Abstract
Background
Esophageal squamous cell carcinoma (ESCC) stands as a significant global health challenge, distinguished by its aggressive progression from the esophageal epithelium. Central to this malignancy is sphingolipid metabolism, a critical pathway that governs key cellular processes, including apoptosis and immune regulation, thereby influencing tumor behavior. The advent of single-cell and transcriptome sequencing technologies has catalyzed significant advancements in oncology research, offering unprecedented insights into the molecular underpinnings of cancer.
Methods
We explored sphingolipid metabolism-related genes in ESCC using scRNA-seq data from GEO and transcriptome data from TCGA. We assessed 97 genes in epithelial cells with AUCell, UCell, and singscore algorithms, followed by bulk RNA-seq and differential analysis to identify prognosis-related genes. Immune infiltration and potential immunotherapeutic strategies were also investigated, and tumor gene mutations and drug treatment strategies were analyzed.
Result
Our study identified distinct gene expression patterns, highlighting ARSD, CTSA, DEGS1, and PPTQ's roles in later cellular stages. We identified seven independent prognostic genes and created a precise nomogram for prognosis.
Conclusion
This study integrates single-cell and transcriptomic data to provide a reliable prognostic model associated with sphingolipid metabolism and to inform immunotherapy and pharmacotherapy for ESCC at the genetic level. The findings have significant implications for precision therapy in esophageal cancer.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Esophageal cancer, arising from the malignant transformation of esophageal mucosal epithelial or glandular epithelial cells, is a formidable global health concern, ranking as the eighth most prevalent cancer and the sixth leading cause of cancer-related mortality worldwide [1,2,3,4]. This malignancy comprises two principal histological subtypes: esophageal squamous cell carcinoma (ESCC) and esophageal adenocarcinoma (EAC) [5]. Geographically, ESCC is predominantly observed in developing nations, whereas EAC predominates in developed regions, further compounding the complexity of this disease [6]. Esophageal cancer is characterized by a high fatality rate, often diagnosed at an advanced stage, underscoring the challenges in achieving a favorable prognosis [7]. Its multifaceted pathogenesis involves a confluence of genetic, environmental, and lifestyle factors [8]. The particularly grim prognosis associated with ESCC is closely linked to its late-stage diagnosis [9]. The traditional therapeutic approach for ESCC generally includes surgical or endoscopic resection and radiotherapy. However, these modalities often offer limited curative potential, particularly in advanced stages of the disease [10, 11]. Consequently, there is a pressing need to explore novel therapeutic avenues to enhance the quality of life for ESCC patients. Immunotherapy has emerged as a promising frontier in addressing this clinical challenge, offering renewed optimism through recent advancements in therapeutic strategies and immunotherapeutic interventions [12, 13]. The integration of precision medicine with immunotherapy has markedly enhanced both the treatment efficacy and prognosis for patients with ESCC [14]. However, the field confronts persistent difficulties and challenges, making the quest for novel biomarkers of paramount importance for refining treatment strategies and prognostic predictions.
Sphingolipid metabolism is pivotal in numerous cellular processes, encompassing apoptosis, cell signaling, and immune regulation [15], involving intricate mechanisms including cell proliferation, migration, and differentiation [16]. Extensive research has elucidated the intricate connection between sphingolipid metabolism and the onset and progression of cancer. In particular, sphingomyelin biosynthesis within this metabolic pathway plays a pivotal role in driving the growth and metabolic reprogramming of cancer cells [17]. Irregularities within sphingolipid metabolism have been linked to a spectrum of disorders, spanning metabolic anomalies, neurological afflictions, and specific cancer subtypes [18]. Recent findings suggest that sphingolipid-metabolizing enzymes could emerge as promising targets for novel anti-cancer therapies [19]. Consequently, our study will meticulously investigate the involvement of sphingolipid metabolism in the progression of esophageal cancer, aiming to illuminate the genesis and advancement of this malignancy.
The advent of advanced sequencing technologies has facilitated the discovery of diverse biomarkers associated with esophageal cancer [20, 21]. Notably, the synergistic application of scRNA-seq technology in conjunction with transcriptome sequencing has yielded significant breakthroughs within the realm of oncology [12, 22]. As the pursuit of personalized and precision-based tumor therapeutics gains momentum, the amalgamation of scRNA-seq and bulk RNA-seq data analysis emerges as a pivotal strategy for comprehensive and in-depth investigations into the intricacies of ESCC pathogenesis. This integrated approach not only enriches our understanding of ESCC but also holds promise for advancing the elucidation of its underlying molecular mechanisms.
In this investigation, we initiated our study by delineating multiple cellular subpopulations through the annotation of single-cell data pertaining to ESCC. Subsequently, we conducted a comprehensive analysis of intercellular communication at the single-cell level, thereby affording us insights into the intricate immune microenvironment characterizing the tumor. To further scrutinize ESCC with respect to distinct cell types, we amalgamated the sphingolipid metabolism gene set with three single-cell scoring methods, enabling the identification of epithelial cells exhibiting significant differences between tumor and normal cohorts. Given the epithelial origin of ESCC, we leveraged these disparities across cell subpopulations to extract genes specifically from epithelial cells, subsequently subjecting them to differential analysis. The resulting differential gene set was subsequently integrated with bulk RNA-seq data, and a rigorous screening process was undertaken to identify genes with prognostic relevance for ESCC, employing ten distinct machine learning algorithms. Building upon the identified prognostic-related genes, we proceeded to construct prognostic models for ESCC. In tandem with this, we conducted a thorough exploration of the immune microenvironment in ESCC, with a particular focus on potential immunotherapeutic avenues, accomplished through immune infiltration analysis. Our efforts culminated in the identification of seven genes deemed prognostically significant in ESCC. These collective endeavors serve as a foundation for elucidating the regulatory mechanisms governed by sphingolipid metabolism-related genes in ESCC and offer a theoretical framework for the development of both immunotherapeutic modalities and targeted treatment strategies for this malignancy.
2 Materials and methods
2.1 Raw data sources
Single-cell sequencing data pertinent to esophageal squamous cell carcinoma (ESCC) were sourced from the GEO database, specifically GSE188900, comprising two primary untreated ESCC samples and a singular normal sample. Subsequently, 97 genes related to sphingolipid metabolism were obtained from the InnateDB website (https://www.innatedb.com/redirect.do?go=searchMols). For ESCC bulk RNA sequencing, data were accessed through Xena's Web site (https://xena.ucsc.edu/). This dataset encompassed 173 esophageal cancer samples from the TCGA cohort, featuring gene expression profiles alongside comprehensive clinical information, encompassing parameters such as time to survival, survival status, gender, and TNM stage. Additionally, the TCGA dataset furnished data on disease-free survival. As a validation set, we procured the GSE53624 dataset from the GEO database, which comprised 119 disease samples after exclusion of normal samples. Notably, transcriptome data from each sample were subjected to preliminary correction in preparation for the construction of our prognostic model. These multifaceted datasets collectively underpinned our comprehensive analyses and model development endeavors.
2.2 Single-cell data processing
In the initial phase of our analysis, we subjected the ESCC single-cell RNA sequencing data to rigorous examination using the ‘‘Seurat’’ R package, prioritizing data quality and reliability [23]. To this end, we implemented a quality control step, which entailed the exclusion of cells exhibiting excessive mitochondrial gene expression exceeding 5% and cells demonstrating either over 2500 or under 200 expressed genes, effectively mitigating the inclusion of low-quality cells. Subsequently, to mitigate potential batch effects originating from sample sources, we harnessed the ‘‘harmony’’ R package for robust batch correction, thereby ensuring that our subsequent analytical endeavors remained minimally influenced by sample origins. We proceeded with data normalization using the ‘‘Seurat’’ R package, a crucial step that homogenizes expression levels across cells to enable accurate comparisons and integration. Principal component analysis (PCA) was conducted for dimensionality reduction, leveraging the ‘‘RunPCA’’ function within ‘‘Seurat’’ to discern principal components characterized by higher variance ratios, thereby facilitating subsequent cell clustering analyses. Utilizing the ‘‘FindNeighbors’’ and ‘‘FindClusters’’ functions in tandem with the t-distributed stochastic neighbor embedding (tSNE) nonlinear dimensionality reduction method, we effectively visualized the resulting cellular clusters, affording greater insights into the structural and relational aspects of single-cell data. To discern specifically expressed genes within each cell cluster, the ‘‘FindAllMarkers’’ and "FindMarkers" functions within the ‘‘scran’’ R package facilitated Wilcoxon tests, pinpointing genes exhibiting significant differential expression across diverse cell populations, thus advancing our comprehension of cell types and their respective functions. Cell type annotations were predicated on data obtained from the CellMarker online resource (http://xteam.xbio.top/CellMarker/index.jsp), which furnished valuable guidance for the precise identification of cell types within individual clusters in the single-cell data, enriching the accuracy of our analytical outcomes.
2.3 Single-cell scoring
In evaluating the significance of a gene set encompassing 97 genes associated with sphingolipid metabolism, we employed three established algorithms, namely Aucell, Ucell, and singscore, to glean insights from expression data. Firstly, Aucell quantified the enrichment of the gene set within the top 5% of genes across the single-cell data, ascertained through area under the curve calculations. This metric illuminated the extent of gene set enrichment relative to the ranked gene expression in individual samples, thereby elucidating variations in gene set expression across the sample cohort. Secondly, UCell, an unsupervised cell type identification tool, leveraged gene expression patterns within single-cell transcriptome data to discern and categorize cells [24]. Employing techniques such as clustering and dimensionality reduction, it provided cell type assignments, thereby advancing our understanding of cellular identity and function. Lastly, singscore facilitated the assessment of the activity level of specific biological functions within individual samples or cells. Anchored in gene sets and their respective weights and orientations, this method derived cell state scores, allowing for the quantification of biological function activity. These tools and approaches hold pivotal roles in the domain of single-cell transcriptomics and genome enrichment analysis. They serve as instrumental means to unveil cellular functions, biological processes, and alterations associated with diseases, thereby enhancing our comprehension of the intricacies inherent to biological systems. The simultaneous application of diverse assessment methods for specific gene sets mitigates scoring inaccuracies and biases, furnishing a more comprehensive and nuanced understanding of biological phenomena. This multifaceted approach bears immense significance within the realm of bioinformatics research.
2.4 Cellular communication analysis
To comprehensively investigate intercellular signaling in ESCC and gain deeper insights into the immune cell functions, we employed the R software package ‘‘CellChat’’ for our analytical endeavors [25]. ‘‘CellChat’’ harnesses gene expression data and incorporates existing knowledge regarding interactions among signaling molecules, receptors, and cofactors to model diverse modes of intercellular communication. This approach facilitates an exploration of intricate cellular interactions and communication mechanisms. In the context of epithelial cells, we conducted differential gene expression analysis employing the ‘‘limma’’ package in R to identify pivotal genes. Subsequently, we turned to the ‘‘Monocle’’ R package, a tool tailored for the analysis of single-cell RNA sequencing data, enabling pseudo-temporal analysis of cell developmental trajectories within the epithelial cell population [26, 27]. ‘‘Monocle’’ empowers the study of dynamic changes and state transitions during biological processes, thereby unveiling temporal sequences underlying tumorigenesis and progression. This analytical framework highlights dynamic shifts in key genes and signaling pathways, providing invaluable insights into the developmental trajectories of tumor tissues. Furthermore, we harnessed the "CellCall" R software package, distinguished for its capacity to integrate intracellular and intercellular signaling to infer intercellular communication networks and internal regulatory signals [28]. An outstanding feature of ‘‘CellCall’’ is its amalgamation of intercellular ligand-receptor communication with intracellular transcription factor expression to form the Ligand-Receptor-Transcription Factor (L-R-TF) axis. Coupled with pathway activity analysis, this approach delves into the receptor cell pathway alterations resulting from two intercellular communications. This comprehensive toolkit augments our understanding of intercellular communication and signaling networks, facilitating in-depth investigations into cellular functions and interactions. These methodologies and tools find essential applications within the realm of bioinformatics research, enabling the elucidation of complex cellular relationships and signaling mechanisms.
2.5 Screening for ESCC prognosis-related genes
In our quest to identify prognostically relevant genes in ESCC patients, we executed a Weighted Gene Co-Expression Network Analysis (WGCNA). WGCNA, a potent methodology for constructing and modularizing diverse gene networks, facilitates the discernment of underlying patterns and relationships within the data. Initially, we subjected the samples to cluster analysis to identify any potential outliers and ensure data integrity. Subsequently, leveraging WGCNA, we crafted co-expression networks, a process that inherently segments the genome into distinct modules. Hierarchical clustering and dynamic tree-cutting techniques were deployed to precisely delineate and define these modules. Employing module affiliation attributes (MM) and gene significance (GS), we gauged the modules’ associations with clinical characteristics. Of particular interest were core co-expression modules exhibiting robust module affiliation (MM > 0.8) and clinical significance (GS > 0.2). These modules exhibited heightened intra-gene connectivity and demonstrated statistically significant correlations with clinical features. Our focal attention rested on core co-expression modules displaying the strongest correlations with tumor groups, thereby revealing potential co-expressed genes linked to tumorigenesis. To further winnow down genes associated with ESCC prognosis, we conducted an intersection analysis involving core module genes and differentially expressed genes identified via WGCNA, employing a battery of ten machine learning algorithms.A variety of machine learning paradigms were applied, such as Cox Boost, Lasso, Elastic Net (Enet), Ridge, Survival-Support Vector Machine (Survival-SVM), Generalized Boosted Regression Modeling (GBM), Cox Partial Least Squares Regression (plsRcox), Stepwise Cox, Random Survival Forest (RSF), and Supervised Principal Component Analysis (SuperPC). From these techniques, 101 different algorithmic combinations were developed. And we selected the method with the highest Mean C-index for genetic screening.This meticulous approach culminated in the identification of genes underpinning ESCC prognosis, bolstered by multiple analytical methodologies, thereby furnishing a catalog of pivotal candidate genes meriting further exploration. This comprehensive analytical strategy affords a more holistic understanding of the intricate gene-prognosis relationships, thus underpinning the rationale for personalized treatment strategies and disease management.
2.6 Construction and evaluation of predictive models
To enhance the precision of prognostic assessments in ESCC, we employed the R package ‘‘Rms’’ in conjunction with key feature genes to construct a predictive model termed a Nomogram [29, 30]. The Nomogram represents a graphical tool wherein each predictor is assigned a corresponding score within a delineated table. Accumulating these scores for individual factors yields a total score for each patient. The primary aim of this Nomogram model is to facilitate more accurate prognostic predictions and clinical assessments for ESCC patients. By amalgamating multiple predictors, this model furnishes a comprehensive information framework that augments medical decision-making and personalized treatment planning [31]. To ascertain the predictive efficacy and clinical utility of the Nomogram model, we leveraged calibration curves and decision curves (DCA) for visualization and analysis. Calibration curves afford insights into the model's predictive accuracy, while DCA enables the assessment of its clinical applicability at varied thresholds. These visualization tools collectively gauge the effectiveness and value of the Nomogram model in clinical practice, thereby elevating the precision of prognostic evaluations and clinical management for ESCC patients. This integrated analytical approach underscores the pivotal role of bioinformatics research in clinical translational applications.
2.7 Gene set enrichment analysis
To glean insights into the expression patterns of our selected feature genes within distinct functional pathways, we harnessed the R package ‘‘clusterProfiler’’ to conduct Gene Set Enrichment Analysis (GSEA) at the individual gene level [32,33,34]. This analytical approach entailed independent GSEA for each feature gene, affording comprehensive exploration of their contributions across various biological pathways. Of particular focus were the top 5 Kyoto Encyclopedia of Genes and Genomes (KEGG) pathways distinguished by their heightened enrichment significance. The salient enrichment information pertaining to these pathways is herein presented. KEGG pathways represent a valuable resource for unraveling the intricate interplay between genes and fundamental biological processes, metabolic pathways, and diseases. The deliberate selection of these pathways augments our comprehension of the functional roles played by the featured genes and the biological phenomena they influence. This analytical paradigm effectively unveils the potential roles of feature genes in cellular function and regulatory mechanisms, thus offering crucial insights that pave the way for subsequent in-depth investigations in the realm of biology.
2.8 Immune infiltration and TME analysis
Utilizing the CIBERSORT algorithm, which leverages gene expression matrices, we conducted a robust assessment of immune cell composition, a pivotal facet in ESCC research [35]. This powerful tool facilitated the quantification of relative proportions of 22 distinct immune cell types, effectively discriminating between ESCC patients and normal subjects. Presentation of these data was achieved through visually informative lollipop plots depicting the expression profiles of individual genes. Subsequently, employing the R package ‘‘corrplot,’’ we embarked on a comprehensive analysis of correlations among the 22 immune cell types, elucidating these intricate relationships via a heatmap visualization [36, 37]. Additionally, we employed box plots to compare expression disparities across 22 immune cell types and 10 immune functions across diverse ESCC subtypes. Spearman rank correlation analysis further deepened our understanding of the associations between immune cells, gene expression, and immune functions. To unravel the nuanced variances in immune cell infiltration across diverse risk groups, we harnessed heatmaps as effective visual aids, facilitating the discernment of subtle differentiations within immune cell populations. Moreover, the specialized functionalities offered by the "estimate" R software package enabled the meticulous quantification of immunological scores, stromal scores, and ESTIMATE scores in ESCC patients. This strategic approach provided a robust assessment of the tumor microenvironment and its potential implications. In pursuit of the identification of promising chemotherapeutic agents tailored to distinct risk groups, we turned to the predictive capacities afforded by the "oncoPredict" R software package. This tool empowered us to conduct intricate prognostic analyses, thereby informing the development of more tailored therapeutic strategies, attuned to the unique needs of individual patients. This comprehensive and integrated approach affords profound insights into the intricate interplay between immune cells and the tumor microenvironment within the domain of ESCC research, thereby offering indispensable guidance for the advancement of personalized treatment modalities.
2.9 Mutation analysis
Employing the R package ‘‘maftools,’’ we conducted an exhaustive exploration to scrutinize the prevalence and distribution patterns of somatic mutations across distinct genes, a pivotal pursuit in unraveling the pathogenesis of ESCC [38]. This endeavor facilitated the identification of genes potentially wielding substantial influence in ESCC etiology, discerned through their notable mutation frequencies. Additionally, the ‘‘maftools’’ software package furnished an array of visualization tools, enabling the depiction of co-occurrence and mutual exclusion patterns among mutations. These visual representations engendered a more lucid comprehension of potential synergistic or antagonistic interplays between different genetic alterations. Furthermore, we categorized TCGA-ESCC patients into four distinct cohorts predicated on median risk scores and median tumor mutation loads (TMB), setting the stage for a comparative analysis accentuating survival disparities among these groups. Elaborate scrutiny, predicated on median risk scores and TMB values, afforded an encompassing grasp of the survival characteristics characterizing distinct patient subpopulations, thereby enriching the pool of insights available for personalized treatment strategies and informed clinical decision-making. These analytical outcomes bear paramount significance for advancing our comprehension of the molecular underpinnings of ESCC and identifying prospective targeted therapeutic avenues.
2.10 Statistical analysis
Our comprehensive statistical analyses and data processing were meticulously executed within the R programming environment, specifically employing R version 4.2.0. To ascertain the statistical significance of our survival analyses, we harnessed Kaplan–Meier curves and conducted log-rank tests. For the generation of comprehensive survival curves, we adeptly leveraged the capabilities conferred by the ‘‘Survminer’’ R software package. To enhance data visualization and facilitate the exploration of potential underlying patterns, we judiciously availed the specialized functionalities offered by the ‘‘pheatmap’’ R package, enabling the construction of informative figures. In instances where variables conformed to a normal distribution, differences were quantified employing two-tailed t-tests or, when suitable, one-way ANOVA. Conversely, for data sets exhibiting non-normal distribution characteristics, we opted for the Wilcoxon test or the Kruskal–Wallis test to assess variations. Notably, we maintained stringent adherence to a p-value threshold of less than 0.05 throughout our analyses.
3 Results
3.1 Marker gene-based single-cell data dimensionality reduction sentence eh clustering and cell type annotation
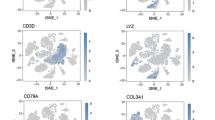
We acquired single-cell sequencing data pertaining to ESCC from the dataset GSE188900, encompassing two tumor samples derived from pathologically confirmed ESCC patients (GSM5691642, GSM5691643), along with one distal normal sample (GSM5691649) for comparative single-cell analysis. Subsequently, we harnessed the Seurat software package, a widely adopted tool in the realm of single-cell RNA sequencing data analysis, to initiate the initial processing and exploration of the obtained single-cell data. Within this framework, we meticulously curated Seurat objects, incorporating pivotal pre-processing steps. This included the culling of cells evincing over 2500 or less than 200 expressed genes, as well as those demonstrating mitochondrial gene content ratios surpassing 5%. This rigorous curation served to expunge potentially aberrant or subpar cells, ensuring data integrity for subsequent analyses. Following this, a multifaceted sequence encompassing normalization, identification of highly variable genes, and matrix normalization transpired, culminating in the initiation of dimensionality reduction clustering. Through PCA, the cellular distribution within normal and tumor group samples was delineated in PCA space (Fig. 1A), while an ElbowPlot was instrumental in determining the optimal number of dimensions, ultimately selecting the top ten for subsequent dimensionality reduction analyses. Employing tSNE, we partitioned each cell within the tumor and normal groups into 25 discrete clusters (Fig. 1B), subsequently facilitating their alignment within comparable spaces. Marker genes for pancreatic tissue cells, sourced from pertinent literature and the CellMarker website (http://xteam.xbio.top/CellMarker/index.jsp), were instrumental in characterizing the distribution of specific genes across cellular clusters (Fig. 1C) and their expression patterns within the tSNE maps (Fig. 1D). With the completion of cell type annotation, we expounded upon the distribution of diverse cell types within normal and tumor groups, employing tSNE visualization (Fig. 1E). Leveraging the ‘‘slingshot’’ package, we artfully delineated cell differentiation profiles and undertook pseudo-temporal inference based on single-cell sequencing data, unveiling the overarching structure of cell clusters in a ‘‘pseudo-temporal’’ context (Fig. 1F). Expression profiles of marker genes within distinct cell types were showcased in Fig. 1G, substantiating the accuracy and reliability of cell type annotation through violin plots. Finally, tSNE plots and histograms (Fig. 1H and I) adeptly depicted the distribution and proportional representation of various cell types in both normal control samples and tumor tissue samples, effectively elucidating their percentages. This comprehensive suite of analytical endeavors culminated in the successful dimensionality reduction clustering of single-cell data and the precise annotation of cell types.
(A) The upper panel illustrates the cell distribution within the normal and tumor group samples through PCA, while the lower panel depicts the distribution of the top 20 ranked Principal Components (PCs). (B) Employing non-linear dimensionality reduction, tSNE was utilized for clustering all single-cell data into 25 distinct cell clusters. (C) Presented are bubble plots elucidating the correlation between marker genes and the 25 cell clusters identified via tSNE clustering. The size of the bubbles signifies the magnitude of gene expression, with color denoting the average gene expression. (D) Expression dot plots for the 9 marker genes on the tSNE graph, wherein darker shades represent elevated gene expression levels. (E) The tSNE figure with annotations designates distinct cell types with corresponding colors. (F) Global cell clustering relationships are portrayed through a line graph, with each color representing a unique cell type. The line graph encompasses pseudotime results, and nodes within the graph correspond to pseudotime developmental branching points. (G) Marker gene violin plots exhibit gene expression profiles across various cell types. The horizontal axis delineates marker genes, while the vertical axis designates cell types. (H) The tSNE figure delineates cell distributions in both normal control tissue samples and tumor tissue samples, with each color representing a specific cell type. (I) Bar charts elucidate the proportions of cells across various cell types in the samples, with each color signifying a cell type, and the length of the bars representing cell proportion. (J) The cellular communication landscape is visualized through cell–cell interactions. Line thickness and accompanying numbers indicate the count of cell–cell interactions. (K) Dot plots showcase input and output intensities for nine cells involved in cell–cell interactions. (L) Factor calculations for non-negative matrix decomposition are depicted through a line graph. (M) A river diagram illustrates patterns of cellular communication
3.2 CellChat deciphers cellular communication in the tumor microenvironment
To gain deeper insights into the functional roles of diverse cell populations within the tumor microenvironment, we harnessed the CellChat software package, an invaluable tool for predicting and scrutinizing intercellular communication networks. Our meticulously annotated tumor tissue cells, initially extracted from Seurat objects, were subsequently integrated into the human ligand-receptor database furnished by the software package. CellChat harnesses a simulation-based approach, leveraging gene expression data in tandem with established knowledge regarding signaling ligands, receptors, and their associated cofactors to probabilistically model intercellular communication. This model is adeptly implemented through a diverse array of modes of action, ultimately facilitating the quantification of interactions between various cell types based on a priori information (Fig. 1J). The hierarchy plot offered by CellChat serves as an invaluable visualization tool, offering insights into the relative positions and connectivity profiles of distinct cells within the signaling network (Fig. 1K). Leveraging non-negative matrix factorization (NMF), we embarked on the discernment of communication patterns among cells of varied types. This commenced with the judicious determination of the optimal number of factorization components (Fig. 1L) and culminated in the stratification of diverse cell types into subgroups. The resultant river plot effectively portrayed the communication dynamics characterizing distinct cell types. This comprehensive approach afforded an intricate understanding of the interplay between cells, thereby shedding light on the complex network of intercellular communication underpinning the tumor microenvironment.
3.3 Screening of sphingolipid metabolism-related cells by combining three scoring methods (AUCell, UCell, Scoring)
To evaluate the expression activity of sphingolipid metabolism-related genes within esophageal squamous cell carcinoma (ESCC) tumor tissues, we adopted a gene set scoring approach based on single-cell sequencing data. In this endeavor, we employed a trio of robust scoring tools, namely AUCell, UCell, and Scoring, to ensure the reliability of our results. These scoring methods assessed each cell type, annotated based on a predefined set of glycosyltransferase-related genes. The outcomes from these three scoring techniques were standardized, normalized, and aggregated to yield the final score. Subsequently, we visualized the variation in gene set scores across diverse cell types through mapping onto the tSNE plot (Fig. 2A). Figure 2B aptly illustrates the scores assigned to each cell type within both normal control and tumor samples by each of the three scoring methods, along with the cumulative total scores. Bubble plots effectively convey the scores of distinct cell types (Fig. 2C). Furthermore, violin plots (Fig. 2D) vividly portray the final scores of various cell types in the normal and tumor groups. Notably, T and NK cells, macrophages, epithelial cells, fibroblasts, monocytes, and endothelial cells exhibited statistically significant score differences between the tumor and normal groups. Specifically, sphingolipid metabolism-related genes in macrophages, epithelial cells, monocytes, and endothelial cells displayed increased expression levels in tumor tissues compared to normal samples. To elucidate the expression patterns of key sphingolipid metabolism-related genes across distinct cell types, we employed heatmaps (Fig. 2E). Given the conspicuous distinctions observed in epithelial cells in the context of esophageal squamous cell carcinoma, we meticulously isolated this cell subpopulation for subsequent in-depth analysis. Employing differential analysis, we pinpointed genes exhibiting differential expression in epithelial cells between normal control and tumor samples, thereby laying the foundation for further investigations. Through the multifaceted assessment of gene sets and a deliberate focus on sphingolipid metabolism-related genes, our study zeroed in on the pivotal role of epithelial cells in the context of esophageal squamous cell carcinoma.
(A) A heatmap illustrates the scoring scores for the entire gene set on the tSNE plot, with brighter colors indicating higher gene set scores. (B) Violin plots depict scores for different cell types in normal control and tumor samples for the three scoring modalities, along with violin plots presenting the final scores. Normal control samples are represented in pink, while tumor tissue samples are denoted in cyan. (C) Bubble plots illustrate the correlation between the three single-cell scoring methods and the total score across the nine cell types. Bubble size reflects the magnitude of the correlation, transitioning from blue to red indicates the shift from negative to positive correlation. (D) Violin plots facilitate a comparison of scores among cells of the nine cell types in normal control and tumor tissue samples, with significance levels denoted as follows: *p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001. (E) A heatmap showcases the expression levels of seven sphingolipid metabolism-related genes across various cell types, with color gradients from grey to green signifying increased epitope levels
3.4 Cellular communication analysis and pseudotime analysis of epithelial cells based on single-cell data
To gain deeper insights into single-cell epithelial cell behavior, particularly their association with sphingolipid metabolism, we conducted comprehensive analyses encompassing cell communication and pseudotime analysis within the epithelial cell population isolated from tumor tissues, utilizing Seurat objects. Employing the cellchat package, we initially performed cell communication analysis, elucidating the quantitative exchange of signaling events by epithelial cells (Fig. 3A). Notably, our analysis revealed the pivotal involvement of the PTN signaling pathway in intercellular communication within the epithelial cell milieu. Visualized through chordal plots (Fig. 3B), the influence of the PTN signaling pathway became evident. We further delineated the significance of four distinct ligand-receptor pairs within this pathway, with PTN-NCL exhibiting the highest prominence (Fig. 3C). To enhance our comprehension of this specific ligand-receptor pair, we constructed a circular diagram (Fig. 3D) to portray its communication dynamics. In Fig. 3E, we ascertained the multifaceted roles played by epithelial cells in the PTN signaling pathway, encompassing their roles as senders, receivers, mediators, and influencers. To unveil the differentiation and evolutionary trajectory of these cells at the single-cell level, we embarked on pseudotime analysis and cell trajectory analysis to recapitulate their developmental progression. Figure 3F depicts the chronological evolution of sphingolipid metabolism gene expression in epithelial cells, revealing distinct temporal patterns. Notably, ARSD, CTSA, DEGS1, and PPTQ exhibited peak expression towards the end of cellular development, while PSAP and HEXA were prominently expressed in the early stages. Utilizing the ‘‘monocle’’ package, we observed the pseudotime distribution of tumor cells, shedding light on their developmental trajectory, as showcased in Fig. 3G-I. To investigate the impact of sphingolipid metabolism on epithelial cell communication with other cellular populations, we stratified epithelial cells into high-expression (SMhigh) and low-expression (SMlow) groups for CellCall cell communication analysis. Leveraging a priori knowledge of ligand-receptor-transcription factor (L-R-TF) interactions, we discerned intercellular communication patterns based on ligand and receptor expression levels and downstream TF activities. This comprehensive analysis elucidated key pathways involved in intercellular communication and depicted the strength of communication between high and low sphingolipid metabolizing epithelial cells and various cell types through bubble plots (Fig. 3J). Detailed communication scores of L-R interactions among different cell types were visualized in heatmaps (Fig. 3K). In summary, our findings underscore the nuanced cellular communication dynamics within the tumor microenvironment, driven in part by sphingolipid metabolism variations among epithelial cells.
(A) The upper panel displays the frequency with which epithelial cells receive signals from various cell types, while the lower panel illustrates the frequency of signals sent by epithelial cells to other cell types. (B) Depiction of the interactions within the PTN signaling pathway among diverse cell types, with line thickness indicating the frequency of interactions. (C) A bar graph demonstrates the contribution of four ligand-receptor pairs in the PTN signaling pathway. (D) A string diagram illustrates interactions within the PTN-NCL ligand-receptor pathway among different cell types, with line thickness indicating the strength of these interactions. (E) A heatmap reveals the roles of various cell types in cellular communication, with different cell types on the horizontal axis and four communication role categories on the vertical axis. Darker colors signify a higher significance of a particular cell type in a given role. (F) A heatmap showcases the expression profiles of key genes related to sphingolipid metabolism in pseudotemporal order. The horizontal axis represents the pseudotemporal order from pseudotime analysis, while the vertical axis lists the 13 key sphingolipid metabolism-related genes in epithelial cells. Colors from blue to red indicate gene expression levels, and the clustering tree on the left groups genes with similar expression patterns. (G) tSEN (t-distributed stochastic neighbor embedding) plot of epithelial cells in tumor tissues, with each color indicating a cluster of cells that share similar characteristics. (H) A diagram depicting the trajectories of pseudotime analysis, where each color corresponds to a cluster of cells, and black circular dots represent branching points in cellular development. (I) Pseudotime analysis trajectory plot, where each colored band signifies a different stage of cell development, and circular black dots denote branching nodes in the trajectory. (J) A correlation bubble diagram illustrates cellular communication pathways between various cell types and epithelial cells with high or low expression of sphingolipid metabolism-related genes. Bubble size corresponds to the strength of the correlation, with color transitioning from blue to red indicating the shift from negative to positive correlation. (K) A heatmap depicts the correlation between cell–cell ligand-receptor interactions
3.5 KEGG enrichment analysis of epithelial cells
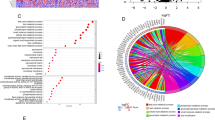
To gain deeper insights into the gene expression patterns of epithelial cells within esophageal squamous cell carcinoma tumor tissues, we employed the Gene Set Enrichment Analysis (GSEA) method, focusing on KEGG pathway enrichment analysis. The results illuminated pronounced enrichments at the apex of the gene set, distinguishing tumor samples from normal control samples. Specifically, pathways related to phagosomes, cell adhesion molecules, lysosomes, and antigen processing and presentation exhibited prominent enrichment (Fig. 4A, B, C, and F). Conversely, pathways such as glycolysis/gluconeogenesis and tyrosine metabolism were situated at the nadir of the gene set's enrichment landscape (Fig. 4D, E).
A GSEA was conducted using KEGG gene sets, where NES represents the normalized enrichment score and FDR denotes the p-value adjusted using the False Discovery Rate method. Each dot on the plot corresponds to an enrichment score, with red indicating high expression and blue representing low expression. Notably, phagolysosomes were found to be significantly upregulated and enriched in epithelial cells. B Upregulated enrichment of cell adhesion molecules. C Enrichment of lysosomes exhibiting upregulation. D Downregulated enrichment associated with glycolysis/glycogenesis. E Downregulated enrichment in tyrosine metabolism. F Upregulated enrichment of antigen processing and presentation pathways
3.6 Epithelial cell genes combined with machine learning to construct prognostic model
Following the extraction of differentially expressed genes within epithelial cells, we performed a clustering analysis utilizing WGCNA to delineate distinct gene modules. Subsequently, we amalgamated modules with analogous expression profiles, thereby yielding a comprehensive clustering of distinct modules between normal and tumor states. From these modules, we selected the top 100 genes in each of the most statistically significant modules, denoted as the yellow, green, brown, and blue modules, thus constituting the core gene set (Fig. 5A, B). In the ensuing steps, we partitioned the TCGA dataset into training and validation sets in a randomized 3:1 ratio, and augmented it with the GSE53624 dataset for external validation. Employing ten distinct machine learning algorithms on this gene set and evaluating their performance using the C-index metric, the RSF + Lasso algorithm emerged as the optimal choice. This algorithm yielded a selection of seven genes (ARL6IP5, COX17, EIF1B, PPP1R15A, SNRPB, TUBB2A, VOPP1), forming the basis for the construction of the prognostic model (Fig. 5C). In a final exploration, we elucidated the positive and negative expression relationships that exist among these seven genes (Fig. 5D).
A–B Utilization of weighted gene co-expression network analysis (WGCNA) to cluster and identify core genes within epithelial cells. C Application of ten distinct machine learning algorithms for model construction, with subsequent selection of the most optimal method for prognostic modeling. D Examination of correlation patterns among the model genes
3.7 Model construction and test of predictive value of disease
To construct our prediction model, we meticulously partitioned the samples into distinct training and test groups, wherein TCGA samples were internally subdivided for this purpose, and the GSE53624 dataset was employed as an external test set to ensure robustness and generalization. The training group facilitated the model’s development, while the test group was instrumental in rigorously validating its accuracy. We subsequently ranked the risk scores assigned to esophageal cancer patients across all three datasets and thoughtfully visualized them in scatter plots alongside their corresponding survival status, unveiling a pronounced positive correlation between heightened risk scores and increased mortality rates (Fig. 6A–C). Leveraging the R package ‘‘pheatmap,’’ we skillfully generated heatmaps (Fig. 6A–C) to illuminate the distribution of expression patterns exhibited by the seven OS-related genes among high-risk and low-risk patient groups. Within the training group, two genes (VOPP1 and ARL6IP5) exhibited strong positive correlations in the low-risk subgroup and similarly in the high-risk subgroup, thus designating them as low-risk genes. In contrast, the remaining five genes (TUBB2A, SNRPB, PPP1R15A, EIF1B, and COX17) displayed high positive correlations in the high-risk subgroup and strikingly negative correlations in the low-risk subgroup, categorizing them as risk genes. To gauge the prognostic significance of our meticulously constructed risk scoring model, we meticulously assessed the relationship between survival prognosis and time. This assessment segregated esophageal cancer patients into high-risk and low-risk groups. Subsequently, we ascertained the model's effectiveness by generating ROC curves, providing a quantitative measure of predictive accuracy. The ROC analysis indicated excellent performance, with AUC values for 1 year, 3 year, and 5 year survival of esophageal cancer patients in the Train (TCGA) dataset registering at 0.913, 0.957, and 1.000, respectively (Fig. 6D). This high accuracy and sensitivity underscored the model's efficacy in prognosticating patient outcomes. To validate our ROC results and eliminate potential bias, we randomly partitioned the acquired Test (TCGA) test subset and the Test (GSE53624) test subset, employing them for internal and external validation, respectively. This thorough validation consistently demonstrated the model’s strong predictive accuracy, with AUC values of 0.474, 0.821, NA, and 0.601, 0.659, and 0.712 after 1, 3, and 5 years for the Test (TCGA) test subset and the Test (GSE53624) test subset, respectively, further affirming its robust performance (Fig. 6D–F). Additionally, we conducted Kaplan–Meier analysis to compare the predictive capabilities of the model and delineated survival curves for the Train (TCGA) dataset, the Test (TCGA) test subset, and the Test (GSE53624) test subset, revealing statistically significant differences (P < 0.05) between the high-risk and low-risk groups in all three datasets (Fig. 6G–I). These findings underscore the higher overall survival rate among esophageal cancer patients in the low-risk group compared to the high-risk group, and illustrate that survival rates in both groups steadily decrease over time.
A–C Segmentation of TCGA (training set), TCGA (test set), and GSE53624 (test set) samples into high and low-risk categories based on the risk score median, with presentation of risk level distributions, survival statuses, and model gene expressions. D–F Performance evaluation of the prognostic model through time-related ROC curve analysis for the three groups, including TCGA (training set), TCGA (test set), and GSE53624 (test set). G–I Generation of survival time difference curves for the three groups, depicting disparities between high and low-risk cohorts
3.8 Correlation analysis between clinicopathologic features of models with clinical subgroup analysis
To explore the association between high- and low-risk groups and various clinical characteristics, we generated correlation heatmaps elucidating the relationships between these groups and gender, stage, T-stage, N-stage, M-stage, and risk scores (Fig. 7A). This comprehensive heatmap encompassed data from all TCGA esophageal cancer patient samples. We delved deeper into the distinctions in the proportions of high- and low-risk groups within different clinical characteristics, including gender, stage, T, N, and M. Remarkably, we observed variations in the patient count across different clinicopathological characteristics between the high- and low-risk groups (Fig. 7B–F). Recognizing the substantial disparities in individual clinical characteristics and overall survival (OS) between these groups, we conducted a focused investigation to ascertain whether patient prognoses diverged within distinct clinical subgroups. We categorized esophageal cancer patients into five unique subgroups based on their clinical attributes, and subsequently assessed differences in survival curves between high- and low-risk groups (Fig. 7B–F). Notably, our analysis revealed that high-risk patients exhibited significantly diminished overall survival compared to their low-risk counterparts in all subgroups, with the exception of female patients and those at the M1 stage. This underscored the advantageous survival benefits conferred upon low-risk patients. In light of these findings, we assert that our meticulously constructed model stands as a robust and dependable clinical prognostic tool, adept at accurately predicting the prognosis of esophageal cancer patients across diverse clinical subgroups.
The outcomes of the correlation analysis between clinicopathologic features and clinical subgroups are illustrated as follows: A A heatmap displaying the relationship between risk scores and clinicopathologic characteristics. Additionally, B–F present the distinctions in patient distribution between the high-risk and low-risk groups across various pathological attributes, along with the findings from the clinical subgroup analysis encompassing B gender, C staging, D T-staging, E N-staging, and F M-staging
3.9 Independent prognostic analysis of clinical characteristics with nomograms
Considering the robust correlation between the risk model we meticulously constructed and unfavorable prognostic outcomes, we conducted both univariate and multivariate independent prognostic analyses, assessing the potential of these seven genes as autonomous prognostic factors for each esophageal cancer patient. The results of the univariate analysis unveiled that stage (p < 0.001), M stage (p < 0.001), and N stage (p < 0.001) exhibited significant associations with the prognoses of esophageal cancer patients (Fig. 8A). Subsequently, following an extensive multifactorial Cox analysis, we discerned that the sole clinical characteristic exhibiting independent prognostic significance was the risk score (p < 0.001). These outcomes underscored the reliability and autonomy of the risk score as a prognostic predictor (all p-values < 0.05) (Fig. 8B). In an endeavor to broaden the clinical applicability and utility of our meticulously constructed risk model, we devised a column-line plot integrating N stage, M stage, and risk score, which functioned as a tool for predicting 1-, 3-, and 5 year prognostic survival probabilities (Fig. 8C). The risk score wielded the most substantial impact on OS prediction, thereby intimating that esophageal cancer prognoses could be more precisely predicted through the utilization of this risk model. The calibration curves emphasise the consistency between predicted and observed values associated with the 1-, 3- and 5 year OS probabilities, and have almost identical trends to the diagonal, suggesting that the Nomogram is highly stable (Fig. 8D). Furthermore, the c-index values of the risk score, Net Benefit, and area under the ROC curve eclipsed those of other clinical features, providing further testament to the superior predictive accuracy of our constructed model concerning patient survival (Fig. 8E, F).
Independent prognostic analyses were conducted as follows: A Univariate independent prognostic analyses were performed with various clinical indicators. B Subsequently, multifactorial Cox analyses were carried out with different clinical indicators. C A nomogram plot was constructed by integrating clinical indicators. D Calibration curves were generated for 1, 2, and 3 years to assess the accuracy of the nomogram. E Furthermore, a comparison was made among risk scores, sexes, stages, and C-index values of M and N. F Decision curve analysis (DCA) was performed, assessing different clinical traits such as D, T, E, N, and N for their potential in clinical decision-making
3.10 Analysis of differential and modelled gene enrichment between high and low risk groups
To decipher the biological processes under the regulatory influence of the identified model genes, subsequently impacting patient prognostic outcomes, we conducted a comprehensive GSEA analysis targeting these model genes. The analysis yielded intriguing results, highlighting the up-regulation of pathways such as drug metabolism cytochrome P450, glutathione metabolism, and olfactory transduction in the high-risk group, while revealing down-regulation of pathways such as cell adhesion molecules (CAMs), dilated cardiomyopathy, and focal adhesion in the low-risk group (Fig. 9A, B). Furthermore, our quest for pathway enrichment among differentially expressed genes between the high and low-risk groups led us to the observation that these genes were significantly enriched in the Transcriptional misregulation in cancer pathway (Fig. 9C). To gain a more comprehensive understanding of the pathway enrichment distinctions in the high and low-risk groups, we conducted GSVA analysis, unveiling that the low-risk group exhibited high expression in pathways including glycosphingolipid biosynthesis—ganglio series, other glycan degradation, intestinal immune network for IgA production, dorsoventral axis formation, phosphatidylinositol signaling system, and inositol phosphate metabolism, whereas the remaining KEGG pathways displayed elevated expression in the high-risk group (Fig. 9D).
To elucidate the differences in biological processes influenced by the identified model genes and their impact on patient prognosis, comprehensive enrichment analyses were conducted. A–B Firstly, GSEA analysis was employed to discern the up- and down-regulated pathways in both high and low risk groups. C Subsequently, KEGG enrichment analysis was performed to investigate the functional pathways enriched with differential genes in the high and low risk groups. D Furthermore, GSVA analysis was utilized to ascertain the differential expression of various KEGG pathways in these high and low risk groups. These multifaceted analyses provided insights into the potential mechanisms underlying the observed differences in patient outcomes associated with the model genes
3.11 Immunological relevance of modelling genes and the tumor microenvironment
Our investigation delved into the correlation between each of the model genes and various immune cell types, revealing noteworthy associations. Specifically, ARL6IP5 exhibited a positive correlation with a diverse array of T cells, while COX17 displayed a negative correlation with multiple T cell subtypes. Moreover, EIF1B showed a positive correlation with Eosinophils, SNRPB exhibited a negative correlation with Mast cells, and TUBB2A displayed a negative correlation with T helper cells, all of which warrant further scrutiny (Fig. 10A–G). Simultaneously, as we explored the interplay between genes and immune cells, we initiated a correlation analysis among different cell types within esophageal cancer patients. Notably, a negative correlation emerged between Macrophages and Endothelial cells, as well as B cells. Furthermore, the negative correlation between TUBB2A, SNRPB, and PPP1R15A with B cells appeared to be more pronounced, suggesting potential associations of these genes with Macrophages and Endothelial cells (Fig. 10H, J). Disparities in the expression of immune checkpoints between the high- and low-risk groups were evident (Fig. 10I). Additionally, variations in Stromal Score, Immune Score, and the composite score of the tumor microenvironment were observed between these groups (Fig. 10K). These distinctions may underlie the differences in survival outcomes between high- and low-risk groups.
Immunological correlation of modelling genes and tumor microenvironment A–G Correlation analysis of seven modelling genes with different immune cells H Degree of correlation between different immune cells and positive and negative regulatory relationship between modelling genes and immune cells (I) Differential expression of different immune checkpoints in high and low risk groups (J) Correlation analysis of modelling genes with immune cells (K) Differences in scores of the tumor microenvironment in the high and low risk groups * p < 0.05; * p < 0.01; *** p < 0.001
3.12 Comparison of somatic mutations in the low and high risk groups
Leveraging somatic mutation frequency data from TCGA pertaining to oesophageal cancer, we conducted a comparative analysis of gene mutations within the high and low-risk groups. Our findings revealed notable distinctions, with MUC16 exhibiting a higher mutation frequency in the low-risk group (24%) compared to the high-risk group (19%), a trend similarly observed for SYNE1 (Fig. 11A, B). Notably, single nucleotide polymorphisms (SNPs) emerged as the predominant mutation category in oesophageal cancer, with TP53, TTN, and MUC16 ranking as the top three genes with the highest mutation frequencies (Fig. 11C). To delve into the relationship between the expression of model genes and TMB, we discerned a direct correlation whereby increased ARL6IP5 expression corresponded to an elevated TMB fraction (Fig. 11D). Further unveiling the connection between high and low mutation loads and patient survival, we conducted survival curve analysis. This analysis unveiled that patients with higher TMB exhibited poorer prognostic outcomes compared to those with lower TMB. Notably, patients with the bleakest prognoses were those with higher TMB levels coupled with elevated risk scores, underscoring the significance of tumor mutation load and risk score in shaping patient prognostic outcomes (Fig. 11E, F).
The mutational landscape of esophageal cancer in both high- and low-risk groups was meticulously examined, unveiling critical insights into the genomic alterations associated with risk stratification and signature genes. A, B Comparative analysis of the mutation status and the identity of mutated genes between the high- and low-risk groups was carried out. C This analysis was situated within the broader context of the overall mutational profile of esophageal cancer. D The association between the expression levels of the ARL6IP5 gene and the tumor mutation load was elucidated, shedding light on potential relationships between this gene and genomic instability. E Furthermore, the impact of tumor mutation load levels on patient survival time was assessed. F Notably, the patients' survival time was evaluated following a combined stratification based on both tumor mutation load and the risk score, offering a comprehensive view of their prognostic implications. These mutational analyses provide valuable insights into the genetic underpinnings of risk stratification and potential drivers of patient outcomes in esophageal cancer
3.13 Differential analysis of drug sensitivity
Risk scores hold the potential for extensive applications in the realm of immunotherapy for esophageal cancer patients, facilitating both in-depth analyses of treatment efficacy and the fine-tuning of drug dosages. Within a cohort encompassing nine immunotherapeutic agents prescribed for esophageal cancer treatment, a statistically significant disparity in drug sensitivity emerged when comparing the high- and low-risk groups (P < 0.05). Notably, one specific drug, ERK_6604, exhibited a comparatively lower IC50 in the high-risk group, indicating heightened sensitivity within this subgroup (Fig. 12C). In contrast, the remaining eight drugs (Afuresertib, AT13148, ERK6604, GSK269962A, JQ1, Nutlin-3a, PD173074, and RO-3308) demonstrated elevated IC50 values within the high-risk group, signifying diminished sensitivity in comparison to the low-risk group (Fig. 12A, B, D, E, F, G, H, I). These findings underscore the pivotal role of risk scores in advancing our comprehension of immunotherapy responses among esophageal cancer patients, ultimately enhancing the precision and efficacy of therapeutic interventions.
4 Discussion
Esophageal cancer represents a highly aggressive malignancy within the digestive tract, characterized by a notably elevated propensity for metastasis, and regrettably, patients confronting invasive and metastatic disease frequently experience diminished survival prospects. This somber reality underscores esophageal cancer's standing as a leading contributor to global cancer-related mortality [39,40,41]. Current therapeutic approaches for esophageal cancer encompass a spectrum of modalities, including surgical resection [42], radiation therapy [43], chemotherapy [44], targeted therapy [45], and immunotherapy [46]. Notably, despite documented efficacy in certain cases, such as surgical, chemotherapeutic, and radiotherapeutic interventions, the overall prognosis for esophageal cancer patients remains disheartening [47]. These circumstances underscore the imperative for the development of novel and more effective strategies aimed at ameliorating the prognostic landscape for individuals afflicted by esophageal cancer. Furthermore, the significance of early detection and intervention in the context of esophageal cancer cannot be overstated, as progression to advanced stages frequently entails infiltration into neighboring lymph nodes and tissues, markedly complicating treatment endeavors.
Single-cell analysis has emerged as a pivotal tool in the comprehensive exploration of tumor biology, offering invaluable insights into the intricate attributes, functionalities, and intricate interplay among diverse cell subtypes within neoplastic lesions. Its utility is particularly pronounced in investigations pertaining to intra-tumor heterogeneity, the delineation of cellular differentiation trajectories, and the dissection of intercellular communication networks [48]. In the realm of ESCC research, single-cell analysis stands as an indispensable asset, furnishing an unparalleled depth of high-resolution data at the cellular stratum. This has significantly advanced our comprehension of the underpinnings of ESCC pathogenesis, informed the development of therapeutic paradigms, and enhanced the precision of prognostic assessments [49].
In this comprehensive study, we harnessed the power of scRNA-seq data and meticulously curated a set of genes closely associated with epithelial cells to delve into the prognostic landscape of ESCC. Our investigation entailed a meticulous comparative analysis of the single-cell profiles between ESCC tumor tissues and their normal counterparts at the single-cell level, employing a refined single-cell scoring methodology. Leveraging Cox one-way regression analysis, we successfully winnowed down a roster of ESCC epithelial cell-related genes with notable prognostic significance. Subsequently, through a systematic exploration of 101 combinations of machine learning algorithms, we meticulously identified the optimal amalgamation of machine learning algorithms that underpinned the construction of our predictive model. This culminated in the identification of seven genes, each demonstrating robust independent prognostic potential, thus constituting the core variables within our modeling framework. Utilizing these molecular features, we derived a risk score scheme that stratified ESCC patients into distinct cohorts characterized as high risk and low risk. Notably, our meticulous examination of prognostic survival curves and immune infiltration patterns unearthed a pronounced disparity between these two groups, with the low-risk cohort demonstrating notably favorable prognoses, a disparity that reached statistical significance (P < 0.05). Furthermore, our scrutiny extended to pathway enrichment analyses, uncovering a predilection for pathways linked to drug metabolism, cytochrome activity, glutathione metabolism, olfactory transduction, ribosomal processes, and insulin signaling within the high-risk group, shedding light on potential mechanistic underpinnings of their unfavorable prognosis. Importantly, our correlation analysis of clinicopathological attributes and subgroup-specific prognostic assessments underscored the clinical relevance and applicability of our risk model, presenting clinicians with a robust tool for making precise therapeutic decisions tailored to ESCC patients and ultimately enhancing the overall quality of patient survival.
Sphingolipid metabolism, a complex and multifaceted cellular process, exerts profound influences on vital cellular mechanisms encompassing apoptosis, cell signaling, and immune modulation, thereby intricately governing cell behaviors such as proliferation, migration, and differentiation [50]. Extensive research endeavors have illuminated the conspicuous interplay between sphingolipid metabolism and cancer metastasis, with a particular emphasis on the pivotal role of sphingomyelin synthesis in dictating the growth and metabolic dynamics of cancer cells [51]. Perturbations in sphingolipid metabolism have been inexorably linked to an array of pathological conditions, spanning metabolic disorders, neurological ailments, and several cancer types, prompting the exploration of sphingomyelin-metabolizing enzymes as promising candidates for targeted therapeutic interventions against cancer [52]. In the context of this study, our meticulous examination unveiled a pronounced and notable correlation amongst sphingolipid metabolism-related genes within epithelial cells. This intriguing observation underscores the tantalizing prospect that epithelial cell genes intricately intertwined with sphingolipid metabolism could conceivably serve as novel and potent prognostic biomarkers germane to ESCC.
The genesis of ESCC can be traced back to the esophageal squamous epithelium, where a cascade of early events unfolds, marked by the abnormal proliferation and genetic variations of epithelial cells, as evidenced by prior research [2]. These initial molecular perturbations set the stage for the development of malignant phenotypes characterized by unbridled cellular proliferation, evasion of apoptosis, and the promotion of invasive and metastatic behaviors. In the context of ESCC, the mutation-induced damage incurred by epithelial cells precipitates a functional compromise in the mucosal barrier, rendering esophageal tissues more vulnerable to harm inflicted by deleterious factors [53]. The ensuing malignancy in ESCC is typified by the capacity of malignant cells to breach the mucosal barrier, infiltrate deeper layers of tissue, and subsequently embark on metastatic journeys to distant organs, including lymph nodes and the stomach [54]. These transformations in cell behavior, notably the malignant transition and heightened invasive propensity exhibited by epithelial cells, stand as pivotal and orchestrating events in this intricate process. Consequently, in-depth research centered on the dynamics of epithelial cells holds substantial promise in unraveling the underlying mechanisms, thereby furnishing valuable insights that can be harnessed for the prevention, early detection, and treatment of ESCC. Ultimately, these endeavors aim to ameliorate the survival rates and enhance the quality of life for ESCC patients.
COX17 is involved in the assembly and function of cytochrome c oxidase and plays a key role in maintaining mitochondrial function and energy metabolism [55]. It has been reported that deletion of COX17 acetylation disrupts mitochondrial morphology and ultrastructure [56]. COX17 may influence cellular energy supply and growth by regulating mitochondrial function, and indirectly affects adaptation and viability of metabolic pathways, such as sphingolipid metabolism, in tumors. ARL6IP5 encodes a protein involved in cell membrane stability and endocytic trafficking. It has been shown that ARL6IP5 inhibits tumor DNA repair, promotes apoptosis and autophagy [57, 58]. EIF1B is part of the translation initiation factors involved in the initiation of mRNA translation [59, 60]). In cancer cells, EIF1B may regulate key phases of the cell cycle and influence cell proliferation and apoptosis processes, thereby promoting tumor growth and progression. As a regulator of protein phosphatase 1, PPP1R15A is involved in the regulation of cellular stress response and cell cycle [61]. Its abnormality may lead to dysregulation of the stress response, thereby affecting tumor cell response and drug resistance to therapy [62]. SNRPB is a component of the spliceosome and is involved in the splicing and processing of precursor mRNAs [63]. It is essential for the regulation of gene expression.Abnormalities in SNRPB may lead to altered expression patterns of cancer-related genes, which in turn affects the biological behavior of cancer cells [64]. TUBB2A encodes β-microtubule protein, a key factor in microtubule assembly and cytoskeleton maintenance, which plays a role in cell division and migration [65].TUBB2A also serves as a novel biomarker for the prediction of distant metastatic breast cancer [66].
TMB, quantifying the genomic mutations within tumor cells [67], exerts a pivotal influence on tumor progression and the responsiveness to conventional therapeutic modalities, thereby emerging as a promising avenue in cancer treatment strategies [68, 69]. Significantly, TMB scores exhibited a statistically significant distinction in immunization scores between the high-risk and low-risk cohorts, with their combined scores effectively stratifying the prognosis of ESCC. This compelling observation suggests that individuals in the high-risk category, characterized by elevated TMB levels, may demonstrate heightened receptivity to immunotherapeutic interventions. In contrast, those in the low-risk category, coupled with a low TMB profile, exhibit a more favorable prognostic outlook.
While our study has yielded a prognostic model founded upon ESCC epithelial cell-related genes that holds substantial implications for ESCC prognosis and immune responses, offering novel insights into prospective prognostic biomarker development, as well as unraveling ESCC histiocyte-related pathways, cellular differentiation trajectories, and inter-cellular communication at the single-cell resolution, we acknowledge the inherent limitations within our research. Primarily, our utilization of publicly available databases resulted in relatively limited and modest sample sizes within our validation cohorts. Secondly, the inability of single-cell sequencing technology to fully capture the heterogeneity of the tumor microenvironment, as well as data sampling bias may have biased the findings. Furthermore, our study underscores the imperative need for subsequent experimental investigations to comprehensively elucidate the biological functions and putative mechanisms of action attributed to the identified candidate genes. In summary, the prognostic model established in this study holds promise as a pivotal tool for prognosticating ESCC patient outcomes. Simultaneously, it paves the way for substantial prospects in the integration of immunotherapeutic modalities within clinical ESCC management.
5 Conclusion
Our investigation adeptly integrates single-cell analysis with high-throughput transcriptomics, culminating in the development of a robust prognostic framework linked to sphingolipid metabolism. This pivotal advancement harbors extensive ramifications, particularly in the realms of precision therapeutic interventions and customized immunotherapeutic approaches catering to individuals afflicted by esophageal carcinoma.
Data availability
The datasets analyzed in the current study are available in the GEO repository (https://www.ncbi.nlm.nih.gov/geo/)and Xena’s Web site (https://xena.ucsc.edu/). The datasets used and/or analyzed during the current study are available from the corresponding author upon reasonable request.
References
Chen C, Yang C, Tian X, Liang Y, Wang S, Wang X, Shou Y, Li H, Xiao Q, Shu J, Sun M, Chen K. Downregulation of miR-100-5p in cancer-associated fibroblast-derived exosomes facilitates lymphangiogenesis in esophageal squamous cell carcinoma. Cancer Med. 2023;12:14468–83.
Yang D, Li R, Wang H, Wang J, Li Y, Wang H, Wang W, Liu Z. Clinical significance of tumor necrosis factor receptor 2 in middle and lower thoracic esophageal squamous cell carcinoma. Oncol Lett. 2018;16:2971–8.
Wang Y, Cheng J, Xie D, Ding X, Hou H, Chen X, Er P, Zhang F, Zhao L, Yuan Z, Pang Q, Wang P, Qian D. NS1-binding protein radiosensitizes esophageal squamous cell carcinoma by transcriptionally suppressing c-Myc. Cancer Commun. 2018;38:33.
Ren Q, Zhang P, Zhang S, Chen W, Chi H, Wang W, Zhang W, Lin H, Yu Y. A SARS-CoV-2 related signature that explores the tumor microenvironment and predicts immunotherapy response in esophageal squamous cell cancer. Aging. 2023;15:10501–23.
Sung H, Ferlay J, Siegel RL, Laversanne M, Soerjomataram I, Jemal A, Bray F. Global cancer statistics estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2021;71:209–49.
Arnold M, Soerjomataram I, Ferlay J, Forman D. Global incidence of oesophageal cancer by histological subtype in 2012. Gut. 2015;64:381–7.
Then EO, Lopez M, Saleem S, Gayam V, Sunkara T, Culliford A, Gaduputi V. Esophageal cancer: an updated surveillance epidemiology and end results database analysis. World J Oncol. 2020;11:55–64.
Uhlenhopp DJ, Then EO, Sunkara T, Gaduputi V. Epidemiology of esophageal cancer: update in global trends, etiology and risk factors. Clin J Gastroenterol. 2020;13:1010–21.
da Costa NM, Soares Lima SC, de Almeida Simão T, Ribeiro Pinto LF. The potential of molecular markers to improve interventions through the natural history of oesophageal squamous cell carcinoma. 2013. Biosci Rep. https://doi.org/10.1042/BSR20130063.
Rustgi A, El-Serag HB. Esophageal carcinoma. N Engl J Med. 2015;372:1472–3.
Motoyama S, Hosaka M, Kamei T, Ueno M, Watanabe M, Ohkura YU, Mine S, Tanabe S, Toyokawa T, Wakita A, Nishikawa K, Ninomiya I, Fujita K, Kanda M, Hirai M, Hoshi M, Chiu SW, Takata M, Yamaguchi T, Miura M. A multi-institutional study to diagnose the risk of lymph node metastasis using a CRP genetic polymorphism test kit in pT1, cN0 thoracic esophageal squamous cell carcinoma. Anticancer Res. 2022;42:6105–12.
Zhang Y, Wang D, Peng M, Tang L, Ouyang J, Xiong F, Guo C, Tang Y, Zhou Y, Liao Q, Wu X, Wang H, Yu J, Li Y, Li X, Li G, Zeng Z, Tan Y, Xiong W. Single-cell RNA sequencing in cancer research. J Experim Clin Cancer research : CR. 2021;40:81.
Potter JW, Jones KB, Barrott JJ. Sarcoma-the standard-bearer in cancer discovery. Crit Rev Oncol Hematol. 2018;126:1–5.
Xia C, Su J, Liu C, Mai Z, Yin S, Yang C, Fu L. Human microbiomes in cancer development and therapy. MedComm. 2023;4: e221.
Ogretmen B. Sphingolipid metabolism in cancer signalling and therapy. Nat Rev Cancer. 2018;18:33–50.
Gault CR, Obeid LM, Hannun YA. An overview of sphingolipid metabolism: from synthesis to breakdown. Adv Exp Med Biol. 2010;688:1–23.
Voelkel-Johnson C, Norris JS, White-Gilbertson S. Interdiction of sphingolipid metabolism revisited: focus on prostate cancer. Adv Cancer Res. 2018;140:265–93.
Parveen F, Bender D, Law SH, Mishra VK, Chen CC, Ke LY. Role of ceramidases in sphingolipid metabolism and human diseases. Cells. 2019;8:1573.
Beckham TH, Elojeimy S, Cheng JC, Turner LS, Hoffman SR, Norris JS, Liu X. Targeting sphingolipid metabolism in head and neck cancer: rational therapeutic potentials. Expert Opin Ther Targets. 2010;14:529–39.
Yang PW, Lin MC, Huang PM, Wang CP, Chen TC, Chen CN, Tsai MH, Cheng JC, Chuang EY, Hsieh MS, Lou PJ, Lee JM. Risk factors and genetic biomarkers of multiple primary cancers in esophageal cancer patients. Front Oncol. 2020;10: 585621.
Li D, Zhang L, Liu Y, Sun H, Onwuka JU, Zhao Z, Tian W, Xu J, Zhao Y, Xu H. Specific DNA methylation markers in the diagnosis and prognosis of esophageal cancer. Aging. 2019;11:11640–58.
Shlyakhto EV. Scientific basics of personalized medicine: realities and opportunities. Her Russ Acad Sci. 2022;92:671–82.
Hao Y, Hao S, Andersen-Nissen E, Mauck WM 3rd, Zheng S, Butler A, Lee MJ, Wilk AJ, Darby C, Zager M, Hoffman P, Stoeckius M, Papalexi E, Mimitou EP, Jain J, Srivastava A, Stuart T, Fleming LM, Yeung B, Rogers AJ, McElrath JM, Blish CA, Gottardo R, Smibert P, Satija R. Integrated analysis of multimodal single-cell data. Cell. 2021;184:3573-3587.e29.
Andreatta M, Carmona SJ. UCell: Robust and scalable single-cell gene signature scoring. Comput Struct Biotechnol J. 2021;19:3796–8.
Jin S, Guerrero-Juarez CF, Zhang L, Chang I, Ramos R, Kuan CH, Myung P, Plikus MV, Nie Q. Inference and analysis of cell-cell communication using Cell Chat. Nat Commun. 2021;12:1088.
Qiu X, Hill A, Packer J, Lin D, Ma YA, Trapnell C. Single-cell mRNA quantification and differential analysis with Census. Nat Methods. 2017;14:309–15.
Qiu X, Mao Q, Tang Y, Wang L, Chawla R, Pliner HA, Trapnell C. Reversed graph embedding resolves complex single-cell trajectories. Nat Methods. 2017;14:979–82.
Zhang Y, Liu T, Hu X, Wang M, Wang J, Zou B, Tan P, Cui T, Dou Y, Ning L. Cell call: integrating paired ligand–receptor and transcription factor activities for cell–cell communication. Nucl Acids Res. 2021;49(2021):8520–34.
Chi H, Chen H, Wang R, Zhang J, Jiang L, Zhang S, Jiang C, Huang J, Quan X, Liu Y, Zhang Q, Yang G. Proposing new early detection indicators for pancreatic cancer: combining machine learning and neural networks for serum miRNA-based diagnostic model. Front Oncol. 2023;13:1244578.
Song B, Chi H, Peng G, Song Y, Cui Z, Zhu Y, Chen G, Wu J, Liu W, Dong C, Wang Y, Xu K, Yu Z, Song B. Characterization of coagulation-related gene signature to predict prognosis and tumor immune microenvironment in skin cutaneous melanoma. Front Oncol. 2022;12: 975255.
Song B, Wu P, Liang Z, Wang J, Zheng Y, Wang Y, Chi H, Li Z, Song Y, Yin X, Yu Z, Song B. A novel necroptosis-related gene signature in skin cutaneous melanoma prognosis and tumor microenvironment. Front Genet. 2022;13: 917007.
Shen Y, Chi H, Xu K, Li Y, Yin X, Chen S, Yang Q, He M, Zhu G, Li X. A novel classification model for lower-grade glioma patients based on pyroptosis-related genes. Brain Sci. 2022. https://doi.org/10.3390/brainsci12060700.
Huang X, Chi H, Gou S, Guo X, Li L, Peng G, Zhang J, Xu J, Nian S, Yuan Q. An aggrephagy-related LncRNA signature for the prognosis of pancreatic adenocarcinoma. Genes. 2023. https://doi.org/10.3390/genes14010124.
Huang J, Liu M, Chen H, Zhang J, Xie X, Jiang L, Zhang S, Jiang C, Zhang J, Zhang Q, Yang G, Chi H, Tian G. Elucidating the Influence of MPT-driven necrosis-linked LncRNAs on immunotherapy outcomes, sensitivity to chemotherapy, and mechanisms of cell death in clear cell renal carcinoma. Front Oncol. 2023;13:1276715.
Jiang C, Zhang S, Jiang L, Chen Z, Chen H, Huang J, Tang J, Luo X, Yang G, Liu J, Chi H. Precision unveiled: Synergistic genomic landscapes in breast cancer-Integrating single-cell analysis and decoding drug toxicity for elite prognostication and tailored therapeutics. Environ Toxicol. 2024;39:3448–72.
Zhang S, Jiang C, Jiang L, Chen H, Huang J, Zhang J, Wang R, Chi H, Yang G, Tian G. Uncovering the immune microenvironment and molecular subtypes of hepatitis B-related liver cirrhosis and developing stable a diagnostic differential model by machine learning and artificial neural networks. Front Mol Biosci. 2023;10:1275897.
Jiang L, Ren X, Yang J, Chen H, Zhang S, Zhou X, Huang J, Jiang C, Gu Y, Tang J, Yang G, Chi H, Qin J. Mitophagy and clear cell renal cell carcinoma: insights from single-cell and spatial transcriptomics analysis. Front Immunol. 2024;15:1400431.
Jiang L, Liu J, Zhang S, Jiang C, Huang J, Chen H, Zhou X, Fu Y, Yang Z, Wang R, Yang G, Chi H, Li B. Role of glycosylation-related gene MGAT1 in pancreatic ductal adenocarcinoma. Front Immunol. 2024;15:1438935.
Qiao L, Liang N, Xie J, Luo H, Zhang J, Deng G, Li Y, Zhang J. Gene silencing of galectin-3 changes the biological behavior of Eca109 human esophageal cancer cells. Mol Med Rep. 2016;13:160–6.
Zhao Y, Xu L, Wang X, Niu S, Chen H, Li C. A novel prognostic mRNA/miRNA signature for esophageal cancer and its immune landscape in cancer progression. Mol Oncol. 2021;15:1088–109.
Jin W, Yang Q, Chi H, Wei K, Zhang P, Zhao G, Chen S, Xia Z, Li X. Ensemble deep learning enhanced with self-attention for predicting immunotherapeutic responses to cancers. Front Immunol. 2022;13:1025330.
Chen BF, Zhu CC, Wang CG, Ma DH, Lin J, Zhang B, Kong M. Clinical comparative study of minimally invasive esophagectomy versus open esophagectomy for esophageal carcinoma. Zhonghua wai ke za zhi [Chin J Sur]. 2010;48:1206–9.
Choi KH, Kim J, Lee SW, Kang YN, Jang H. Dosimetric comparison between modulated arc therapy and static intensity modulated radiotherapy in thoracic esophageal cancer: a single institutional experience. Radiat Oncol J. 2018;36:63–70.
Ilson DH, Minsky BD, Ku GY, Rusch V, Rizk N, Shah M, Kelsen DP, Capanu M, Tang L, Campbell J, Bains M. Phase 2 trial of induction and concurrent chemoradiotherapy with weekly irinotecan and cisplatin followed by surgery for esophageal cancer. Cancer. 2012;118:2820–7.
Power DG, Ilson DH. Integration of targeted agents in the neo-adjuvant treatment of gastro-esophageal cancers. Ther Adv Med Oncol. 2009;1:145–65.
Sadanand S. Immunotherapy for esophageal cancer. Nature Med. 2021. https://doi.org/10.1038/d41591-021-00022-8.
Song W, Wang H, Tian Y, Liu S, Chen X, Cui J, Zhao Y. Refractory solitary cervical lymph node metastasis after esophageal squamous cell carcinoma surgery and its successful treatment with immune checkpoint inhibitor: a case report and literature review. Medicine. 2020;99: e19440.
Radpour R, Forouharkhou F. Single-cell analysis of tumors: creating new value for molecular biomarker discovery of cancer stem cells and tumor-infiltrating immune cells. World J Stem Cells. 2018;10:160–71.
Guo G, Luc S, Marco E, Lin TW, Peng C, Kerenyi MA, Beyaz S, Kim W, Xu J, Das PP, Neff T, Zou K, Yuan GC, Orkin SH. Mapping cellular hierarchy by single-cell analysis of the cell surface repertoire. Cell Stem Cell. 2013;13:492–505.
Mu X, Ji C, Wang Q, Liu K, Hao X, Zhang G, Shi X, Zhang Y, Gonzalez FJ, Wang Q, Wang Y. Non-targeted metabolomics reveals diagnostic biomarker in the tongue coating of patients with chronic gastritis. J Pharm Biomed Anal. 2019;174:541–51.
Liu XY, Liu YB, Xu JC, Zhang YF, Ruan YY, Zhao Y, Wu LF, Hu JW, Zhang Z, He MJ, Chen TY, Xu XY, Zhang JW, Zhang YQ, Zhou PH. Single-cell transcriptomic analysis deciphers key transitional signatures associated with oncogenic evolution in human intramucosal oesophageal squamous cell carcinoma. Clin Transl Med. 2023;13: e1203.
Kok JW, Sietsma H. Sphingolipid metabolism enzymes as targets for anticancer therapy. Curr Drug Targets. 2004;5:375–82.
Kubala SA, Patil SU, Shreffler WG, Hurley BP. Pathogen induced chemo-attractant hepoxilin A3 drives neutrophils, but not eosinophils across epithelial barriers. Prostaglandins Other Lipid Mediat. 2014;108:1–8.
Kuwano H, Baba K, Ikebe M, Kitamura K, Adachi Y, Mori M, Sugimachi K. Gastric involvement of oesophageal squamous cell carcinoma. Br J Surg. 1992;79:328–30.
Naletova I, Greco V, Sciuto S, Attanasio F, Rizzarelli E. ionophore ability of carnosine and its trehalose conjugate assists copper signal in triggering brain-derived neurotrophic factor and vascular endothelial growth factor activation in vitro. Int J Mol Sci. 2021;22:13504.
Guhathakurta S, Erdogdu NU, Hoffmann JJ, Grzadzielewska I, Schendzielorz A, Seyfferth J, Mårtensson CU, Corrado M, Karoutas A, Warscheid B, Pfanner N, Becker T, Akhtar A. COX17 acetylation via MOF-KANSL complex promotes mitochondrial integrity and function. Nat Metab. 2023;5:1931–52.
Kim JY, Bahar E, Lee JY, Chang S, Kim SH, Park EY, Do SI, Yoon H, Kim HS. ARL6IP5 reduces cisplatin-resistance by suppressing DNA repair and promoting apoptosis pathways in ovarian carcinoma. Cell Death Dis. 2022;13:239.
Siddique I, Kamble K, Gupta S, Solanki K, Bhola S, Ahsan N, Gupta S. ARL6IP5 ameliorates α-synuclein burden by inducing autophagy via preventing ubiquitination and degradation of ATG12. Int J Mol Sci. 2023;24:10499.
Anisimova AS, Kolyupanova NM, Makarova NE, Egorov AA, Kulakovskiy IV, Dmitriev SE. Human tissues exhibit diverse composition of translation machinery. Int J Mol Sci. 2023;24:8361.
R. Grosely, C. Alvarado, I.P. Ivanov, O.B. Nicholson, J.D. Puglisi, T.E. Dever, and C.P. Lapointe, eIF1 and eIF5 dynamically control translation start site fidelity. bioRxiv (2024).
Jiang L, Xu L, Zheng L, Wang Y, Zhuang M, Yang D. Salidroside attenuates sepsis-associated acute lung injury through PPP1R15A mediated endoplasmic reticulum stress inhibition. Bioorg Med Chem. 2022;71: 116865.
Wang R, Zhang Y, Guo S, Pei S, Guo W, Wu Z, Wang H, Wang M, Li Y, Zhu Y, Meng LH, Lang J, Jin G, Xiao Y, Hu L, Kong X. Single-cell RNA sequencing reveals the suppressive effect of PPP1R15A inhibitor Sephin1 in antitumor immunity. Science. 2023;26:105954.
Li Y, Chen Z, Peng J, Yuan C, Yan S, Yang N, Li P, Kong B. The splicing factor SNRPB promotes ovarian cancer progression through regulating aberrant exon skipping of POLA1 and BRCA2. Oncogene. 2023;42:2386–401.
Wu J, Lu F, Yu B, Wang W, Ye X. The oncogenic role of SNRPB in human tumors: a pan-cancer analysis. Front Mol Biosci. 2022;9: 994440.
Bittermann E, Abdelhamed Z, Liegel RP, Menke C, Timms A, Beier DR, Stottmann RW. Differential requirements of tubulin genes in mammalian forebrain development. PLoS Genet. 2019;15: e1008243.
Shin D, Park J, Han D, Moon JH, Ryu HS, Kim Y. Identification of TUBB2A by quantitative proteomic analysis as a novel biomarker for the prediction of distant metastatic breast cancer. Clin Proteom. 2020;17:16.
Liu X, Shi X, Guo W, Ke Y, Li Y, Pan S, Li X, Liu M, Liu M, Wang Y, Ruan Q, Ma H. A promising esophageal cancer prognostic signature of ferroptosis-related LncRNA to predict immune scenery and immunotherapy response. Int J General Med. 2021;14:5845–62.
Yu C, Liu X, Yang J, Zhang M, Jin H, Ma X, Shi H. Combination of immunotherapy with targeted therapy: theory and practice in metastatic melanoma. Front Immunol. 2019;10:990.
Guo Z, Yan X, Song C, Wang Q, Wang Y, Liu XP, Huang J, Li S, Hu W. FAT3 mutation is associated with tumor mutation burden and poor prognosis in esophageal cancer. Front Oncol. 2021;11: 603660.
Funding
This study was not funded.
Author information
Authors and Affiliations
Contributions
XL, JL, and RH conceived the study. RH, JT, HL, TZ, LD, SW, PZ and GT drafted the manuscript. RH, JT, HL, and TZ performed the literature search and collected the data. RH and JT analyzed and visualized the data. XL and JL helped with the final revisions of this manuscript. All authors made significant, direct, and intellectual contributions to this study and consented to its publication.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
He, R., Tang, J., Lai, H. et al. Deciphering the role of sphingolipid metabolism in the immune microenvironment and prognosis of esophageal cancer via single-cell sequencing and bulk data analysis. Discov Onc 15, 505 (2024). https://doi.org/10.1007/s12672-024-01379-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s12672-024-01379-1