Abstract
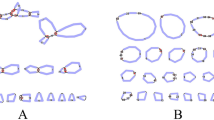
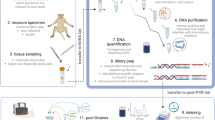
Just a few years ago, genomic studies of non-model organisms were extremely time and cost prohibitive, but recent advances in next-generation sequencing have made it possible to generate hundreds to thousands of informative single nucleotide polymorphisms (SNPs) for non-model organisms. We generated a SNP panel using restriction site-associated DNA (RAD) sequencing for species of the genus Buteo (Falconiformes: Accipitridae) with the aim of creating genomic resources in the absence of a reference genome. Using blood and feather samples from three different Buteo species, we used HiSeq Illumina sequencing to generate raw reads. We built 29 162 contigs and discovered 89 921 SNPs using paired-end single-digest RAD data. Using Bayesian clustering and principal componenets analysis, this SNP panel successfully assigned individuals of three Buteo (B. swainsoni, B. jamaicensis, B. lagopus) species correctly to species. We then scanned the SNPs for association with variation in coloration within the three species. This de novo reference will have broad application to the study of Buteo genomic diversity, analysis of population structure within species, identification of hybrids, and other applied studies of the avian genome.


Similar content being viewed by others
References
Ali OA, O’Rourke SM, Amish SJ, Meek MH, Luikart G, Jeffres C, Miller MR (2016) RAD capture (Rapture): Flexible and efficient sequence-based genotyping. Genetics 202:289–400
Andrews KR, Good JM, Miller MR, Luikard G, Hohenlohe PA (2016) Harnessing the power of RADseq for ecological and evolutionary genomics. Nat Rev Genet 17:81–92
Canavelli SB, Bechard MJ, Woodbridge B, Kochert MN, Maceda JJ, Zaccagnini ME (2003) Habitat use by Swainson’s hawks on their austral wintering grounds in Argentina. J Raptor Res 37:125–134
Davey JW, Hohenlohe PA, Etter PD, Boone JQ, Catchen JM, Blaxter ML (2011) Genome-wide genetic marker discovery and genotyping using next-generation sequencing. Nat Rev Genet 12:499–510
Ellegren H (2014) Genome sequencing and population genomics in non-model organisms. Trends Ecol Evol 29:51–63
Fisher RJ, Wellicome TI, Bayne EM, Poulin RG, Todd LD, Ford AT (2015) Extreme precipitation reduces reproductive output of an endangered raptor. J Appl Ecol 52:1500–1508
Fumagalli M, Vieira FG, Linderoth T, Nielsen R (2014) ngsTools: methods for population genetics analyses from next-generation sequencing data. Bioinformatics 30:1486–1487
Hull JM, Anderson R, Bradbury M, Estep JA, Ernest HB (2007) Population structure and genetic diversity in Swainson’s Hawks (Buteo swainsoni): implications for conservation. Conserv Genet 9:305–316
Hull JM, Strobel BN, Boal CW, Hull AC, Dykstra CR, Irish AM, Fish AM, Ernest HB (2008a) Comparative phylogeography and population genetics within Buteo lineatus reveals evidence of distinct evolutionary lineages. Mol Phylogenet Evol 49:988–996
Hull JM, Hull AC, Sacks BN, Smith JP, Ernest HB (2008b) Landscape characteristics influence morphological and genetic differentiation in a widespread raptor (Buteo jamaicensis). Mol Ecol 17:810–824
Hull JM, Mindell DP, Talbot SL, Kay EH, Hoekstra HE, Ernest HB (2010) Population structure and plumage polymorphism: the intraspecific evolutionary relationships of a polymorphic raptor, Buteo jamaicensis harlani. BMC Evol Biol 10:224
Kjellén N, Roos G (2000) Population trends in Swedish raptors demonstrated by migration counts at Falsterbo, Sweden 1942–97. Bird Study 47:195–211
Korneliussen TS, Albrechtsen A, Nielsen R (2014) ANGSD: analysis of next generation sequencing data. Bioinformatics 15:356
Kross SM, Tylianakis JM, Nelson XJ (2012) Effects of introducing threatened falcons into vineyards on abundance of Passeriformes and bird damage to grapes. Conserv Biol 26:142–149
Li H, Durbin R (2009) Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25:1754–1760
Li H, Handsaker B, Wysoker A, Fennell T, Ruan J, Homer N, Marth G, Abecasis G, Durbin R (1000) Genome Project Data Processing Subgroup (2009) The Sequence alignment/map (SAM) format and SAMtools. Bioinformatics 25:2078–2079
Madden T (2002) The BLAST sequence analysis tool. [Updated 2003 Aug 13]. In: McEntyre J, Ostell J (eds). The NCBI Handbook [Internet]. Bethesda (MD): National Center for Biotechnology Information (US); 2002-. Chapter 16. Available from:http://www.ncbi.nlm.nih.gov/books/NBK21097/
Martín B, Ferrer M (2013) Assessing biodiversity distribution using diurnal raptors in Andalusia, southern Spain. Ardeola 60:15–28
Miller MR, Dunham JP, Amores A, Cresko WA, Johnson EA (2007) Rapid and cost-effective polymorphism identification and genotyping using restriction site associated DNA (RAD) markers. Genome Res 17:240–248
Miller MR, Brunelli JP, Wheeler PA, Liu S, Rexroad CE, Palti Y, Doe CQ, Thorgaard GH (2012) A conserved haplotype controls parallel adaptation in geographically distant salmonid populations. Mol Ecol 21:237–249
Narum SR, Buerkle CA, Davey JW, Miller MR, Hohenlohe PA (2013) Genotyping-by-sequencing in ecological and conservation genomics. Mol Ecol 22:2841–2847
Nikolic Z, Laube B, Weber RG, Lichter P, Kioschis P, Poustka A, Mülhardt C, Becker CM (1998) The human glycine receptor subunit α3 Glra3 gene structure, chromosomal localization, and functional characterization of alternative transcripts. J Biol Chem 273:19708–19714
Preston CR, Beane RD (1993) Red-tailed Hawk (Buteo jamaicensis). In: Poole A, Gill F (eds) The Birds of North America, no. 52. Academy of Natural Sciences, Philadelphia, and American Ornithologists’ Union, Washington, D.C
Purcell S, Neale B, Todd-Brown K, Thomas L, Ferreira MAR, Bender D, Maller J, Sklar P, de Bakker PIW, Daly MJ, Sham PC (2007) PLINK: a tool set for whole-genome association and population-based linkage analysis. Am J Hum Genet 81:559–575
Quinlan AR, Hall IM (2010) BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26:841–842
Riesling MJ, Kruckenhauser L, Gamauf A, Haring E (2003) Molecular phylogeny of the genus Buteo (Aves: Accipitridae) based on mitochondrial marker sequences. Mol Phylogenet Evol 27:328–342
Roulin A, Wink M (2004) Predator–prey relationships and the evolution of colour polymorphism: a comparative analysis in diurnal raptors. Biol J Linn Soc 81:565–578
Sala OE, Chapin FS, Armesto JJ, Berlow E, Bloomfield J, Dirzo R, Huber-Sanwald E, Huenneke LF, Jackson RB, Kinzig A, Leemans R (2000) Global biodiversity scenarios for the year 2100. Science 287:1770–1774
Skotte L, Korneliussen TS, Albrechtsen A (2013) Estimating individual admixture proportions from next-generation sequencing data. Genetics 195:693–702
Terraube J, Villers A, Ruffino L, Iso-Iivari L, Henttonen H, Oksanen T, Korpimaki E (2015) Coping with fast climate change in northern ecosystems: mechanisms underlying the population-level response of a specialist avian predator. Ecography 38:690–699
Wickham H (2009) ggplot2: elegant graphics for data analysis. Springer-Verlag: New York
Acknowledgments
We would like to thank those who contributed samples to this study, including the Golden Gate Raptor Observatory, Christopher Briggs, and Frank Nicoletti. We would also like to acknowledge the Genomic Variation Laboratory and Alisha Goodblah for her guidance in laboratory procedures, and Ismael Saglam and the Miller lab for training in next-generation sequencing bioinformatics. Financial support came from the National Science Foundation Graduate Research Fellowship Program (Grant Number 1148897) and the Jastro Sheilds fellowship through the University of California, Davis.
Data accessibility
The 89,921 SNP data set used in this study is available from GenBank accession KX722542-KX751703.
Author contributions
M. A. M. designed the study, conducted laboratory work including extractions and library preparation, conducted the bioinformatics analyses, and wrote the manuscript. J. M. H. conceived the study and was involved in study design, provided technical supervision and guidance, and contributed to manuscript writing.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Mayo, M.A., Hull, J.M. Discovery and preliminary multi-species evaluation of single nucleotide polymorphism resources for genus Buteo developed from restriction site-associated DNA paired-end data. Conservation Genet Resour 9, 151–156 (2017). https://doi.org/10.1007/s12686-016-0619-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12686-016-0619-7