covered by a grant from Junta de Castilla y León cofinanced by the European Regional Development Fund (IES161P20). J.P.P.’s wage was partly funded by Instituto de Salud Carlos III’s Thematic Network of Cooperative Research in Health—RETICS ARADyAL grant number RD16/0006/0019.
Author information
Authors and Affiliations
Contributions
Conceptualization, I.D., M.I.-G., M.E., A.G.-S., and C.S.; methodology, M.E. A.G.-S., M.J.M., J.P.-P., M.I.-G, and C.S.; investigation, M.E., A.G.-S., M.J.M., J.P.-P., and C.S.; validation, M.E., A.G.-S., M.J.M., J.P.-P., and C.S.; formal analysis, M.E., A.G.-S., M.J.M., J.P.-P., I.D., M.I.-G., and C.S.; visualization, M.E., J.P.-P., and C.S; writing—original draft preparation, M.E., I.D., and C.S.; writing—review and editing, M.E., A.G.-S., M.J.M., J.P.-P., I.D., M.I.-G., and C.S.; supervision, I.D., M.I.-G., and C.S.; funding acquisition, I.D., M.I.-G., and C.S. All authors have read and agreed to the published version of the manuscript. The authors declare that all data were generated in-house and that no paper mill was used.
Corresponding author
Ethics declarations
Conflict of interest
I.D. reports personal fees from Novartis, Sanofi, GSK, Astra-Zeneca, LETI, and Teva, and grant and personal fees from Thermo Fisher outside the submitted work. The rest of the authors declare no conflict of interest.
Disclaimer
The funders had no role in the design of the study, in the collection, analyses, or interpretation of data, in the writing of the manuscript, or in the decision to publish the results.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Key Points
- Cell activation increases hY3 (RNY3) levels in the Jurkat cell line.
- Higher levels of hY3 produce increased levels of IL-13 mRNA upon cell stimulation in the Jurkat cell line.
- Higher levels of hY3 increase the proliferative rate of Jurkat cells upon stimulation.
Supplementary Information
Below is the link to the electronic supplementary material.
13105_2022_920_MOESM1_ESM.jpg
Supplementary file1 (JPG 542 KB) Supplementary Figure 1. Protein extracts of purified exosomes. Protein extracts of purified exosomes were subjected to WB to assess exosome purification. Lanes 1-3 Jurkat-derived exosomes of non-transfected cells, lanes 4-6 Jurkat-derived exosomes of hY3 transfected cells. Western blots were hybridized with polyclonal antibodies against CD9, CD81 and CD63 as specified by the manufacturer of the kit.
13105_2022_920_MOESM2_ESM.jpg
Supplementary file2 (JPG 631 KB) Supplementary Figure 2. Cell stimulation in the presence of purified exosomes. (a) RNY3 RNA levels after transfection, (b) RNY3 RNA levels in Jurkat-derived exosomes after transfection, (c) Comparison of proliferation rates of cells in the presence of RNY3 enriched exosomes and control exosomes upon stimulation and in normal conditions. No significative differences were found associated with hY3 content exosomes (p = 0.317 and 0.252 in control and stimulation conditions, respectively). RNA levels were assessed by RT-qPCR. Proliferation was assessed by BrdU incorporation. (AU, arbitrary units; *, p < 0.05; **, p < 0.01; ***, p < 0.001)
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Estravís, M., García-Sánchez, A., Martin, M.J. et al. RNY3 modulates cell proliferation and IL13 mRNA levels in a T lymphocyte model: a possible new epigenetic mechanism of IL-13 regulation. J Physiol Biochem 79, 59–69 (2023). https://doi.org/10.1007/s13105-022-00920-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13105-022-00920-6