Abstract
Although previous research has shown that inflammation is associated with development of colorectal cancer (CRC), questions remain about whether inflammatory factor-secreting bacteria play a crucial role in CRC development. The potential role of gut microbiota in secreting inflammatory factors involved in the carcinogenesis of CRC among Chinese patients was explored in this study. 16S rRNA sequencing was utilized to evaluate the distinct microbial characteristics between patients with CRC and colorectal adenoma. The serum levels of TNF-α, IL-6 and IL-10 were measured using Enzyme-linked immunosorbent assay (ELISA), while the expression of LRG1 and TGF-β1 in tissues was evaluated by immunohistochemistry. The correlation between gut microbiota and inflammatory factor signaling was analyzed. Compared with the adenoma group, CRC patients exhibit distinct pathologies. Moreover, elevated levels of CEA, erythrocytes and haemoglobin in the blood of CRC patients were found. In addition, CRC patients have significantly higher levels of TNF-α, IL-6, IL-10, LRG1 and TGF-β1. Spearman correlation analysis revealed that LRG1 was positively related to IL-6 and TNF-α, respectively. The correlation analysis results of TGF-β1 were consistent with the above. The abundance of Blautia and Streptococcus was lower in CRC patients, while the relative abundance of Alistipes, Peptostreptococcus and Porphyromonas was significantly elevated. Moreover, positive correlations between Alistipes and inflammatory factor signaling were also found. Our results suggest that gut commensal Alistipes is a key bacterium with pro-inflammatory properties in the CRC carcinogenesis. TNF-α and IL-6 associated with Alistipes might activate LRG1/TGF-β1 signaling which contributed to the carcinogenesis of CRC.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Globally, colorectal cancer (CRC) ranks the third in cancer incidence (10%) and is the second most common cause of cancer-related mortality (9.4%). The incidence of CRC, however, varies, with the highest number of cases reported in Asia (52.3%), followed by Europe (26.9%) and North America (9.3%) [1]. With the rising trend of mortality from CRC in Asia, there's a critical demand to discover the underlying mechanism for more efficient strategies to treat CRC, including early identification and prevention.
Microorganisms in the colon maintain colonic homeostasis [2, 3]. Disruption of this balance can lead to gut microbiota dysbiosis, which can potentially cause the development of CRC [4,5,6,7]. However, little is known about the causal involvement of these alterations in the development of CRC. Previous studies have shown that certain microbial species may promote cancer development [8]. Adhering to and infiltrating colonic epithelial cells, Fusobacterium nucleatum induces chronic inflammation and immune evasion via its lipopolysaccharides, thereby promoting a pro-inflammatory tumor microenvironment [9]. Additionally, it modulates host signaling pathways, like the Wnt pathway, fostering tumorigenesis. In contrast, Bacteroides fragilis secretes the Bacteroides fragilis toxin, which induces cellular proliferation, apoptosis disruption, and inflammation [10]. This chronic inflammation and disruption of gut microbiota balance can lead to a compromised intestinal barrier and further facilitate cancer progression. Furthermore, Porphyromonas gingivalis accelerate CRC development by secreting butyric acid in mice [11]. Apart from these, are there any other causative mechanisms that have yet to be identified. Therefore, further research is needed to understand the interaction between gut bacteria and CRC development.
The presence of microbiota is tied to the onset and advancement of CRC through its impact on intestinal inflammation related to tumors [12, 13]. An imbalance in gut microbiota can trigger an abnormal immune system response, which can induce inflammation and subsequently increase the levels of inflammatory factors [14]. A large number of studies have shown a correlation between inflammation and tumors [15]. Certain cytokines, i.e., tumor necrosis factor (TNF-α) and interleukin(IL)-6, are known for regulating inflammation which can result in CRC development [16, 17]. Transforming growth factor-beta (TGF-β) from monocyte macrophages, platelets and glial cells, also regulate cell proliferation, migration, differentiation and immuno-inflammatory responses, which can further enhance carcinogenesis. Among several subtypes of TGF-β, TGF-β1 is a key factor in inducing epithelial-mesenchymal transition (EMT) and angiogenesis in tumor cells [18]. TGF-β1 can signal through two different type I receptors in endothelial cells: activin receptor-like kinase (ALK)1 and ALK5. ALK1 promotes angiogenesis by activating SMAD1/5/8, possibly in association with endoglin (ENG). ALK5, on the other hand, is more involved in suppressing angiogenesis and promoting fibrosis through the SMAD2/3 pathway [19, 20].
Human leucine-abundant α-2-glycoprotein-1 (LRG1) is another potential indicator of CRC activity [21, 22]. Aberrant LRG1/TGF-β1 signaling is implicated in diseases such as cancer. LRG1 modulates TGF-β1 signaling, particularly in endothelial cells, and plays a role in vascular development and repair. LRG1 amplifies the angiogenic impact of TGF-β1 by altering the signaling route from SMAD2/3 to SMAD1/5/8 [20]. Studies suggest that LRG1 can be induced by certain cytokines. The activation of LRG1 is triggered by IL-1β and TNF-a, leading to the activation of NF-kB, which plays an important role in immune response [23]. IL-6 plays a role in vascular instability by activating LRG1 transcription via STAT3 phosphorylation [24].
Since the presence of inflammatory factor-secreting bacteria in the gut has not been fully identified in the current study, the potential mechanisms also require further experimental exploration. To identify gut microbiota linked to CRC development, we selected patients with colorectal adenoma as the control group. Colorectal adenoma is a benign tumor of the colon. It is regarded as a precancerous lesion and is a common disease especially among elders. Study shows that 95% of CRC is composed of adenocarcinoma that progresses from adenomatous polyp or adenoma [25]. Thus, the focus of this research is to study among CRC patients, what bacteria are involved in the development of CRC through inflammatory factors. This will help us understand the underlying mechanisms and further search for potential therapeutic targets.
2 Materials and methods
2.1 Patients and sample collection
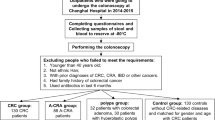
A total of 20 CRC patients, including 12 men and 8 women (aged 42–69 years), underwent surgery at the Anhui No.2 Provincial People’s Hospital from January 2023 to December 2023. In the adenoma group, there were 13 men and 7 women (aged 46–69 years). We collected intestinal mucosal tissue and serum samples of patients with colorectal adenoma and CRC. All patients with CRC included in this study were confirmed through pathological diagnosis. All patients with colorectal adenoma underwent total colonoscopy to confirm the absence of CRC. Patients were excluded if they received neoadjuvant treatment before tissue harvesting, or had been treated with antibiotics for a minimum of 4 weeks before sampling. We also examined fecal samples from patients who were included. These fecal specimens were gathered 3 ~ 14 days prior to bowel cleanse of colonoscopy. Intact fresh feces were collected in sterile boxes and stored at −80 °C on arrival to the laboratory. We obtained informed consent from all patients involved in this study. We followed regulations according to the Declaration of Helsinki and received approval from Medical Ethics Committee of Anhui No. 2 Provincial People’s Hospital. (No. R2023-013, China).
2.2 Reagents and materials
Rabbit anti-human LRG1 (13224-1-AP) and TGF-β1 (21898-AP) were purchased from Proteintech, China. Immunohistochemical kits were purchased from Beijing Zhongshan Jinqiao Company, China. Enzyme-linked immunosorbent assay (ELISA) kits (human TNF-α, human IL-6 and human IL-10) were purchased from Jianglai Biotechnology, China.
2.3 ELISA measurement of cytokines
Blood samples were used to extract blood serum. Blood samples were clotted at 25 °C for 30 min before centrifugation (1500 g, 4 °C, 15 min), then collected and stored at −80 °C until further analysis.The quantities of various cytokines, including human TNF-α, IL-6, and IL-10, were measured with ELISA kit, adhering to the guidelines provided by the manufacturer.
2.4 Fecal microbial community analysis
Stool samples were collected from patients without prior bowel preparation and under non-fasting conditions using a standard 5 ml stool collection kit. Samples were excluded if they were watery or if the donor had undergone pre-operative chemotherapy or radiotherapy. Fecal samples, each weighing no less than 6 g, were collected from the study subjects and immediately stored at −80 °C. Total genomic DNA was extracted from the fecal samples following the protocol provided by the Fast Pure Stool DNA Isolation Kit [26]. DNA quality and concentration measurements were conducted using 1.0% agarose gel electrophoresis and NanoDrop® ND-2000 spectrophotometer (Thermo Scientific Inc., USA). Amplification of the bacterial 16S rRNA within V3-V4 areas was achieved using primers 336F and 806R. Sequencing was completed using the standard protocols of Majorbio BioPharm Technology Co., Ltd. (Shanghai, China).
2.5 Immunohistochemical staining
Tissue segments embedded in paraffin underwent dewaxing, rehydration, immersion in citric acid, followed by incubation in a microwave setup for antigen recovery. Tissue samples were examined using primary antibodies including rabbit anti-LRG1 (1:200) and rabbit anti-TGF-β1 (1:200). Sections of the colon underwent incubation using secondary antibodies. The technique of immunohistochemistry was carried out with an immunohistochemistry kit. Hematoxylin was used to stain the cell nuclei for a duration of 30 s.
Differentiation of the tissues was performed by mixing hydrochloric acid in ethanol for 30 s, followed by a 15 min water rinse and dehydration using aqueous alcohol and xylene. The immunohistochemical images were visualized by a microscope (Olympus, Japan).
2.6 Statistical analysis
We compared two groups (patients with adenoma/CRC) with t-test by calculating mean and standard deviation. Chi-Square and Fischer exact test were used to compare qualitative data between groups. The Wilcoxon rank test used to test the significance of bacterial percent abundance between groups (threshold > 0.05%). *P < 0.05 was considered to indicate statistically significant differences. Every statistical evaluation was calculated with SPSS (SPSS, Chicago, IL, USA). We used Graph Pad Prism 8.0 (Graph Pad Software, USA) to faciliate the graphical handling.
3 Result
3.1 Clinical pathological differences between patients with CRC and colorectal adenoma
A total of 20 patients with colorectal adenoma (mean patient age was 57.25 ± 7.23 years) and 20 patients with CRC (mean patient age was 57.97 ± 8.12 years) were included. There was no statistical significance among the two groups with respect to sex and age (P > 0.05).
Figure 1 shows the histological differences between adenoma and CRC (Fig. 1A, B). The pathological manifestations of tumor cells in the colonic tissues of CRC patients include a solid striated arrangement, occasional glandular structures, infiltrative growth, and penetration into the muscularis propria of the mucosa, along with mucus components visible within the cytoplasm. These features align with the typical pathological characteristics of CRC. Carcinoembryonic antigen (CEA) is used clinically as a tumor marker for CRC. Hemoglobin and erythrocytes are also used to monitor CRC progression since CRC can cause intestinal hemorrhaging.
Clinical characteristics of CRC patients. A Hematoxylin and eosin staining of colonic tissues of patients with the adenoma control (magnification × 200). B Hematoxylin and eosin staining of colonic tissues of CRC patient (magnification × 200). C–E Relative protein expression of CEA, Erythrocyte and Hemoglobin in blood of patients with CRC and adenoma controls (n = 20). F–H levels of TNF-α, IL-6 and IL-10 in the serum were measured by ELISA among the two groups (n = 20). Data are shown as the mean ± SD. *P < 0.05; **P < 0.01; ***P < 0.001 versus adenoma controls
From our study, we found that among CRC patients, CEA is significantly higher, while erythrocyte count and hemoglobin levels are significantly lower when comparing to adenoma group (P < 0.05) (Fig. 1C–E). TNF-α (P < 0.001), IL-6 (P < 0.001) and IL-10 (P < 0.01) were markedly higher in CRC patients (Fig. 1F–H).
3.2 Positive correlation between pro-inflammatory cytokines and LRG1 protein expression
LRG1 and TGF-β1 are significant molecular markers in CRC. LRG1 has been shown to be involved in angiogenesis and tumor progression, while TGF-β1 is a pivotal factor inis a key regulator of cell growth, differentiation, and immune response. In CRC, increased levels of these molecules are often connected with poor prognosis and advanced disease stages.
The expression of LRG1 and TGF-β1 in CRC is significantly higher than in patients with adenomas (P < 0.01) (Fig. 2). Under immunohistochemical staining, LRG1 is highly expressed in CRC tissues, suggesting that LRG1 is involved in the progression of CRC.
Representative immunohistochemical staining of LRG1 and TGF-β1 in human samples of adenoma tissue and CRC tissue. A–E LRG1 was overexpressed in CRC tissues. A Immunohistochemical analysis of LRG1 in the adenoma tissues (magnification × 200) and B magnification × 400. C Immunohistochemical analysis of LRG1 in the CRC tissues (magnification × 200) and D magnification × 400. E Quantification of the integral optical density of LRG1 in two samples. F–J TGF-β1 was overexpressed in CRC tissues. F Immunohistochemical analysis of TGF-β1 in the adenoma tissues (magnification × 200) and G magnification × 400. H Immunohistochemical analysis of TGF-β1 in the CRC tissues (magnification × 200) and I magnification × 400. J Quantification of the integral optical density of TGF-β1 in two samples. *P < 0.05; **P < 0.01; ***P < 0.001 versus adenoma control
Figure 3 depicts the correlation analysis of pro-inflammatory cytokines with the expression of LRG1 and TGF-β1. LRG1 is positively related to IL-6 (r = 0.846, P < 0.001); TGF-β1 is positively correlated with IL-6 (r = 0.496, P < 0.05); LRG1 is positively correlated with TNF-α (r = 0.734, P < 0.001); TGF-β1 is positively correlated with TNF-α (r = 0.529, P < 0.05).
3.3 Distinct microbial characteristics in patients with CRC and colorectal adenoma
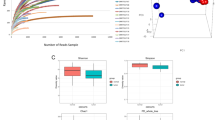
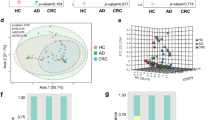
We performed 16S rRNA sequencing to assess the composition of gut microbiota of CRC. The Venn diagram reveals two sets of partly overlapping operational taxonomic units (OTUs), suggesting a difference between OTUs in CRC and adenoma patients. There were 421 and 777 distinct OTUs in adenoma and CRC patients, respectively, while the number of OTUs common to both samples was 750 (Fig. 4A). CRC patients exhibited a greater number of OTUs compared to the adenoma controls. Consequently, these two groups possessed unique OTUs. The phylogenetic tree categorized the gut microbiota from two groups into two principal branches (Fig. 4B). Our findings suggest that despite substantial variations observed between the two groups, we still observed mostly the same microbial community patterns in individual groups. Subsequently, we conducted weighted UniFrac PCoA evaluation to depict the comparative resemblance in the gut microbiota composition. In addition, PC1 and PC2 scores showed significant differences in flora composition between the 2 groups (R = 0.3327, P < 0.01) (Fig. 4C). PCoA analysis shows that adenoma and CRC clusters are visibly distinct. We performed species composition analyses to further investigate the distribution of gut microbiota. The histograms of gut microbiota community structure displayed the relative abundance of microbiota at the genus level (Fig. 4D). At the genus level, the top 10 genera at the taxonomic level for both sample groups were Blautia, Escherichia-Shigella, Streptococcus, Bacteroides, Bifidobacterium, Faecalibacterium, Ruminococcus_torques_group, Akkermansia, Prevotella and Collinsella. The proportions of Blautia and Streptococcus in the CRC patients are less compared to the adenoma control. The proportions are significant for Blautia (P < 0.01) and Streptococcus (P < 0.05) (Fig. 4E). However, the proportions of Alistipes, Peptostreptococcus, and Porphyromonas in CRC patients are higher than those in adenoma group (P < 0.05) (Fig. 4E).
Analysis of the composition of bacterial microbiota for samples from CRC and adenoma patients. A Venn diagram that shows the overlapping operation taxonomic units in the gut microbiota of the two groups. B Beta diversity for adenoma and CRC patients assessed using cluster tree. C Principal coordinates analysis (PCoA) of the two groups based on weighted UniFrac distances. Each plot represents one sample. D Histogram of the relative abundance of the dominant bacterial genus. E Species difference analysis at the genus level (comparing CRC to adenoma). F LDA scores (only taxa with an LDA score threshold of > 3.5 are shown). Data are shown as the mean ± SD. *P < 0.05; **P < 0.01; ***P < 0.001 (control: adenoma). The bacterial distribution at various taxonomic levels among the two groups was compared using the Wilcoxon rank test
We further performed linear discriminant analysis (LDA) on the effect size (LefSe) to identify the primary biomarkers and predominant microbiota between both groups. As shown in Fig. 4F, the abundance of Oscillospiracea, Alistipes, Rikenellaceae, Christensenellaceae, Christensenellales, Porphyromonas, Porphyromonadaceae and Peptostreptococcus is the highest in the CRC patients. Nonetheless, the percentage of Lachnospiraceae was found to be greater in patients with adenoma than those with CRC. Our findings suggest that alterations in these microbial communities may be related to the CRC onset and development.
3.4 Correlation between Alistipes and inflammatory factor signaling
Redundancy and canonical correlation methods (RDA/CCA) mainly reflects the relationship between bacterial microbiota and clinical inflammatory factors. A subset of clinical factors was obtained by using maximum correlation coefficient. We performed CCA analyses according to sample species distribution tables for clinical factors. inflammatory factor signaling we selected for this study included TNF-α, IL-6, IL-10, LRG1 and TGF-β1.
Analysis of the RDA/CAA clinical inflammatory factors reveals that differences in inflammatory signaling were tightly associated with gut microbiota. This analysis further identified that TNF-α, IL-6, LRG1, and TGF-β1 exerted the most significant influence on tumor gut microbiota, with IL-10 following in importance (Fig. 5A).
Environmental factor analysis showing the correlation of bacterial abundance with inflammatory factor signaling. A Redundancy analysis/canonical correlation analysis (RDA/CCA) showing the degree of influence of clinical inflammatory factor signaling on microbiota composition. The blue dots: gut microbes from CRC patients; The red dots:gut microbes from adenoma patients. Arrows: inflammatory factor signaling (TNF-α, IL-6, IL-10, LRG1 and TGF-β1). The longer the arrow, the greater the correlation. The smaller the angle is, the higher the correlation will be. The angle between environmental factors is positive correlation when it is acute, and negative correlation when it is obtuse. B Correlation heatmap between gut microbiota and inflammatory factor signaling. Horizontal axis: inflammatory factor signaling including TNF-α, IL-6, IL-10, LRG1 and TGF-β1. Vertical axis: bacteria identified by 16S rRNA. Colors in the graph indicate the magnitude of the correlation coefficients. *P < 0.05; **P < 0.01; ***P < 0.001
To explore the correlations between gut microbiota and clinical inflammatory factors, we created a correlation heatmap based on the different clinical inflammatory factors in CRC patients. The correlation between microbial classification and clinical inflammatory factor signaling reveals that Alistipes is positively related to TNF-α (r = 0.5204, P < 0.01), IL-6 (r = 0.5107, P < 0.05), LRG1 (r = 0.5172, P < 0.01) and TGF-β1 (r = 0.5164, P < 0.01); Peptostreptococcus is positively related to TNF-α (r = 0.4267, P < 0.05), IL-6 (r = 0.4164, P < 0.05), LRG1 (r = 0.4246, P < 0.05) and TGF-β1 (r = 0.4417, P < 0.05); Porphyromonas is positively related to TNF-α (r = 0.4446 P < 0.05), IL-6 (r = 0.4447, P < 0.05), LRG1 (r = 0.4445, P < 0.05) and TGF-β1 (r = 0.3141, P < 0.05) (Fig. 5B). Comparatively, Blautia is inversely associated with TNF-α (r = −0.5432, P < 0.01), IL-6(r = −0.5320, P < 0.01), LRG1 (r = −0.5391, P < 0.01) and TGF-β1 (r = −0.5159, P < 0.01); Butyricoccus is negatively associated with TNF-α (r = −0.5708, P < 0.01), IL-6 (r = −0.5742, P < 0.01), LRG1 (r = −0.5755, P < 0.01), TGF-β1 (r = −0.5910, P < 0.01) and IL-10 (r = −0.6290, P < 0.001) (Fig. 5B). Our results indicate Alistipes shows a stronger positive association with inflammatory factor signaling compared to Peptostreptococcus and Porphyromonas. Besides, Alistipes has the strongest positive correlation with inflammatory factor signaling.
4 Discussion
A variety of factors affect the development of CRC and the underlying mechanism remains incompletely understood. Here, we profiled the fecal gut microbiota, the serum, and the tissue protein of CRC patients and compared them with those of precancerous colorectal adenoma patients. There are distinct differences in the serological and cytological presentations between the two. CEA is significantly higher, while erythrocyte count and hemoglobin levels are significantly lower when compared to the adenoma group. CRC patients have significantly higher levels of TNF-α, IL-6, IL-10, LRG1 and TGF-β1 when compared with adenoma patients. Spearman correlation analysis revealed that LRG1 was positively related to IL-6 and TNF-α, respectively. The correlation analysis result of TGF-β1 was consistent with the above. Furthermore, we found that Alistipes was significantly elevated in CRC patients. Moreover, the strongest positive correlation between Alistipes and inflammatory factor signaling was also found. Our results suggest that gut commensal Alistipes is a key bacterium with pro-inflammatory properties in the carcinogenesis of CRC. TNF-α and IL-6 associated with Alistipes might activate LRG1/TGF-β1 signaling which contributed to the carcinogenesis mechanisms of CRC.
The inflammatory reactions ultimately lead to the occurrence of colon cancer. In the study of CRC, several aspects of inflammation are involved, including inflammatory factors, the development of immune cells, and the activation of related signaling pathways. Previous studies have suggested that upregulation of mucosal pro-inflammatory cytokines is associated with the development of CRC [3]. Consistent with previous studies, we confirmed the elevation of pro-inflammatory cytokines in patients with CRC compared to adenoma. In our study, all three cytokines, TNF-α, IL-6, and IL-10, were significantly elevated in CRC patients. Although IL-10 is a crucial anti-inflammatory cytokine, it may act as an inflammation-inducing factor in cancer patients. Li et al. [27] showed that IL-10 and IL-18 are highly expressed in the serum of CRC patients, making IL-10 and IL-18 useful as indicators to determine the prognosis of CRC patients. Consistent with these findings, our results indicated elevated IL-10 levels in CRC patients.
LRG1 in the context of development of inflammatory diseases and tumors, mainly through TGF-β1 signaling, has attracted the attention of researchers and has achieved important advances [28]. Research indicates that the TGF-β1 signaling is the key factor for CRC [18]. LRG1, as a preliminary controller of the TGF-β1 signaling pathway, likely plays a crucial role in driving the angiogenic transition in TGF-β1 signaling [29]. Nevertheless, the upstream mechanisms leading to the induction of LRG1 under disease conditions remain poorly understood in current research. There is also evidence that shows pro-inflammatory cytokines such as IL-6 and TNF-α could induce LRG1 [30,31,32]. Consistent with previous studies, we found higher levels of LRG1 and TGF-β1 in CRC tissues compared to adenoma tissues. In this study, there were significant differences in the levels of inflammatory factors (TNF-α, IL-6 and IL-10), LRG1 proteins and TGF-β1 signaling between the two groups. Spearman correlation analysis revealed that LRG1 was positively related to IL-6 and TNF-α, respectively. The correlation analysis result of TGF-β1 was consistent with the above.
This points to the crucial role that inflammatory factor signaling play in the pathogenesis of CRC.
The Venn diagram from 16S rRNA analysis shows that OTU from patients with CRC and adenoma are not completely overlapped, suggesting a discrepancy between these two groups. The number of OTUs is also higher in the CRC patients than in adenoma patients. CRC-associated gut microbiota exhibits increased species richness, reduced presence of potentially protective species, and elevated levels of precarcinogenic species, as indicated by prior research [24]. Consistent with previous research, these results suggest that increased diversity may not be a sign of a healthy gut, but rather the excessive growth of harmful bacteria, which further promotes the development of adenomas and cancers. In β-diversity parameters, PCoA analysis shows that adenoma and CRC clusters are visibly distinct. This indicates a significant difference in gut microbiome composition between the two groups.
Then, we employed bar charts depicting community composition to examine variations in the community structure between the two groups. Our findings revealed that the top 10 genera at the taxonomic level for both sample groups. Although these bacterial species are commonly found in the two groups, fluctuations in their quantities could be linked to the onset of CRC. We performed species difference analyses and LDA on the effect size LefSe to identify the primary biomarkers and predominant microbiota between both groups. Our results showed a lower proportion of Blautia and Streptococcus in CRC, while the proportions of Alistipes, Peptostreptococcus, and Porphyromonas were higher.
Blautia is generally considered to be a beneficial gut bacteria involved in maintaining homeostasis and producing short-chain fatty acids (SCFAs) in the gut, with anti-inflammatory and anti-cancer properties. Several studies have reported a decreased abundance of Blautia in CRC, Blautia producta inhibits colon cancer development by promoting CD8+ T cell tumor immunosurveillance through the degradation of lysophosphatidylglycerol in the tissue microenvironment [33]. The abundance of the gut commensal Streptococcus may vary in CRC. Many studies report an increase in certain species of Streptococcus in patients with CRC, while some suggest a reduction in Streptococcus levels, especially when compared to healthy individuals. Certain Streptococcus species, such as Streptococcus gallolyticus, are thought to contribute to cancer development by promoting chronic inflammatory states in the gut, which create a conducive environment for tumor growth [34]. Peptostreptococcus is a genus of anaerobic bacteria. Several studies have shown an increased abundance of Peptostreptococcus, particularly Peptostreptococcus anaerobius, in CRC patients [35]. Porphyromonas is commonly found in the oral microbiota, particularly Porphyromonas gingivalis, which is closely associated with periodontal disease. However, some studies have found that Porphyromonas may also be present in the gut. Porphyromonas can activate the immune system by releasing toxins (e.g., gingival proteases), triggering a chronic inflammatory response in the gut [36].
Alistipes belongs to the phylum Bacteroidetes and plays a crucial role in inflammation and disease. The role of Alistipes may depend on its surrounding microenvironment, and it may behave differently across different disease states [37]. Alistipes has been found to potentially offer protection against specific diseases, such as liver fibrosis and cardiovascular disease [37]. Reduced Alistipes levels lead to a decrease in SCFAs, potentially resulting in severe fibrosis in individuals with nonalcoholic fatty liver disease [38]. In hypertension, Alistipes is thought to cause inflammation and epithelial changes.
The role of Alistipes may be dual, but in the tumor microenvironment, it may promote inflammation and disease progression. Some researches have found that Alistipes exerts a defensive impact against CRC by modulating the immune response and reducing intestinal inflammation, particularly in healthy populations where a higher abundance of Alistipes is associated with better colorectal health [39]. However, recent research has revealed that Alistipes in CRC correlates with disease progression, suggesting that it may contribute to cancer development [40]. Research has also implicated Alistipes as a pathogen in CRC and as a factor linked to depression [39]. In CRC patients, Alistipes exacerbated chronic inflammation and increased intestinal permeability, suggesting that it may contribute to tumor progression and poor prognosis [41]. Validation experiments showed that A. finegoldii promoted CRC tumor growth [42]. Our study found that Alistipes was significantly elevated in CRC patients compared to adenoma group, suggesting that it may contribute to CRC carcinogenesis. However, it remains unclear how Alistipes influences the cancer-promoting effects in CRC.
In addition, a previous study found that IL-6 and TNF-α, promote the expression of LRG1 [8, 15]. Our study also confirmed significant correlations of IL-6 and TNF-α with both LRG1 and TGF-β1. We further examined the relationship between Alistipes and different inflammatory factor signaling (TNF-α, IL-6, IL-10, LRG1 and TGF-β1), in order to delineate the signaling pathways that may be involved. Analysis of the RDA/CAA environmental factors reveals that differences in inflammatory factor signaling were tightly associated with gut microbiota. Additionally, our results indicate that TNF-α, IL-6, LRG1, and TGF-β1 have a more significant impact on altering the bacterial composition within tumor tissue. Previous research also reveals that Alistipes may enhance the growth and development of mast cells through the activation of IL-6 synthesis [5]. Research indicated that in IL10−/− mice, A. finegoldii efficiently triggered CRC through the IL-6/STAT3 pathways [38, 40, 43]. Iida and colleagues suggested that the binding of the pro-inflammatory gram-negative bacterium A. shahii to TLR4 triggers TNF production [39]. Researchers suggested a direct link between Alistipes and the function of TLR4-priming/TNF generation [20]. These studies suggest that Alistipes is associated with pro-inflammatory processes. In our study, correlation between microbial classification and clinical inflammatory factor signaling reveals that Alistipes has the strongest positive correlation with inflammatory factor signaling. These results suggest that the gut commensal Alistipes is a key bacterium with pro-inflammatory properties in the carcinogenesis mechanisms of CRC.
This study preliminarily explored the role of Alistipes in the early carcinogenesis mechanisms of CRC. The identification of specific microbes for CRC in this study could lead to the development of more personalized strategies, especially in individuals at risk of carcinogenesis. To further elucidate the role of Alistipes in CRC and its underlying mechanisms, we will co-culture Alistipes with normal colon epithelial cells to observe changes in cell proliferation, apoptosis, invasion, and migration. Additionally, we will knock down or mutate suspected oncogenes in the bacteria and observe changes in their oncogenic potential to identify specific oncogenes or pathways in CRC animal models. Further research will also investigate other bacterial strains that may be involved in CRC and their interactions with inflammatory factors.
5 Conclusion
Our investigation focuses on the interplay between LRG1 and Alistipes in disease, as well as their possible roles in the carcinogenesis of CRC. In the present study, we confirm that the inflammatory response is exacerbated in individuals with CRC, as indicated by elevated TNF-α and IL-6 levels detected in their serum. Gut commensal Alistipes is a key bacterium with pro-inflammatory properties in the carcinogenesis of CRC. TNF-α and IL-6 associated with Alistipes might activate LRG1/TGF-β1 signaling which contributed to the early carcinogenesis mechanisms of CRC. The discovery of this differential microorganism and the study of its potential mechanisms will provide a theoretical foundation for exploring the carcinogenesis of CRC.
Data availability
The data used and/or analyzed during the current study are available from the corresponding author upon reasonable request.
References
Alhhazmi AA, et al. Gut microbial and associated metabolite markers for colorectal cancer diagnosis. Microorganisms. 2023;11(8):2037.
Chen CB, et al. Identification of intestinal microbiomeassociated with lymph-vascular invasion in colorectal cancer patients and predictive label construction. Front Cell Infect Microbiol. 2023;13:1098310.
Liu X, et al. Alterations of the predominant fecal microbiota and disruption of the gut mucosal barrier in patients with early-stage colorectal cancer. Biomed Res Int. 2020;2020:2948282.
Li N, et al. The relationship between gut microbiome features and chemotherapy response in gastrointestinal cancer. Front Oncol. 2021;11:781697.
Liu JG, et al. Identifcation of colorectal cancer progression-associated intestinal microbiome and predictive signature construction. J Transl Med. 2023;21(1):373.
Wang ZK, et al. Colorectal cancer and gut microbiota studies in China. Gut Microbes. 2023;15(1):2236364.
Feng Q, et al. The gut microbiota is believed to be directly involved in colorectal carcinogenesis. Nat Commun. 2015;6:6528.
Chen HY, et al. Urea cycle activation triggered by host- microbiota maladaptation driving colorectal tumorigenesis. Cell Metab. 2023;35(4):651–66.
Wang N, Fang JY. Fusobacterium nucleatum, a key pathogenic factor and microbial biomarker for colorectal cancer. Trends Microbiol. 2023;31(2):159–72.
Quaglio AEV, et al. Gut microbiota, inflammatory bowel disease and colorectal cancer. World J Gastroenterol. 2022;28(30):4053–60.
Shintaro O, et al. Gut bacteria identified in colorectal cancer patients promote tumourigenesis via butyrate secretion. Nat Commun. 2021;12(1):5674.
Park J, et al. Fecal microbiota and gut microbederived extracellular vesicles in colorectal cancer. Front Oncol. 2021;11:650026.
Niccolai E, et al. Significant and conflicting correlation of il-9 with prevotella and bacteroides in human colorectal cancer. Front Immunol. 2021;11:573158.
Hajjar R, et al. Gut microbiota influence anastomotic healing in colorectal cancer surgery through modulation of mucosal proinflammatory cytokines. Gut. 2023;72(6):1143–54.
Yang L, et al. Intratumoral microbiota: roles in cancer initiation, development and therapeutic efficacy. Signal Transduct Target Ther. 2023;8(1):35.
Sharma BR, Kanneganti TD. Inflammasome signaling in colorectal cancer. Transl Res. 2023;252:45–52.
Yamamoto T, et al. IL-6 levels correlate with prognosis and immunosuppressive stromal cells in patients with colorectal cancer. Ann Surg Oncol. 2023;30(8):5267–77.
Massague J, Sheppar D. TGF-β signaling in health and disease. Cell. 2023;186(19):4007–37.
Jin Z, et al. The prognostic impact of leucine-rich α-2-glycoprotein-1 in cholangiocarcinoma and its association with the IL-6/TGF-β1 axis. J Surg Res. 2020;252:147–55.
Wang X, et al. LRG1 promotes angiogenesis by modulating endothelial TGF-β signalling. Nature. 2013;499(7458):306–11.
Zhang JJ, et al. LRG1 modulates epithelial-mesenchymal transition and angiogenesis in colorectal cancer via HIF-1α activation. J Exp Clin Cancer Res. 2016;35:29.
Yasuda R, et al. Serum leucine-rich alpha-2 glycoprotein and calprotectin in children with inflammatory bowel disease: a multicenter study in Japan. J Gastroenterol Hepatol. 2023;38(7):1131–9.
Wang YY, et al. TNF-α-induced LRG1 promotes angiogenesis and mesenchymal stem cell migration in the subchondral bone during osteoarthritis. Cell Death Dis. 2017;8(3):e2715.
Wong SH, Yu J. Gut microbiota in colorectal cancer: mechanisms of action and clinical applications. Nat Rev Gastroenterol Hepatol. 2019;16(11):690–704.
Sawicki T, et al. A review of colorectal cancer in terms of epidemiology, risk factors, development, symptoms and diagnosis. Cancers (Basel). 2021;13(9):2025.
Sheng KL, et al. Grape seed proanthocyanidin extract ameliorates dextran sulfate sodium-induced colitis through intestinal barrier improvement, oxidative stress reduction, and inflammatory cytokines and gut microbiota modulation. Food Funct. 2020;11(9):7817–29.
Dritsoula A, et al. Angiopathic activity of LRG1 is induced by the IL-6/STAT3 pathway. Sci Rep. 2022;12(1):4867.
Iida N, et al. Commensal bacteria control cancer response to therapy by modulating the tumor microenvironment. Science. 2013;342(6161):967–70.
Li ZF, et al. Exosomal leucine-rich-alpha2-glycoprotein 1 derived from non-small-cell lung cancer cells promotes angiogenesis via TGF-β signal pathway. Mol Ther Oncolytics. 2019;14:313–22.
Ohta H, et al. Gene expression of leucine-rich alpha-2 glycoprotein in the polypoid lesion of inflammatory colorectal polyps in miniature dachshunds. J Vet Med Sci. 2020;82(10):1445–9.
Zhou Y, et al. LRG1 promotes proliferation and inhibits apoptosis in colorectal cancer cells via RUNX1 activation. PLoS ONE. 2017;12(4): e0175122.
Zhang Q, et al. Leucine-rich alpha-2-glycoprotein-1 is up-regulated in colorectal cancer and is a tumor promoter. Onco Targets Ther. 2018;11:2745–52.
Zhang XS, et al. Tissue-resident Lachnospiraceae family bacteria protect against colorectal carcinogenesis by promoting tumor immune surveillance. Cell Host Microbe. 2023;31(3):418–32.
Sonja OH, et al. Streptococcus gallolyticus abrogates anti-carcinogenic properties of tannic acid on low-passage colorectal carcinomas. Sci Rep. 2020;10(1):4714.
Long XH, et al. Peptostreptococcus anaerobius promotes colorectal carcinogenesis and modulates tumour immunity. Nat Microbiol. 2019;4(12):2319–30.
Koliarakis I, et al. Oral bacteria and intestinal dysbiosis in colorectal cancer. Int J Mol Sci. 2019;20(17):4146.
Parker BJ, et al. The genus Alistipes: gut bacteria with emerging implications to inflammation, cancer, and mental health. Front Immunol. 2020;11:906.
Gupta H, et al. Gut microbiome in non-alcoholic fatty liver disease: from mechanisms to therapeutic role. Biomedicines. 2022;10(3):550.
Fong W, et al. Exploring the role of the gut microbiota in modulating colorectal cancer immunity. Gut. 2023;72(12):2272–85.
Moschen AR. Lipocalin 2 protects from inflammation and tumorigenesis associated with gut microbiota alterations. Cell Host Microbe. 2016;19(4):455–69.
Oana LP, et al. An overview of gut microbiota and colon diseases with a focus on adenomatous colon polyps. Int J Mol Sci. 2020;21(19):7359.
Kang X, et al. Altered gut microbiota of obesity subjects promotes colorectal carcinogenesis in mice. EBioMedicine. 2023;93:104670.
Feng Q, et al. Gut microbiome development along the colorectal adenoma-carcinoma sequence. Nat Commun. 2015;6:6528.
Funding
This research was funded by Anhui Medical University Research Foundation (Jingjing Fu, Grant No. 2022xkj124), the Key Program of Natural Science Research of Anhui Provincial Education Department (Lei Jiang, Grant No. 2023AH053381) and the Wu Jieping Medical Foundation (Lei Jiang, Grant No. 320.6750.2022-20-30).
Author information
Authors and Affiliations
Contributions
JF: conceptualization, designed the experiment, writing-original draft preparation and funding acquisition. GL: conceptualization, designed the experiment, writing-original draft preparation. XL, LC and BR: collected the materials and conducted quantitative data analysis. SS: formal analysis and software. LJ: supervision, conceptualization, writing-review & editing and funding acquisition. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
The study was conducted in accordance with the Declaration of Helsinki and approved by the Medical Ethic Committee of Anhui No.2 Provincial People’s Hospital (No. R2023-012, China). The written informed consent was obtained from each patient.
Consent for publication
All authors approved the content and submission.
Competing interests
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Fu, J., Li, G., Li, X. et al. Gut commensal Alistipes as a potential pathogenic factor in colorectal cancer. Discov Onc 15, 473 (2024). https://doi.org/10.1007/s12672-024-01393-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s12672-024-01393-3