Abstract
There are five grizzly bear (Ursus arctos horribilis) populations in the lower 48 states of the United States. My goal in this Commentary was to ascertain whether genetic diversity is being lost from the isolated GYE grizzly bear population and to better understand any viability implications. I reviewed the scientific literature, including two key genetic studies that the US Fish and Wildlife Service (USFWS) relied upon for their 2007 and current 2017 GYE grizzly bear genetics policy. I discovered that some studies reveal a loss of heterozygosity in the GYE bear population, both historically and in recent decades. Some had a statistically significant depletion rate. My review took place periodically between 2010 and 2021 and indicates that the genome of the GYE grizzly bear population is too small for long-term adaptation. The paper includes a discussion about evolutionary adaptation which invokes time frames rarely considered by nature conservation planners. I also examined genetic statements in the USFWS’s 2017 GYE grizzly bear delisting regulations and highlighted those that seem incongruent with current scientific thought. If this paper is read by some scientists, land managers, administrators, environmentalists, and others with some genetics background, they will better understand some USFWS decisions and policy statements. This case study illustrates that land management agencies can provide a one-sided treatment of some science when writing regulations about genetics.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
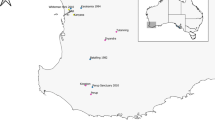
At present, there are four distinct grizzly bear populations in the U.S. lower 48 states, consisting of the Greater Yellowstone Ecosystem (GYE; 20,000 km2 with 737 bears), the U.S. portion of the Northern Continental Divide Ecosystem (NCDE; 25,000 km2 with 1068 bears), the U.S portion of the Selkirk Mountains area (3021 km2 with 53 bears), the Cabinet-Yaak area (6700 km2 with 55–60 bears) and the Northern Cascades Ecosystem (25,000 km2 with 0 bears) (USFWS 2021) (Fig. 1). Additionally, wildlife biologists familiar with the Selkirk-Bitterroot-Frank Church River-of-No-Return Wilderness Complex (SBFW; 15,054 km2) acknowledged sightings of two transient bears over the last 13 years with another five confirmed within 15 km or less of this recovery area (personal communication, Montana Fish, Wildlife & Parks, and USFWS). There exists 358,000 km2 of human-dominated, mostly non-protected land separating the GYE, NCDE, and SBFW. The distance between the GYE and NCDE grizzly bear populations is approximately 110 km (White et al. 2017).
The GYE grizzly bear population was removed from the “threatened” list on July 31, 2017, on the basis of provisions of the 1973 US ESA (16 U.S.C. 1531–1544, 87 Stat. 884, as amended). However, the grizzly bear was returned to the ESA listing by a Federal District Court judge on September 24, 2018. This was the second time a federal judge refuted all the US Fish and Wildlife Service (USFWS) reasons for delisting this bear population. The final USFWS decision to delist the GYE grizzly bear on April 30, 2007, and again on July 31, 2017, was based primarily on demographic population data; however, genetics data were also taken into account. For more information about the study area, US grizzly bear distribution and population status, and ESA history, see Part I of the case study.
This work, spanning 2010 to 2021, addresses my concern about whether the GYE grizzly bear population is losing genetic diversity, how much, and what is the viability implications. This suspicion of potential ongoing genetic problems followed from an understanding that habitat loss and fragmentation generally result in a decrease in the number of alleles as well as a reduction in observed and also expected heterozygosity in mammals (Lino et al. 2019). To also address my question in a policy context, I assessed the USFWS’s treatment of genetics in its proposed 2007 and final 2017 regulations to remove the GYE grizzly bear Distinct Population Segment from the Endangered Species Act (ESA) “threatened” list (USFWS 2007, 2017). I also reviewed USFWS responses to public comments on the regulations since they also reflect USFWS thinking. With only a modest familiarity with genetics, some scientists, bear managers, administrators, environmentalists, and others who try to review the USFWS 2017 federal delisting ruling, may be stymied by the jargon and the complexity of the pertinent genetic studies. The message that Schonewald-Cox et al. (1983) conveyed 38 years ago remains relevant: technical literature must be explained so practitioners and others can understand more of it.
I proceeded by reviewing the scientific literature and then evaluated USFWS statements in their 130-page, 2017 final grizzly bear regulations (USFWS 2017) that pertain to genetics. This review indicates that land management or regulatory agencies can provide a one-sided treatment of some information when it comes to genetics. This paper is therefore warranted to see how some federal genetics policies for the GYE grizzly bear can be misrepresented and what impact that might have on the bear. This paper is a continuation of work started in Shafer (2013).
Since many non-geneticists understandably have a limited grasp of population genetics and evolution, this review might be described as “translational ecology” (Schlesinger 2010). That is, translating the scientific literature so that others with different or less technical backgrounds can understand it. In the words of Groom et al. (2006, p. 626), “It is impossible to integrate conservation science into environmental policy if no one but the scientist understands the science.”
Genetics background
It is important to be somewhat familiar with certain aspects of conservation genetics to better understand this paper. What follows are highlights.
Habitat fragmentation negatively impacts individual genetic fitness and the overall population viability of many species (Young and Clarke 2000; Lindenmayer and Fischer 2006). Studies have reported a positive correlation between a population’s heterozygosity (H) level and its fitness (Allendorf and Leary 1986; Mitton 1993; Reed 2005); however, see the Genetic Fitness Section in the Appendix for newer insights. Loss of genetic variation resulting from decreases in H and allelic diversity (P) (Avise 1994), is caused by habitat fragmentation and population isolation (Keyghobadi 2007).
Corridors are presumed to reduce the negative impacts of habitat fragmentation, such as the loss of genetic variation (Christie and Knowles 2015). Some studies have documented the use of corridors by animals (Gilbert-Norton et al. 2010; Resasco 2019).
Inbreeding results in increased homozygosity or decreased heterozygosity (H). The negative influence reduced heterozygosity has on populations is well documented (Frankham 2010). Inbreeding is found in some outbreeding plants and animals (Lacy 1997) and for example, in brown bears in Nordic zoos (Laidre et al. 1996). Inbreeding depression (F) is the negative outcome of inbreeding. A rapid decline in fitness has been observed in zoo populations (Ralls and Ballou 1983) due to inbreeding and has also been detected in wild animals (Keller and Waller 2002). When measured in various wild mammal populations, F was moderate to high (Crnokrak and Roff 1999). Small population size encourages inbreeding, increasing the number of individuals that are homozygous for deleterious recessive alleles (Wright 1977) while larger populations have greater genetic diversity (Frankham 1996).
Gene flow between isolated populations reduces the frequency of homozygous genotypes (Wahlund 1929) and therefore the potential impact of deleterious alleles. The continuous infusion of new alleles, even in large populations, helps to ensure that enough genetic variation is maintained to prevent extirpation or extinction (Frankham et al. 2010).
Genetic erosion has a negative influence on most species declining toward extinction (Spielman et al. 2004) and can often lead to their demise (Frankham 1995b). The probability of species extirpation or extinction due to genetics was deemed to be a minor concern in the past (Lande 1988; Caughley 1994), but many do not hold this viewpoint today (e.g., Frankham 2005c; Brook 2008). An “extinction spiral” accompanies a decline in genetic diversity and can result in local population extirpation (Gilpin and Soulé 1986).
Model approaches for assessing the genetic diversity of a population
Adequate population size
In 1995, the US District Court for the District of Columbia remanded (sent back) the USFWS’s treatment of genetics isolation on grizzly bear populations to the agency for further study. However, by September 21, 2009, the Court deferred to USFWS’S expertise in judging “adequate population size” (Greater Yellowstone Coalition v. Christopher Servheen 2011). Thus, the determination of adequate population size was left to agency discretion. The USFWS used effective population size (Ne) in its consideration of what an “adequate population” would be for GYE bears.
Effective population size
Effective population size (Ne), an idea introduced by Wright (1931), is the best concept for predicting the rate of loss of genetic variation in isolated populations. Ne, therefore, is the key factor used to monitor or manage a population’s genetic variation. It is defined as “the number of individuals that would result in the same loss of genetic diversity, inbreeding, or genetic drift if they behaved in the manner of an idealized population” (Frankham et al. 2010, p. 530). That is, “ideal” refers to a hypothetical population with a constant size, equal sex ratio, and no immigration, emigration, mutation, or selection. Simply, Ne is an indicator of a population’s evolutionary potential and risk from inbreeding (Franklin and Frankham 1998).
Nc is always larger than Ne except in very unusual situations. The larger the Ne, the smaller the loss of genetic variation. In other words, a larger Ne retains more genetic diversity, since the rate of genetic drift and the loss of genetic diversity will be lower at a larger Ne value. Effective population size (Ne) (the number of breeding individuals needed for long-term adaptation) is therefore different from the total, or census population size (Nc). Wang (2005) warned that Ne is “notoriously difficult to estimate” and described various concepts of Ne: inbreeding effective size (Nei), variance effective size (Nev), eigenvalue mutation effective size, and coalescent theory effective size (Ewens 1990; Gregorious 1991; Caballero 1994; Whitlock and Barton 1997; Charlesworth et al. 2003). Nei and Nev are the most used, where Nei indicates how much H changes over time due to inbreeding, while Nev is a measure of changes in allele frequency due to genetic drift. Wang (2005), Wang et al. (2010), and Palstra and Ruzzante (2008) reviewed the various approaches to estimate Ne. Despite recent advances in measuring Ne (Luikart et al. 2010), there remain various pitfalls in its calculation (Hare et al. 2011; Palstra and Fraser 2012).
The 50/500 rule: population size needed to diminish genetic erosion
The 50/500 rule was derived from a literature review by Franklin (1980) and Soulé (1980). Franklin (1980) recommended a Ne of at least 50 individuals for short-term genetic fitness for single and independent populations, such as vigor, fecundity, fertility, and disease resistance. This number was derived from the work of animal breeders who found that fewer than 50 individuals caused an unacceptable 1% loss of H per generation due to inbreeding. Franklin (1980) also recommended a Ne of 500 individuals to prevent the erosion of genetic variation and allow for long-term adaptation, as did Soulé (1980). Franklin’s estimate was derived from work on Drosophila bristle number (Lande 1976). It was also based on the assumption that the loss of genetic variation at Ne = 500 will be balanced by a gain through mutation, an assumption that has been questioned (Lacy 1992). Soulé (1987, p. 175) later, and with some reluctance, reaffirmed his guess at Nc: the order of magnitude lower population boundary would be “low thousands” for a 95% chance of persistence, without any fitness loss, for several 100 years.
The 50/500 rule was accepted by some scientists (Newmark 1985; Wilcox 1986) but later fell into disfavor with others (Lande 1988; Boyce 1997). Frankham et al. (2014) recommended revising the 50/500 rule to 100/1000. Franklin et al. (2014) retorted that the Frankham et al. argument was unconvincing. Jamieson and Allendorf (2012) recommended 2000 for long-term evolutionary fitness but this all depends on the adopted ratio for Ne/Nc. Therefore, after more than three decades, the Ne = 500 value for long-term fitness has not been abandoned by some scientists (Franklin et al. 2014; Jamieson and Allendorf 2012) though boosted upward by others. A few authors mentioned connectivity in their Ne discussions (Frankham et al. 2014; Jamieson and Allendorf 2012) but none explicitly stated that Ne applies to both single and connected populations. Fifty years ago, Frankel (1970, p. 166) suggested that healthy population sizes demand “a thousand rather than hundreds.”
Genetic variation in GYE grizzly bears
Proctor et al. (2005), Proctor et al. (2012)) used a likelihood-based test to find area-specific allele frequencies and employed 15-locus microsatellite genotyping to find an average expected heterozygosity (He) in GYE bears of 0.64 to 0.67. They confirmed the genetic distinctiveness of the grizzly bear populations in the western US and Canada and attributed it to isolation.
Paetkau et al. (1998) used approximations for eight mtDNA loci that revealed 69% H in the NCDE population, but only 55% in the GYE population. There was a 15–20% reduction in H for the GYE population during one hundred years and an F of 1–4%. They observed that the GYE population exhibited among the lowest genetic diversities of any brown bear population in continental North America.
Waits et al. (1998) used both mtDNA and nuclear microsatellite analysis for 220 brown bears over North America. Using samples from DNA depositories (GYE 53 bears, NCDE 49 bears), they found a genetic diversity (h) of 0.611 for NCDE bears which dropped to 0.240 for GYE bears. Note: h, defined by the authors as genetic diversity, equals H for nuclear genes but not for mtDNA. This drop was not statistically significant in the GYE though the authors characterized it as “considerably lower” than for the NCDE.
Miller and Waits (2003) discovered a slight but not statistically significant decline in H from the 1910s (0.580) to the 1990s (0.560). Allelic diversity grew from 4.50 (1910s), to 4.63 (1960s) to 4.88 (1990s). Historical samples used eight dinucleotide microsatellites. Contemporary data came from Paetkau et al. (1998) and other sources.
Haroldson et al. (2010) used nuclear genetic protocols. They adopted 16 microsatellites obtained from 424 bears sampled from 1983 to 2007, including eight identified by Miller and Waits (2003). They found a 67% H for NCDE bears which declined to 60% for GYE bears. This 7% H drop was less than the 14% H drop derived by Miller and Waits (2003).
Kamath et al. (2015), whose authors included Interagency Grizzly Bear Study Team (IGBST) members, used three methods to determine Ne. They examined 20 microsatellites, including the eight used by Miller and Waits (2003), but came to a very different conclusion: no changes in H or P from 1985 to 2010.
Except for Miller and Waits (2003), all these workers assumed that the genetic diversity in GYE and NCDE bear populations was once the same until the effects of isolation manifested. The Miller and Waits (2003) and Kamath et al. (2015) studies will now be examined in detail because of the influence they had on USFWS policy.
The scientific basis for USFWS genetics policy
Miller and Waits (2003): initial findings, the debate, and follow-up policy
Miller and Waits (2003) examined tissue samples from 110 bears in museum collections deposited from 1912 to 1920 (28), from 1959 to 1981 (72), and from 1992 to 1999 (136). Using these historical samples, Miller and Waits (2003, p. 4338) found that the H of GYE grizzly bears “declined slightly” from the 1910s to the 1990s. This drop was later judged as statistically significant (Allendorf et al. 2006). They also estimated that P was 4.50 in the 1910s, 4.63 in the 1960s, and 4.88 in the 1990s so P increased slightly over this period. They also calculated H at 55% for the GYE which was lower than the NCDE’s 69%. They characterized genetic diversity in the GYE bears as “significantly lower” than in the NCDE population (Miller and Waits 2003, p. 4334). In addition, they found an inbreeding rate of 2.3% per generation.
They reported very low estimates of Ne (~ 85 for 1910–the 1960s and ~ 75–89 for 1960–1990) and indicated the upper and lower bounds of the mean Nc from 1965 to 1995 to be 180 and 454, respectively. Based on evidence that Nc was at least 400, and using earlier models and other studies, Miller and Waits (2003) estimated Ne to be 108 in 2003 and thought their data supported the notion that the lower genetic diversity in the GYE grizzly bears resulted from the population drop they experienced in the early 1970s (Allendorf et al. 2006). They used a Ne/Nc of around 0.3. Based on their findings, in 2007, the USFWS planned to implement translocations by 2020 and every 10 years thereafter (USFWS 2007); which apparently never took place.
Miller and Waits (2003) asserted that genetic factors were unlikely to have a significant impact on the viability of the GYE grizzly bear population over the next several decades but they also said gene flow would be beneficial. They later clarified that the projection timeline for needed genetic infusion to be “20–30 years” (Allendorf et al. 2006).
Another scientist, John Ballou, using models, predicts only a minor H loss over time. With a current Ne of say 80, inbreeding will increase at only about 0.0006 per generation [1/ (2*Ne)] (J. Ballou, Smithsonian Institution’s National Zoological Park [retired], personal communication, October 2016). Yet another scientist also predicted minor H loss. Allendorf et al. (2019) calculated a GYE grizzly bear H loss of 0.005 every 10 years given a population of 500 grizzly bears and assuming a Ne of > 100. But two other scientists argued, “there is no safe amount of inbreeding for normally outbred organisms” (Frankel and Soulé 1981, p. 72). H varies from 0.0 to 0.26 in mammals, with a mean of 0.04 (Sinclair et al. 2006).
Based on Miller and Waits (2003), the USFWS asserted that the GYE grizzly bear population required a Ne of only 100 bears to maintain their fitness, despite estimates by some other geneticists that Nc in the range of 2000–4000 bears would be needed to maintain genetic variation for future adaptation (Lynch and Lande 1998; Sinclair et al. 2006). Miller and Waits (2003) indicated that if the GYE grizzly bear population Ne ever rose to 500, and assuming a Ne/Nc of 0.27, then an Nc > 1850 would be required for long-term adaptation. Miller and Watts, therefore, made both short-term recommendations (Ne as 100, which would demand an Nc of > 400) and long-term recommendations (Nc = 1850). However, the USFWS focused on the short-term recommendation.
Scientists expressed their concern regarding the USFWS’s basis for judging the required Ne and signed a letter to the USFWS on March 20, 2006. Those scientists believed that the GYE grizzly bear population needed a Ne of 500 (or an Nc of 2000–3000 bears) before enough genetic diversity would be available to withstand random, regional-scale events (Craighead et al. 2006). Furthermore, in 2002, the consensus of the contributors to the book The Role of Genetics in Population Viability Analysis (Allendorf and Ryman 2002) decided that an Nc of 500 individuals was too small to support the long-term genetic fitness of a population.
In addition, commenters regarding the early USFWS 2007 proposed rule (2007 Federal Register notice) suggested that the goal for the GYE grizzly bear population (Nc) should also consist of 2000–3000 bears. If not, they argued that movement corridors connecting the GYE population with other bear populations should be created (USFWS 2007). Despite these suggestions and those of the above scientists, the USFWS decided that an Nc of 500 individuals was sufficient for delisting. The USFWS also believed that the GYE grizzly bear population was “close to carrying capacity inside the DMA (demographic monitoring area)” which to them was 700 bears (USFWS 2017, p. 30,571). Yellowstone National Park resource managers adopted the Miller and Waits (2003) recommendations (United Nations Educational, Scientific and Cultural Organization 2011).
The above recommendations are not the largest for Nc. Some scientists are convinced that estimates for Ne/Nc are 0.1 for species in general (Nunney and Campbell 1993; Vucetich et al. 1997; Frankham 1995c; Frankham et al. 2010). If so, any estimate for Nc would be 10 × higher than Ne, not just 4 × higher using a Ne/Nc ratio of 0.25. Lande (1995) estimated Ne as 5000 to retain evolutionary potential, which Frankham et al. (2010) thought was too high. Lynch and Lande (1998) estimated that Ne would vary between 1000 and 5000 bears. Allendorf and Ryman (2002) reviewed these recommendations in detail.
Kamath et al. (2015): findings and revised genetics policy
The USFWS in 2007 indicated they would provide updated guidance for GYE grizzly bear management if future genetic data challenged its policy (USFWS 2007). New information from a study instigated and conducted by the IGBST surfaced in 2015. Kamath et al. (2015) used three methods to determine Ne. Using a single sample nuclear approach, they examined 20 microsatellites in 729 live bears, including the same microsatellites used by Miller and Waits (2003) and others. They also used two multiple sample approaches to estimate variance Ne (Nev) and inbreeding Ne (Nei).
One of the techniques used was estimation by parentage assignment (EPA), as described by Wang et al. (2010). Kamath et al. (2015) recalculated Miller and Waits’ (2003) historical Ne estimates and found that Nei had greatly increased from ~ 80 during 1910–1960 to ~ 280 at that time. They estimated that from 1984 to 2007, Ne/Nc was 0.42–0.66, a fourfold increase. Their adopted 0.42–0.66 ratio was very different from a computer simulation value of 0.24–0.32 (Harris and Allendorf 1989; Allendorf et al. 1991). Kamath et al. (2015) believed that Nei for the GYE grizzly bear population increased from 102 in 1982 to 469 in 2010, and pointed out that H levels measured by other scientists actually increased: 0.55 in 1998 (Paetkau et al. 1998), to 0.56 in 2003 (Miller and Waits 2003), and then to 0.60 in 2010 (Haroldson et al. 2010). However, the results were not statistically significant. During the 2017 Federal Register public comment period, some argued that the new EPA method of genetic assessment needed more review (USFWS 2017b, p. 30609 ).
Kamath et al. (2015) found no changes in H or P from 1985 to 2010. They calculated an inbreeding coefficient (F) of only 0.2% for 1985–2010, in contrast to an F of 2.3% per generation for 1985–1999 obtained by Miller and Waits (2003). The USFWS subsequently questioned whether the changes in H were “biologically meaningful” (USFWS 2017, p. 30,623) and proposed withdrawing their translocation plans in 2016 (USFWS 2016). Other scientists were not hesitant to comment on the genetic diversity differences they observed in these studies. Based on data from allozyme loci, microsatellite loci, and mtDNA, Allendorf et al. (2006) concluded that the GYE grizzly bears had “substantially less” genetic variation compared to NCDE bears and the drop was statistically significant. These judgments diverge from the manner the USFWS portrayed the Miller and Waits (2003) findings: “there is no evidence for a ‘shrinking gene pool’” (USFWS 2017, p. 30,610). Kamath and colleagues said the GYE grizzly bear population would benefit from gene flow, as did Paetkau et al. (1998), Waits et al. (1998), Miller and Waits (2003), and Haroldson et al. (2010).
The USFWS embraced the findings of Kamath et al. (2015). As a result, the US Geological Survey (USGS) issued a press release indicating that the GYE grizzly bear population had not suffered a significant loss in genetic diversity (US Geological Survey 2015) and would therefore not suffer any important loss in genetic variation with a population size of 640–797 individuals. It asserted that the number of individuals was “sufficient to assure long-term genetic viability.” Reading the same Kamath and colleagues’ paper, an NGO issued a press release that stated that GYE grizzly bear genetic diversity had been stable since the 1980s, with an inbreeding rate of only 0.2%, so translocation was unnecessary (Anonymous 2015). The USFWS accepted the conclusions of various scientists that natural dispersal would benefit the GYE bears genome (USFWS 2017, p. 30,536). However, the USFWS nevertheless indicated that translocation would only occur if there were demonstrable negative impacts to the bears because of lowered H. But after citing Miller and Waits (2003) and Kamath et al. (2015), the USFWS continued to assert that “genetic concerns are not a threat to the GYE grizzly bear population” (USFWS 2017, p. 30,536). The US Forest Service (USFS) (2006, p. 68) believed the GYE grizzly bear population was already viable. The USFWS supported this USFS position: “there is more than enough [GYE] habitat to support a viable population” (USFWS 2017, p. 30,511).
Of key importance, however, is that government scientists and managers working in the GYE by 2003 had adopted calculations that address only short-term genetic fitness and not long-term genetic vigor. Furthermore, some regulations and other statements indicate that the USFWS does not understand Ne. For example, a statement in a USFWS strategy report (USFWS 2016) indicates the agency misapplied the Kamath et al. (2015) study. After referring to the Kamath et al. (2015) claim that Nc is 460, the agency believed this value was close to the 500-value espoused by Franklin (1980) for long-term evolutionary fitness. However, Franklin (1980) was referring to Ne, not Nc, so an Nc of 500 bears is much too small to achieve long-term genetic fitness. The criticism about misunderstanding Ne was a common complaint in the USFWS 2017 rules comments. “Other commentators took issue with our calculation and analysis of effective population size” (USFWS 2017, p. 30,609). More examples of misinterpretation follow.
Based on Kamath et al. (2015), the USFWS maintained that Ne for GYE grizzly bears is more than 4 × the minimum Ne calculated by Miller and Waits (2003). However, Miller and Waits (2003) were discussing short-term fitness when they said that existing Ne is likely to be “near or more than > 100.” The USFWS then mischaracterized what Miller and Waits (2003) said and suggested using Ne = 100 as the new yardstick for overall genetic fitness (USFWS 2017). However, Miller and Waits (2003) and Kamath et al. (2015) clearly observed that the GYE grizzly bears do not meet the population standard for long-term genetic fitness. The current USFWS’s goal is to maintain a total population of 500 bears in the DMA based on short term genetic fitness (USFWS 2017b, p. 30,610). But long-term genetic fitness is the ideal.
A planning time frame of a hundred years is not uncommon for species viability analyses as the USFWS pointed out (USFWS 2017, p. 30,561). A several 100-year time frame would be moving in a direction to retain genetic variation long term if the bear population size provided enough genetic diversity for adaptation. The agency mentioned minimum viable population size (e.g., Trail et al. 2007) only once in the regulations but without definition, discussion or projection. One scientist deemed it impossible to do a population viability analysis for even the best-studied bear species in the world, the grizzly bear (Boyce et al. 2001).
There is one inconvenient truth about these studies that warrant mention. No agreement about the accuracy of microsatellite data exists (Balloux and Lugon-Moulin 2002) and there are limitations to such methods (Putman and Carbone 2014). Although molecular markers are assumed to be substitutes for quantitative genetic variation, that correlation is weak (Reed and Frankham 2001). Storfer (1996) and others suggest that the use of quantitative variation would be a better approach. These grizzly bear genetic studies used neutral markers, such as microsatellites, which do not code for functional proteins and therefore do not consist of the genetic material used in natural selection. Therefore, we should not unquestionably accept these study results based on neutral genetic variation (McKay and Latta 2002). Nevertheless, that is all we have to guide us for now. Allendorf (2016) treats the history of various approaches used to determine the genetic composition of individuals and populations.
Summation
Compared to the NCDE, the GYE population is losing H (Paetkau et al. 1998; Waits et al. 1998; Miller and Waits 2003; Haroldson et al. 2010). This drop is statistically significant (e.g. Miller and Waits 2003, cited in (Allendorf 2006). However, Kamath et al. (2015) found no decrease in H or P from 1985 to 2010. Some models predict only a minor loss in H due to inbreeding. When comparing H differences between GYE and NCDE bears, some IGBST members (White et al. 2017) simply said the differences were “relatively large.”
However, H is not the most important factor here, even though it was emphasized by the USFWS and the IGBST. For example, the agency thinks low H decreases the ability to adapt and evolve (USFWS 2017, p. 30535, 30515). H loss is less significant than the loss of P (Petit et al. 1996), and the latter will be lost more quickly (Keyghobadi 2007). Genetic drift will cause a loss of alleles and possibly retain (fix) the deleterious ones. Heterozygosity does not tell us much about allelic diversity (Allendorf 1986), so reliance on H values indicates little about the potential for long-term adaptation. Management for H by itself is misguided because the removal of P is more critical to a genetically diverse population (Fuerst and Maruyama 1986). A review of fragmentation studies found the most marked result was the loss of alleles, while measures of H and the number of polymorphic loci were less prominent (Schlaepfer et al. 2018). One rules commentator made a reasonable suggestion: create a model of the rate of allele loss at different population sizes resulting from genetic drift (USFWS 2017, p. 30,609).
Perhaps in the future, we can calculate how much genetic diversity will be retained based on population sizes or habitat area/protected area configuration (Méndez et al. 2014). For now, if the GYE remains isolated, numerous scientists indicate it is too small to retain grizzly bear genetic variation over the long term.
As this case study outlines, some USFWS statements in the 2017 regulations are of concern. As a review, the USFWS (2017) remarked that: Ne was 100 for GYE bears and Nc was 469 (p. 30,610), there is uncertainty about whether the GYE bears “differ markedly” from the NCDE population (p. 30,519), H values indicate that GYE bears will continue to remain “healthy” (p. 30,535) and translocation into the GYE would occur only as a “last resort” (p. 30,536). Thus, according to the USFWS, genetic isolation is “not a threat” to the GYE grizzly bears in the “foreseeable future” (p. 30610). Not surprisingly, one rules commenter asked the USFWS to define the foreseeable future (USFWS 2017, p. 30,607).
For an elaboration on four key-related genetics concepts and their relationship to USFWS genetic policy statements, see the “Appendix”. We now will expand our “time frame of concern” (Frankel 1974).
A neglected concern: evolutionary adaptation
There are claims that fast genetic changes have taken place in a variety of short-generation species like birds (e.g., date of arrival at nesting grounds) due to changes in climate (Rice and Emery 2003; Bradshaw and Holzapfel 2006; Scheffers et al. 2016). Resistance to antibiotics and insecticides represent another example (Georghiou 1986). However, adaptation usually requires long periods of time (Barnovsky and Kraatz 2007). Regardless, the potential for evolution is a function of the amount of usable genetic diversity (Fisher 1930). More specifically, quantitative genetic variation is the key ingredient for adaptation albeit the least known and hardest to measure (Frankham et al. 2004). Endangered species generally have less genetic diversity than species that are not endangered (Frankham 1995a). One scientist (Weeks et al. 2011) made a distinction between translocations for increasing the fitness of a small population and those to maintain its adaptive potential. A rules commentator suggested making projections about the evolutionary health of the GYE grizzly bears (USFWS 2017, p. 30,609).
Disregarding the need for genetic diversity to support the evolutionary process represents only a short-term, and arguably a shortsighted management perspective. That short-term perspective is typically the need to access and counter the effects of inbreeding. A long-term perspective involves the need to retain evolutionary potential and yet management planning by US Federal land managing agencies is often guided by documents designed for 15–20-year time-spans. This approach is illogical today. Hence, movement corridors or translocations in perpetuity are needed for what Pulsford et al. (2015) called “evolutionary process connectivity.”
Addressing future climate change, for example, demands that we ensure that populations have ample genetic diversity to allow for future genetic modifications (Holt 1990; Hoffman and Sgrò 2011; Beston et al. 2015; Sgrò et al. 2011; Weeks et al. 2011). But we also need to be aware of Holt’s (1990, p. 314) warning: “Predicting the microevolutionary consequences of climate change for even a single species is dauntingly complex.”
Land managers need to incorporate more long-term evolutionary biology concepts into their management strategies (Lankau et al. 2011; Cook and Sgrò 2017a). There is a lack of knowledge among today’s land stewards on evolutionary theory (Smith et al. 2014; Cook and Sgrò 2017b; Taylor et al. 2017; Cook and Sgró 2019).
The global challenges we currently face demand that we do not ignore evolutionary biology principles (Carroll et al. 2014). Such insufficient knowledge was observed in this case study. Understanding the importance of genetics and evolutionary potential is critical to ensure successful biodiversity conservation, and thus we must facilitate collaboration with genetic experts or encourage training opportunities for managers charged with genetic decision making. As Frankel (1974) admonished, our society should “acquire evolutionary responsibility.”
Management implications
The population size goal for the GYE grizzly bear population should be at least the lower threshold value needed to provide ample genetic variation for the long term. This corresponds with a planning framework of hundreds of years (Shaffer 1992; Bader (2000). The continual infusion of new alleles into even a large population will help ensure ample genetic variation and prevent extinction (Allendorf et al. 2013). If grizzly bear dispersal between the GYE and NCDE seems impossible, a translocation strategy may have the same result as maintaining a much larger Nc in one area (Weeks et al. 2011), for example, 2000–5000 bears in the GYE. An Nc of 2000–5000 individuals will likely be sufficient over the long-term assuming a Ne/Nc ratio of around 0.25 (Jamieson and Allendorf 2012; Frankham et al. 2014; Allendorf et al. 2019). However, the USFWS said, “We disagree with the suggestion that there must be 2500 to 5000 grizzly bears throughout the 48 states” (USFWS 2017, p. 30,558). Many scientists cited in this case study disagree with the USFWS on this point, even if one only considers the GYE population.
As of 2018, the IGBST regards the GYE grizzly bear ecological carrying capacity in YNP to be full (Van Manen et al. 2019) which reflects an earlier research judgment (Schwartz et al. 2006). Based on my calculations here, while using the Nc estimates by genetic experts (Nc = 2000–5000 bears), and assuming a carrying capacity of 700 bears (Haroldson et al. 2010), the required number of individuals in the GYE for optimal genetics over the long terms is roughly 3–7 times greater than Haroldson and colleagues’ 700 individuals.
Therefore, the following USFWS statement is decidedly not true: “suitable habitat, including fragmented and unfragmented areas, contains the habitat necessary for a healthy and viable grizzly bear population in the long term” (USFWS 2017, p. 30,588). The proximity of other bear populations and habitat corridors/connectivity are critical components of any long-term conservation strategy. Another rules commentator said what should now be obvious: “connectivity or lack thereof has the potential to impact this population’s [GYE] genetic fitness” (USFWS 2017, p. 30,579). A long-term genetic perspective (i.e., 100 years and more) may sound like an unrealistic “ivory tower” recommendation, but a similar one is needed today for climate change. Hannah and Salm (2005) recommended that protected area management plans for climate change include 30–50 and 80–100-time horizons. As a well-known National Park Service (NPS) historian remarked, NPS “natural resources management seems usually not to have been thought of in truly far-reaching time spans” (Sellars 1997, p. 308).
The periodic infusion of new genetic material through translocation may be our best conservation option for retaining genetic diversity in the GYE grizzly bear population. However, we should nevertheless continue efforts to retain or recreate habitat connectivity between the GYE, NCDE, and the SBFW. As Frankham et al. (2004, p. 134) remarked, “Genetic management of fragmented populations is the greatest unmet challenge in conservation genetics.”
Lest we forget, Frankel (1970) and later Franklin (1980) recommended that our primary management goal should be the recovery of populations of a size that allows evolution by natural selection to continue. The GYE grizzly bear population provides an opportunity to manage these bears relying on such early genetic insight and all that has followed.
To repeat, the following guideline for long-term genetic fitness is based on the advice of some respected conservation geneticists (Jamieson and Allendorf 2012; Frankham et al. 2014; Allendorf et al. 2019): an Nc of 2000–5000 individuals will likely be sufficient over the long-term assuming a Ne/Nc ratio of around 0.25. Land managers need to seek out meta-analyses, consult geneticists regarding periodic regional analyses, or ramp up their own genetic expertise.
Data availability
Any data not in the manuscript are available from the author.
References
Allendorf FW (2016) Genetics and the conservation of natural populations: allozymes to genomes. Mol Ecol 26(2):420–430. https://doi.org/10.1111/mec.13948
Allendorf FW, Leary RF (1986) Heterozygosity and fitness in natural populations of animals. In: Soulé ME (ed) Conservation biology: the science of scarcity and diversity. Sinauer, Sunderland, pp 57–76
Allendorf FW, Ryman N (2002) The role of genetics in population viability. In: Beissinger SR, McCullough DR (eds) Population viability analysis. University of Washington Press, Seattle, pp 50–85
Allendorf FW, Servheen C (1986) Genetics and conservation of grizzly bears. Trends Ecol Evol 1:88–89. https://doi.org/10.1016/0169-5347(86)90030-3
Allendorf FW, Harris RB, Metzgar LH (1991) Estimation of effective population size of grizzly bears by computer simulation. In: Dudley EC (ed) The unity of evolutionary biology. In: Proceedings of the Fourth International Congress of Systematic and Evolutionary Biology. Dioscorides, Portland, pp 650–654
Allendorf FW, Miller CR, Waits LP (2006) Genetics and demography of grizzly bear populations. In: Groom MJ, Meffe GK, Carroll CR (eds) Principles of conservation biology, 3rd edn. Sinauer, Sunderland, pp 404–407
Allendorf FW, Luikart G, Aitken SN (2013) Conservation and the genetics of populations, 2nd edn. Wiley-Blackwell, Oxford
Allendorf FW, Metzgar LH, Horejsi BL, Mattson DJ, Craighead FL (2019) The status of the grizzly bear and conservation of biological diversity in the Northern Rocky Mountains. A compendium of expert statements. Flathead-Lolo-Bitterroot Citizen Task Force, Missoula, Montana. Available at: https://www.montanaforestplan.org/images/in-the-news/The-Status-of-the-Grizzly-Bear-and-Conservation-of-Biological-Diversity-in-the-Northern-Rocky-Mountains-October-2019.pdf. Accessed Dec 2019
Anonymous (2015) Genetic study confirms growth of Yellowstone grizzly bear population. Outdoor News Bulletin 69 (11) November 16, 2015. https://wildlifemanagement.institute/outdoor-news-bulletin/november-2015/genetic-study-confirms-growth-yellowstone-grizzly-bear. Accessed Jan 2020
Ariew A, Lewontin RC (2004) The confusion of fitness. Br J Philos Sci 55:347–263
Avise JC (1994) Molecular markers, natural history and evolution. Chapman Hall, New York
Bader M (2000) Distribution of grizzly bears in the U.S Northern Rockies. Northwest Sci 74:325–334
Balloux F, Amos W, Coulson T (2004) Does heterozygosity estimate inbreeding in real populations? Mol Ecol 13:3021–3031. https://doi.org/10.1111/j.1365-294x.2004.02318.x
Balloux F, Lugon-Moulin N (2002) The estimation of population differentiation with microsatellite markers. Mol Ecol 11:155–165. https://doi.org/10.1046/j.0962-1083.2001.01436.x
Barnovsky AD, Kraatz BP (2007) The role of climatic change in the evolution of mammals. Bioscience 57(6):523–532. https://doi.org/10.1641/B570615
Barton NH, Hewitt GM (1989) Adaptation, speciation and hybrid zones. Nature 341:497–503. https://doi.org/10.1038/341497a0
Beston E, Clobert J, Cole JE (2015) Dispersal response to climate change: scaling down to intraspecific variation. Ecol Lett 18:1226–1233. https://doi.org/10.1111/ele.12502
Boyce MS (1997) Population viability analysis: adaptive management for threatened and endangered species. In: Boyce MS, Hanley A (eds) Ecosystem management: applications for sustainable forest and wildlife resources. Yale University Press, New Haven, pp 226–236
Boyce MS, Knight RR, Blanchard BM, Servheen C (2001) Population viability for grizzly bears: a critical review. Interagency Association of Bear Research and Management Series 34 https://www.researchgate.net/publication/230694923
Bradshaw WE, Holzapel CM (2006) Evolutionary response to rapid climate change. Science 312:1477–1478. https://doi.org/10.1126/science.1127000
Brook BW (2008) Demographics versus genetics in conservation biology. In: Carroll SP, Fox CW (eds) Conservation biology: evolution in action. Oxford University Press, New York, pp 35–49
Caballero A (1994) Developments in the prediction of effective population size. Heredity 73:657–679. https://doi.org/10.1038/hdy.1994.174
Carroll SP, Jørgensen PS, Kinnison MT, Bergstrom CT, Denison RF, Gluckman PT, Smith B, Strauss SY, Tabashnik BE (2014) Applying evolutionary biology to address global challenges. Science 346:e1245993. https://doi.org/10.1126/science.1245993
Caughley G (1994) Directions in conservation biology. J Appl Ecol 63:215–244. https://doi.org/10.2307/5542
Chapman JR, Nakagawa S, Coltman DW, Slate J, Sheldon BC (2009) A quantitative review of heterozygosity-fitness correlations in animal populations. Mol Ecol 18:2746–2765. https://doi.org/10.1111/j.1365-294X.2009.04247.x
Charlesworth B, Charlesworth D, Barton NH (2003) The effects of genetic and geographic structure on neutral variation. Annu Rev Ecol Syst 34:99–125. https://doi.org/10.1038/hdy.1994.174
Christie MR, Knowles LL (2015) Habitat corridors facilitate genetic resistance irrespective of species dispersal abilities or population sizes. Evol Appl 8(5):454–463. https://doi.org/10.1111/eva.12255
Cook CN, Sgrò CM (2017a) Aligning science and policy to achieve evolutionarily enlightened conservation. Conserv Biol 31:501–512. https://doi.org/10.1111/cobi.12863
Cook CN, Sgrò CM (2017b) Understanding managers’ and scientists’ perspectives on opportunities to achieve more evolutionarily enlightened management in conservation. Evol Appl 11:1371–1388. https://doi.org/10.1111/eva.12631
Cook CN, Sgró CM (2019) Conservation practitioners’ understanding of how to manage evolutionary processes. Conserv Biol 33:993–1001. https://doi.org/10.1111/cobi.13306
Craighead L, Gilbert B, Olenicki T (2006) Comments submitted to the US Fish and Wildlife Service regarding delisting of the Yellowstone Grizzly Bear DPS. Fed Reg 70:69853–69884. www.craigheadresearch.org/uploads/7/6/9/0/7690832/craighead_gilbert_olenicki_final_comments.pdf. Accessed Apr 2022
Crnokrak P, Roff DA (1999) Inbreeding depression in the wild. Heredity 75:530–540. https://doi.org/10.1038/sj.hdy.6885530
David P (1998) Heterozygosity–fitness correlations: new perspectives on old problems. Heredity 80:531–537. https://doi.org/10.1046/j.1365-2540.1998.00393.x
De Barba M, Waits LP, Garton EO, Genovesi P, Randi E, Mustoni A, Groff C (2010) The power of genetic monitoring for studying demography, ecology and genetics of a reintroduced brown bear population. Mol ecol 19(18):3938–3951. https://doi.org/10.1111/j.1365-294X.2010.04791.x
Denniston CD (1978) Small population size and genetic diversity: implications for endangered species. In: Temple SA (ed) Endangered birds: management techniques for preserving threatened species. University of Wisconsin Press, Madison, pp 281–289
DeWoody JA, Harder AM, Mathur S, Willoughby JR (2021) The long-standing significance of genetic diversity in conservation. Mol Ecol 30(17):4147–4154. https://doi.org/10.1111/mec.16051
Dobzhansky T (1948) Genetics of natural populations. XVIII. Experiments on chromosomes of Drosophila pseudoobscura from different geographic regions. Genetics 33:588–602
Edmunds S (2007) Between a rock and a hard place: evaluating the relative risks of inbreeding and outbreeding for conservation and management. Mol Ecol 16:463–475. https://doi.org/10.1111/j.1365-294X.2006.03148.x
Ewens WJ (1990) Population genetics theory-the past and the future. In: Lessard S (ed) Mathematical and statistical developments of evolutionary theory, vol 299. NATO Science Series, pp 177–227
Fisher RA (1930) The genetical theory of natural selection. Clarendon Press, Oxford
Frankel OH (1970) Variation–the essence of life. Sir William Macleay memorial lecture. P Linn Soc NSW 5:158–169
Frankel OH (1974) Genetic conservation: our evolutionary responsibility. Genetics 78:53–65
Frankel OH, Soulé ME (1981) Conservation and evolution. Cambridge University Press, New York
Frankham R (1995a) Conservation genetics. Annu Rev Genet 29:305–327. https://doi.org/10.1146/annurev.ge.29.120195.001513
Frankham R (1995b) Inbreeding and extinction: a threshold effect. Conserv Biol 9:792–799. https://doi.org/10.1046/j.1523-1739.1995.09040792.x
Frankham R (1995c) Effective population size/adult population size ratios in wildlife: a review. Genet Res 66:97–109. https://doi.org/10.1017/s00166
Frankham R (1996) Relationship of genetic variation to population size in wildlife. Conserv Biol 10:1500–1508. https://doi.org/10.1046/j.1523-1739.1996.10061500.x
Frankham R (2005) Genetics and extinction. Biol Conserv 126:131–140. https://doi.org/10.1016/j.biocon.2005.05.002
Frankham R (2010) Challenges and opportunities of genetic approaches to biological conservation. Biol Conserv 143:1919–1927. https://doi.org/10.1016/j.biocon.2010.05.011
Frankham R, Lees K, Montgomery ME, England PR, Lowe E, Briscoe DA (1999) Do population bottlenecks reduce evolutionary potential? Anim Conserv 2:255–260. https://doi.org/10.1111/j.1469-1795.1999.tb00071.x
Frankham R, Ballou JD, Briscoe DA (2004) A primer of conservation genetics. Cambridge University Press, New York
Frankham R, Ballou JD, Briscoe DA (2010) Introduction to conservation genetics, 2nd edn. Cambridge, New York
Frankham R, Ballou JD, Eldridge MDB, Lacy RC, Ralls K, Dudash MR, Fenster CB, Gulinck H, Matthysen E (2011) Predicting the probability of outbreeding depression. Conserv Biol 25:465–475. https://doi.org/10.1111/j.1523-1739.2011.01662.x
Frankham R, Bradshaw CJA, Brook BW (2014) Genetics in conservation management: revised recommendations for the 50/500 rules, Red List criteria and population viability analyses. Biol Conserv 170:56–63. https://doi.org/10.1016/j.biocon.2013.12.036
Frankham R, Ballou JD, Ralls K, Eldridge MDB, Dudash MR, Fenster B, Lacy RC, Sunnucks P (2017) Genetic management of fragmented animal and plant populations. Oxford University Press, Oxford
Franklin IR (1980) Evolutionary change in small populations. In: Soulé ME, Wilcox BA (eds) Conservation biology: an evolutionary-ecological perspective. Sinauer Associates, Sunderland, pp 135–149
Franklin IR, Frankham R (1998) How large must populations be to retain evolutionary potential. Anim Conserv 1:69–70. https://doi.org/10.1111/j.1469-1795.1998.tb00228.x
Franklin IR, Allendorf FW, Jamieson IG (2014) The 50/500 rule is still valid-Reply to Frankham et al. Biol Conserv. https://doi.org/10.1016/j.biocon.2014.05.004
Fuerst PA, Maruyama T (1986) Considerations on the conservation of alleles and of genic heterozygosity in small populations. Zoo Biol 5:171–179. https://doi.org/10.1002/zoo.1430050211
Georghiou GP (1986) The magnitude of the resistance problem. In: Georghiou P, Taylor CE (eds) Pesticide resistance: strategies and tactics for management. National Academy Press, Washington, pp 14–43
Gilbert-Norton L, Wilson R, Stevens JR, Beard KH (2010) A meta-analytic review of corridor effectiveness. Conserv Biol 24:660–668. https://doi.org/10.1111/j.1523-1739.2010.01450.x
Gilpin ME, Soulé ME (1986) Minimum viable populations: processes of species extinction. In: Soulé ME (ed) Conservation biology: the science of scarcity and diversity. Sinauer Associates, Sunderland, pp 19–34
Greater Yellowstone Coalition v. Christopher Servheen (665 F. 3d 1015 9th Cir. 2011, p 33)
Gregorious HR (1991) On the concept of effective number. Theoret Popul Biol 40:269–283. https://doi.org/10.1016/0040-5809(91)90056-1
Groom MJ, Meffe GK, Carrol CR (2006) Principles of conservation biology, 3rd edn. Sinauer Associates, Sunderland
Hannah L, Salm R (2005) Protected areas management in a changing climate. In: Lovejoy TE, Hannah L (eds) Climate change and biodiversity. Yale University Press, New Haven, pp 363–371
Hansson B, Westerberg L (2002) On the correlation between heterozygosity and fitness in natural populations. Mol Ecol 11:2467–2474. https://doi.org/10.1046/j.1365-294X.2002.01644.x
Hare M, Nunney L, Schwartz MK, Ruzzante DE, Burford M, Waples RS, Ruegg K, Spira T (2011) Understanding and estimating effective population size for practical application in marine species management. Conserv Biol 25:438–449. https://doi.org/10.1111/j.1523-1739.2010.01637.x
Haroldson MA, Schwartz CC, Kendall KC, Gunther KA, Moody DS, Frey KD, Paetkau D (2010) Genetic analysis of individual origins supports isolation of grizzly bears in the Greater Yellowstone ecosystem. Ursus 21:1–13. https://doi.org/10.2192/09gr022.1
Harris RB, Allendorf FW (1989) Genetically effective population size of large mammals: an assessment of estimators. Conserv Biol 3:181–191. https://doi.org/10.1111/j.1523-1739.1989.tb00070.x
Hoffmann AA, Sgrò CM (2011) Climate change and evolutionary adaptation. Nature 470:479–485. https://doi.org/10.1038/nature09670
Holt RD (1990) The microevolutionary consequences of climate change. Trends Ecol Evol 5:311–315. https://doi.org/10.1016/0169-5347(90)90088-U
Jamieson IG (2009) Loss of genetic diversity and inbreeding in New Zealand’s threatened bird species. Science for Conservation 293, Department of Conservation, Wellington
Jamieson IG, Allendorf FW (2012) How does the 50/500 rule apply to MVPs? Trends Ecol Evol 27:578–584. https://doi.org/10.1016/j.tree.2012.07.001
Kamath L, Haroldson MA, Luikart G, Paetkau D, Whitman C, Van Manen FT (2015) Multiple estimates of effective population size for monitoring a long-lived vertebrate: an application to Yellowstone grizzly bears. Mol Ecol 24:5507–5521. https://doi.org/10.1111/mec.13398
Keller LF, Waller DM (2002) Inbreeding effects in wild populations. Trends Ecol Evol 17:230–241. https://doi.org/10.1016/S0169-5347(02)02489-8
Keyghobadi N (2007) The genetic implications of habitat fragmentation for animals. Can J Zool 85:1049–1064. https://doi.org/10.1139/Z07-095
Lacy RC (1987) Loss of genetic diversity from managed populations: interacting effects of drift, mutation, immigration, selection, and population subdivision. Conserv biol 1(2):143–158
Lacy RC (1992) The effects of inbreeding on isolated populations. Are minimum viable population sizes predictable? In: Fieldler PL, Jain SK (eds) The theory and practice of nature conservation preservation and management. Chapman and Hall, New York, pp 277–296
Lacy RC (1997) Importance of genetic variation to the viability of mammalian populations. J Mamm 78:320–335. https://doi.org/10.2307/1382885
Laidre L, Andrén RH, Larsson O, Ryman N (1996) Inbreeding depression in brown bears. Biol Conserv 76:69–72. https://doi.org/10.1016/0006-3207(95)00084-4
Lamb CT, Ford AT, Proctor MF, Royle JA, Mowat G, Boutin S (2019) Genetic tagging in the Anthropocene: scaling ecology from alleles to ecosystems. Ecol Appl 29(4):e01876. https://doi.org/10.1002/bes2.1540
Lande R (1976) The maintenance of genetic variability by mutation in a polygenic character with mixed loci. Genet Res 26:221–235. https://doi.org/10.1017/s001667230001603
Lande R (1988) Genetics and demography in biological conservation. Science 241:1455–1460. https://doi.org/10.1126/science.3420403
Lande R (1995) Mutation and conservation. Conserv Biol 9:782–791. https://doi.org/10.1046/j.1523-1739.1995.09040782.x
Lankau R, Jørgensen PS, Harris DJ, Sih A (2011) Incorporating evolutionary principles into environmental management and policy. Evol Appl 4:315–325. https://doi.org/10.1111/j.1752-4571.2010.00171.x
Leberg P (2005) Genetic approaches for estimating the effective size of populations. J Wildl Manag 69:1385–1399. https://www.jstor.org/stable/380350
Lindenmayer DB, Fischer J (2006) Habitat fragmentation and landscape change: an ecological and conservation synthesis. Island Press, Washington
Linløkken AN (2018) Genetic diversity of small populations. In: Liu K (ed) Genetic diversity and disease susceptibility. IntechOpen. https://doi.org/10.5772/intechopen.7692343-56
Lino A, Fonseca C, Rojas D, Fischer E, Pereira MJR (2019) A meta-analysis of the effects of habitat loss and fragmentation on genetic diversity in mammals. Mamm Biol 94:69–76. https://doi.org/10.1016/j.mambio.2018.09.006
Love Stowell SM, Pinzone CA, Martin AP (2017) Overcoming barriers to active interventions for genetic diversity. Biodiv Conserv 26:753–1765. https://doi.org/10.1007/s10531-017-1330-z
Luikart G, Ryman N, Tallmon DA, Schwartz MK, Allendorf FW (2010) Estimation of census and effective population sizes: the increasing usefulness of DNA-based approaches. Conserv Genet 11:355–373. https://doi.org/10.1007/s10592-010-0050-7
Lynch M (1996) Quantitative genetics in conservation. In: Avise JC, Hamrick JL (eds) Conservation genetics: case histories from nature. Chapman and Hall, New York, pp 471–501
Lynch M, Lande R (1998) The critical effective size for a genetically secure population. Anim Conserv 1:70–72. https://doi.org/10.1111/j.1469-1795.1998.tb00229.x
Lynch M, Conery J, Bürger R (1995) Mutational meltdown in sexual populations. Evolution 49:1067–1080. https://doi.org/10.2307/2410432
McKay JK, Latta RG (2002) Adaptive population divergence: markers, QTL and traits. Trends Ecol Evol 17:285–291. https://doi.org/10.1016/S0169-5347(02)02478-3
Méndez M, Vögeli M, Tella JL, Godoy JA (2014) Joint effects of population size and isolation on genetic erosion in fragmented populations: finding fragmentation thresholds for management. Evol Appl 7:506–518. https://doi.org/10.1111/eva.12154
Miller CR, Waits L (2003) The history of effective population size and genetic diversity in the Yellowstone grizzly (Ursus arctos): implications for conservation. Proc Natl Acad Sci USA 100:4334–4339. https://doi.org/10.1073/pnas.0735531100
Mills LS, Allendorf FW (1996) The one-migrant-per-generation rule in conservation and management. Conserv Biol 10:1509–1518. https://doi.org/10.1046/j.1523-1739.1996.10061509.x
Mitton JB (1993) Theory and data pertinent to the relationship between heterozygosity and fitness. In: Thornhill NW (ed) The natural history of inbreeding and outbreeding: theoretical and empirical perspectives. University of Chicago Press, Chicago, pp 17–41
Newmark WD (1985) Legal and biotic boundaries of western North American national parks: a problem of congruence. Biol Conserv 33:197–208. https://doi.org/10.1016/0006-3207(85)90013-8
Nunney L, Campbell KA (1993) Assessing minimum viable population size: demography meets population genetics. Trends Ecol Evol 8:234–239. https://doi.org/10.1016/0169-5347(93)90197-W
Paetkau DH, Waits LP, Clarkson PL, Craighead L, Vyse E, Strobeck C (1998) Variation in genetic diversity across the range of North American brown bears. Conserv Biol 12:418–429. https://doi.org/10.1111/j.1523-1739.1998.96457.x
Paige KN (2017) Conservation genetics: founding principles, primary concerns. In: Brawn J, Meine C, Robinson S (eds) Foundations of conservation biology. University of Chicago Press, Chicago
Palstra F, Fraser DJ (2012) Effective/census population size ratio estimation: compendium and appraisal. Ecol Evol 2:2357–2365. https://doi.org/10.1002/ece3.329
Palstra F, Ruzzante DE (2008) Genetic estimates of contemporary effective population size: what can they tell us about the importance of genetic stochasticity for wild population persistence? Mol Ecol 17(5):3428–3447. https://doi.org/10.1111/j.1365-294X.2008.03842.x
Petit RJ, Mousadik AE, Pons O (1996) Identifying populations for conservation on the basis of genetic markers. Conserv Biol 12(4):844–855. https://doi.org/10.1111/j.1523-1739.1998.96489.x
Proctor MF, McLellan BN, Strobeck C, Barclay RMR (2005) Genetic analysis reveals demographic fragmentation of grizzly bears yielding vulnerably small populations. Proc Biol Sci 272(1579):2404–2419. https://doi.org/10.1098/rspb.2005.3246
Proctor MF et al (2012) Population fragmentation and inter-ecosystem movements of grizzly bears in Western Canada and the Northern United States. Wildl Monogr 180:1–46. https://doi.org/10.1002/wmon.6
Pulsford I, Lindenmayer D, Wyborn C, Lausche B, Worboys GL, Vasilijević M, Lefroy T (2015) Connectivity conservation management. In: Worboys GL, Lockwood MA, Kothari A, Feary S, Pulsford I (eds) Protected area governance and management. ANU, Canberra, pp 851–888
Putman AI, Carbone I (2014) Challenges in analysis and interpretation of microsatellite data for population genetic studies. Ecol Evol 4:4399–4428. https://doi.org/10.1002/ece3.1305
Ralls K, Ballou JD (1983) Extinction: lessons from zoos. In: Schonewald-Cox CM, Chambers SM, MacBryde B, Thomas WL (eds) Genetics and conservation: a reference for managing wild animal and plant populations. Benjamin/Commins, Menlo Park, pp 164–184
Reed DH (2005) Relationship between population size and fitness. Conserv Biol 19:563–568. https://doi.org/10.1111/j.1523-1739.2005.00444.x
Reed DH (2010) Albatrosses, eagles and newts, oh my! Exceptions to the prevailing paradigm concerning genetic diversity and population viability? Anim Conserv 13:448–457. https://doi.org/10.1111/j.1469-1795.2010.00353.x
Reed DH, Frankham R (2001) How closely correlated are molecular markers and quantitative measures of genetic variation? Evolution 55(6):1095–1103. https://doi.org/10.1111/j.0014-3820.2001.tb00629.x
Reed DH, O’Grady JJ, Brook BW, Ballou JD, Frankham R (2003) Estimates of minimum viable population sizes for vertebrates and factors influencing those estimates. Biol Conserv 113:23–34. https://doi.org/10.1016/S0006-3207(02)00346-4
Resasco J (2019) Meta-analysis on a decade of testing corridor efficacy: what have we learned? Current Landsc Ecol Rep 4:61–69. https://doi.org/10.1007/s40823-019-00041-9
Rice KJ, Emery NC (2003) Managing microevolution: restoration in the face of global change. Front Ecol Environ 1(9):469–478. https://doi.org/10.1890/1540-9295(2003)001
Scheffers BR et al (2016) The broad footprint of climate change from genes to biomes to people. Science 354(6313):Aaf7671. https://doi.org/10.1126/science.Aaf7671
Schlaepfer DH, Braschler B, Rusterholz H-P, Bauer B (2018) Genetic effects of anthropogenic habitat fragmentation on remnant animal and plant populations: a meta-analysis. Ecosphere 9(10):e02488. https://doi.org/10.1002/ecs2.2488
Schlesinger WH (2010) Translational ecology. Science 329:609. https://doi.org/10.1126/science.1195624
Schonewald-Cox CM, Chambers SM, McBride B, Thomas WL (eds) (1983) Genetics and conservation: a reference for managing wild animal and plant populations. Benjamin-Cummins, Menlo Park
Schwartz CC, Haroldson MA, White GC, Harris RB, Cherry S, Keating KA, Moody D, Servheen C (2006) Temporal, spatial, and environmental influences on the demographics of grizzly bears in the Greater Yellowstone Ecosystem. Wild Monogr 161(1):1–8. https://doi.org/10.2193/0084-0173(2006)161[1:TSAEIO]2.0.CO;2
Sellars RW (1997) Preserving nature in the national parks: a history. With a new preface and epilogue. Yale University Press, New Haven
Servheen C, Waller JS, Sandstrom P 2002 Identification and management of linkage zones for grizzly bears between large blocks of public land in the Northern Rocky Mountains. In: Irwin CL, Garrett P, McDermott K (eds) Proceedings of the 2001 International Conference on Ecology and Transportation, Center for Transportation and Environment, North Carolina State University, pp 161–179. http://repositories.cdlib.org/jmie/roadeco/Servheen2001a. Accessed July 2021.
Sgró CM, Lowe AJ, Hoffman AA (2011) Building evolutionary resilience for conserving biodiversity under climate change. Evol Appl 4:326–337. https://doi.org/10.1111/j.1752-4571.2010.00157
Shafer CL (1990) Nature reserves: Island theory and conservation practice. Smithsonian Institution Press, Washington
Shafer CL (2001) Inter-reserve distance. Biol Conserv 100:215–227. https://doi.org/10.1016/S0006-3207(01)00025-8
Shafer CL (2013) Grizzly bear emigration and land use: an interdisciplinary case study of the greater yellowstone ecosystem [Dissertation] George Mason University, Fairfax, Virginia
Shaffer ML (1992) Keeping the grizzly bear in the American West. The Wilderness Society, Washington
Sinclair ARE, Fryxell JM, Caughley G (2006) Wildlife ecology, conservation, and management, 2nd edn. Blackwell, Oxford
Smith TB, Kinnison MT, Strauss SY, Fuller TL, Carroll S (2014) Prescriptive evolution to conserve and manage biodiversity. Annu Rev Ecol Evol Syst 45:1–22. https://doi.org/10.1146/annurev-ecolsys-120213-091747
Soulé ME (1980) Thresholds for survival: maintaining fitness and evolutionary potential. In: Soulé ME, Wilcox BA (eds) Conservation biology: an evolutionary-ecological perspective. Sinauer Associates, Sunderland, pp 151–169
Soulé ME (1987) Where do we go from here? In: Soulé ME (ed) Viable populations for conservation. Cambridge University Press, New York. https://doi.org/10.1017/cbo9780511623400.011
Spielman D, Brook BW, Frankham R (2004) Most species are not driven to extinction before genetic factors impact them. Proc Natl Acad Sci USA 101:15261–15264. https://doi.org/10.1073/pnas.0403809101
Storfer A (1996) Quantitative genetics: a promising approach for the assessment of genetic variation in endangered species. Trends Ecol Evol 11:343–348. https://doi.org/10.1016/0169-5347(96)20051-5
Storfer A (1999) Gene flow and endangered species translocations: a topic revisited. Biol Conserv 87:173–180. https://doi.org/10.1016/s0006-3207(98)00066-4
Tallmon DA, Luikart G, Waples RS (2004) The alluring simplicity and complex reality of genetic rescue. Trends Ecol Evol 19:489–496. https://doi.org/10.1016/j.tree.2004.07.003
Taylor HR, Dussex N, van Heezik Y (2017) Bridging the conservation genetics gap by identifying barriers to implementation for conservation practitioners. Global Ecol Conserv 10:231–242. https://doi.org/10.1016/j.gecco.2017.04.001
Templeton AR (1986) Coadaptation and outbreeding depression. In: Soulé ME (ed) Conservation biology: the science of scarcity and diversity. Sinauer, Sunderland, pp 105–111
Trail LW, Bradshaw CJA, Brook BW (2007) Minimum viable population size: a meta-analysis of 30 years of published estimates. Biol Conserv 139:159–166. https://doi.org/10.1016/j.biocon.2007.06.011
United Nations Educational, Scientific and Cultural Organization (UNESCO) 2011 State party’s report on the state of conservation of its property inscribed on the World Heritage List. Expert meeting on the global state of conservation challenges for World Heritage properties 13–15 April 2011. https://whc.unesco.org/en/events/740. Accessed Jan 2013
U.S. Fish and Wildlife Service (2007) Final rule designating the Greater Yellowstone Area population of grizzly bears as a distinct population segment; Removing the Yellowstone distinct population segment of grizzly bears from the Federal List of Endangered and Threatened Wildlife. Fed Reg 72:14866–14938. https://doi.org/10.5962/bhl.title.115837
U.S. Fish and Wildlife Service (2016) US endangered and threatened wildlife and plants; Removing the Greater Yellowstone Ecosystem population of grizzly bears from the federal list of endangered and threatened wildlife; Proposed rule. Fed Reg 81:13174–13227. https://www.govinfo.gov/cont-30/pdf/2017-13160.pdf
U.S. Fish and Wildlife Service (2017) Endangered and threatened wildlife and plants; Removing the Greater Yellowstone Ecosystem population of grizzly bears from the federal list of endangered and threatened wildlife. Fed Reg 82:30502–30633. https://www.govinfo.gov.content/pkg/FR-2017-06-30/pdf/2017-13160.pdf
U.S. Fish and Wildlife Service (2021) Biological report for the grizzly bear (Ursus arctos horribilis) in the Lower-48 states. Version 1.1. January 31, 2021, Montana. https://www.fws.gov/mountain-prairie/es/species/mammals/grizzly/20210131_V1.1_SSA_for_grizzly_bear_in_the_lower-48_States.pdf
U.S. Forest Service (2006) Forest plan amendment for grizzly bear habitat conservation in the Greater Yellowstone Area national forests, final environmental impact statement, U.S. Forest Service, Washington, D.C. https://www.fs.usda.gov/Internet/FSE_DOCUMENTS/stelprdb5187773.pdf. Accessed July 2021
U.S. Geological Survey (2015) Genetic study confirms growth of Yellowstone grizzly bear population, October 30. https://www.usgs.gov/news/genetic-study-confirms-growth-yellowstone-grizzly-bear-population. Accessed Oct 2016
Van Manen FT, Haroldson MA, Karabensh BE (2019) Yellowstone grizzly bear investigations: annual report of the Interagency Grizzly Bear Study Team, U.S. Geological Survey, Bozeman, Montana 2018. https://prd-wret.s3.us-west-2.amazonaws.com/assets/palladium/production/atoms/files/2018_AnnualReport_FINAL%28508%29.pdf. Accessed Jan 2020
Varvio S, Chakraborty R, Nei M (1986) Genetic variation in subdivided populations and conservation genetics. Heredity 57:189–198. https://doi.org/10.1038/hdy.1986.109
Vergeer P, Ouborg NJ, Hendry AP (2018) Genetic considerations of introduction efforts. In: Carroll SP, Fox CW (eds) Conservation biology: evolution in action. Oxford University Press, New York, pp 116–129
Vilá C et al (2003) Rescue of a severely bottlenecked wolf (Canis lupus) population by a single immigrant. Philos T Roy Soc B 270:91–97
Vucetich JA, Waite TA (2000) Is one migrant per generation sufficient for the genetic management of fluctuating populations? Anim Conserv 3:261–266. https://doi.org/10.1111/j.1469-1795.2000.tb00111.x
Vucetich JA, Waite TA, Nunney L (1997) Fluctuating population size and the ratio of effective to census population size. Evolution 51:2017–2021. https://doi.org/10.2307/2411022
Wahlund S (1929) Zusammensetzung von population und korrelationserscheinung vom stand-punkt der vererbungslehre aus betrachtet. Hereditas 11:65–106. https://doi.org/10.1111/j.1601-5223.1928.tb02483.x
Waits LL, Paetkau C, Strobeck C, Ward RH (1998) A comparison of genetic variability in brown bears (Ursus arctos) from Alaska, Canada, and the lower 48 states. Ursus 10:307–314
Wang J (1984) Application of the one-migrant-per-generation rule to conservation and management. Conserv Biol 18(2):332–342. https://doi.org/10.1111/j.1523-1739.2004.00440.x
Wang J (2005) Estimation of effective population size from data on genetic markers. Phil T Roy Soc B 360(1459):1395–1409. https://doi.org/10.1098/rstb.2005.1682
Wang J, Brekke P, Huchard E, Knapp LA, Cowlishaw G (2010) Estimation of parameters of inbreeding and genetic drift in populations with overlapping generations. Evolution 64:1704–1718. https://doi.org/10.1111/j.1558-5646.2010.00953.x
Weeks AR et al (2011) Assessing the benefits and risks of translocations in changing environments: a genetic perspective. Evol Appl 4:709–725. https://doi.org/10.1111/j.1752-4571.2011.00192.x
Westemeier RL et al (1998) Tracking the long-term decline and recovery of an isolated population. Science 282(5394):1695–1698. https://doi.org/10.1126/science.282.5394.1695
White PJ, Gunther KA, Van Manen FT (2017) Yellowstone grizzly bears: ecology and conservation of an icon of wildness. Yellowstone National Park, Yellowstone Forever and U.S. Geological Survey, Northern Rocky Mountain Science Center, Montana
Whiteley AR, Fitzpatrick SW, Funk WC, Tallmon DA (2015) Genetic rescue to the rescue. Trends ecol evol 30(1):42–49. https://doi.org/10.1016/j.tree.2014.10.009
Whitlock MC, Barton NH (1997) The effective size of a subdivided population. Genetics 146:427–441. https://doi.org/10.1093/genetics/146.1.427
Wilcox BA (1986) Extinction models and conservation. Trends Ecol Evol 1:47–48. https://doi.org/10.1016/0169-5347(86)90073-X
Wright S (1931) Evolution in Mendelian populations. Genetics 16:97–159
Wright S (1977) Evolution and the genetics of populations. University of Chicago Press, Chicago
Young AG, Clarke GM (2000) Genetics, demography and viability of fragmented populations. Cambridge University Press, New York
Acknowledgements
J. A. Shafer provided feedback on all paper drafts. C. M. Schonewald was a sounding board for many of my ideas. I thank my university committee for support: E.C. Parsons, L. M. Talbot, T. E. Lovejoy, and J. C. Kozlowski. Other university professors and classmates commented on the paper: V. A. Buckley-Beason, R. G. Kroner, F. B. Herta, and C.L. Romelo. I thank J. D. Ballou for reading an earlier version of the manuscript. Many anonymous reviewers provided helpful comments on the paper and my gratitude to them is profound.
Funding
No funding.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The author(s) declare that they have no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Appendix: Elaboration on four genetic concepts
Appendix: Elaboration on four genetic concepts
Genetic drift
Genetic variation is maintained in a population due to the interaction of four factors: gene flow, mutation, genetic drift, and natural selection (Lacy 1987). A concern not emphasized by USFWS is the impact of genetic drift on the loss of genetic diversity. Genetic drift can overcome natural selection and result in the slow assembly of deleterious mutations (Hare et al. 2011). This is called genetic load (Wallace 1970). If mutation continues, it is referred to as mutational meltdown (Lynch et al. 1995). Theory predicts that allelic diversity in small populations will be lost at the rate of half the Ne per generation (Wright 1931), but must be accompanied by numerous unrealistic assumptions about the population. The effect of inbreeding in small populations occurs more slowly than does the impact of genetic drift (Leberg 2005), nor does inbreeding result in a loss of alleles (Jamieson 2009). Allendorf (1986) agreed that P is more significant for future evolutionary potential than is H. Gene flow remains necessary to support genetic diversity for long-term adaptation. Natural selection is probably happening and at a very much reduced rate in the GYE population since it is so small. Whatever genetic changes happen to the population will be due to genetic drift—random changes—not natural selection. This would be an argument for translocation sooner rather than later (J. Ballou, personal communication, November 2016). The fixation of alleles through genetic drift could manifest even in populations of moderate Ne (Avise 1994) or for populations of any size (Allendorf et al. 2013). As Allendorf and Ryman (2002) explained, inbreeding is not always essential to cause a loss in fitness. However, genetic drift can cause alleles to be lost and reduce H (Leberg 2005).
Translocation
When and if grizzly bear translocations (a form of genetic rescue) are conducted by the USFWS, approximately 1–10 migrants per generation who successfully breed will be needed to compensate for any genetic variation loss due to small population size (Mills and Allendorf 1996) which Wang (1984) supported. An earlier 1986 USFWS workshop (Allendorf and Servheen 1986) recommended at least one migrant per generation. Some workers indicate that more than ten individuals may be needed (Vucetich and Waite 2000). At this level of migration, the loss of P and H will be minimal while still allowing for intrapopulation divergence in allele frequencies. After an isolated population of greater prairie chicken (Tympanuchus cupido) declined from 2000 in 1962 to < 50 by 1994, genetic diversity and egg production dropped. But after birds from a large genetically diverse population were translocated into the remnant population, egg viability returned (Westemeier et al. 1998). A single wolf (Canis lupus) immigrant infused genetic diversity into a genetically depauperate population (Vilá et al. 2003).
The migration of one or more breeding individuals per generation can achieve a balance between the four genetic diversity factors (gene flow, mutation, genetic drift, and natural selection) whatever the size of the receiving population. This counter-intuitive result indicates that integration of new adaptive alleles into the receiving population’s gene pool will occur rather quickly (Barton and Hewitt 1989), though Varvio et al. (1986) warned that it may take decades. Translocation is required to maintain the genetic diversity of the GYE bears and to allow for possible adaptation to climate change and other factors that may threaten their long-term survival. Thus, the purpose of translocations is to increase genetic diversity and allow for adaptation (Weeks et al. 2011).
Genetic fitness
The USFWS believes that the GYE grizzly bear population is fit due to observed normal litter sizes, no apparent evidence of disease, high rates of survivorship, an equal sex ratio, normal body size and physical characteristics, and a stable-to-increasing population size (USFWS 2017, p. 30535). However, such USFWS observations of perceived bear fitness do not ensure that the population is not weakened as a result of reduced genetic variation. For example, some subtle adaptations would be difficult to observe such as behavioral changes, more hemoglobin to compensate for increasing altitude, or new enzymes to adjust to for different diets (Frankham et al. 2017). Fitness has been defined as including survival, disease resistance, growth and developmental rate, and developmental stability (Allendorf and Leary 1986). But there is disagreement by other scientists on what constitutes fitness (Ariew and Lewontin 2004). Furthermore, the expression of survival abnormalities due to genetic impoverishment can manifest long after genetic changes occur (Lacy 1997). Thus, there is a need for genetic monitoring to measure changes in H and sub-population introgression (De Barba et al. 2010) which the USFWS is doing using genetic tags like described by Lamb et al. (2019).
However, some scientists are skeptical of the relationship between H and fitness (Lynch 1996; Paige 2017). A meta-analysis by Reed et al. (2003) showed that the relationship explained only 15–20% of variation in fitness. Another animal meta-analysis by Chapman et al. (2009) revealed only a small positive relationship. Other results were inconsistent (David 1998; Hansson and Westberg 2002). Correlations between heterozygosity and inbreeding were strong only when around 200 loci were examined (Balloux et al. 2004). In spite of such results suggesting a declining positive relationship, DeWoody et al. (2021) emphatically argued that genetic diversity is tied to overall fitness. Some populations have survived for hundreds of generations with low genetic variation (Reed 2010), but they lack the alleles for adaptation (Linløkken 2018). Population bottlenecks often lose rare alleles (Denniston 1978). Bottlenecks of short duration may have little influence on heterozygosity but can cause a severe reduction in alleles (Allendorf 1986).
Outbreeding depression
The concept of outbreeding was first described by Dobzhansky (1948). A genetic dilemma remains because we do not always know whether translocation is the best strategy for sustaining genetic variation. Although a population can benefit from the infusion of genetic diversity, the introduction of new alleles can also dilute existing local adaptations and taxonomic integrity and result in outbreeding depression. Genetic swamping can occur when locally adapted genes are disrupted by gene flow (Shafer 1990, 2001; Storfer 1999; Tallmon et al. 2004). Weeks et al. (2011) developed an outline to assist in decision-making. The dilemma described resulted in one scientist concluding that “no unassailable generalizations are possible” (Vergeer et al. 2018), while another described the conflict as “being between a rock and a hard place” (Edmunds 2007, p. 463). This long-standing problem warrants further research (Whiteley et al. 2015). Nevertheless, according to Frankham (2010) and Love Stowell et al. (2017), the fear of diluting the genetic integrity of a population or species through outbreeding depression could provide land stewards with an excuse for avoiding genetic management actions whose outcomes were uncertain. In situations where the probability of outbreeding depression was low, when inbred populations were outcrossed there was a positive genetic outcome in 92.9% of the 156 cases reviewed (Frankham et al. 2011). Hence, the fear of outbreeding depression appears to have been overstated (Hoffmann and Sgrò 2011).
As a result of their review, Frankham et al. (2011) suggested that if two populations are genetically similar, occupy similar habitats, and have been isolated for less than 500 years, then the probability of outbreeding depression is minor. One laboratory experiment illustrated that the negative effect of losing genetic diversity was not due to inbreeding, but rather was based on a decreased ability to adapt (Frankham et al. 1999). Therefore, the probability of harmful outbreeding depression occurring after translocating grizzly bears between the GYE and NCDE is not the concern that Templeton (1986), for example, thought it was for species in general. Outbreeding depression is often temporary (Frankham et al. 2017).
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Shafer, C.L. A greater yellowstone ecosystem grizzly bear case study: genetic reassessment for managers. Conservation Genet Resour 14, 331–345 (2022). https://doi.org/10.1007/s12686-022-01262-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12686-022-01262-7