Abstract
Background
Polygenic scores (PGS) have been developed for cancer risk-estimation and show potential as tools to prompt earlier referral for high-risk individuals and aid risk-stratification within cancer screening programmes. This review explores the potential for using PGS to identify individuals at risk of the most common cancers seen in primary care.
Methods
Two electronic databases were searched up until November 2023 to identify quantitative, qualitative, and mixed methods studies that reported on the acceptability and clinical impact of using PGS to identify individuals at highest risk of breast, prostate, colorectal and lung cancer in primary care. The Mixed Methods Appraisal Tool (MMAT) was used to assess the quality of included studies and a narrative synthesis was used to analyse data.
Results
A total of 190 papers were identified, 18 of which were eligible for inclusion. A cancer risk-assessment tool incorporating PGS was acceptable to the general practice population and their healthcare providers but major challenges to implementation were identified, including lack of evidence for PGS in non-European ancestry and a need for healthcare provider education in genomic medicine. A PGS cancer risk-assessment had relatively limited impact on psychosocial outcomes and health behaviours. However, for prostate cancer, potential applications for its use in primary care were shown.
Conclusions
Cancer risk assessment incorporating PGS in primary care is acceptable to patients and healthcare providers but there is a paucity of research exploring clinical impact. Few studies were identified, and more research is required before clinical implementation of PGS can be recommended.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Cancer is the second leading cause of death globally, accounting for 10 million deaths in 2020 (Ritchie et al. 2018). The commonest cancers, lung (LC), breast (BC), prostate (PC) and colorectal (CRC), together account for nearly half of all cancers. By 2040, worldwide cancer rates are projected to increase by 55% with the greatest increase in cancer burden being seen in low and middle income countries (LMIC) (International Agency for Research on Cancer 2023). 30-50% of cancers are thought to be preventable (World Health Organisation 2022) and this highlights the role for cancer prevention and screening.
UK data from 2015-16, showed that most cases of cancer (61%) were diagnosed by GP referral, 19% as hospital emergencies and only 6% detected by national screening programmes (Cancer Research UK 2018). Primary care, therefore, plays a crucial role in the timely diagnosis of cancer. However, patients often present with non-specific symptoms and complex symptomatology. There is a need to develop methods to support primary care to detect cancer early.
Polygenic scores (PGS) reflect an individual’s estimated genetic predisposition to complex common diseases such as cancer. PGS are calculated by the aggregation of hundreds-to-thousands of single nucleotide polymorphisms (SNPs) with weak-to-moderate cancer associations which are weighted according to effect size (Uffelmann et al. 2021). The overall scores for an individual are compared to the general population. PGS have therefore been identified as potential tools to make more precise and personalised predictions about cancer risk (Pharoah et al. 2002). Applications include the development of stratified health screening programmes and enhanced cancer risk-prediction tools (Huntley et al. 2023).
PGS have been developed for many common complex diseases, but they show wide variability in application and reporting. This poses a barrier to implementation in clinical care. At the time of writing, a search of the PGS Catalog (PGS Catalog 2023) reveals 120, 66, 56 and 22 PGS associated with BC, PC, CRC and LC respectively. PGS Reporting Standards have been developed (Wand et al. 2021) and the “Evaluation of Polygenic Score Applications” PHG Foundation report (Moorthie et al. 2023) has been written to aid researchers and commercial companies developing PGS to overcome some of these barriers.
As genome sequencing becomes more affordable, genomic testing in primary care will become increasingly accessible. However it is important that before PGS are incorporated into clinical practice, their acceptability and clinical utility are demonstrated (Moorthie et al. 2021). In the context of cancer risk-estimation, PGS have potential to add value to existing cancer risk models. However, to be clinically useful, superiority needs to be demonstrated against existing practice. The benefits of their use need to outweigh harms, the setting of their use defined, thresholds for intervention set and cost-effectiveness demonstrated (Usher-Smith et al. 2015)
Given the potential of PGS in cancer risk prediction, this review aims to establish the current evidence for the acceptability and clinical impact of available PGS to identify individuals at highest risk of BC, PC, CRC or LC in the primary care setting.
Methods
The Preferred Reporting Items for Systematic Reviews and Meta-analysis (PRISMA) 2020 systematic review guidelines informed the development and reporting of this study (Page et al. 2021).
Search strategy
Medline and EMBASE electronic databases were searched from 2000 to November 2023 using a combination of title and abstract search terms and MeSH terms incorporating breast, colorectal, lung or prostate cancer, polygenic score, primary care and outcome. The full search strategy is presented in supporting information, S1.
Study selection
Inclusion criteria
-
Peer-reviewed primary research (English language)
-
Quantitative, qualitative, or primary mixed methods studies
-
BC, PC, CRC or LC risk-assessments
-
Primary care population or primary healthcare providers
-
Intervention focused on PGS or SNP cancer risk-alleles
-
Outcomes included clinical feasibility, utility and acceptability of the genetic risk tool
Exclusion criteria
-
High-risk individuals (including those with monogenic cancer risk susceptibility genes, recurrent or relapsing cancer or current cancer diagnosis)
-
Population-based health screening cohorts without specific relevance to primary care
-
Conference abstracts, letters, correspondence, editorials, comments or reviews
One reviewer (FD) conducted the database searches, removed duplicates and reviewed title and abstracts of all citations. A second reviewer (JT) independently screened a random sample of all the citations (10%) for inclusion and discrepancies were resolved in a consensus meeting. At least two reviewers (FD, JU, LT, JT) reviewed all citations eligible for full text review. Studies excluded by all reviewers were considered ineligible and discrepancies were again resolved at consensus. Two reviewers (FD & JU) agreed the final citations for inclusion. Further citations were identified from alternative source searches, such as through reference lists, Google and peer recommendation.
Quality assessment
Two reviewers (FD and LT) independently completed quality assessment for all eligible papers using the Mixed Methods Appraisal Tool (MMAT) (Hong et al. 2018), which is appropriate for assessing the quality of qualitative, quantitative and mixed-methods research. No studies were excluded based on quality. Discrepancies, of which there were only a few, were discussed at consensus meetings. The final quality assessment is presented in supplementary information, S2.
Data extraction and synthesis
Data extraction was conducted by FD for all eligible studies using a standardized form developed and piloted with a small number of studies. It included author, year, country, cancer type(s), primary aim, sample size, demographic characteristics, study design, recruitment, type of PGS, PGS uptake, data collection and outcomes and is included in supplementary information, S3. The same terminology used in the original papers in relation to ancestry, race and ethnicity was also used in the results discussion.
Outcomes were assessed according to categories shown in Table 1.
Results
Study selection
After removal of duplicates, the search generated 190 citations. Of these, 145 citations were excluded after title and abstract review. The commonest reason for exclusion being that the intervention did not include PGS. A further 32 citations were excluded after full text screening. An additional five papers were identified using alternative search methods, resulting in a total of 18 papers eligible for inclusion (Leventhal et al. 2013; Graves et al. 2013; Nusbaum et al. 2013; Conran et al. 2021; Saya et al. 2022; Saya et al. 2020; Butrick et al. 2014; Kirkegaard et al. 2018; Fredsoe et al. 2020a, 2020b; Green et al. 2022; Benafif et al. 2022; Kerman et al. 2023; Ayoub et al. 2023; Archer et al. 2020, 2023; Myers et al. 2015; Weinberg et al. 2014) (Fig. 1).
Study characteristics
Variation was seen in study design and reported outcome measures, and this precluded meta-analysis or quantitative synthesis. The study characteristics and authors’ primary conclusions are summarized in Tables 2 and 3. Most studies explored the attitudes of patients to PGS testing (n=14, 78%), others explored HCP’s attitudes (n=4, 22%). Most studies considered PGS in relation to one cancer type (BC, PC or CRC). No eligible studies were identified for LC. Only four high-income countries were represented: eight USA (Leventhal et al. 2013; Graves et al. 2013; Nusbaum et al. 2013; Butrick et al. 2014; Conran et al. 2021; Kerman et al. 2023; Weinberg et al. 2014; Myers et al. 2015), five UK (Archer et al. 2020, 2023; Ayoub et al. 2023; Green et al. 2022; Benafif et al. 2022), three Denmark (Fredsoe et al. 2020a, 2020b; Kirkegaard et al. 2013) and two Australia (Saya et al. 2020, 2022). The majority of studies used an observational quantitative design (Graves et al. 2013; Saya et al. 2020; Fredsoe et al. 2020a, 2020b; Green et al. 2022; Benafif et al. 2022; Ayoub et al. 2023; Kerman et al. 2023; Conran et al. 2021; Weinberg et al. 2014), four used qualitative methods (Archer et al. 2020, 2023; Nusbaum et al. 2013; Kirkegaard et al. 2018) and four mixed methods (Leventhal et al. 2013; Saya et al. 2022; Butrick et al. 2014; Myers et al. 2015). Data were collected using demographic questionnaires, national surveys, baseline questionnaires, focus groups and semi-structured interviews. Three studies used case studies and simulated patients for HCP interpretation (Archer et al. 2020, 2023; Kerman et al. 2023).
Data analysis
Patient specific outcomes
PGS characteristics
Variations were seen in how PGS was calculated and how cancer risk was categorised, Table 4. Some studies used comparative measures of association (Conran et al. 2021; Green et al. 2022; Benafif et al. 2022; Weinberg et al. 2014; Myers et al. 2015) whilst others used absolute risk thresholds (Graves et al. 2013; Nusbaum et al. 2013; Saya et al. 2020; Butrick et al. 2014; Kirkegaard et al. 2018; Fredsoe et al. 2020a, 2020b; Archer et al. 2020, 2023). The number of SNPs used to calculate the risk score ranged from two to 313. Some risk scores were integrated with other health information. The CanRisk tool (CanRisk v2.3.5 2023) was the most comprehensive, integrating PGS with rare pathogenic risk variants and a wide range of non-genomic risk factors for breast cancer (Archer et al. 2020, 2023). The type of cancer, country of study and study type may have influenced these differences. Saya et al. (Australia) (Saya et al. 2020) converted their PGS into an absolute 10y risk in alignment with Australian screening guidance. In the UK, Green et al. (Green et al. 2022) categorized five risk thresholds between 1-5% for PC, in line with NICE guidance for suspected cancer referral where a referral is recommended for PPV ≥ 3% risk of cancer (National Institute for Health and CareExcellence 2015). In contrast the BARCODE1 pilot study (UK) defined high-risk for PC as individuals in the top 90th centile conferring a 2.7 relative risk (Benafif et al. 2022). The studies by Fredsoe et al. (Denmark) (Fredsoe et al. 2020a, 2020b) expressed lifetime PC risk as either low or high. Finally, Archer et al. (UK) calculated lifetime or 10y risk of BC with thresholds defined as near population, moderate or high-risk (aligned with NICE guidance) (Archer et al. 2023). PGS variation meant that limited direct comparisons between studies could be made and a quantitative synthesis was not possible.
PGS test uptake
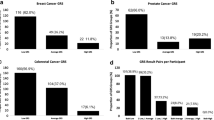
Five publications considered the uptake of PGS testing (calculated as a percentage of participants who were offered the test and agreed to take it) for PC and CRC (Graves et al. 2013; Saya et al. 2020; Butrick et al. 2014; Benafif et al. 2022; Saya et al. 2022), Table 5. In all studies, test uptake rates were high, 84-96%. Despite race-specific limitations of testing in minority groups being explicitly mentioned within the study documentation, Butrick et al reported high uptake in a subgroup of minority patients, 95% (Butrick et al. 2014).
Motivations for testing
Four studies reported on motivations for testing, Table 6. All studies reported that personal information gathering (Leventhal et al. 2013; Nusbaum et al. 2013; Saya et al. 2022; Kirkegaard et al. 2018) was a main reason for doing the test. Other reasons included gathering information for family members (Leventhal et al. 2013; Nusbaum et al. 2013), altruistic reasons (Leventhal et al. 2013; Nusbaum et al. 2013; Saya et al. 2022), on HCP’s recommendation (Leventhal et al. 2013) and due to personal risk factors or family history CRC (Leventhal et al. 2013; Saya et al. 2022). In the study by Kirkegaard et al. (Kirkegaard et al. 2013) participants were interested in knowing their PC risk but not specifically the genetic lifetime risk (Kirkegaard et al. 2018). Leventhal et al. reported that a subgroup of cancer survivors (n=5) were less interested in the test than non-cancer survivors (Leventhal et al. 2013).
Barriers to testing
Three studies explored patient barriers to having a PGS test (Leventhal et al. 2013; Nusbaum et al. 2013; Saya et al. 2022), Table 7. Privacy and insurance concerns and the low perceived value of risk information were common to all (Leventhal et al. 2013; Nusbaum et al. 2013; Saya et al. 2022). Others included doubt that risk assessment would impact care (Leventhal et al. 2013), worry (Nusbaum et al. 2013; Saya et al. 2022) and a preference for knowing current risk rather than potential future risk (Saya et al. 2022).
Patient understanding of the test / informed decision making
Four studies explored participant’s understanding of the genetic test (Fredsoe et al. 2020a; Saya et al. 2022; Saya et al. 2020; Nusbaum et al. 2013), Table 8. Notably, nearly half the participants in the study by Fredsoe et al. did not recall having a genetic test and significantly fewer high-risk individuals (75%) correctly identified their risk of PC compared to average-risk individuals (98.5%) (p<0.001) (Fredsoe et al. 2020a). The other three studies reported that participants generally understood the test (Nusbaum et al. 2013; Saya et al. 2022; Saya et al. 2020) with one reporting adequate knowledge in 83% participants (Saya et al. 2020). Saya et al. (Saya et al. 2022) reported that adequate knowledge was significantly higher in women, higher educated and primarily English speakers. Those participants opting for testing and higher educated individuals scored significantly higher on informed choice measures (Saya et al. 2022).
Psychosocial impact of the test
Six studies reported on the impact of having a PGS test on cancer worry and cancer related anxiety, reporting mixed results, Table 9. Three studies reported increased worry and distress (Myers et al. 2015; Conran et al. 2021; Saya et al. 2020). Myers et al. reported that for individuals at elevated risk of CRC, irrespective of race, participants had significantly increased worry and distress (Myers et al. 2015). Similarly, Conran et al. reported a significant increase in worry about risk of developing CRC in women and younger patients. Furthermore, those with a least one high PGS were significantly more worried about their risk of developing cancer (p=0.01) (Conran et al. 2021). Saya et al. reported less worry in patients with average-risk scores and significant increased cancer specific anxiety in patients with moderate-risk scores (p=0.01) (Saya et al. 2020). The qualitative study by Leventhal et al. noted a range of emotional responses (Leventhal et al. 2013). In contrast, the studies by Nusbaum et al. and Graves et al. (Graves et al. 2013; Nusbaum et al. 2013) reported no or low levels of distress or worry and no correlation between risk score and perceived risk (Graves et al. 2013). Nusbaum et al. reported that established risk factors such as personal or family history of cancer had a greater impact on perceived CRC risk (Nusbaum et al. 2013).
Results delivery and sharing of information
Only two studies considered the method of results delivery (Leventhal et al. 2013; Nusbaum et al. 2013). Both indicated a preference for in-person delivery of results information. Participants in two studies (Nusbaum et al. 2013; Graves et al. 2013) frequently indicated intention to share results with family members but less frequently with HCPs.
Impact on health behaviours (future screening, PSA testing and lifestyle intent)
Eight studies reported on participant cancer screening intent, Table 10. Although four studies reported increased intentions for cancer screening (Conran et al. 2021; Graves et al. 2013; Nusbaum et al. 2013; Saya et al. 2020), objective measures of increased screening post-test were not seen in three studies (Graves et al. 2013; Weinberg et al. 2014; Saya et al. 2020). In contrast, one study by Benafif et al. demonstrated high prostate cancer screening uptake rates of 72% for individuals in the top 10% of PGS distribution (Benafif et al. 2022). Qualitative studies reported mixed views on screening intent (Leventhal et al. 2013) and differential views towards screening post-test between white versus non-white participants (Myers et al. 2015). Only two studies reported on lifestyle behaviour change, both reporting positive intent (Graves et al. 2013; Nusbaum et al. 2013).
Two studies reported on the number of repeat PSA tests done 2y after a PGS risk assessment for PC (Fredsoe et al. 2020a, 2020b). Overall there were no differences in PSA testing between the intervention and control practices even after adjusting for confounders (Fredsoe et al. 2020b). However subgroup analysis of the intervention practices showed that men with an average genetic risk had a significantly lower chance of having a future PSA test (OR = 0.62, p<0.01) and men with high genetic risk a higher chance of having a future PSA test (OR 8.94, p<0.01) when compared to men who did not have the genetic risk assessment (Fredsoe et al. 2020b). Furthermore a high genetic risk score was found to be the most significant predictor of a repeat PSA test (HR = 5.99, p=0.007) (Fredsoe et al. 2020a) and that higher percentages of elevated PSA test were found in men with high genetic risk compared with men who had no genetic test (OR = 3.31, p<0.01) (Fredsoe et al. 2020b).
Cancer diagnosis
Two publications studied the relationship between PGS and PC diagnosis (Green et al. 2022; Benafif et al. 2022). Green et al. (retrospective study) determined incidence of PC within a 2y period for men presenting to their GP with lower urinary tract symptoms. They reported an association between the integrated PGS/age risk-score and incidence of PC with higher scores being seen in cases compared to controls (p=3.5e-30) (Green et al. 2022). They demonstrated that a PGS/age risk-score could be used to identify men at lowest risk of PC (<1% incidence of PC).
Benafif et al. (prospective study) used PGS to identify men at highest risk of PC. Men in the top 10% PGS score were offered screening with MRI and prostate biopsy. 8.75% of men were identified in this high-risk group. The PC diagnosis rate in this group was 26.9%, all 7 patients were placed under active surveillance programmes (Benafif et al. 2022).
Primary healthcare providers attitudes and opinions
Four studies explored primary HCP’s attitudes to PGS for cancer risk assessment (Kerman et al. 2023; Ayoub et al. 2023; Archer et al. 2020, 2023), Table 11. All three studies which explored PGS in the context of a personalized BC risk assessment, demonstrated that HCPs attitudes were positive towards the concept but a need for education and training in genomics was highlighted (Archer et al. 2020, 2023; Ayoub et al. 2023). Ayoub et al. reported that nearly half (49%) were not familiar with PGS and 23% reported that GPs were ‘not confident’ in explaining polygenic inheritance. A range of preferences for HCP learning formats were identified (Ayoub et al. 2023).
Archer et al. (Archer et al. 2020, 2023) reported on an integrated breast cancer risk assessment tool that incorporated PGS but there was little focus on the PGS component itself. Although participants reported confidence in communication skills regarding multi-factorial risk, they were concerned about their limited genetics knowledge and highlighted a need for additional training on interpreting genetic risk (Archer et al. 2020). Other more generalised concerns included the time needed for assessment, the importance of integrating the tool within clinical work flow and electronic medical record systems and the importance of providing clinical management information as part of the risk assessment (Archer et al. 2023).
Kerman et al. (Kerman et al. 2023) explored PC risk and reported that the majority of HCPs reported no additional genetics training beyond medical school.
Two studies considered use of a PGS tool in the context of ethnicity (Ayoub et al. 2023; Kerman et al. 2023). The study by Kerman et al. explored the influence of race on screening recommendations for PC and compared this to the patient’s risk using a PGS (Kerman et al. 2023). It was shown that physicians were more likely to recommend PC screening to black than white patients even for the same PGS (OR =1.58, p=0.025). There were no significant associations between the HCP’s self-reported race and ethnicity on PC screening recommendations. These results suggest that PGS did not remove the influence of race on clinical decision making.
In the study by Ayoub et al. (Ayoub et al. 2023) HCPs raised concerns that the use of a PGS tool that was validated in white European women would widen health inequality (Ayoub et al. 2023).
Discussion
Key findings
This review shows that the use of PGS to determine risk of common cancers in general practice is acceptable to patients and HCPs. High uptake rates of PGS testing amongst patients were seen despite concerns about privacy, insurance and limitations of testing in different ancestry groups. Patient acceptability is further supported by the relatively limited impact of PGS testing that was demonstrated on psychosocial outcomes. HCPs were also positive about the potential for PGS in cancer risk estimation. However, they highlighted a need for education and training in genomics and polygenic inheritance and raised concerns that using a PGS test with race-specific limitations could widen health inequality.
The impact on screening and health behaviour was limited. Although self-reported screening intentions were stated in several studies, this was not seen in objective outcomes. The impact of PGS testing for PC risk was explored in 3 studies which highlighted potential applications in primary care, including impact on future PSA testing rates (Fredsoe et al. 2020a, 2020b) and the potential to risk stratify individuals at risk of PC. (Green et al. 2022; Benafif et al. 2022).
Strengths and limitations
A comprehensive search strategy with robust and clear definitions for key terminology was used including a range of terms for PGS. However, publications that used alternative terms for genetic risk, such as ‘CanRisk tool’ may have been missed. The search was limited to the top four most common cancers (BC, PC, CRC and LC). Excluding other cancer types appears justified given that no eligible studies were identified for LC. The adoption of a mixed methods approach enabled a comprehensive review of the current evidence available.
This review has several limitations. Firstly, there was a high degree of overlap seen in the eligible studies with 13 being published by five common research groups or patient cohorts (Butrick et al. 2014; Nusbaum et al. 2013; Graves et al. 2013; Leventhal et al. 2013; Weinberg et al. 2014; Myers et al. 2015; Kirkegaard et al. 2018; Fredsoe et al. 2020a, 2020b; Fredsoe et al. 2020a, 2020b; Saya et al. 2022; Saya et al. 2020; Archer et al. 2020, 2023), introducing potential source bias. Despite this, all studies were included as each reported on different qualitative and quantitative outcomes. Where indicated, overlap between studies have been highlighted within the tables, as seen for PGS uptake.
Secondly, all included studies were conducted in high-income countries and recruitment was often limited to Caucasian individuals (due to disparity in PGS accuracy across ancestry groups). This limits the transferability of these findings to LMICs and different ancestry groups. The lack of diversity in genetic studies is well recognised (Fatumo et al. 2022) and different methods are being developed to construct PGS from distinct ancestry groups to improve prediction and accuracy (Weissbrod et al. 2022; Ruan et al. 2022; Cavazos and Witte 2021). More balanced multi-ancestry genomic data sets are also being developed and include the ‘All of Us Program (USA)’ (National Institutes of Health (NIH) 2024), the ‘Million Veteran Program (USA)’ (US Department of Veterans Affairs (VA) 2024) and ‘Our Future Health (UK)’ (+ Our Future Health 2022).
Thirdly, the heterogeneity of the included studies meant we were unable to perform a quantitative synthesis or meta-analysis. Heterogeneity was seen across the study design and different cohorts as well as variations in the different uses of PGS and different outcomes. Differences were seen in the number of SNPs incorporated in the PGS and the way the PGS risk information was delivered to patients in some studies. This may have impacted individuals understanding and risk perception. It has been proposed that the communication of risk information is standardised to optimize interpretation and understanding (Brockman et al. 2021).
Comparison with existing literature
Existing studies exploring PGS for risk-stratification in population-based cancer screening programmes demonstrate patient acceptability and are consistent with the findings in our study (Taylor et al. 2023; Laza-Vásquez et al. 2022). Overall public optimism was reported for risk-stratified screening programmes for breast, ovarian, kidney, prostate and all cancer types in a systematic review (Taylor et al. 2023) with the majority of studies related to risk-stratified breast and ovarian cancer screening. Similar acceptability was seen in the ‘DECIDO’ study for use of PGS to risk-stratify women for breast cancer screening (Laza-Vásquez et al. 2022). Although more research is needed in this area, the results from two large studies ‘WISDOM’ (USA) (Esserman 2017) and ‘MyPeBS’ hope to provide further evidence for acceptability for breast cancer risk-stratification incorporating PGS (MyPeBS 2022).
The absence of impact of PGS on screening and lifestyle behaviours seen in our study has also been seen in other common complex diseases including cardiovascular disease (Hollands et al. 2016; Silarova et al. 2019). Furthermore a review of systematic reviews found no evidence that presenting highly personalised risk information to individuals alone produced strong effects on lifestyle health-related behaviours (French et al. 2017).
The results presented here regarding HCPs attitudes to genetic cancer risk assessment are also consistent with existing literature. For example, a systematic review which explored clinicians’ attitudes to genetic cancer risk-assessment using family history tools in primary care (Laforest et al. 2019) showed that overall, general practitioners viewed their potential role in genetic risk assessment as important but found assessing risk difficult due to lack of knowledge and confidence in interpreting and explaining genetic test results. A further systematic review by Hamilton et al (Hamilton et al. 2017) explored primary HCPs knowledge and attitudes regarding genetic tests for individuals at risk of hereditary cancer syndromes and direct-to-consumer tests in relation to BC, PC and CRC. They highlighted a need for educational interventions due to low levels of genetic knowledge. Vassy et al (Vassy et al. 2023b) explored perceived benefits and barriers to implementing precision preventative care. Primary HCPs generally endorsed the use of PGS in preventative medicine but identified concerns regarding lack of understanding, potential patient anxiety and lack of evidence for use in non-European populations. In another study, similar barriers were seen in implementing risk-prediction tools for cancer in general (including those without genomic factors) (Usher-Smith et al. 2015).
Implications for policy, practice (GP) and research
This review clearly demonstrates enthusiasm amongst HCPs and patients for PGS in cancer risk prediction, but further research and debate is needed before clinical application in general practice can be widely recommended.
The first priority is to develop and validate PGS in diverse populations. Although efforts are ongoing to improve performance of PGS across ancestry groups, the feasibility of using a population structure-adjusted PGS for multiple common diseases, including BC, PC and CRC, has recently been demonstrated (Hao et al. 2022). A further study “GenoVA study” (Vassy et al. 2023a) aims to use this tool to show how PGS might be equitably integrated into the busy primary care context whilst using a RCT trial design to compare the impact of PGS implementation versus usual care on patient outcomes. Risk assessment tools also need to be acceptable to hard-to-reach groups and for use in LMICs, thus ensuring equity of access.
Although the costs of genetic testing are falling and PGS cancer risk assessment tools have been shown to be acceptable to patients and HCPs in high income countries (HICs), these tools are unlikely to yet be a priority for LMICs (Pramesh et al. 2022). Very few LMIC have successful, sustainable cancer screening programs (Sullivan et al. 2021) with several barriers to implementation including sociocultural acceptance (Pak et al. 2021).
Currently most evidence exploring the potential of PGS for cancer risk-estimation is related to BC and PC. For BC risk-stratification using PGS, many large scale studies are underway including ‘MyPeBS’ (Europe) (MyPeBS 2022), ‘WISDOM’ (USA) (Esserman 2017) and ‘PROCAS’ (UK) (McWilliams et al. 2022). For PC the ‘ProCaRis’ study (University of Aarhus 2022) aims to determine if PGS can be used to better identify men at low risk of PC and reduce unsystematic PSA screening. Whilst preliminary results from the ’BARCODE1’ study have shown that PGS can be used to identify men at highest risk of significant PC (McHugh et al. 2022).
If primary HCPs are to play a pivotal role in genetic risk-assessment in the future, then the reported lack of knowledge and confidence surrounding genetic risk-assessment and interpretation highlights the urgent need for workforce training. In the UK, the government funded, NHS England Genomics Education Programme has developed a range of educational tools to upskill the workforce(Genomics Education Programme 2023). However global collaboratives will be needed to help support LMIC to participate in this space.
Further research should also focus on how to integrate genomic results with clinical decision support tools within the electronic healthcare record and to determine the best report format to use to aid risk communication with patients. This will enhance acceptability amongst HCPs. Results from the ‘eMERGE’ study, which aims to integrate genetic risk information with clinical care and lifestyle recommendations, may clarify how best to approach this (Linder et al. 2023).
Other innovations, such as multi-cancer early detection tests (MCEDs), show promise to support primary care to detect cancer early (Schrag et al. 2023) and the results of the NHS-Galleri clinical trial, using MCEDs in asymptomatic primary care populations, are awaited (Neal et al. 2022). Sensitivity of MCEDs has been shown to be poor though for some cancer types, including PC and BC (Klein et al. 2021), and this highlights the potential for a complimentary role for PGS particularly in the risk-assessment of PC and BC.
Finally, although PGS show promise and potential to improve risk prediction in cancer it is important that the focus on genetic risk does not detract from other contributors of disease such as environmental and lifestyle risks that may have potentially greater impact on cancer outcomes (Sud et al. 2023). For maximum benefit integrated PGS that include non-genomic risk factors should ultimately be used so that preventative strategies can be targeted at individuals at highest risk of cancer.
Data availability
No datasets were generated or analysed during the current study.
References
+ Our Future Health (2022) Seven things you need to know about Our Future Health. 30 June 2022. Our Future Health. https://ourfuturehealth.org.uk/news/seven-things-you-need-to-know-about-the-uks-largest-ever-health-research-programme/ [Accessed: 14 July 2023].
Archer S, Babb de Villiers C, Scheibl F, Carver T, Hartley S, Lee A, Cunningham AP, Easton DF, McIntosh JG, Emery J, Tischkowitz M, Antoniou AC, Walter FM (2020) Evaluating clinician acceptability of the prototype CanRisk tool for predicting risk of breast and ovarian cancer: a multi-methods study. PLoS One 15(3):e0229999. https://doi.org/10.1371/journal.pone.0229999
Archer S, Donoso FS, Carver T, Yue A, Cunningham AP, Ficorella L, Tischkowitz M, Easton DF, Antoniou AC, Emery J, Usher-Smith J, Walter FM (2023) Exploring the barriers to and facilitators of implementing CanRisk in primary care: a qualitative thematic framework analysis. Br J Gen Pract. https://doi.org/10.3399/BJGP.2022.0643
Ayoub A, Lapointe J, Nabi H, Pashayan N (2023) Risk-stratified breast cancer screening incorporating a polygenic risk score: a survey of uk general practitioners’ knowledge and attitudes. Genes 14(3):732. https://doi.org/10.3390/genes14030732
Benafif S, Ni Raghallaigh H, McGrowder E, Saunders EJ, Brook MN et al (2022) The BARCODE1 Pilot: a feasibility study of using germline single nucleotide polymorphisms to target prostate cancer screening. BJU Int 129(3):325–336. https://doi.org/10.1111/bju.15535
Brockman DG, Petronio L, Dron JS, Kwon BC, Vosburg T, Nip L, Tang A, O’Reilly M, Lennon N, Wong B, Ng K, Huang KH, Fahed AC, Khera AV (2021) Design and user experience testing of a polygenic score report: a qualitative study of prospective users. BMC Med Genet 14(1):238. https://doi.org/10.1186/s12920-021-01056-0
Butrick MN, Vanhusen L, Leventhal K-G, Hooker GW, Nusbaum R, Peshkin BN, Salehizadeh Y, Pavlick J, Schwartz MD, Graves KD (2014) Discussing race-related limitations of genomic testing for colon cancer risk: implications for education and counseling. Soc Sci Med (1982) 114:26–37. https://doi.org/10.1016/j.socscimed.2014.05.014
Cancer Research UK (2018) Cancer in the UK 2018.p.20. https://www.cancerresearchuk.org/sites/default/files/state_of_the_nation_apr_2018_v2_0.pdf.
CanRisk v2.3.5 (2023) CanRisk. 12 June 2023. https://www.canrisk.org [Accessed: 14 September 2023].
Cavazos TB, Witte JS (2021) Inclusion of variants discovered from diverse populations improves polygenic risk score transferability. Hum Genet Genom Adv 2(1):100017. https://doi.org/10.1016/j.xhgg.2020.100017
Conran CA, Shi Z, Resurreccion WK, Na R, Helfand BT, Genova E, Zheng SL, Brendler CB, Xu J (2021) Assessing the clinical utility of genetic risk scores for targeted cancer screening. J Transl Med 19(1):41. https://doi.org/10.1186/s12967-020-02699-w
Esserman LJ (2017) The WISDOM Study: breaking the deadlock in the breast cancer screening debate. npj. Breast Cancer 3(1):1–7. https://doi.org/10.1038/s41523-017-0035-5
Fatumo S, Chikowore T, Choudhury A, Ayub M, Martin AR, Kuchenbaecker K (2022) A roadmap to increase diversity in genomic studies. Nat Med 28(2):243–250. https://doi.org/10.1038/s41591-021-01672-4
Fredsoe J, Kirkegaard P, Edwards A, Vedsted P, Sorensen KD, Bro F (2020a) A genetic risk assessment for prostate cancer influences patients’ risk perception and use of repeat psa testing: a cross-sectional study in danish general practice. BJGP Open 4(2):1039. https://doi.org/10.3399/BJGPOPEN20X101039
Fredsoe J, Koetsenruyter J, Vedsted P, Kirkegaard P, Vaeth M, Edwards A, Orntoft TF, Sorensen KD, Bro F (2020b) The effect of assessing genetic risk of prostate cancer on the use of PSA tests in primary care: a cluster randomized controlled trial. PLoS Med 17(2):e1003033. https://doi.org/10.1371/journal.pmed.1003033
French DP, Cameron E, Benton JS, Deaton C, Harvie M (2017) Can communicating personalised disease risk promote healthy behaviour change? a systematic review of systematic reviews. Ann Behav Med 51(5):718–729. https://doi.org/10.1007/s12160-017-9895-z
Genomics Education Programme (2023) NHS England Genomics Education - Delivering genomics education, training and experience for the healthcare workforce. 13 October 2023. HEE Genomics Education Programme. https://www.genomicseducation.hee.nhs.uk/ [Accessed: 19 October 2023].
Graves KD, Leventhal K-G, Nusbaum R, Salehizadeh Y, Hooker GW, Peshkin BN, Butrick M, Tuong W, Mathew J, Goerlitz D, Fishman MB, Shields PG, Schwartz MD (2013) Behavioral and psychosocial responses to genomic testing for colorectal cancer risk. Genomics 102(2):123–130. https://doi.org/10.1016/j.ygeno.2013.04.002
Green H.D., Merriel S.W.D., Oram R.A., Ruth K.S., Tyrrell J., Jones S.E., Thirlwell C., Weedon M.N., & Bailey S.E.R. (2022) Applying a genetic risk score for prostate cancer to men with lower urinary tract symptoms in primary care to predict prostate cancer diagnosis: a cohort study in the UK Biobank. medRxiv. ((Green) Exeter Centre of Excellence for Diabetes Research (EXCEED), University of Exeter, Medical School, St Luke’s Campus, University of Exeter, Heavitree Road, Devon, Exeter EX1 2LU, United Kingdom). 10.1101/2022.01.21.22269629.
Hamilton JG, Abdiwahab E, Edwards HM, Fang M-L, Jdayani A, Breslau ES (2017) Primary care providers’ cancer genetic testing-related knowledge, attitudes, and communication behaviors: a systematic review and research agenda. J Gen Intern Med 32(3):315–324. https://doi.org/10.1007/s11606-016-3943-4
Hao L, Kraft P, Berriz GF, Hynes ED, Koch C et al (2022) Development of a clinical polygenic risk score assay and reporting workflow. Nat Med 28(5):1006–1013. https://doi.org/10.1038/s41591-022-01767-6
Hollands GJ, French DP, Griffin SJ, Prevost AT, Sutton S, King S, Marteau TM (2016) The impact of communicating genetic risks of disease on risk-reducing health behaviour: systematic review with meta-analysis. The BMJ 352:i1102. https://doi.org/10.1136/bmj.i1102
Hong QN, Fàbregues S, Bartlett G, Boardman F, Cargo M, Dagenais P, Gagnon M-P, Griffiths F, Nicolau B, O’Cathain A, Rousseau M-C, Vedel I, Pluye P (2018) The Mixed Methods Appraisal Tool (MMAT) version 2018 for information professionals and researchers. Educ Inf 34(4):285–291. https://doi.org/10.3233/EFI-180221
Huntley C, Torr B, Sud A, Rowlands CF, Way R, Snape K, Hanson H, Swanton C, Broggio J, Lucassen A, McCartney M, Houlston RS, Hingorani AD, Jones ME, Turnbull C (2023) Utility of polygenic risk scores in UK cancer screening: a modelling analysis. Lancet Oncol 24(6):658–668. https://doi.org/10.1016/S1470-2045(23)00156-0
International Agency for Research on Cancer, W. (2023) Cancer Tomorrow- a tool that predicts future cancer incidence and mortality burden worldwide form the current estimates in 2020 up until 2040. https://gco.iarc.fr/tomorrow/en.
Kerman BJ, Brunette CA, Harris EJ, Antwi AA, Lemke AA, Vassy JL (2023) Primary care physician use of patient race and polygenic risk scores in medical decision-making. Genet Med 25(4):100800. https://doi.org/10.1016/j.gim.2023.100800
Kirkegaard P, Edwards A, Nielsen TLO, Orntoft TF, Sorensen KD, Borre M, Bro F (2018) Perceptions about screening for prostate cancer using genetic lifetime risk assessment: a qualitative study. BMC Fam Pract 19(1):32. https://doi.org/10.1186/s12875-018-0717-6
Kirkegaard P, Vedsted P, Edwards A, Fenger-Gron M, Bro F (2013) A cluster-randomised, parallel group, controlled intervention study of genetic prostate cancer risk assessment and use of psa tests in general practice-the ProCaRis study: study protocol. BMJ Open 3(3):002452. https://doi.org/10.1136/bmjopen-2012-002452
Klein EA, Richards D, Cohn A, Tummala M, Lapham R, Cosgrove D, Chung G, Clement J, Gao J, Hunkapiller N, Jamshidi A, Kurtzman KN, Seiden MV, Swanton C, Liu MC (2021) Clinical validation of a targeted methylation-based multi-cancer early detection test using an independent validation set. Ann Oncol 32(9):1167–1177. https://doi.org/10.1016/j.annonc.2021.05.806
Laforest F, Kirkegaard P, Mann B, Edwards A (2019) Genetic cancer risk assessment in general practice: systematic review of tools available, clinician attitudes, and patient outcomes. Br J Gen Pract 69(679):e97–e105. https://doi.org/10.3399/bjgp18X700265
Laza-Vásquez C, Martínez-Alonso M, Forné-Izquierdo C, Vilaplana-Mayoral J, Cruz-Esteve I, Sánchez-López I, Reñé-Reñé M, Cazorla-Sánchez C, Hernández-Andreu M, Galindo-Ortego G, Llorens-Gabandé M, Pons-Rodríguez A, Rué M (2022) Feasibility and acceptability of personalized breast cancer screening (DECIDO Study): a single-arm proof-of-concept trial. Int J Environ Res Public Health 19(16):10426. https://doi.org/10.3390/ijerph191610426
Leventhal K-G, Tuong W, Peshkin BN, Salehizadeh Y, Fishman MB, Eggly S, FitzGerald K, Schwartz MD, Graves KD (2013) ‘Is it really worth it to get tested?’: primary care patients’ impressions of predictive SNP testing for colon cancer. J Genet Couns 22(1):138–151. https://doi.org/10.1007/s10897-012-9530-x
Linder JE, Allworth A, Bland HT, Caraballo PJ, Chisholm RL et al (2023) Returning integrated genomic risk and clinical recommendations: The eMERGE study. Genet Med 25(4). https://doi.org/10.1016/j.gim.2023.100006
McHugh JK, Raghallaigh HN, Bancroft E, Kote-Jarai Z, Benafif S, Eeles RA (2022) The BARCODE1 study in primary care: early results targeting men with increased genetic risk of developing prostate cancer-examining the interim data from a community-based screening program using polygenic risk score to target screening. J Clin Oncol 40(6 SUPPL). https://doi.org/10.1200/JCO.2022.40.6_suppl.231
McWilliams L, Evans DG, Payne K, Harrison F, Howell A, Howell SJ, French DP (2022) Implementing risk-stratified breast screening in england: an agenda setting meeting. Cancers 14(19):4636. https://doi.org/10.3390/cancers14194636
Moorthie, S., Hall, A., Janus, J., Brigden, T., Babb de Villiers, C., Blackburn, L., Johnson, E. & Kroese, M. (2021) Polygenic Scores and clinical utility. https://www.phgfoundation.org/media/35/download/polygenic-scores-and-clinical-utility.pdf?v=1&inline=1.
Moorthie, S., Janus, J., Turner, H., Babb de Villiers, C., Mitchell, C. & Kroese, M. (2023) Evaluation of polygenic score applications. https://www.phgfoundation.org/media/880/download/Evaluation%20of%20polygenic%20score%20applications.pdf?v=1&inline=1.
Myers RE, Ruth K, Manne SL, Cocroft J, Sifri R, Ziring B, Burgh D, Ross E, Weinberg DS (2015) Effects of genetic and environmental risk assessment feedback on colorectal cancer screening adherence. J Behav Med 38(5):777–786. https://doi.org/10.1007/s10865-015-9626-5
MyPeBS (2022) International randomized study comparing personalized, risk-stratified to standard breast cancer screening in women aged 40-70. https://clinicaltrials.gov/ct2/show/NCT03672331.
National Institute for Health and CareExcellence, N. (2015) Overview | Suspected cancer: recognition and referral | Guidance | NICE. https://www.nice.org.uk/guidance/ng12.
National Institutes of Health (NIH) (2024) The All of Us Research Programme. 2024. https://gbr01.safelinks.protection.outlook.com/?url=https%3A%2F%2Fallofus.nih.gov%2F&data=05%7C02%7Cfdannhauser%40nhs.net%7C72efe104b70b41c0148f08dc36bc4f05%7C37c354b285b047f5b22207b48d774ee3%7C0%7C0%7C638445429604109248%7CUnknown%7CTWFpbGZsb3d8eyJWIjoiMC4wLjAwMDAiLCJQIjoiV2luMzIiLCJBTiI6Ik1haWwiLCJXVCI6Mn0%3D%7C0%7C%7C%7C&sdata=Nq5HvRw0UetNZpjTxjCAUTY6tZlZEZUsA%2FcVs6gMch4%3D&reserved=0 [Accessed: 7 March 2024].
Neal RD, Johnson P, Clarke CA, Hamilton SA, Zhang N, Kumar H, Swanton C, Sasieni P (2022) Cell-Free DNA-based multi-cancer early detection test in an asymptomatic screening population (NHS-Galleri): design of a pragmatic, prospective randomised controlled trial. Cancers 14(19):4818. https://doi.org/10.3390/cancers14194818
Nusbaum R, Leventhal K-G, Hooker GW, Peshkin BN, Butrick M, Salehizadeh Y, Tuong W, Eggly S, Mathew J, Goerlitz D, Shields PG, Schwartz MD, Graves KD (2013) Translational genomic research: protocol development and initial outcomes following SNP testing for colon cancer risk. Transl Behav Med 3(1):17–29. https://doi.org/10.1007/s13142-012-0149-0
Page MJ, Moher D, Bossuyt PM, Boutron I, Hoffmann TC et al (2021) PRISMA 2020 explanation and elaboration: updated guidance and exemplars for reporting systematic reviews. BMJ 372:n160. https://doi.org/10.1136/bmj.n160
Pak LM, Purad CC, Nadipally S, Rao MP, Mukherjee S, Hegde SKB, Golshan M (2021) Cancer awareness and stigma in rural assam india: baseline survey of the detect early and save her/him (DESH) program. Ann Surg Oncol 28(12):7006–7013. https://doi.org/10.1245/s10434-021-10366-7
PGS Catalog (2023) The Polygenic Score (PGS) Catalog. 22 June 2023. https://www.pgscatalog.org [Accessed: 28 June 2023].
Pharoah PDP, Antoniou A, Bobrow M, Zimmern RL, Easton DF, Ponder BAJ (2002) Polygenic susceptibility to breast cancer and implications for prevention. Nat Genet 31(1):33–36. https://doi.org/10.1038/ng853
Pramesh CS, Badwe RA, Bhoo-Pathy N, Booth CM, Chinnaswamy G et al (2022) Priorities for cancer research in low- and middle-income countries: a global perspective. Nat Med 28(4):649–657. https://doi.org/10.1038/s41591-022-01738-x
Ritchie, H., Spooner, F. & Roser, M. (2018) Causes of death. Our World in Data. https://ourworldindata.org/causes-of-death.
Ruan Y, Lin Y-F, Feng Y-CA, Chen C-Y, Lam M, Guo Z, He L, Sawa A, Martin AR, Qin S, Huang H, Ge T (2022) Improving Polygenic Prediction in Ancestrally Diverse Populations. Nat Genet 54(5):573–580. https://doi.org/10.1038/s41588-022-01054-7
Saya S, McIntosh JG, Winship IM, Clendenning M, Milton S, Oberoi J, Dowty JG, Buchanan DD, Jenkins MA, Emery JD (2020) A genomic test for colorectal cancer risk: is this acceptable and feasible in primary care? Public Health Genomics 23(3–4):110–121. https://doi.org/10.1159/000508963
Saya S, McIntosh JG, Winship IM, Milton S, Clendenning M, Kyriakides M, Oberoi J, Buchanan DD, Jenkins MA, Emery JD (2022) Informed choice and attitudes regarding a genomic test to predict risk of colorectal cancer in general practice. Patient Educ Couns 105(4):987–995. https://doi.org/10.1016/j.pec.2021.08.008
Schrag D, Beer TM, McDonnell CH, Nadauld L, Dilaveri CA, Reid R, Marinac CR, Chung KC, Lopatin M, Fung ET, Klein EA (2023) Blood-based tests for multicancer early detection (PATHFINDER): a prospective cohort study. Lancet 402(10409):1251–1260. https://doi.org/10.1016/S0140-6736(23)01700-2
Silarova B, Sharp S, Usher-Smith JA, Lucas J, Payne RA, Shefer G, Moore C, Girling C, Lawrence K, Tolkien Z, Walker M, Butterworth A, Di Angelantonio E, Danesh J, Griffin SJ (2019) Effect of communicating phenotypic and genetic risk of coronary heart disease alongside web-based lifestyle advice: the INFORM Randomised Controlled Trial. Heart 105(13):982–989. https://doi.org/10.1136/heartjnl-2018-314211
Sud A, Horton RH, Hingorani AD, Tzoulaki I, Turnbull C, Houlston RS, Lucassen A (2023) Realistic expectations are key to realising the benefits of polygenic scores. BMJ (Clinical research ed) 380:e073149. https://doi.org/10.1136/bmj-2022-073149
Sullivan BG, Qazi A, Senthil M (2021) Cancer screening programs in low- and middle-income countries: strategies for success. Ann Surg Oncol 28(12):6918–6919. https://doi.org/10.1245/s10434-021-10509-w
Taylor LC, Hutchinson A, Law K, Shah V, Usher-Smith JA, Dennison RA (2023) Acceptability of risk stratification within population-based cancer screening from the perspective of the general public: A mixed-methods systematic review. Health Expect 26(3):989–1008. https://doi.org/10.1111/hex.13739
Uffelmann E, Huang QQ, Munung NS, de Vries J, Okada Y, Martin AR, Martin HC, Lappalainen T, Posthuma D (2021) Genome-wide association studies. Nat Rev Dis Primers 1(59):1–21
University of Aarhus (2022) Prostate cancer risk assessment using genetic markers in general practice. https://clinicaltrials.gov/study/NCT01739062.
US Department of Veterans Affairs (VA) (2024) Million Veteran Program. 31 January 2024. https://gbr01.safelinks.protection.outlook.com/?url=https%3A%2F%2Fwww.mvp.va.gov%2Fpwa%2F&data=05%7C02%7Cfdannhauser%40nhs.net%7C72efe104b70b41c0148f08dc36bc4f05%7C37c354b285b047f5b22207b48d774ee3%7C0%7C0%7C638445429604115515%7CUnknown%7CTWFpbGZsb3d8eyJWIjoiMC4wLjAwMDAiLCJQIjoiV2luMzIiLCJBTiI6Ik1haWwiLCJXVCI6Mn0%3D%7C0%7C%7C%7C&sdata=BMmv6dBJFEkbYx8Bvhm0WJYf9hU8L4tXC%2BDpCcbqQ74%3D&reserved=0 [Accessed: 7 March 2024].
Usher-Smith J, Emery J, Hamilton W, Griffin SJ, Walter FM (2015) Risk prediction tools for cancer in primary care. Br J Cancer 113(12):1645–1650. https://doi.org/10.1038/bjc.2015.409
Vassy JL, Brunette CA, Lebo MS, MacIsaac K, Yi T et al (2023a) The GenoVA study: Equitable implementation of a pragmatic randomized trial of polygenic-risk scoring in primary care. Am J Hum Genet 110(11):1841–1852. https://doi.org/10.1016/j.ajhg.2023.10.001
Vassy JL, Kerman BJ, Harris EJ, Lemke AA, Clayman ML, Antwi AA, MacIsaac K, Yi T, Brunette CA (2023b) Perceived benefits and barriers to implementing precision preventive care: results of a national physician survey. Eur J Hum Genet 1–8. https://doi.org/10.1038/s41431-023-01318-8
Wand H, Lambert SA, Tamburro C, Iacocca MA, O’Sullivan JW et al (2021) Improving reporting standards for polygenic scores in risk prediction studies. Nature 591(7849):211–219. https://doi.org/10.1038/s41586-021-03243-6
Weinberg DS, Myers RE, Keenan E, Ruth K, Sifri R, Ziring B, Ross E, Manne SL (2014) Genetic and environmental risk assessment and colorectal cancer screening in an average-risk population: a randomized trial. Ann Intern Med 161(8):537–545. https://doi.org/10.7326/M14-0765
Weissbrod O, Kanai M, Shi H, Gazal S, Peyrot WJ, Khera AV, Okada Y, Martin AR, Finucane H, Price AL (2022) Leveraging fine-mapping and multi-population training data to improve cross-population polygenic risk scores. Nat Genet 54(4):450–458. https://doi.org/10.1038/s41588-022-01036-9
World Health Organisation (2022) Cancer - Fact sheet. https://www.who.int/news-room/fact-sheets/detail/cancer.
Acknowledgements
The authors would like to thank Veronica Phillips and Eleanor Barker of the University of Cambridge Medical Library for guidance on developing the search strategy.
Funding
The authors did not receive support from any organisation for the submitted work.
Juliet A. Usher-Smith is supported by an NIHR Advanced Fellowship (NIHR300861).
Author information
Authors and Affiliations
Contributions
FD conceptualised the review, performed the literature search and wrote the original draft. FD and JU agreed the final citations for inclusion. All authors were involved in data analysis, quality assessment and read and reviewed the final manuscript. All authors have read and agreed to the published version of the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Dannhauser, F.C., Taylor, L.C., Tung, J.S. et al. The acceptability and clinical impact of using polygenic scores for risk-estimation of common cancers in primary care: a systematic review. J Community Genet 15, 217–234 (2024). https://doi.org/10.1007/s12687-024-00709-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12687-024-00709-8