Abstract
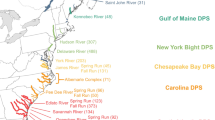
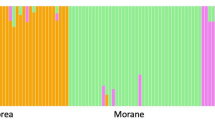
This study provides the first standardized global microsatellite database for a shark species, the tiger shark (Galeocerdo cuvier). Genotyping of reference individuals was used to develop and apply a calibration key for data from eight microsatellite loci data produced by three different laboratories, thereby allowing merging of genotypes into a single dataset. The unified data helped to elucidate the global population structure of the species and provided improved statistical power, through allowing a higher number of samples per location compared to the original studies from which the samples were obtained. Pairwise FST estimates and PCA plots showed significant genetic differentiation between Atlantic and Indo-Pacific samples, confirming previous findings by identifying the presence of a strong genetic break between tiger sharks inhabiting the two ocean basins. In turn, the standardized database (n = 799) also allowed archived historical samples to be genotyped and assigned back to their population (ocean basin) of origin. We demonstrate how calibration tests in population structure studies using microsatellite data is important as it simply provides more data to single studies. Importance factors for successful assignment analysis is discussed, as well as how the possibility of assigning historical samples of unknown origin back to the population, increases sample value. Our results demonstrate that global calibration of microsatellite and other genetic datasets can improve the statistical power and resolution of population structure analysis; an approach applicable not only when working with highly mobile globally distributed species such as the tiger shark, but with any species for which multiple genetic datasets exist.



Similar content being viewed by others
References
Adamack AT, Gruber B (2014) PopGenReport: simplifying basic population genetic analyses in R. Methods Ecol Evol 5(4):384–387. https://doi.org/10.1111/2041-210X.12158
Bernard AM, Feldheim KA, Shivji MS (2015) Isolation and characterization of polymorphic microsatellite markers from a globally distributed marine apex predator, the tiger shark (Galeocerdo cuvier). Conserv Genet Resour 7(2):509–511. https://doi.org/10.1007/s12686-014-0408-0
Bernard AM, Feldheim KA, Heithaus MR, Wintner SP, Wetherbee BM, Shivji MS (2016) Global population genetic dynamics of a highly migratory, apex predator shark. Mol Ecol 25(21):5312–5329. https://doi.org/10.1111/mec.13845
Bernatchez L, Wellenreuther M, Araneda C, Ashton DT, Barth JMI, Beacham TD, Withler RE (2017) Harnessing the power of genomics to secure the future of seafood. Trends Ecol Evol 32(9):665–680. https://doi.org/10.1016/j.tree.2017.06.010
Bi K, Linderoth T, Vanderpool D, Good JM, Nielsen R, Moritz C (2013) Unlocking the vault: next-generation museum population genomics. Mol Ecol 22(24):6018–6032
Burrell AS, Disotell TR, Bergey CM (2015) The use of museum specimens with high-throughput DNA sequencers. J Hum Evol 79:35–44
Carmo CB, Ferrette BLS, Camargo SM, Roxo FF, Coelho R, Garla RC, Mendonça FF (2019) A new map of the tiger shark (Galeocerdo cuvier) genetic population structure in the western Atlantic Ocean: hypothesis of an equatorial convergence centre. Aquat Conserv 29(5):760–772. https://doi.org/10.1002/aqc.3029
Corrigan S, Lowther AD, Beheregaray LB, Bruce BD, Cliff G, Duffy CA, Rogers PJ (2018) Population connectivity of the highly migratory Shortfin Mako (Isurus oxyrinchus) and implications for management in the southern hemisphere. Front Ecol Evol. https://doi.org/10.3389/fevo.2018.00187
Ebert DA, Fowler S, Compagno LJV (2013) Sharks of the world: a fully illustrated guide. ISBN: 9780957394605|528 pages. Wild Nature Press, Plymouth.
Ellis JS, Griffiths ÁAM, Stevens JR, Gilbey J, Armstrong ÁA, Cauwelier ÁE, Olafsson ÁK (2011) Microsatellite standardization and evaluation of genotyping error in a large multi-partner research programme for conservation of Atlantic salmon (Salmo salar L.). Genetica 139:353–367. https://doi.org/10.1007/s10709-011-9554-4
Erbe C, McPherson C (2013) Acoustic characterisation of bycatch mitigation pingers on shark control nets in Queensland, Australia. Endanger Species Res. https://doi.org/10.3354/esr00467
Fenderson LE, Kovach AI, Llamas B (2020) Spatiotemporal landscape genetics: investigating ecology and evolution through space and time. Mol Ecol 29(2):218–246. https://doi.org/10.1111/mec.15315
Ferreira LC, Simfendorfer CC (2019) Galeocerdo cuvier. The IUCN red list of threatened species 2019. IUCN Red List Threatened Species, 8235. https://doi.org/10.2305/IUCN.UK.20191.RLTS.T39378A2913541.en
Ferreira LC, Thums M, Meeuwig JJ, Vianna GMS, Stevens J, McAuley R, Meekan MG, Castonguay M (2015) Crossing latitudes—long-distance tracking of an apex predator. PLOS ONE 10(2):e0116916
Fischer MC, Rellstab C, Leuzinger M, Roumet M, Gugerli F, Shimizu KK, Widmer A (2017) Estimating genomic diversity and population differentiation - an empirical comparison of microsatellite and SNP variation in Arabidopsis halleri. BMC Genomics 18(1):1–15. https://doi.org/10.1186/s12864-016-34597
Francis RM (2017) pophelper: an R package and web app to analyse and visualize population structure. Mol Ecol Resour 17(1):27–32. https://doi.org/10.1111/1755-0998.12509
Gubili C, Bilgin R, Kalkan E, Karhan SÜ, Jones CS, Sims DW, Noble LR (2011) Antipodean white sharks on a Mediterranean walkabout? Historical dispersal leads to genetic discontinuity and an endangered anomalous population. Proc Royal Soc B Biol Sci 278(1712):1679–1686. https://doi.org/10.1098/rspb.2010.1856
Haaland ØA, Glover KA, Seliussen BB, Skaug HJ (2011) Genotyping errors in a calibrated DNA register: implications for identification of individuals. BMC Genet 12(1):36. https://doi.org/10.1186/1471-2156-12-36
Hale ML, Burg TM, Steeves TE (2012) Sampling for microsatellite-based population genetic studies: 25 to 30 individuals per population is enough to accurately estimate allele frequencies. PLoS ONE. https://doi.org/10.1371/journal.pone.0045170
Hammerschlag N, Gallagher AJ, Wester J, Luo J, Ault JS (2012) Don’t bite the hand that feeds: assessing ecological impacts of provisioning ecotourism on an apex marine predator. Funct Ecol 26(3):567–576. https://doi.org/10.1111/j.1365-2435.2012.01973.x
Hansen MM, Olivieri I, Waller DM, Nielsen EE (2012) Monitoring adaptive genetic responses to environmental change. Mol Ecol 21(6):1311–1329. https://doi.org/10.1111/j.1365294X.2011.05463.x
Heithaus MR, Wirsing AJ, Dill LM, Heithaus LI (2007) Long-term movements of tiger sharks satellite-tagged in Shark Bay, Western Australia. Mar Biol 151(4):1455–1461. https://doi.org/10.1007/s00227-006-0583-y
Hellberg ME, Burton RS, Neigel JE, Palumbi SR (2002) Genetic assessment of connectivity among marine populations. Bull Mar Sci 70(1 SUPPL.):273–290
Henriksen O (2015) Genetic insights into the population composition of two regional inshore mixed stocks of Atlantic cod (Gadus morhua) in West Greenland. Master thesis. Technical University of Denmark, Silkeborg, Denmark, p 82
Holmes BJ, Williams SM, Otway NM, Nielsen EE, Maher SL, Bennett MB, Ovenden JR (2017) Population structure and connectivity of tiger sharks (Galeocerdo cuvier) across the Indo-Pacific Ocean basin. Royal Society Publishing 4:170309. https://doi.org/10.1098/rsos.170309
Jombart T (2008) Adegenet: a R package for the multivariate analysis of genetic markers. Bioinformatics 24(11):1403–1405. https://doi.org/10.1093/bioinformatics/btn129
Karlsson S, Moen T, Lien S, Glover KA, Hindar K (2011) Generic genetic differences between farmed and wild Atlantic salmon identified from a 7K SNP-chip. Mol Ecol Resour 11(SUPPL. 1):247–253. https://doi.org/10.1111/j.1755-0998.2010.02959.x
Koljonen M-L, King TL, Nielsen EE (2007) Genetic identification of individuals and populations. Atlantic Salmon Genet Conserv Manag. https://doi.org/10.1002/9780470995846.ch9
Larson SE, Daly-Engel TS (2017) Review of current conservation genetic analyses of Northeast Pacific Sharks. Adv Mar Biol. https://doi.org/10.1016/bs.amb.2017.06.005
Lea JSE, Wetherbee BM, Queiroz N, Burnie N, Aming C, Sousa LL, Shivji MS (2015) Repeated, long-distance migrations by a philopatric predator targeting highly contrasting ecosystems. Sci Rep 5(1):11202. https://doi.org/10.1038/srep11202
Manel S, Gaggiotti OE, Waples RS (2005) Assignment methods: matching biological questions with appropriate techniques. Trends Ecol Evol 20(3):136–142. https://doi.org/10.1016/j.tree.2004.12.004
Moran P, Teel DJ, Lahood ES, Drake J, Kalinowski S (2006) Standardising multi-laboratory microsatellite data in Pacific salmon: an historical view of the future. Ecol Freshw Fish. https://doi.org/10.1111/j.1600-0633.2006.00201.x
Nance HA, Klimley P, Galván-Magaña F, Martínez-Ortíz J, Marko PB, Goldstien SJ (2011) Demographic processes underlying subtle patterns of population structure in the scalloped hammerhead shark, Sphyrna lewini. PLoS ONE 6(7):e21459
Nazareno AG, Bemmels JB, Dick CW, Lohmann LG (2017) Minimum sample sizes for population genomics: an empirical study from an Amazonian plant species. Mol Ecol Resour 17(6):1136–1147. https://doi.org/10.1111/1755-0998.12654
Nei M (1978) Estimation of average heterozygosity and genetic distance from a small number of individuals. Genetics 89(3):583–590
Nielsen EE, Morgan JAT, Maher SL, Edson J, Gauthier M, Pepperell J, Ovenden JR (2017) Extracting DNA from ‘jaws’: high yield and quality from archived tiger shark (Galeocerdo cuvier) skeletal material. Mol Ecol Resour 17(3):431–442. https://doi.org/10.1111/1755-0998.12580
Nielsen EE, Cariani A, Aoidh EM, Maes GE, Milano I, Ogden R, Carvalho GR (2012) Geneassociated markers provide tools for tackling illegal fishing and false eco-certification. Nature Commun 3:851. https://doi.org/10.1038/ncomms1845
Nielsen EE, Hansen MM, Meldrup D (2006) Evidence of microsatellite hitch-hiking selection in Atlantic cod (Gadus morhua L.): implications for inferring population structure in nonmodel organisms. Mol Ecol 15(11):3219–3229. https://doi.org/10.1111/j.1365-294X.2006.03025.x
Nielsen EE, Hemmer-Hansen J, Poulsen NA, Loeschcke V, Moen T, Johansen T, Carvalho GR (2009) Genomic signatures of local directional selection in a high gene flow marine organism; The Atlantic cod (Gadus morhua). BMC Evol Biol 9(1):276. https://doi.org/10.1186/1471-2148-9-276
Nielsen A, Eg E (2016) Population or point-of-origin identification. Seafood Authenticity and Traceability. Elsevier, Amsterdam. https://doi.org/10.1016/B978-0-12-801592-6.00008-5
Ogden R (2008) Fisheries forensics: the use of DNA tools for improving compliance, traceability and enforcement in the fishing industry. Fish Fish 9(4):462–472. https://doi.org/10.1111/j.1467-2979.2008.00305.x
O’Leary SJ, Feldheim KA, Chapman DD (2013) Novel microsatellite loci for white, Carcharodon carcharias and sandtiger sharks, Carcharias taurus (order Lamniformes). Conserv Genet Resour 5(3):627–629. https://doi.org/10.1007/s12686-013-9866-z
O’Leary SJ, Feldheim KA, Fields AT, Natanson LJ, Wintner S, Hussey N, Chapman DD (2015) Genetic diversity of white sharks, Carcharodon carcharias, in the Northwest Atlantic and Southern Africa. J Hered 106(3):258–265. https://doi.org/10.1093/jhered/esv001
Ovenden JR (2013) Crinkles in connectivity: combining genetics and other types of biological data to estimate movement and interbreeding between populations. Marine Freshw Res 64(3):201–207. https://doi.org/10.1071/MF12314
Ovenden JR, Berry O, Welch DJ, Buckworth RC, Dichmont CM (2015) Ocean’s eleven: a critical evaluation of the role of population, evolutionary and molecular genetics in the management of wild fisheries. Fish Fish 16(1):125–159. https://doi.org/10.1111/faf.12052
Pembleton LW, Cogan NOI, Forster JW (2013) StAMPP: an R package for calculation of genetic differentiation and structure of mixed-ploidy level populations. Mol Ecol Resour 13(5):946–952. https://doi.org/10.1111/1755-0998.12129
Pirog A, Jaquemet S, Ravigné V, Cliff G, Clua E, Holmes BJ, Magalon H (2019) Genetic population structure and demography of an apex predator, the tiger shark Galeocerdo cuvier. Ecol Evol 9(10):5551–5571. https://doi.org/10.1002/ece3.5111
Piry S (2004) GENECLASS2: a software for genetic assignment and first-generation migrant detection. J Hered 95(6):536–539. https://doi.org/10.1093/jhered/esh074
Potvin C, Bernatchez L (2001) Lacustrine spatial distribution of landlocked Atlantic salmon populations assessed across generations by multilocus individual assignment and mixed-stock analysis. Mol Ecol 10(10):2375–2388. https://doi.org/10.1046/j.0962-1083.2001.01374.x
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155(2):945–959
Rannala B, Mountain JL (1997) Detecting immigration by using multilocus genotypes. Proc Nat Acad Sci 94(17):9197–9201
Reeb CA, Arcangeli L, Block BA (2000) Structure and migration corridors in Pacific populations of the Swordfish Xiphius gladius, as inferred through analyses of mitochondrial DNA. Mar Biol 136(6):1123–1131. https://doi.org/10.1007/s002270000291
Renshaw MA, Saillant E, Broughton RE, Gold JR (2006) Application of hypervariable genetic markers to forensic identification of “wild” from hatchery-raised red drum, Sciaenops ocellatus. Forensic Sci Int 156(1):9–15. https://doi.org/10.1016/j.forsciint.2005.05.038
Rousset F (2008) GENEPOP’007: a complete re-implementation of the GENEPOP software for Windows and Linux. Mol Ecol Resour 8(1):103–106. https://doi.org/10.1111/j.1471-8286.2007.01931.x
Schrey AW, Heist EJ (2003) Microsatellite analysis of population structure in the shortfin mako (Isurus oxyrinchus). Can J Fish Aquat Sci 60(6):670–675. https://doi.org/10.1139/F03-064
Seeb LW, Antonovich A, Banks MA, Beacham TD, Bellinger MR, Blankenship SM, Smith CT (2011) Development of a standardized DNA database for Chinook Salmon. Fisheries. https://doi.org/10.1577/1548-8446(2007)32[540:DOASDD]2.0.CO;2
Spaet JLY, Jabado RW, Henderson AC, Moore ABM, Berumen ML (2015) Population genetics of four heavily exploited shark species around the Arabian Peninsula. Ecol Evol 5(12):2317–2332
Stephenson JJ, Campbell MR, Hess JE, Kozfkay C, Matala AP, McPhee MV, Wenburg JK (2009) A centralized model for creating shared, standardized, microsatellite data that simplifies interlaboratory collaboration. Conserv Genet 10(4):1145–1149. https://doi.org/10.1007/s10592-0089729-4
Weir, Cockerham (2009) Estimating F-statistics for the analysis of population structure. Society 38(6):1358–1370
Yoav B, Yosef H (1995) Controlling the false discovery rate: a practical and powerful approach to multiple testing. J Royal Stat Soc Series B (Methodological) 57(1):289–300
Acknowledgements
We wish to thank all the researchers, institutes and museums that provided us with samples that were crucial for the completion of this study: Smithsonian Institution National Museum of Natural History, Florida Museum of Natural History, Virginia Institute of Marine Science, National Oceanic and Atmospheric Administration (NOAA, Rhode Island), Harvard Museum of Comparative Zoology, Dr. Austin Gallagher (Beneath the Waves), Dr. Javier Quesada (Natural Sciences Museum of Barcelona), Dr. Peter Rask Møller (Natural History Museum of Denmark), and Dr. Stefan Merker (State Museum of Natural History Stuttgart). We also wish to thank Romina Henriques for valuable comments on a previous draft. Finally, we are grateful to all of the laboratory technicians at DTU (Maj-Britt Jacobsen and Camilla Kristensen) for their help in carrying out the lab work.
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Sort, M., Manuzzi, A., Jiménez-Mena, B. et al. Come together: calibration of tiger shark (Galeocerdo cuvier) microsatellite databases for investigating global population structure and assignment of historical specimens. Conservation Genet Resour 13, 209–220 (2021). https://doi.org/10.1007/s12686-021-01197-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12686-021-01197-5