Abstract
Gastric cancer (GC) is highly heterogeneous and prone to metastasis, which are obstacles to the effectiveness of treatment. The basement membrane (BM) acts as a barrier to tumor cell invasion and metastasis. It is critical to investigate the relationship between BM status, metastasis, and patient prognosis. In several large cohorts, we investigated BM gene expression-based molecular classification and risk-prognosis models for GC, examined tumor microenvironment (TME) differences among different molecular subtypes, and developed risk models in predicting prognosis, immunotherapy effectiveness, and chemotherapy resistance. Three GC subtypes (BMclusterA/B/C) based on BM gene expression status were discovered. Each of the three GC subtypes has unique immune infiltration and activated oncogenic signals. Moreover, a 6-gene score (BMscore) predictive model was developed. The low BMscore group had a high tumor mutation burden, high immunogenicity, and low RHOJ expression levels, implying that individuals with GC in this category may be more susceptible to immunotherapy and treatment. The EMT subtype showed a considerably higher BMscore than the other subtypes in the Asian Organization for Research on Cancer (ACRG) molecular classification. Endothelial cells, smooth muscle cells, and fibroblasts may be engaged in regulating BM reorganization in GC progression, according to single-cell transcriptome analyses. In conclusion, we defined a novel molecular classification of GC based on BM genes, developed a prognostic risk model, and elucidated the cell subpopulations involved in BM remodeling at the single-cell level. This study has deepened the understanding of the relationship between GC metastasis and BM alterations, achieved prognostic stratification, and guided therapy.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
GC is a major cause of cancer death, and patients can be found all over the globe, with a particular concentration in the densely populated Asian and European continents. Due to dietary habits and genetic factors, China has the highest number of GC cases and deaths in the world [1]. Helicobacter pylori infection is a greater contributor to GC in East Asia [2]. A multicenter prospective cohort analysis in Chinese GC patients discovered that H. pylori infection accounted for 78.5% of non-cardia and 62.1% of cardia gastric cancers [3]. Immunotherapy is a cancer treatment that has recently been developed, and an increasing number of studies/clinical trials have demonstrated the efficacy and power of immunotherapy [4,5,6,7]. However, there are still some patients who do not respond to immunotherapy. If the effects of immunotherapy could be predicted before treatment utilizing molecular markers, it would be of considerable value to individualize patient care.
The majority of human tissue is surrounded by BM, a thin, dense layer of the extracellular matrix that lies at the base of epithelial cells. The laminin network and the type IV collagen network, which are two distinct polymer networks, are joined by extracellular matrix proteins to form the BM's lamellar structure [8]. BM serves as a barrier for the cell. It has been demonstrated that cells pass through the BM in a variety of ways. Different cells may use different methods to cross the BM depending on where they are located in the body, their physiological status, and even their stage of development [9,10,11,12]. Forming invasive pseudopods, down-regulating adhesion receptors, physically forcing a breakthrough, or inducing cellular remodeling across the basement membrane are all common ways for normal cells to do this. To properly metastasize, tumor cells must additionally penetrate their outer layer as well as the BM of the metastatic tissue. According to current research, there are four basic ways for tumor cells to break through the BM: mechanical forces created by cell proliferation; selective invasion driven by telangiectatic precursor cells; facilitation of invasion by cancer-associated fibroblasts in the extracellular matrix; and inhibition of tumor cell invasion by myoepithelial cells [13, 14]. The BM, in addition to affecting the movement of different cell types, can also perform immunomodulatory actions. BM laminin fractions can directly limit T cell activation and proliferation in mice by reducing T cell receptor signaling [15]. It is suggested that changes in BM state may indicate tumor cell malignancy and pathological grading.
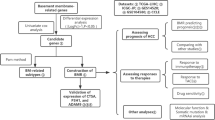
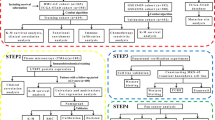
Changes in the BM status may reflect the degree of malignancy and pathological grade of cancer. However, there is still a lack of research on the relationship between the basement membrane and the pathological grade of gastric cancer. In this study, molecular subtypes of gastric cancer based on BM gene expression were analyzed by consensus clustering. The differences in TMB, immune infiltration, tumor purity, and cancer-related signaling pathways between different subtypes were studied. Univariate COX regression, Lasso regression, and multivariate COX regression were used to construct a predictive model of survival risk based on BM genes, and further, evaluate the differences in immunotherapy efficacy and chemotherapy resistance between high and low-risk groups. In addition, cell subsets that may be involved in BM remodeling in gastric cancer were investigated and characterized at the single-cell level.
2 Materials and methods
2.1 Data sources and processing
Gene expression data and clinical annotation information were collected from GEO and TCGA databases. This study included seven eligible GC cohorts (GSE15459, GSE34942, GSE57303, GSE62254, GSE84437, GSE183904 and TCGA-STAD) (Table S1) [16,17,18,19,20]. We downloaded the original 'CEL' files for the first four microarray data from the GPL570 platform on Affymetrix® chips and used the RMA algorithm of affy package to perform background correction, normalization, and integration of each dataset in turn. We used the sva (Surrogate Variable Analysis) package’s “ComBat” algorithm to correct for batch effects caused by non-biotechnical bias between the four datasets. The four datasets were combined and given the name Meta cohort. We directly downloaded the normalized matrix file for the microarray data GSE84437 from the GPL6947 platform. The openly available RNA sequencing data (TPM values) from the TCGA platform were downloaded directly from the TCGA website using the search term “TCGA-STAD”. To facilitate subsequent analysis, we converted the TPM values to log2(TPM + 1). We removed genes that had no expression in any of the samples in each dataset. In cases where multiple probes were enriched to the same gene, we chose the probe with the highest expression value as the corresponding gene's expression value. TCGA provided data on somatic mutations and copy number variation (CNV). To analyze somatic mutation data, the maftools package was used [21]. The GEO database’s GSE183904 dataset was used to obtain single-cell data. We downloaded the single-cell matrix file and integrated the single-cell data from 36 of these patients with primary gastric cancer. For batch correction, the Harmony algorithm was used. We used the FindVariableFeature function to find the top 3000 highly variable genes, the ScaleData function to scale all genes, and the RunPCA function to downscale the top 3000 highly variable genes that were filtered. The tSNE method was used to find cell clusters by downscaling using the "FindNeighbors" and "FindCluster" functions and a resolution of 0.5. The cell clusters were identified using GC cell subpopulation marker genes. R (version 4.1.1) was used for all data analysis. For data visualization, the R packages ggplot2 and pheatmap were used.
2.2 Unsupervised clustering analysis based on BM genes
Ranjay Jayadev et al. conducted a study that yielded the BM gene set [22]. Considering the different numbers of BM genes detected by different sequencing platforms, we took the intersection to obtain 203 common BM genes for this study (Table S2). The Meta cohort, TCGA cohort, and GSE84437 cohort were all included in the clustering analysis, which was carried out using the K-means algorithm from the ConsensuClusterPlus package [23]. We selected 80% of the samples, and 80% of the genes for repeat sampling, with the maximum K value set to 6 and the number of repeat samples set to 1000.
2.3 Differential gene analysis
The limma package was used to examine genes that were expressed differently in the two groups [24]. Genes with |logFC|> 1.2 and P-adjust < 0.05 were considered differentially expressed between groups.
2.4 Function enrichment analysis
For gene function enrichment analysis, both Gene Ontology (GO) and Gene Set Variation Analysis (GSVA) were used. The GO analysis is based on differentially expressed genes, which may miss some important but less differentially expressed genes in the pathways. The GSVA algorithm computes the cumulative density function of all genes in each pathway of a sample, ranks the genes, and computes the GSVA score based on its ranking order and scoring rules. The progeny package was used to calculate the targeted cancer signaling pathway score of the samples. Androgen, EGFR, Estrogen, Hypoxia, JAK-STAT, MAPK, NFB, PI3K, P53, TNFa, Trail, TGF, VEGF, and WNT are the 14 cancer signaling pathways built into the current version of progeny. For GO, GSVA, and cancer-related signaling pathway analyses, the ClusterProfiler, GSVA [25], and progeny packages [26] were used, respectively.
2.5 Tumor microenvironment (TME) analysis
The ssGSEA algorithm in the GSVA package was used to analyze the differences in the expression levels of 29 TME tags in GC patients [27]. MCPcounter was used to assess variations in the degree of infiltration between the eight immune cell types and the two stromal cell types (fibroblasts and endothelial cells). To assess tumor purity, Estimate package was utilized, with greater stromal and immune scores indicating poorer tumor purity.
2.6 Regression analysis
This study comprised univariate COX regression analysis, multivariate COX regression analysis, and LASSO regression analysis. The dependent variables in the COX regression analysis were survival time and survival status. To deal with the problem of covariance produced by too many independent variables, the LASSO regression was applied, in which the dependent variable was survival status and the independent variable was the gene's expression value. The risk models were built using multivariate COX regression. Multivariate COX regression was used to construct risk models to characterize the association between the BMscore and the prognosis of GC patients. The R package survival and survminer were used for univariate COX regression analysis and KM survival analysis. The R package glmnet (Generalized Linear Models via Coordinate Descent) was used for LASSO regression and multivariate COX regression analysis [28]. BMscore formula: (− 0.753) * FBN3 + (− 0.226) * ADAM9 + (0.088) * FREM2 + (0.155) * GPC3 + (0.294) * LOXL4 + (0.428) * ITGB5.
2.7 Single-cell analysis
The main single-cell data analysis was performed using the R package Seurat, and the built-in AddModuleScore function was used to calculate the fraction of distinct BM components concentrated in the cells [29]. iTALK was used to investigate intercellular communication.
2.8 Statistical analysis
Statistical analysis was performed with the use of R software. T-test was used for comparison between two groups, and the ANOVA test was used for comparison between three or more groups. Pearson correlation coefficients were utilized to analyze correlations between gene expression values, and mantel tests were performed to analyze correlations between BM gene expression matrices and endothelial, fibroblast score matrices. The Kaplan–Meier method was used to perform survival analysis, with the log-rank test used to evaluate significance.
3 Results
3.1 Unsupervised clustering analysis reveals three BM-associated GC subtypes
Meta cohort, TCGA cohort, and GSE84437 cohort were included in the cluster analysis. The sample distribution of the meta cohort before and after debatching was examined by t-SNE, and the results showed that the sample distribution was uniform after debatching (Fig. S1A, B). A total of 203 basement membrane genes were included in this study, including 52 cell surface receptor genes, 22 collagen genes, 58 glycoprotein genes, 16 proteoglycan genes, 38 regulatory factor genes, 4 secretion factor genes, 4 basement membrane accessory protein genes, and 9 other protein genes (Table S2, Fig. S1C). The transcripts of these 203 BM genes were extracted from the Meta cohort, TCGA cohort, and GSE84437 cohort, and then clustered based on the K-means unsupervised clustering algorithm (Fig. 1A–C). According to the background of the matrix heatmap and the descending slope of the CDF curve under different K values, the clustering results of the three cohorts were comprehensively considered, and finally, gastric cancer could be divided into three subtypes (Table S3).
Subtype analysis of GC based on BM gene expression. A Unsupervised clustering results of the Meta cohort. The eight subgraphs, in order from left to right, top to bottom, are Subplots 1–5 show the matrix heat map of the meta-queue for k = 2–6, subplot 6 shows the consistent cumulative distribution function (CDF) plot, where the value of k with a small decreasing slope of the CDF is usually chosen, subplot 7 is the Delta Area Plot, which shows the relative change of area under the CDF curve compared with k and k-1. A larger value indicates that the clustering effect under this k value is more obvious than that under K-1, subplot 8 is the tracking plot, which shows the classification of the samples attributed to different values of k. Different color blocks represent different classifications, and the samples whose color is often changed before and after taking different K values represent classification instability. B Unsupervised clustering results for the TCGA-STAD cohort. The subplot arrangement is consistent with the Meta cohort. C Unsupervised clustering results for the GSE84437 cohort. The subplot arrangement is consistent with the Meta cohort
By comparing the differences in expression levels of basement membrane genes and prognosis, we defined the subtype with the best prognosis and high expression of BM genes as BMclusterA. The subtype of gastric cancer with poor prognosis and moderate expression of BM genes was defined as BMclusterB. The subtype with the worst prognosis and low expression of BM genes was defined as BMclusterC (Fig. 2A–C, Fig. S2A–C). To verify the reliability of the clustering results, we used tSNE downscaling to compare the heterogeneity of sample distribution of BMclusterA, B, and C subtypes. The results showed that the sample distribution of different subtypes could be effectively distinguished in all three cohorts (Fig. S2D–F). To investigate the molecular basis for survival disparities among three BM-associated GC subtypes, we calculated enrichment scores for 50 pathways per sample using GSVA based on the h.all.v2022.1.Hs.symbols.gmt gene set from the MSigDB database. In subtype C, the signaling pathways related to tumor cell metastasis such as the epithelial cell polarization pathway, epithelial cell mesenchymal transition pathway, and angiogenesis signaling pathway were highly activated, while the signaling pathways related to tumor proliferation such as oxidative phosphorylation, glycolysis, and MYC pathway were inhibited (Fig. 2D). Subtype A is activated in exactly the opposite way to subtype C in these signaling pathways, and subtype B is in an intermediate state. These results suggest that BMclusterA is characterized by low metastasis and high proliferation, BMclusterC is characterized by high metastasis and low proliferation, and BMclusterB is in the transition state between A and C subtypes. To compare the expression levels of basement membrane genes among the three subtypes, we screened the differentially expressed basement membrane genes between tumor tissues and normal tissues in the TCGA data set (Fig. 2E, F). The distribution of these differential BM genes in the three subtypes A, B, and C showed a continuous hierarchical pattern, and the expression of most of them increased gradually (Fig. 2G, Fig.S2G, H), reflecting that the three GC subtypes may represent different stages of tumor development.
The differences in prognosis among the three subtypes of gastric cancer. A KM survival analysis of BMclusterA/B/C subtypes in the Meta cohort. B KM survival analysis of BMclusterA/B/C subtypes in the GSE84437 cohort. C KM survival analysis curves for BMclusterA, B, C subtypes in TCGA cohort. D GSVA enrichment analysis shows the activation status of biological signaling pathways in BMclusterA/B/C subtypes. Red represents relatively activated pathways and blue represents relatively inhibited pathways. E Volcano plot used to demonstrate BM genes that are differentially expressed between GC tumor and normal tissue. Blue dots represent genes that are down-regulated in tumor tissue and red dots represent genes that are up-regulated in tumor tissue. F Bar graph demonstrating the classification of 46 differentially expressed BM genes. G Heatmap demonstrating the expression levels of the 46 differentially expressed BM genes in the three GC subtypes compared to normal tissue
3.2 The immune infiltration and activation of oncogenic signals are different among the three GC subtypes
Tumor ecosystems are very complex, containing not only malignant cells but also a range of host cells such as endothelial cells, fibroblasts, and different immune cells that influence tumor development and invasion. We used the estimate and MCPcounter package to study the immune infiltration and tumor purity in the tumor microenvironment of the three BM-associated GC subtypes. In the BMclusterC subtype, there are abundant stromal cells and immune cells infiltration, and the purity of tumor cells is the lowest, but CD8 + T cells and NK cells, which are the main tumor killer cells, are the lowest infiltration in this subtype (Fig. 3A). This may account for the poor prognosis of BMclusterC. The infiltration level of fibroblasts, the main cell type for extracellular matrix formation, maintenance, and remodeling, was gradually increased in BMclusterA, B, and C subtypes (Fig. 3A, Fig. S2B). We also analyzed the degree of enrichment of cancer-related signaling pathways among different subtypes. The enrichment of P53, Trail, Andogen, Hypoxia, and TGFβ signaling pathways in BMclusterA, B, and C gradually increased, while the enrichment of VEGF, EGFR, and MAPK signaling pathways gradually decreased (Fig. 3B). P53 is capable of reversing oncogenic function by mutation, resulting in a gain of function that promotes tumor invasion and EMT [30]. Mutation landscape analysis of some core genes in these signaling pathways revealed that a significant proportion of patients in these three gastric cancer subtypes had missense mutations in TP53, indicating that the biological function of P53 may be altered (Fig. 3C). Furthermore, using the GSE62254 dataset with complete pathological information, we compared the association of BMclusterA, B and C subtypes with other gastric cancer pathological types. It can be seen that the proportion of GC diffuse type and EMT type in BMclusterA, B, and C subtypes increased in turn, and the recurrence rate of treatment also increased gradually, but EBV virus status seems to be unrelated to each subtype. (Fig. 3D). BM mutation profiles did not demonstrate significant differences across the three subtypes at the single gene variation level, and the overall incidence of BM gene mutations in gastric cancer was modest, with a comparatively high rate of COL12A1 mutations (Fig. 3E). Most of the chromosomal regions containing BM genes were amplified at the genomic level (Fig. 3F).
The immune infiltration and activation of oncogenic signals are different among BMclusterA/B/C subtypes of gastric cancer. A Heatmap of TME cell infiltration and tumor purity between BMclusterA/B/C subtypes in the Meta cohort. B Heatmap of the distribution of enriched cancer-related signaling pathways among BMclusterA/B/C subtypes in the Meta cohort. C Mutational landscape of core genes in the enriched cancer signaling pathways. D Statistical histogram of pathological information among BMclusterA/B/C subtypes in the GSE62254 dataset. E Mutation landscape of 46 differential basement membrane genes. F CNV frequency map of the 46 differential basement membrane genes
3.3 A correlation between TME status and BM genes expression among three GC subtypes
We performed gene expression analysis between BMclusterA/B/C and normal groups, respectively, and performed GO enrichment analysis for differentially expressed genes (Fig. 4A). A high number of B cell-mediated signaling pathways are enriched in the BMclusterA subtype, indicating that B cell-mediated immunological processes are active (Fig. 4B). The lymphatic vascular development signaling pathway is abundant in the BMclusterB subtype (Fig. 4C). This signaling pathway is thought to be associated with tumor cell metastasis [31]. Positive endothelial cell migration and vascular endothelial growth factor signaling pathways are abundant in the BMclusterC subtype (Fig. 4D). These results suggest that each BM-associated GC subtype has distinct biological characteristics. We hypothesized that the gene expression profiles in BMclusterA, B, and C subtypes were correlated with TME status, so the scores of 29 TME tags in each GC subtype were calculated (Fig. 5A). From BMclusterA to -B and -C subtypes, levels of infiltration of cancer-associated fibroblasts, tumor-associated macrophages, and pro-tumor cytokine production rose gradually (Fig. 5B). BM remodeling is more active in the BMclusterB subtype (Fig. 5B), which may experience BM breakdown. The BM is a barrier, and changes in its permeability have a direct impact on cellular transit. Transport by macrophages, dendritic cells, and myeloid-derived immune cells is progressively increased from BMclusterA to -B and -C, consistent with the BM remodeling (Fig. 5B), while transport by Treg and TH2 is reduced, suggesting the possibility of other regulatory mechanisms. Matrix metalloproteinases (MMPs) are a crucial protein class in matrix destruction and remodeling [32], and we discovered that the expression levels of most MMPs were highest in the BMclusterB subtype, indicating active BM remodeling (Fig. 5C, D). These results suggest that the three BM-related subtypes of gastric cancer may reflect the three states of gastric cancer progression, among which BM remodeling occurs in the BMclusterB subtype, which is a key stage for tumor cells to be able to infiltrate and metastasis.
The altered biological pathways in the three GC subtypes relative to normal tissues. A Venn diagram showing the distribution of differential genes in BMclusterA/B/C subtypes compared to normal samples. B GO analysis of differentially expressed genes in BMclusterA showed the top 20 enriched pathways. C GO analysis of differentially expressed genes in BMclusterB showed the top 20 enriched pathways. D GO analysis of differentially expressed genes in BMclusterC showed the top 40 enriched pathways
The TME tags of the three GC subtypes correlated with BM gene expression levels. A The heatmap shows the scores of 29 TME tags calculated by ssGSEA in the BMclusterA/B/C subtypes of the Meta cohort. B Boxplots show the scores of 9 TME tags in the BMclusterA/B/C subtypes of the Meta cohort. Statistical differences were tested using t ‘test. C Expression of 16 MMPs and NTN4 in the BMclusterA/B/C subtypes of Meta cohort. The upper and lower ends of the boxes represent the interquartile range of values. Lines in the boxes represent median values and black dots represent outliers. Asterisks represent statistical P values (*P < 0.05; **P < 0.01; ***P < 0.001, ****P < 0.0001). Statistical differences between the three subtypes were tested using the ANOVA test. D Expression of 16 MMPs and NTN4 in the BMclusterA/B/C subtypes of TCGA cohort. The upper and lower ends of the boxes represent the interquartile range of values. Lines in the boxes represent median values and black dots represent outliers. Asterisks represent statistical P values (*P < 0.05; **P < 0.01; ***P < 0.001, ****P < 0.0001). Statistical differences between the three subtypes were tested using the ANOVA test
3.4 Construction of BM genes-based prognostic risk model in gastric cancer and its predictive effect on immunotherapy and metastasis
The above analysis revealed significant differences in the prognosis of GC patients with different BM statuses. Based on the 203 basement membrane genes included in the analysis, the prognosis risk prediction model of GC based on BM genes was constructed. All samples from the Meta cohort were randomly assigned in a 1:1 ratio and used as the training set and internal validation set for the model. GSE84437 cohort and TCGA cohort were used for external validation. First, univariate regression analysis was used to screen genes with significant effects on the prognosis of GC. A total of 63 prognostic BM genes were obtained by univariate regression, and then Lasso regression analysis was used to exclude variables with coefficient zero (Fig. S3A, B). Finally, the genes that met the requirements were included in the training set for multivariate COX proportional hazards regression to construct the model. The median BMscore was used to classify the high and low-risk groups, and the receiver operating characteristic (ROC) curve was used to test the predictive ability of the model. The results showed that the model could effectively evaluate the survival risk of patients in both the training set, the internal validation set, and the two external validation sets (Fig. 6A–H). Further, we constructed a Nomogram to predict patient survival by combining clinical information and BMscore (Fig. S3C, D). The expression of FBN3, FREM2, LOXL4, and ITGB5 in GC tissues was verified by HPA(Human Protein Atlas) database (Fig. S4A). GPC3, LOXL4, and ITGB5 were independent predictive risk variables among the six genes utilized to build the model (Fig. 6I). We explored the relationship between GC subtypes and BMscore, showing that BMscore gradually increased from BMclusterA to -B and—C subtypes (Fig. 6J). The BMscore in the four ACRG subtypes of GC was also calculated, and the EMT subtype had a considerably higher BMscore than the other three subtypes (Fig. 6K), suggesting that the prognostic model based on BM genes can effectively evaluate the EMT subtypes with the least favorable treatment response. TMB and IPS scores calculated for high and low BMscore groups revealed that the low-risk group had a greater tumor mutation burden as well as immunogenicity, implying that the low-risk group is more likely to benefit from immunotherapy. Recent research has demonstrated that enhanced RHOJ gene expression can promote actin remodeling in tumor cells as well as trigger DNA repair mechanisms to repair chemotherapy-induced DNA damage and mediate chemo-resistance [33]. RHOJ expression was substantially lower in the low-risk group than in the high-risk group, indicating that the low-risk group may be more sensitive to chemotherapeutic treatments (Fig. 6L).
Construction of BM prognostic risk model for GC. A, KM survival curves for the high and low BMscore groups in the training set of the Meta cohort. B ROC curves for the training set of Meta cohort. C KM survival curves for the high and low BMscore groups in the validation set of the Meta cohort. D ROC curves for the validation set of Meta cohort. E KM survival curves for the high and low BMscore groups in the GSE84437. F ROC curves of GSE84437.G: KM survival curves for the high and low BMscore groups in the TCGA cohort. H ROC curves of TCGA cohort. I the Hazard ratio for the six genes used to construct the multivariate COX proportional risk regression model. J BMscore for BMclustrA/B/C subtypes in the Meta cohort, GSE84437 cohort, and TCGA cohort. K BMscore for the four ACRG subtypes of GC, including EMT, MSS/TP53 + , MSS/TP53-, and MSI
3.5 Analysis of cell subpopulations involved in BM remodeling and their intracellular communications
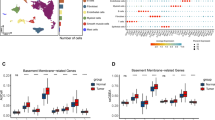
The above results suggest the transition from low- to high-risk GC subpopulations may involve BM remodeling. To elucidate the cell subpopulations involved in BM remodeling, we performed a single-cell transcriptome analysis. Using standard procedures, the GSE183904 dataset from the GEO database was processed, downscaled, and annotated to produce 11 cell subsets (plasma cells, B cells, mast cells, myeloid immune cells, T/NK cell subsets, endocrine cells, epithelial cells, endothelial cells, fibroblasts, and smooth muscle cells) (Fig. 7A). To ensure the accuracy of the cell subpopulations annotation, we assigned numerous marker genes to each cell subtype (Fig. 7B). Further, the 203 BM genes were classified according to molecular function, and then the cell subpopulations in which each class of BM genes was predominantly expressed were analyzed (Fig. 7C). Most of BM genes were discovered to be expressed largely in fibroblasts, endothelial cells, and smooth muscle cells in GC tissue, whereas the four affiliated BM genes were found to be expressed primarily in epithelial cells. Here, epithelial cells are GC cells expressing epithelial markers. Therefore, the major cells involved in BM remodeling include fibroblasts, endothelial cells, tumor cells, and smooth muscle cells. The proteoglycan fraction was found to be mainly expressed in fibroblasts subpopulation (Fig. 7C). We further subdivided fibroblasts and found that the expression level of proteoglycan in fibroblasts gradually increased with the pathological progression of gastric cancer (Fig. 7D–F), suggesting that proteoglycan components may be an important factor secreted by fibroblasts and involved in the regulation of gastric cancer progression.
Single-cell analysis of BM gene expression in each cell subpopulation in GC. A The tSNE dimension reduction map for single cell subpopulations in GC. B Expression maps of marker genes in single-cell subpopulations of GC. C Expression of different BM components in different cell subpopulations of GC. D A UMAP projection of the fibroblasts from GC. E Distribution of fibroblasts among patients with different pathological stages of GC. F Expression of proteoglycan components in different subpopulations of fibroblasts
Furthermore, the cell subpopulations expressing the 46 differentially expressed BM genes mentioned above were analyzed. It can be seen that the cell subpopulations mainly expressed these BM genes are consistent with the above results, but the expression level in tumor tissues is higher than that in normal tissues (Fig. 8A). We correlated the fibroblast and endothelial cell score matrix in the Meta cohort calculated by MCPcounter with 46 differentially expressed BM gene expression matrices and found specific BM gene clusters, such as BGN, COL18A1, THBS2, COL4A1 cluster, and ADAMTS2, NID2, LAMD1 cluster were highly correlated with fibroblasts (Fig. 8B). These results indicate that different cells have different functions and may require cooperation to achieve BM remodeling. To understand how these cells cooperate to regulate the progression of gastric cancer, we analyzed cell communication in four GC pathological stages. It can be seen that with the progression of gastric cancer, the ligand signaling pathway between collagen components and integrin ITGB1 in smooth muscle cells gradually increased (Fig. 8C). It may be that with the erosion of muscle layer by cancer cells, smooth muscle cells, and other cells also undergo adaptive changes and participate in processes such as BM remodeling to regulate tumor progression. Ultimately, the CCLE database (Cancer Cell Line Encyclopedia) was used to thoroughly assess the status of genes related with the BM in 41 GC cell lines (Fig. S4B). Different GC tumor cell lines showed comparable BM gene expression patterns. These cell lines scarcely expressed collagen-related genes and matrix metalloproteinase family-related (MMPs) genes, but they did express the transmembrane protein MMP14. When combined with the prior differential analysis, the expression of MMP14 was highly increased in GC tissues, with the greatest expression occurring in the BMclusterC subtype, which had the worse prognosis. This shows that GC cells may control BM breakdown through MMP14, influencing tumor growth.
Cellular communication between cell subpopulations of GC. A Expression of 46 differentially expressed BM genes in different cell subpopulations between normal and tumor tissues. B Correlation analysis between 46 differentially expressed BM genes and endothelial cells and fibroblasts. C Circle diagram of cell communication pattern in four pathological stages of GC. The arrow points to the receptor (black short line) and the arrow starts as the ligand (green short line)
4 Discussion
In this study, a molecular classification of GC was constructed based on BM genes. Three BM-related subtypes, BMclusterA/B/C, were found in gastric cancer by unsupervised clustering analysis, and the classification results were verified in multiple datasets. The prognosis, BM gene transcriptome patterns, TME, and cancer-related signaling pathways of each GC subtype were compared and analyzed. The differences among these three subtypes reflected the gradual evolution of gastric cancer from subtype A to subtype C. In the progression of gastric cancer, the constraint role of the basement membrane is gradually weakened, the EMT pathway is activated, and the ability to invasion and metastasis of GC cells is enhanced. Moreover, the composition of immune cells and stromal cells in the TME of BMclusterA/B/C gastric cancer subtypes were different, and the activation of carcinogenic signals also showed significant differences. These differences in microenvironmental ecosystems may have potential effects on tumor progression and treatment efficacy. Based on this, we constructed a survival risk prediction model of GC to characterize the predictive role of BM status in the prognosis, immunotherapy efficacy, and chemotherapy resistance of patients, and extend the practical application of BM-associated molecular classification of GC. Several previous studies have also identified various molecular subtypes of gastric cancer [17, 34, 35]. In contrast, our classification approach focused on the association between BM gene expression and metastatic progression in gastric cancer. Based on the concept of BM remodeling, the continuity of gastric cancer subtypes is clarified, and a new molecular classification concept of gastric cancer is proposed.
The basement membrane is a barrier that surrounds the outside of the tumor. The state of the basement membrane affects the entry and exit of immune cells into and out of tumor tissues, as well as the metastasis and colonization of tumor cells themselves [36, 37]. The cellular ecosystem in TME also plays a role in BM remodeling, and they interact with each other [38, 39]. The mechanism by which various stroma or immune cells in tumor tissue are transported through the basement membrane is not well understood. How the basement membrane of tumor tissue is dynamically regulated, and the cell subsets and molecular mechanisms involved in the remodeling of basement membrane also need to be further studied [40, 41]. Our study found that most of the matrix metalloproteinases were highly expressed in the BMclusterB subtype, suggesting that BM remodeling occurs during the progression of GC. The remodeling of the BM is not only limited to degradation [42], but the alteration of pore size as well as BM rigidity on BM also deserves attention [43]. Net4, the protein product of the NTN4 gene, can generate larger pores in BM and reduce the rigidity of the BM by opening the laminin node complex [44]. The expression of NTN4 increased gradually in BMclusterA/B/C. According to single-cell transcriptome analysis, tumor cells, fibroblasts, endothelial cells, and smooth muscle cells may all have a role in BM remodeling in GC. The communication between these cell subpopulations expressing BM components and between them and tumor cells is an important part of the regulation of tumor microenvironment, which has the potential to be used as a target for intervention.
The shortcoming of our study is that our analysis is based on the transcriptional level, and post-transcriptional regulation also affects the protein level, which may cause incomplete agreement with the actual BM protein level [45]. In this study, multiple data sets were used for mutual validation, which could compensate for this deficiency to some extent. The efficiency of cellular transport through the BM is limited by the barrier effect of the BM itself and also depends on the body’s requirement [46]. Therefore, the calculated basement membrane dynamics of each GC subtype based on transcriptional data may not completely reflect the true physiological state.
In conclusion, three molecular subtypes of GC (BMclusterA/B/C) based on BM gene expression were identified by unsupervised cluster learning in this study, and a prognostic model of GC based on six BM genes (FBN3, ADAM9, FREM2, GPC3, LOXL4, and ITGB5) was constructed. Through comprehensive analysis of the biological characteristics of each subtype, it is pointed out that they represent a continuous process of GC progression, accompanied by BM reorganization. We also analyzed the cell subpopulations that may be involved in BM remodeling at the single-cell level, indicating that fibroblasts, endothelial cells, tumor cells, and smooth muscle cells may all play a role in this process. This study will help to understand the important role of BM in the GC progression and provide a new molecular classification and prognostic prediction method for gastric cancer.
Availability of data and materials
Publicly available datasets were analyzed in this study. These data can be found here: https://portal.gdc.cancer.gov/, http://www.ncbi.nlm.nih.gov/geo, https://www.proteinatlas.org/, https://sites.broadinstitute.org/ccle/.
Abbreviations
- ACRG:
-
Asian Organization for Research on Cancer
- BM:
-
Basement membrane
- GC:
-
Gastric cancer
- CNV:
-
Copy number variation
- OS:
-
Overall survival
- TCGA:
-
The cancer genome atlas
- GEO:
-
Gene expression omnibus
- EMT:
-
Epithelial mesenchymal transition
- MMPs:
-
Matrix metalloproteinases
- TME:
-
Tumor microenvironment
- TMB:
-
Tumor mutation burden
- NTN4:
-
Netrin 4
- RHOJ:
-
Ras homolog family member J
- ROC:
-
Receiver operating characteristic
References
Wong MCS, Huang J, Chan PSF, Choi P, Lao XQ, Chan SM, et al. Global incidence and mortality of gastric cancer, 1980–2018. JAMA Netw Open. 2021;4(7):e2118457.
Huang J, Lucero-Prisno DE 3rd, Zhang L, Xu W, Wong SH, Ng SC, et al. Updated epidemiology of gastrointestinal cancers in East Asia. Nat Rev Gastroenterol Hepatol. 2023;20(5):271–87.
Yang L, Kartsonaki C, Yao P, de Martel C, Plummer M, Chapman D, et al. The relative and attributable risks of cardia and non-cardia gastric cancer associated with Helicobacter pylori infection in China: a case-cohort study. Lancet Public Health. 2021;6(12):e888–96.
Takei S, Kawazoe A, Shitara K. The new era of immunotherapy in gastric cancer. Cancers. 2022;14(4):1054.
Shitara K, Van Cutsem E, Bang YJ, Fuchs C, Wyrwicz L, Lee KW, et al. Efficacy and safety of pembrolizumab or pembrolizumab plus chemotherapy vs chemotherapy alone for patients with first-line, advanced gastric cancer: the KEYNOTE-062 phase 3 randomized clinical trial. JAMA Oncol. 2020;6(10):1571–80.
Chung HC, Bang YJ, C SF, Qin SK, Satoh T, Shitara K, et al. First-line pembrolizumab/placebo plus trastuzumab and chemotherapy in HER2-positive advanced gastric cancer: KEYNOTE-811. Future Oncol. 2021;17(5):491–501.
Qi C, Gong J, Li J, Liu D, Qin Y, Ge S, et al. Claudin18.2-specific CAR T cells in gastrointestinal cancers: phase 1 trial interim results. Nat Med. 2022;28(6):1189–98.
Jayadev R, Sherwood DR. Basement membranes. Curr Biol: CB. 2017;27(6):R207–11.
Song J, Zhang X, Buscher K, Wang Y, Wang H, Di Russo J, et al. Endothelial basement membrane laminin 511 contributes to endothelial junctional tightness and thereby inhibits leukocyte transmigration. Cell Rep. 2017;18(5):1256–69.
Kurz AR, Pruenster M, Rohwedder I, Ramadass M, Schäfer K, Harrison U, et al. MST1-dependent vesicle trafficking regulates neutrophil transmigration through the vascular basement membrane. J Clin Investig. 2016;126(11):4125–39.
Rohwedder I, Kurz ARM, Pruenster M, Immler R, Pick R, Eggersmann T, et al. Src family kinase-mediated vesicle trafficking is critical for neutrophil basement membrane penetration. Haematologica. 2020;105(7):1845–56.
Bahr JC, Li XY, Feinberg TY, Jiang L, Weiss SJ. Divergent regulation of basement membrane trafficking by human macrophages and cancer cells. Nat Commun. 2022;13(1):6409.
Chang J, Chaudhuri O. Beyond proteases: basement membrane mechanics and cancer invasion. J Cell Biol. 2019;218(8):2456–69.
Kelley LC, Lohmer LL, Hagedorn EJ, Sherwood DR. Traversing the basement membrane in vivo: a diversity of strategies. J Cell Biol. 2014;204(3):291–302.
Liu X, Qiao Y, Chen J, Ge G. Basement membrane promotes tumor development by attenuating T cell activation. J Mol Cell Biol. 2022;14(2):mjac006.
Chia NY, Deng N, Das K, Huang D, Hu L, Zhu Y, et al. Regulatory crosstalk between lineage-survival oncogenes KLF5, GATA4 and GATA6 cooperatively promotes gastric cancer development. Gut. 2015;64(5):707–19.
Cristescu R, Lee J, Nebozhyn M, Kim KM, Ting JC, Wong SS, et al. Molecular analysis of gastric cancer identifies subtypes associated with distinct clinical outcomes. Nat Med. 2015;21(5):449–56.
Ooi CH, Ivanova T, Wu J, Lee M, Tan IB, Tao J, et al. Oncogenic pathway combinations predict clinical prognosis in gastric cancer. PLoS Genet. 2009;5(10):e1000676.
Qian Z, Zhu G, Tang L, Wang M, Zhang L, Fu J, et al. Whole genome gene copy number profiling of gastric cancer identifies PAK1 and KRAS gene amplification as therapy targets. Genes Chromosom Cancer. 2014;53(11):883–94.
Yoon SJ, Park J, Shin Y, Choi Y, Park SW, Kang SG, et al. Deconvolution of diffuse gastric cancer and the suppression of CD34 on the BALB/c nude mice model. BMC Cancer. 2020;20(1):314.
Mayakonda A, Lin DC, Assenov Y, Plass C, Koeffler HP. Maftools: efficient and comprehensive analysis of somatic variants in cancer. Genome Res. 2018;28(11):1747–56.
Jayadev R, Morais M, Ellingford JM, Srinivasan S, Naylor RW, Lawless C, et al. A basement membrane discovery pipeline uncovers network complexity, regulators, and human disease associations. Sci Adv. 2022;8(20):2265.
Wilkerson MD, Hayes DN. ConsensusClusterPlus: a class discovery tool with confidence assessments and item tracking. Bioinformatics. 2010;26(12):1572–3.
Ritchie ME, Phipson B, Wu D, Hu Y, Law CW, Shi W, et al. Limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 2015;43(7):e47.
Hänzelmann S, Castelo R, Guinney J. GSVA: gene set variation analysis for microarray and RNA-seq data. BMC Bioinformatics. 2013;14:7.
Schubert M, Klinger B, Klünemann M, Sieber A, Uhlitz F, Sauer S, et al. Perturbation-response genes reveal signaling footprints in cancer gene expression. Nat Commun. 2018;9(1):20.
Bagaev A, Kotlov N, Nomie K, Svekolkin V, Gafurov A, Isaeva O, et al. Conserved pan-cancer microenvironment subtypes predict response to immunotherapy. Cancer Cell. 2021;39(6):845-65.e7.
Friedman J, Hastie T, Tibshirani R. Regularization paths for generalized linear models via coordinate descent. J Stat Softw. 2010;33(1):1–22.
Hao Y, Hao S, Andersen-Nissen E, Mauck WM 3rd, Zheng S, Butler A, et al. Integrated analysis of multimodal single-cell data. Cell. 2021;184(13):3573-87.e29.
Zhang C, Liu J, Xu D, Zhang T, Hu W, Feng Z. Gain-of-function mutant p53 in cancer progression and therapy. J Mol Cell Biol. 2020;12(9):674–87.
Stacker SA, Williams SP, Karnezis T, Shayan R, Fox SB, Achen MG. Lymphangiogenesis and lymphatic vessel remodelling in cancer. Nat Rev Cancer. 2014;14(3):159–72.
Kyprianou C, Christodoulou N, Hamilton RS, Nahaboo W, Boomgaard DS, Amadei G, et al. Basement membrane remodelling regulates mouse embryogenesis. Nature. 2020;582(7811):253–8.
Debaugnies M, Rodríguez-Acebes S, Blondeau J, Parent MA, Zocco M, Song Y, et al. RHOJ controls EMT-associated resistance to chemotherapy. Nature. 2023;616(7955):168–75.
Zhang B, Wu Q, Li B, Wang D, Wang L, Zhou YL. m(6)A regulator-mediated methylation modification patterns and tumor microenvironment infiltration characterization in gastric cancer. Mol Cancer. 2020;19(1):53.
Comprehensive molecular characterization of gastric adenocarcinoma. Nature. 2014;513(7517):202-9
Khalilgharibi N, Mao Y. To form and function: on the role of basement membrane mechanics in tissue development, homeostasis and disease. Open Biol. 2021;11(2):200360.
Morrissey MA, Jayadev R, Miley GR, Blebea CA, Chi Q, Ihara S, et al. SPARC promotes cell invasion in vivo by decreasing type IV collagen levels in the basement membrane. PLoS Genet. 2016;12(2):e1005905.
Sherwood DR. Basement membrane remodeling guides cell migration and cell morphogenesis during development. Curr Opin Cell Biol. 2021;72:19–27.
Cruz-Acuña R, García AJ. Synthetic hydrogels mimicking basement membrane matrices to promote cell-matrix interactions. Matrix Biol. 2017;57–58:324–33.
Chang TT, Thakar D, Weaver VM. Force-dependent breaching of the basement membrane. Matrix Biol. 2017;57–58:178–89.
Fiore VF, Krajnc M, Quiroz FG, Levorse J, Pasolli HA, Shvartsman SY, et al. Mechanics of a multilayer epithelium instruct tumour architecture and function. Nature. 2020;585(7825):433–9.
Milán M. Tissue growth: basement membrane degradation triggers cell proliferation. Curr Biol: CB. 2022;32(6):R276–9.
Walter C, Davis JT, Mathur J, Pathak A. Physical defects in basement membrane-mimicking collagen-IV matrices trigger cellular EMT and invasion. Integr Biol. 2018;10(6):342–55.
Mehlen P, Fattet L. Netrin-4 regulates stiffness and metastasis. Nat Mater. 2021;20(6):722–3.
Giraldo NA, Sanchez-Salas R, Peske JD, Vano Y, Becht E, Petitprez F, et al. The clinical role of the TME in solid cancer. Br J Cancer. 2019;120(1):45–53.
Diwanji N, Bergmann A. Basement membrane damage by ROS- and JNK-mediated Mmp2 activation drives macrophage recruitment to overgrown tissue. Nat Commun. 2020;11(1):3631.
Acknowledgements
We thank TCGA and GEO databases for providing a platform for data sharing.
Funding
This study was supported by the National Natural Science Foundation of China (81872276), the Hefei Municipal Natural Science Foundation (2022048 & 2022050), the Pharmaceutical Research and Exploration Fund (YJKJJ06), and the foundation of Anhui Province Key Laboratory of Medical Physics and Technology (LMPT201908).
Author information
Authors and Affiliations
Contributions
WY conceived and designed the work. WY, XT, CF developed the experimental methodologies. XT, YL, WY and CF performed the analysis and interpreted the data. WY, XT, YL, CF, JZ discussed the results and wrote the paper. And WY supervised the study. ALL authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Ethics approval and consent to participate
Not applicable.
Informed consent
Not applicable.
Consent for publication
Not applicable.
Competing interests
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Tang, X., Liu, Y., Zhao, J. et al. Subtyping of gastric cancer based on basement membrane genes that stratifies the prognosis, immune infiltration and therapeutic response. Discov Onc 15, 362 (2024). https://doi.org/10.1007/s12672-024-01238-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s12672-024-01238-z