Abstract
Cerebral small vessel disease (SVD) is common in older people and is associated with lacunar stroke, white matter hyperintensities (WMH) and vascular cognitive impairment. Cerebral blood flow (CBF) is reduced in SVD, particularly within white matter.
Here we quantified test–retest reliability in CBF measurements using pseudo-continuous arterial spin labelling (pCASL) in older adults with clinical and radiological evidence of SVD (N=54, mean (SD): 66.9 (8.7) years, 15 females/39 males). We generated whole-brain CBF maps on two visits at least 7 days apart (mean (SD): 20 (19), range 7-117 days).
Test–retest reliability for CBF was high in all tissue types, with intra-class correlation coefficient [95%CI]: 0.758 [0.616, 0.852] for whole brain, 0.842 [0.743, 0.905] for total grey matter, 0.771 [0.636, 0.861] for deep grey matter (caudate-putamen and thalamus), 0.872 [0.790, 0.923] for normal-appearing white matter (NAWM) and 0.780 [0.650, 0.866] for WMH (all p<0.001). ANCOVA models indicated significant decline in CBF in total grey matter, deep grey matter and NAWM with increasing age and diastolic blood pressure (all p<0.001). CBF was lower in males relative to females (p=0.013 for total grey matter, p=0.004 for NAWM).
We conclude that pCASL has high test–retest reliability as a quantitative measure of CBF in older adults with SVD. These findings support the use of pCASL in routine clinical imaging and as a clinical trial endpoint.
All data come from the PASTIS trial, prospectively registered at: https://eudract.ema.europa.eu (2015-001235-20, registered 13/05/2015), http://www.clinicaltrials.gov (NCT02450253, registered 21/05/2015).
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Cerebral small vessel disease (SVD) is common in older people, observed radiologically as focal lacunar infarcts, micro-haemorrhages and diffuse white matter hyperintensities (WMH) on T2-weighted MRI scans [1, 2]. SVD is the main cause of clinical lacunar stroke and a major source of vascular contributions to cognitive impairment and dementia (VCID) [3,4,5,6]. SVD is associated with reduced cerebral blood flow (CBF) [7, 8]. Resting CBF is lower in individuals with SVD compared to healthy aged control subjects, particularly in subcortical white matter [7,8,9,10,11,12,13,14].
Arterial spin labelling is now well developed as an MRI-derived quantitative measure of CBF. As the test–retest reproducibility of this method has not been quantified in people with SVD, it has not entered routine use for clinical assessment or as a clinical trial outcome measure. Here we report whole-brain CBF maps measured by pseudo-continuous arterial spin labelling (pCASL) [15, 16] in a well-characterised cohort of older adults with symptomatic SVD [17]. First, we aimed to test whether pCASL is an effective method for quantitative assessment of CBF in people with SVD, particularly in white matter areas where absolute CBF values are low [7, 10, 18]. Second, we aimed to quantify test–retest reliability in CBF measured by pCASL within this participant group, using CBF maps derived from two successive visits, at least 7 days apart.
Methods
Study population
All data are from a cohort of older adults with radiological and clinical evidence of symptomatic SVD (N=54, demographic details in Table 1). These participants were all recruited as part of a double-blinded, placebo-controlled, phase-II clinical trial, Perfusion by Arterial spin labelling following Single dose Tadalafil In Small vessel disease (PASTIS; European Union Clinical Trials Register number 2015-001235-20, registered 13/05/2015) [17]. The trial received ethical approval from the UK National Research Ethics Service (REC reference: 15/LO/0714). Further details are given in the Supplementary file.
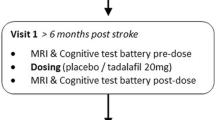
Participants attended an initial screening visit (“visit 0”) and completed an eligibility check and gave informed consent. During the screening visit, education level and Montreal Cognitive Assessment (MoCA) scores were recorded (see Table 1). Following consent, participants attended two study visits (visit 1, visit 2) at least 7 days apart as specified in the study protocol [17]. At each study visit, participants underwent systolic/diastolic blood pressure (SBP/DBP) measurement, a cognitive test battery (see Supplementary file) and brain MRI scanning including pCASL. Participants then received either drug or placebo, according to the cross-over design, after which blood pressure, cognitive and MRI measurements were all repeated. All data reported here are from the set of measurements prior to treatment with drug or placebo.
Inclusion criteria were as follows. 1, radiological evidence of SVD, defined as: MRI evidence of lacunar infarct(s) ≤ 1.5 cm maximum diameter and/or confluent deep white matter leukoaraiosis (≥ grade 2 on the Fazekas scale). 2, Clinical evidence of cerebral small vessel disease defined as either: lacunar stroke syndrome with symptoms lasting >24 hours, occurring at least 6 months prior to visit 1; or: transient ischaemic attack lasting < 24 hours with limb weakness, hemi-sensory loss or dysarthria at least 6 months previously and with MR DWI performed acutely showing lacunar infarction. If MRI was not performed within 10 days of TIA, a lacunar infarction in an anatomically appropriate position as demonstrated on a subsequent MRI was also deemed eligible. 3, Age ≥ 50 years. 4, imaging of the carotid arteries with Doppler ultrasound, CT angiography or MR angiography in the previous 12 months, demonstrating < 70% stenosis in both internal carotid arteries or < 50% stenosis in both internal carotids if measured in previous 12-60 months.
Exclusion criteria included: known diagnosis of dementia; cortical infarction (>1.5 cm maximum diameter); systolic BP < 90 and/or diastolic BP < 50 mmHg; creatinine clearance <30ml/min; stroke or TIA within 6 months. For a full list of exclusion criteria see the published protocol [17].
Blood Pressure measurement
SBP/DBP measurements were taken from each participant for visits 1 and 2, first on arrival after resting and then again after MRI scanning, using a validated Omron MX3 Plus machine.
Magnetic Resonance Image Acquisition
Whole-brain perfusion MRI was acquired using a 3T scanner (Achieva TX MRI scanner, Philips Medical Systems, Eindhoven, Netherlands) at St George’s University Hospitals NHS Foundation Trust. Whole-brain T1-weighted, Fluid Attenuated Inversion Recovery (FLAIR) and pseudo-continuous arterial spin labelling (pCASL) images were acquired. All MRI data were acquired from brain scans performed on a Tuesday or Thursday between the hours of 10:00 and 12:00.
T1-weighted MRI
Whole-brain sagittal 3D T1-weighted images were acquired to enable tissue segmentation with the following protocol: Turbo Field Echo (TFE) sequence with an inversion pre-pulse, TFE factor 240 in multi-shot mode with 3000-ms shot interval, 8° flip angle, TR 7.9 ms, TE 3.8 ms, 1mm×1mm×1.5mm acquired resolution with interpolation to 1 mm isotropic resolution, 1 average and SENSE factor 2 for a 3-minute 47-second acquisition time.
FLAIR MRI
2D T2-weighted axial FLAIR images were acquired to detect WMH using the following protocol: T2 weighted turbo-spin-echo sequences with selective fat suppression (TSE-SPIR), TR 11000 ms, TE 120 ms, TI 2800 ms, 0.65 mm×1.00 mm acquired resolution interpolated to 0.45×0.45 mm over 24-thick slices (5 mm thickness), with 2 averages and a 1.75 SENSE factor for a 3-minute 51-second acquisition time. Examples are shown in Figs. 1, 2.
Example of white matter hyperintensities (WMH) in an older adult with small vessel disease. A, axial FLAIR scan, showing WMH. B, the same axial slice following semi-automated highlighting of WMH using Jim 7.0 software. Participant #022, female aged 77 y, with total WMH volume of 34,510 mm3 (across all scan slices).
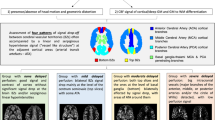
An example of regional anatomical and CBF mapping, with tissue segmentation. A, FLAIR image at full resolution. B, FLAIR image co-registered to the cerebral blood flow map, with voxels re-sized to be comparable with pCASL map. C, cerebral blood flow map, derived from pCASL. The calibration bar shows 0.0 - 80.0 ml/min/100g. D, tissue segmentation map for CBF computation. Each voxel has been defined as either: grey matter (GM), normal appearing white matter (NAWM), white matter hyperintensity (WMH) or cerebrospinal fluid (CSF). E, F: graphs show the probability density functions of cerebral blood flow values in voxels assigned as grey matter (in E) and normal appearing white matter (F). For this participant, median CBF in grey matter was 51.3 mm/min/100g and in NAWM 21.8 ml/min/100g. Participant #023, female, aged 56 y, visit 1.
pCASL MRI
Our pCASL protocol was developed based on the consensus recommendations of the ISMRM Perfusion study group and European consortium for ASL in dementia [15] using the Philips product pCASL sequence in the scanner 5.3 software release. A 64×64 acquisition matrix with 16 slices was used to acquire data with 4mm×4mm×7mm voxel size. Image readout was in 2D Echo-Planar Images (EPI).
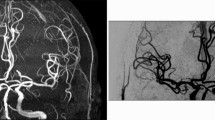
Background tissue suppression was performed based on the ASL consensus recommendations [15]. Two inversion pulses were provided for background suppression as these represent an effective trade-off between tissue suppression and ASL signal. Background suppression pulses were applied at the beginning of the pulse sequence [15]. Spectral Pre-saturation with Inversion Recovery (SPIR) for fat suppression was applied to improve the contrast to noise of the blood perfusion signal. SPIR fat suppression did not add additional time to each slice acquisition. Pairwise acquisition of label and control images was performed. A total of 140 volumes (alternating with and without the spin labelling inversion pulse) were acquired in two separate 10-minute acquisitions using SENSE 2.3, and TE 8ms and TR 4300 ms with a labelling duration (τ) = 1800 ms and post-labelling delay (PLD) = 2000 ms. This was performed twice to increase ASL signal-to-noise ratio in the white matter [19]. This corresponds to a total acquisition time for the pCASL data of 20 min 6 s. A fixed labelling distance of 85 mm from the centre of the imaging block was used with the labelling slice positioned below the cerebellum at an angle perpendicular to the carotid arteries (visualized by time of flight angiography). Proton density-weighted images were acquired, to enable computation of CBF, using the pCASL sequence without the inversion pulse and background suppression, but with fat suppression and an increased TR 5000 ms to minimize T1 weighting (TE 9 ms with 8 averages). Proton density-weighted images were acquired in 40 s.
Computation of CBF maps
The pCASL data acquisitions at each visit were corrected for subject movement using the FMRIB software library (FSL) function eddy_correct (https://fsl.fmrib.ox.ac.uk/FSL) [20]. An average pCASL map was then separately computed for each pCASL data acquisition. The average pCASL maps and the second proton density-weighted image were aligned to the initial proton density-weighted image in each scan session using the FSL Linear Image Registration Tool (flirt) [21]. These transformations were applied to the motion-corrected pCASL data to ensure all proton density-weighted and pCASL images were aligned in the same space. The aligned proton density-weighted images were averaged, and CBF was computed using oxford_asl (part of the FSL-BASIL toolset, https://fsl.fmrib.ox.ac.uk/fsl/fslwiki/BASIL) [22]. Cerebral blood flow in each voxel was calculated in physiological units of ml/min/100g using the standard equation for pCASL with (Equation 1) [15]:
where SIcontrol and SIlabel are the time-averaged signal intensities in the pCASL control and label images, respectively, and SIPD is the signal intensity of a proton density-weighted image. Standard values were inserted into Equation 1 for the brain/blood partition coefficient, λ=0.9 ml/g, the labelling efficiency, α=0.85, longitudinal relaxation time of arterial blood T1,blood=1650 ms at 3T. An example pCASL map is shown in Fig 2C.
White matter hyperintensity (WMH) delineation
WMHs were semi-automatically highlighted on each axial slice of the visit 1 FLAIR images (Fig. 1) using Jim 7.0 software (http://www.xinapse.com/jim-7-software/ Xinapse Systems Ltd, West Bergholt, Essex, UK). WMH were defined as hyperintense regions, which were (1) not due to the presence of blood vessels, and (2) not less than 10 mm2 in size, and (3) not a narrow band, one pixel wide, along the edge of the ventricles. A binary WMH image was generated, and the total WMH volume (in mm3) was computed for each participant. All WMH maps used here were produced by a single operator, blind to treatment allocation and to all clinical details (FAHH). A second, blinded operator (MMHP) also produced maps for a subset of participants (n=51), and interoperator agreement was good (intra-class correlation coefficient for total WMH volume ICC=0.855 [95% confidence interval: 0.760, 0.915], two-way random-effects model).
Tissue Segmentation
For each scan session, T1-weighted images in native space were segmented into grey matter, white matter and cerebrospinal fluid (CSF) tissue probability maps, using a modified form of the standard Statistical Parametric Mapping (SPM) (SPM Version 12, https://www.fil.ion.ucl.ac.uk/spm/) geodesic shooting segmentation and normalisation procedure described in full in our previous papers [23, 24]. This procedure captures population-specific features, e.g. enlarged ventricles, and allows superior delineation of deep grey matter structures compared to the standard SPM pipeline. The binary WMH mask (co-registered into native T1-weighted space) was used to repair the tissue probability maps for misclassification caused by WMHs.
Native space T1-weighted and native space FLAIR were skull-stripped using FSL’s brain extraction tool (https://fsl.fmrib.ox.ac.uk/fsl/fslwiki/BET) [25] and co-registered to the average proton density-weighted image using boundary-based registration (FSL epi-reg) [26]. These 12 parameter linear transformations were used to align the corrected T1-weighted tissue probability maps and the binary WMH map to the CBF maps. A tissue mask in the average proton density-weighted image space was computed assigning each voxel to either grey matter, normal appearing white matter (NAWM), WMH or CSF, based on the maximum tissue probability.
Computation of CBF in whole-brain tissue
For the alignment of the T1-weighted tissue segmentation images to the low-resolution pCASL images, it was necessary to apply a further segmentation step. This tissue segmentation procedure employs a novel application of a tissue segmentation algorithm to CBF maps [27]. It is designed to assign voxels with high CBF values to grey matter and low CBF values to white matter segments. The distribution of CBF values within the grey matter and white matter tissue masks computed in the Tissue Segmentation section (above) was entered as empirical priors to a hidden Markov random field model and segmentation (FMRIB's Automated Segmentation Tool, FAST) [27] to provide an improved segmentation of grey and white matter tissue from the CBF maps. This technique reduces the effects of partial volume and tissue classification errors at the boundary between grey and white matter tissue caused by the large pCASL image voxel size and the relative difference between voxel sizes of the native pCASL and T1-weighted images. In particular, this method assigns voxels with high CBF values at the grey/white matter tissue boundary to the grey matter segment and voxels with low CBF values at the grey/white matter tissue boundary to the white matter segment. To avoid misclassification of CSF and WMH regions, voxels in these regions were not entered into the FAST segmentation step. In our hands this approach was more successful than the more standard co-registration of tissue segmentations from high-resolution T1-weighted images to low-resolution pCASL images (not shown). For each participant at each scan session the median CBF values were calculated for total grey matter, NAWM and WMH. An example of tissue segmentation is shown in Fig. 2.
Computation of CBF in deep grey matter structures
Cerebral deep grey matter structures were segmented on native space T1-weighted images using Freesurfer (Freesurfer Version 5.3.0, https://surfer.nmr.mgh.harvard.edu/fswiki/). The binary segmentations of the caudate, putamen and thalamus were aligned to the CBF maps by application of the affine transformation computed in Tissue Segmentation (above). Median CBF values were calculated for each of these three anatomical deep grey matter structures across the left and right cerebral hemispheres. An average of these three median values is reported for CBF in deep grey matter (DGM).
Statistical Analysis
Statistical analyses were performed using SPSS (version 25.0). Unless otherwise stated data are presented as mean (SD). Test–retest reliability of CBF values (in ml/min/100g) between visit 1 and visit 2 was computed using intra-class correlation coefficients for whole brain, total grey matter, DGM, NAWM and WMH. Within-subjects coefficient of variation (wsCV) was also calculated for CBF values in each of these tissue types. Correlation of CBF values between tissue types was calculated using Pearson’s correlation coefficient. Bland-Altman plots were used to assess bias in CBF data between visit 1 and visit 2. Mean difference and upper and lower limits of agreement defined as ±1.96 standard deviations around the mean difference are reported. ANCOVA models were used to test for associations between: age (years), sex (M/F), blood pressure (SBP, DBP in mmHg) and CBF values. CBF was the dependent variable, sex was a fixed factor and age, SBP and DBP were co-variates. For CBF, SBP and DBP an average of the values for visit 1 and visit 2 was used in these analyses. No corrections were made for multiple comparisons, and p<0.05 was considered significant.
Results
CBF maps were generated using pCASL in a cohort of older adults (age 66.9 (8.7) range: 52-87 years, N=54) all of whom had symptomatic SVD (Table 1, example in Figure 2). All participants had survived a lacunar stroke, and visit 1 occurred at least six months post-stroke. All participants completed visit 1 and visit 2 at least 7 days apart (mean (SD): 20 (19) days, range 7-117 days). Only four participants completed visit 2 more than 30 days after visit 1 (range 54-117 days). If these four participants were excluded, none of the parameters reported below changed significantly (P values in the range: 0.733 to 0.994; not shown).
For each participant CBF data were documented for visit 1 and visit 2 in whole brain and in four tissue types: grey matter (derived from all voxels defined as grey matter), DGM (from grey matter voxels within the caudate-putamen and thalami), NAWM and WMH. Average CBF values are given in Table 2.
To explore internal consistency of the CBF measurement within participants, we compared CBF between visit 1 and visit 2. Scatter plots suggest good agreement between the two measurements (Figure 3). Bland-Altman plots further illustrate limits of agreement within CBF data between visits 1 and 2 (see supplementary Table S1, Figure S1). Intra-class correlation coefficients confirm high test–retest reliability for CBF in total grey matter and NAWM (Table 2) and reasonable test–retest reliability for whole brain, deep grey matter and WMH (Table 2).
Test–retest reliability for CBF measurements (ml/min/100g) between visit 1 and visit 2. A) total grey matter, B) deep grey matter nuclei (caudate-putamen, thalamus), C) normal appearing white matter, D) white matter hyperintensities (WMH), E) whole brain. Each data point represents an individual participant, at study visit 1 (X-axis) relative to visit 2 (Y-axis). Dashed lines show the line of identity.
CBF values were highly correlated between tissue types (Supplementary Table S2). There were positive correlations between total grey matter and deep grey matter, NAWM or WMH (R=0.924, 0.926, 0.642, respectively; p<0.001 for all, Table S2).
Comparing female participants with males, CBF was significantly higher in women in all tissue types (see Table 3, Figure S2). The difference was 5.9 ml/min/100g in grey matter, 4.3 ml/min/100g in DGM, 4.0 ml/min/100g in NAWM and 4.0 ml/min/100g in WMH (Table 3).
ANCOVA models including sex, age and blood pressure (SBP and DBP) showed reasonable fit to the CBF data, albeit with a substantial amount of unexplained variance (R2 = 0.378 or less; p<0.001; see Table 4). Models indicated a significant decline in CBF in total grey matter, DGM and NAWM with increasing age (Table 4, Figure 4). Increasing DBP associated significantly with a decline in CBF in total grey matter, DGM and NAWM (Table 4, Figure 4). The models confirmed a significant association between female sex and higher CBF in all tissue types (Table 4).
CBF with respect to age and diastolic blood pressure (DBP). CBF (ml/min/100g) for total grey matter (panels A, C) and normal appearing white matter (B, D) are plotted with respect to participant age (A, B) and DBP (C, D). CBF data are derived from the average across visit 1 and visit 2 for each participant (N=54). Solid lines show the least-squares linear best fit to the data.
Discussion
We have presented quantitative CBF maps in a well-characterized cohort of older persons with SVD. Test–retest reliability was high for total grey matter (which is dominated by cortical grey matter) and for NAWM, and reasonable for deep grey matter and WMH (Table 2). Based on these findings we consider pCASL a potentially useful tool to follow changes in CBF in older adults with SVD.
The CBF values reported (Table 2) are low relative to textbook values, which are derived from studies in healthy control subjects [28,29,30,31,32,33,34]. CBF values for grey matter are typically in the range 40-60 ml/min/100g for healthy younger adults [28,29,30,31,32,33]. Our CBF values are similar to data from healthy older people in some previous reports [7, 32, 33] and are lower than some others [30, 35, 36]. A recent longitudinal study of 468 older people reported average CBF across the age range 50-90 [37]. The average GM CBF at age 69 is reported as 38.2 ml/min/100g for women and 34.8 ml/min/100g for men [37], both of which are comparable with our values (37.4 ml/min/100g for women, 31.5 ml/min/100g for men, Table 2).
All our participants were older patients with symptomatic SVD, which may explain why our values are in the lower range. Our data are compatible with those reported previously for SVD populations (see a recent systematic review) [14]. In people with SVD, grey matter CBF was typically 20-46 ml/min/100g [7, 10, 12, 38, 39] (our measured value was 33 ml/min/100g). In those studies where white matter CBF was recorded, values were in the range 11-25 ml/min/100g [7, 10, 12, 18, 38, 39] (our value was 13 ml/min/100g).
In common with other ASL studies, owing to the large ASL voxel size, a potential confound in our data is the inclusion of some WM tissue in voxels classified as GM, and vice versa (termed the partial volume effect). Our technique for segmentation of CBF voxels does not provide an explicit partial volume correction within a voxel. This may lead to potential underestimates of grey matter CBF and overestimates of white matter CBF. The data for whole-brain CBF (Figure 3, Table 2) are not subject to this partial volume effect.
The ASL data in this study were all acquired with a single labelling delay, using methods derived from the ASL consensus recommendations [15]. No attempt was made to correct for variation between participants in terms of transit time or haemodynamic effects. This study was formulated to ensure high signal-to-noise ratios for CBF quantification in white matter. Consequently, there was a requirement to obtain sufficient perfusion signal within white matter, and an extensive, single post-labelling delay pCASL scanning protocol was adopted [17] based on methods for perfusion scanning in dementia [15]. The decision to acquire multiple averages with a single post-labelling delay (rather than multiple PLDs, with fewer averages at each PLD) may have led to underestimation of CBF. A recent study using simulations and some grey matter data, suggests that quantification of CBF is underestimated with a single PLD [40]. Despite this, our CBF measurements in human brain grey matter are comparable with those published by others using multiple post-labelling delays [32] or for pCASL using a single PLD [37]. Future studies would benefit from the improvement in accuracy and precision of CBF measurements provided by acquisition with multiple PLDs and quantification of labelling efficiency and blood T1.
Our CBF data for total grey matter and NAWM showed high test–retest reliability (Table 2, Figure 3). It is notable that NAWM had the highest intra-class correlation coefficient (Table 2). This suggests that even in NAWM, with low absolute CBF and low signal/noise ratio, our quantitation of CBF is robust. Reliability within DGM and WMH was reasonable (ICC: 0.771, 0.780, respectively) but lower than in total grey matter or NAWM, likely reflecting the smaller number of voxels sampled. Though our inclusion criteria permit a significant degree of large artery stenosis (up to 70%), this is unlikely to confound the test–retest reliability of CBF measurement within a given participant.
High test–retest reliability has previously been reported in healthy adult controls, comparing ASL-derived CBF measurements between scanners, investigators and time points [28, 29, 31, 36, 41]. In healthy controls (age 20-67 y) the change in CBF on re-scanning at least 6 months later was ± 25% in grey matter and ± 20% in white matter [28]. A recent study using a similar protocol to ours explored test–retest reliability in older people, with scans on average 42 days apart [42]. They reported similar ICC values to ours (0.84 for whole brain, 0.77 for WM) though only a minority of their cohort had significant SVD (8 out of 45, with severe subcortical WMH load) [42]. A study of pCASL in 40 healthy adults (age 18-65) found good consistency across 4 different scanners [29]. There was reasonable test–retest reliability between ASL measurements one week apart, with limits of agreement in grey matter of 25 - 45% (expressed as a fraction of the group average) across the four MRI scanners [29]. The limits of agreement in our data for older SVD patients were similar (28% for grey matter, 33% for NAWM, Table S1). Other groups have observed high test–retest reliability, comparing pCASL measurements less than 1 hour apart (correlation coefficients 0.93-0.96 in young controls, 0.82-0.93 in older controls) [33]. These values give an indication of the intrinsic variability in the measurement system. In light of these, the correlation coefficients we derived (ICC 0.842 for total grey matter, 0.872 for NAWM, Table 2) suggest low within-subject CBF variability over a timescale of 7 days in older adults with SVD.
We found significantly higher CBF in females than in males. This accords with many previous studies [28, 32, 43,44,45,46,47,48] (though not all [34]). This difference may in part reflect differences in circulating blood composition. The physiological range for haematocrit in pre-menopausal women is 10-15% lower than in men, with the gap narrowing above age 55. Female participants in our cohort had 9% lower haematocrit relative to male participants (Table 1), which may contribute to the observed difference in CBF. Other possible explanations include haemodynamic factors [34] and higher circulating levels of female sex hormones in women [45, 46].
In our statistical models, CBF declined with age as expected from previous reports [28, 30, 36, 44, 49, 50]. A large longitudinal study (309 healthy participants, age 20-89) [30] demonstrated a 30% decline in whole-brain CBF between 20 and 80 years of age, with significant association between declining CBF and cognitive impairment in older subjects [30]. In people with overt brain vascular disease, the decline in CBF with age may be more pronounced [11, 18].
In our models DBP had significant negative association with CBF (Table 4, Figure 4). Other groups have reported that hypertensive subjects have lower CBF relative to normotensive controls [51] and that high blood pressure, especially when uncontrolled, associates with declining CBF [52]. This may reflect chronic changes in the cerebral microvasculature of older people [53].
Our study has several limitations. First, the cohort is small (N=54). Though highly significant, our findings on test–retest reliability require validation in larger cohorts. Second, our cohort has an unequal sex distribution, with only 15 (28%) female participants. Even so, sex differences emerged that were highly significant and consistent with previous literature [45, 46]. Third, we did not attempt to validate our pCASL data within subjects against a second CBF measurement modality (either SPECT, PET or DSC-MRI) [16].
In conclusion, we report quantitative CBF mapping using pCASL in a clinically relevant older population with symptomatic SVD. Test–retest data from our study and others [18, 29, 30, 33] suggest that pCASL is well tolerated and may be a technique that can contribute to clinical practice. This method may be applicable for detecting group differences (as endpoints, for comparing interventions) or within-subject changes in longitudinal studies of disease progression. We found higher CBF in women than in men, in all tissue types studied. This highlights the importance of sex-matching in trials with CBF as an endpoint. Our data suggest that pCASL sequences are a robust tool for CBF measurement in clinical and research settings.
Availability of data and material (data transparency)
The data supporting this report are available from the corresponding author upon reasonable request.
Code availability
Not applicable.
References
Wardlaw JM, Smith EE, Biessels GJ, Cordonnier C, Fazekas F, Frayne R, Lindley RI, O'Brien JT, Barkhof F, Benavente OR, Black SE, Brayne C, Breteler M, Chabriat H, DeCarli C, de Leeuw FE, Doubal F, Duering M, Fox NC, Greenberg S, Hachinski V, Kilimann I, Mok V, Oostenbrugge R, Pantoni L, Speck O, Stephan BC, Teipel S, Viswanathan A, Werring D, Chen C, Smith C, van BM, Norrving B, Gorelick PB, Dichgans M. Neuroimaging standards for research into small vessel disease and its contribution to ageing and neurodegeneration. Lancet Neurol. 2013; 12:822-38.
Kynast J, Lampe L, Luck T, Frisch S, Arelin K, Hoffmann KT, Loeffler M, Riedel-Heller SG, Villringer A, Schroeter ML. White matter hyperintensities associated with small vessel disease impair social cognition beside attention and memory. J Cereb Blood Flow Metab. 2018;38:996–1009.
Esiri MM, Wilcock GK, Morris JH. Neuropathological assessment of the lesions of significance in vascular dementia. J Neurol Neurosurg Psychiatry. 1997;63:749–53.
Lawrence AJ, Patel B, Morris RG, Mackinnon AD, Rich PM, Barrick TR, Markus HS. Mechanisms of cognitive impairment in cerebral small vessel disease: multimodal MRI results from the St George's cognition and neuroimaging in stroke (SCANS) study. PLoS One. 2013; 8:e61014.
O’Brien JT, Thomas A. Vascular dementia. Lancet. 2015;386:1698–706.
Jokinen H, Koikkalainen J, Laakso HM, Melkas S, Nieminen T, Brander A, Korvenoja A, Rueckert D, Barkhof F, Scheltens P, Schmidt R, Fazekas F, Madureira S, Verdelho A, Wallin A, Wahlund LO, Waldemar G, Chabriat H, Hennerici M, O’Brien J, Inzitari D, Lotjonen J, Pantoni L, Erkinjuntti T. Global Burden of Small Vessel Disease-Related Brain Changes on MRI Predicts Cognitive and Functional Decline. Stroke. 2020;51:170–8.
Markus HS, Lythgoe DJ, Ostegaard L, O’Sullivan M, Williams SC. Reduced cerebral blood flow in white matter in ischaemic leukoaraiosis demonstrated using quantitative exogenous contrast based perfusion MRI. J Neurol Neurosurg Psychiatry. 2000;69:48–53.
Arba F, Mair G, Carpenter T, Sakka E, Sandercock PA, Lindley RI, Inzitari D, Wardlaw JM. Cerebral White Matter Hypoperfusion Increases with Small-Vessel Disease Burden. Data From the Third International Stroke Trial. J Stroke Cerebrovasc Dis. 2017;26:1506–1513.
Schuff N, Matsumoto S, Kmiecik J, Studholme C, Du A, Ezekiel F, Miller BL, Kramer JH, Jagust WJ, Chui HC, Weiner MW. Cerebral blood flow in ischemic vascular dementia and Alzheimer’s disease, measured by arterial spin-labeling magnetic resonance imaging. Alzheimers Dement. 2009;5:454–62.
Yao H, Sadoshima S, Ibayashi S, Kuwabara Y, Ichiya Y, Fujishima M. Leukoaraiosis and dementia in hypertensive patients. Stroke. 1992;23:1673–7.
Kraut MA, Beason-Held LL, Elkins WD, Resnick SM. The impact of magnetic resonance imaging-detected white matter hyperintensities on longitudinal changes in regional cerebral blood flow. J Cereb Blood Flow Metab. 2008;28:190–7.
van Dalen JW, Moll van Charante EP, Caan MWA, Scheltens P, Majoie CBLM, Nederveen AJ, van Gool WA, Richard E. Effect of Long-Term Vascular Care on Progression of Cerebrovascular Lesions: Magnetic Resonance Imaging Substudy of the PreDIVA Trial (Prevention of Dementia by Intensive Vascular Care). Stroke. 2017; 48:1842-8.
van der Veen PH, Muller M, Vincken KL, Hendrikse J, Mali WP, van der Graaf Y, Geerlings MI, Group SS. Longitudinal relationship between cerebral small-vessel disease and cerebral blood flow: the second manifestations of arterial disease-magnetic resonance study. Stroke. 2015; 46:1233-8
Shi Y, Thrippleton MJ, Makin SD, Marshall I, Geerlings MI, de Craen AJM, van Buchem MA, Wardlaw JM. Cerebral blood flow in small vessel disease: A systematic review and meta-analysis. J Cereb Blood Flow Metab. 2016;36:1653–67.
Alsop DC, Detre JA, Golay X, Gunther M, Hendrikse J, Hernandez-Garcia L, Lu H, MacIntosh BJ, Parkes LM, Smits M, van Osch MJ, Wang DJ, Wong EC, Zaharchuk G. Recommended implementation of arterial spin-labeled perfusion MRI for clinical applications: A consensus of the ISMRM perfusion study group and the European consortium for ASL in dementia. Magn Reson Med. 2015;73:102–16.
Zhang K, Herzog H, Mauler J, Filss C, Okell TW, Kops ER, Tellmann L, Fischer T, Brocke B, Sturm W, Coenen HH, Shah NJ. Comparison of cerebral blood flow acquired by simultaneous [15O]water positron emission tomography and arterial spin labeling magnetic resonance imaging. J Cereb Blood Flow Metab. 2014;34:1373–80.
Pauls MMH, Clarke N, Trippier S, Betteridge S, Howe FA, Khan U, Kruuse C, Madigan JB, Moynihan B, Pereira AC, Rolfe D, Rostrup E, Haig CE, Barrick TR, Isaacs JD, Hainsworth AH. Perfusion by Arterial Spin labelling following Single dose Tadalafil In Small vessel disease (PASTIS): study protocol for a randomised controlled trial. Trials. 2017; 18(1):229.
Bernbaum M, Menon BK, Fick G, Smith EE, Goyal M, Frayne R, Coutts SB. Reduced blood flow in normal white matter predicts development of leukoaraiosis. J Cereb Blood Flow Metab. 2015;35:1610–5.
van Osch MJ, Teeuwisse WM, van Walderveen MA, Hendrikse J, Kies DA, van Buchem MA. Can arterial spin labeling detect white matter perfusion signal? Magn Reson Med. 2009;62:165–73.
Andersson JLR, Sotiropoulos SN. An integrated approach to correction for off-resonance effects and subject movement in diffusion MR imaging. Neuroimage. 2016;125:1063–78.
Jenkinson M, Smith S. A global optimisation method for robust affine registration of brain images. Med Image Anal. 2001;5:143–56.
Woolrich MW, Jbabdi S, Patenaude B, Chappell M, Makni S, Behrens T, Beckmann C, Jenkinson M, Smith SM. Bayesian analysis of neuroimaging data in FSL. Neuroimage. 2009;45:S173-86.
Lambert C, Sam Narean J, Benjamin P, Zeestraten E, Barrick TR, Markus HS. Characterising the grey matter correlates of leukoaraiosis in cerebral small vessel disease. Neuroimage Clin. 2015;9:194–205.
Spilling CA, Jones PW, Dodd JW, Barrick TR. White matter lesions characterise brain involvement in moderate to severe chronic obstructive pulmonary disease, but cerebral atrophy does not. BMC Pulm Med. 2017;17:92.
Smith SM. Fast robust automated brain extraction. Hum Brain Mapp. 2002;17:143–55.
Greve DN, Fischl B. Accurate and robust brain image alignment using boundary-based registration. Neuroimage. 2009;48:63–72.
Zhang Y, Brady M, Smith S. Segmentation of brain MR images through a hidden Markov random field model and the expectation-maximization algorithm. IEEE Trans Med Imaging. 2001;20:45–57.
Parkes LM, Rashid W, Chard DT, Tofts PS. Normal cerebral perfusion measurements using arterial spin labeling: reproducibility, stability, and age and gender effects. Magn Reson Med. 2004;51:736–43.
Almeida JRC, Greenberg T, Lu H, Chase HW, Fournier JC, Cooper CM, Deckersbach T, Adams P, Carmody T, Fava M, Kurian B, McGrath PJ, McInnis MG, Oquendo MA, Parsey R, Weissman M, Trivedi M, Phillips ML. Test-retest reliability of cerebral blood flow in healthy individuals using arterial spin labeling: Findings from the EMBARC study. Magn Reson Imaging. 2018;45:26–33.
De Vis JB, Peng SL, Chen X, Li Y, Liu P, Sur S, Rodrigue KM, Park DC, Lu H. Arterial-spin-labeling (ASL) perfusion MRI predicts cognitive function in elderly individuals: A 4-year longitudinal study. J Magn Reson Imaging. 2018;48:449–58.
Henriksen OM, Larsson HB, Hansen AE, Gruner JM, Law I, Rostrup E. Estimation of intersubject variability of cerebral blood flow measurements using MRI and positron emission tomography. J Magn Reson Imaging. 2012;35:1290–9.
Liu Y, Zhu X, Feinberg D, Guenther M, Gregori J, Weiner MW, Schuff N. Arterial spin labeling MRI study of age and gender effects on brain perfusion hemodynamics. Magn Reson Med. 2012;68:912–22.
Xu G, Rowley HA, Wu G, Alsop DC, Shankaranarayanan A, Dowling M, Christian BT, Oakes TR, Johnson SC. Reliability and precision of pseudo-continuous arterial spin labeling perfusion MRI on 3.0 T and comparison with 15O-water PET in elderly subjects at risk for Alzheimer's disease. NMR Biomed. 2010; 23:286-93.
Smith LA, Melbourne A, Owen D, Cardoso MJ, Sudre CH, Tillin T, Sokolska M, Atkinson D, Chaturvedi N, Ourselin S, Hughes AD, Barkhof F, Jager HR. Cortical cerebral blood flow in ageing: effects of haematocrit, sex, ethnicity and diabetes. Eur Radiol. 2019.
Henriksen OM, Jensen LT, Krabbe K, Guldberg P, Teerlink T, Rostrup E. Resting brain perfusion and selected vascular risk factors in healthy elderly subjects. PLoS One. 2014; 9:e97363.
Jiang L, Kim M, Chodkowski B, Donahue MJ, Pekar JJ, Van Zijl PC, Albert M. Reliability and reproducibility of perfusion MRI in cognitively normal subjects. Magn Reson Imaging. 2010;28:1283–9.
Leidhin CN, McMorrow J, Carey D, Newman L, Williamson W, Fagan AJ, Chappell MA, Kenny RA, Meaney JF, Knight SP. Age-related normative changes in cerebral perfusion: Data from The Irish Longitudinal Study on Ageing (TILDA). Neuroimage. 2021; 229:117741.
Huynh TJ, Murphy B, Pettersen JA, Tu H, Sahlas DJ, Zhang L, Symons SP, Black S, Lee TY, Aviv RI. CT perfusion quantification of small-vessel ischemic severity. AJNR Am J Neuroradiol. 2008;29:1831–6.
Oishi M, Mochizuki Y. Differences in regional cerebral blood flow in two types of leuko-araiosis. J Neurol Sci. 1999;164:129–33.
Bladt P, van Osch MJP, Clement P, Achten E, Sijbers J, den Dekker AJ. Supporting measurements or more averages? How to quantify cerebral blood flow most reliably in 5 minutes by arterial spin labeling. Magn Reson Med. 2020;84:2523–36.
Yen YF, Field AS, Martin EM, Ari N, Burdette JH, Moody DM, Takahashi AM. Test-retest reproducibility of quantitative CBF measurements using FAIR perfusion MRI and acetazolamide challenge. Magn Reson Med. 2002;47:921–8.
Jann K, Shao X, Ma SJ, Cen SY, D'Orazio L, Barisano G, Yan L, Casey M, Lamas J, Staffaroni AM, Kramer JH, Ringman JM, Wang DJJ. Evaluation of Cerebral Blood Flow Measured by 3D PCASL as Biomarker of Vascular Cognitive Impairment and Dementia (VCID) in a Cohort of Elderly Latinx Subjects at Risk of Small Vessel Disease. Front Neurosci. 2021; 15:627627.
Amen DG, Trujillo M, Keator D, Taylor DV, Willeumier K, Meysami S, Raji CA. Gender-Based Cerebral Perfusion Differences in 46,034 Functional Neuroimaging Scans. J Alzheimers Dis. 2017;60:605–14.
Jones K, Johnson KA, Becker JA, Spiers PA, Albert MS, Holman BL. Use of singular value decomposition to characterize age and gender differences in SPECT cerebral perfusion. J Nucl Med. 1998;39:965–73.
Rodriguez G, Warkentin S, Risberg J, Rosadini G. Sex differences in regional cerebral blood flow. J Cereb Blood Flow Metab. 1988;8:783–9.
Esposito G, Van Horn JD, Weinberger DR, Berman KF. Gender differences in cerebral blood flow as a function of cognitive state with PET. J Nucl Med. 1996;37:559–64.
Li ZJ, Matsuda H, Asada T, Ohnishi T, Kanetaka H, Imabayashi E, Tanaka F. Gender difference in brain perfusion 99mTc-ECD SPECT in aged healthy volunteers after correction for partial volume effects. Nucl Med Commun. 2004;25:999–1005.
Shin W, Horowitz S, Ragin A, Chen Y, Walker M, Carroll TJ. Quantitative cerebral perfusion using dynamic susceptibility contrast MRI: evaluation of reproducibility and age- and gender-dependence with fully automatic image postprocessing algorithm. Magn Reson Med. 2007;58:1232–41.
Duara R, Grady C, Haxby J, Ingvar D, Sokoloff L, Margolin RA, Manning RG, Cutler NR, Rapoport SI. Human brain glucose utilization and cognitive function in relation to age. Ann Neurol. 1984;16:703–13.
Chen JJ, Rosas HD, Salat DH. Age-associated reductions in cerebral blood flow are independent from regional atrophy. Neuroimage. 2011;55:468–78.
Dai W, Lopez OL, Carmichael OT, Becker JT, Kuller LH, Gach HM. Abnormal regional cerebral blood flow in cognitively normal elderly subjects with hypertension. Stroke. 2008;39:349–54.
Muller M, van der Graaf Y, Visseren FL, Mali WP, Geerlings MI, Group SS. Hypertension and longitudinal changes in cerebral blood flow: the SMART-MR study. Ann Neurol. 2012; 71:825-33
Toth P, Tarantini S, Csiszar A, Ungvari Z. Functional vascular contributions to cognitive impairment and dementia: mechanisms and consequences of cerebral autoregulatory dysfunction, endothelial impairment, and neurovascular uncoupling in aging. Am J Physiol Heart Circ Physiol. 2017;312:H1–20.
Acknowledgements
We are very grateful to our colleagues in the Department of Neuroradiology, St George’s Hospital, the St George’s Stroke service, the South London CRN Stroke Research Network and St George’s Clinical Research Facility. We thank Dr Robin Young of the Robertson Centre for Biostatistics, University of Glasgow, for helpful comments on statistics. We dedicate this paper to our friend and colleague Debbie Rolfe (1967-2019).
Funding
The PASTIS trial was joint-funded by UK Alzheimer’s Society and Alzheimer’s Drug Discovery Foundation (Grant Ref. 20140901). The funding sources had no involvement in study design; in the collection, analysis and interpretation of data; in the writing of the report; and in the decision to submit the article for publication. The sponsor was St George’s University of London (contact: sponsor@sgul.ac.uk). Mr Anan Shtaya was supported by a NIHR Clinical Lectureship [CL-2015-16-001].
Author information
Authors and Affiliations
Contributions
MMHP, CK, JBM, BM, ER, JDI, TRB, AHH made a substantial contribution to the concept and design. All authors made substantial contribution to the acquisition or analysis or interpretation of data. LRB drafted the article. All authors revised the article critically for intellectual content. All authors approved the version to be published.
Corresponding author
Ethics declarations
Conflicts of interest/Competing interests
LRB and MMHP were employed as part of the PASTIS trial, JDI was Principal Investigator, and AHH was Chief Investigator. CK is a PI on clinical trials with Bristol-Myers-Squibb and Bayer and has received funding from Novo Nordisk, Bayer and Bristol-Myers-Squibb, all not relevant to the present trial. JDI has been a PI on clinical trials funded by Roche, Merck and Lupin Pharmaceuticals and has received funds from Biogen and Roche, none relevant to the present trial. AHH has received honoraria from Eli-Lilly and from NIA, he chairs the Vascular Cognitive Disorders PIA within ISTAART, and he leads MRC-Dementias Platform UK Vascular Experimental Medicine group. All other authors report no relevant disclosures.
Ethics approval for Research involving Human Participants
The trial received ethical approval from the UK National Research Ethics Service (London-Brent research ethics committee, REC reference: 15/LO/0714).
Ethics approval for Research involving Animals
Not applicable.
Consent to participate
Written informed consent was obtained from all participants or their next of kin.
Consent for publication
This paper contains no images of participants and no other identifiable participant data.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Lauren R Binnie and Mathilde M. H. Pauls contributed equally
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Binnie, L.R., Pauls, M.M.H., Benjamin, P. et al. Test–retest reliability of arterial spin labelling for cerebral blood flow in older adults with small vessel disease. Transl. Stroke Res. 13, 583–594 (2022). https://doi.org/10.1007/s12975-021-00983-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12975-021-00983-5