Abstract
CD8+ T lymphocytes are critical in the immune response against neoplasms, yet the prognostic relevance of CD8+ T cell-associated genes in hepatocellular carcinoma (HCC) is not fully understood. We sourced single-cell RNA-sequencing (scRNA-seq) and bulk RNA-seq data for HCC from the GSE98638 dataset and The Cancer Genome Atlas (TCGA) repository. We utilized Weighted Gene Correlation Network Analysis (WGCNA) to identify CD8+ T cell-related genes. A clinical prognostic model for risk stratification was then constructed via Cox-Lasso regression analysis. The Immunophenotypic Score (IPS) was utilized to evaluate the potential of immunotherapeutic interventions in the categorized cohorts. Validation of the expression of CD8+ T cell-associated risk genes was performed using quantitative reverse transcription PCR (qRT-PCR). Integrating scRNA-seq with RNA-seq data, we identified five CD8+ T cell-related signature genes: IKBKE, ATP1B3, MSC, ADA, and BATF. Notably, HCC patients in the high-risk group had markedly decreased overall survival. Elevated infiltration levels of CD8+ T cells, B cells, and macrophages were observed in the high-risk group. Moreover, there was a positive correlation between the risk score and immune checkpoints (ICPs), including PDCD1, CD274, and CTLA4. Patients within the high-risk group subject to PD1 and CTLA4 blockade exhibited higher IPS levels. Additionally, the expression of the five risk genes was upregulated in HCC cell lines and tissues compared to normal cells and tissues. Our findings establish a prognostic signature based on CD8+ T cells, offering a potent predictive model for clinical outcomes and responsiveness to immunotherapy in HCC patients.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Liver cancer is a globally prevalent malignant tumor, with approximately 900,000 new cases and 830,000 fatalities reported annually [1]. Despite its insidious development and lack of obvious clinical symptoms in the early stages, the rapid progression of tumors hampers the effectiveness of treatment methods such as surgery and drugs, resulting in poor prognosis and short survival time [2]. The tumor immune microenvironment (TIME) comprises diverse immune-related components, including immune cells and immune regulatory molecules. Due to its pivotal role in regulating tumor growth and metastasis, it has emerged as a focal point of research in the field of oncology in recent years [3, 4]. CD8+ T cells, as a crucial component, play a significant role in anti-tumor immunity. Receiving stimulation from antigen-presenting cells that carry tumor antigens, CD8+ T cells in their naive state undergo differentiation into cytotoxic CD8+ T cells and migrate towards the tumor location to trigger immune elimination [5]. Furthermore, mounting evidence has established the close association between T cells and prognosis across various cancers such as gastric cancer, lung cancer, and hepatocellular carcinoma (HCC) [6,7,8]. Consequently, the identification of novel T cell markers in HCC and the elucidation of their immunological significance and prognostic value, are anticipated to emerge as new targets within the field of HCC immunotherapy.
Single-cell RNA-sequencing (scRNA-seq) technology offers a comprehensive understanding of tumors from diverse perspectives and levels, encompassing tumor heterogeneity, the tumor microenvironment, tumor immunotherapy, and tumor immune escape [9]. The scRNA-seq enables the acquisition of genetic characteristics specific to cell types, thereby facilitating a more comprehensive understanding of the heterogeneity within the tumor microenvironment (TME) and aiding in formulating personalized treatment strategies. Although the number of scRNA-seq samples is limited, the RNA-seq dataset offers a larger sample size and more comprehensive follow-up information. Consequently, there arises a necessity to integrate scRNA-seq and bulk RNA-seq datasets to identify novel tumor biomarkers and develop prognostic risk stratification models with enhanced accuracy.
In this study, we identified CD8T cell-related genes and constructed a clinical prognostic model for HCC. Moreover, the model's predictive performance was verified through the ICGC cohort. This investigation will establish a solid foundation for identifying novel therapeutic targets in HCC and serve as an initial reference for its treatment strategies.
2 Materials and methods
2.1 Data collection
The GSE98638 dataset, which contains scRNA-seq for T cells of HCC, has been downloaded from the Genome Expression Omnibus (GEO) database. Additionally, the RNA-seq data of 424 patients with HCC were obtained from the Cancer Genome Atlas (TCGA) database. The RNA-seq data of 231 patients with HCC were acquired from the International Cancer Genome Consortium (ICGC) database.
2.2 ScRNA-seq analysis
The scRNA-seq data were filtered and standardized by the "Seurat" R package. Principal component analysis (PCA) and unified manifold approximation and projection (UMAP) were employed for dimensionality reduction, followed by the t-distributed random neighborhood embedding (tSNE) clustering method for data visualization analysis. The FindAllMarkers function was utilized to identify marker genes in T cell subgroups, with the conditions set as min.pct = 0.25, logfc.threshold > 0.25, and adjPvalFilter < 0.05. Additionally, the "celldex" R package and “ref_Human_all.RData” reference data were employed for cell annotation.
2.3 Weighted gene co-expression network analysis
The content matrix of 22 immune cells in the TCGA-LIHC samples was calculated via the CIBERSORT algorithm [10]. The eigengenes of the modules were calculated by the "WGCNA" package, and Pearson’s correlation analysis was conducted to investigate the relationship between each eigengene and 22 immune cell types, to identify modules associated with CD8 T cells. We selected the core genes of the module with the highest coefficient for subsequent GO and KEGG enrichment analysis.
2.4 Screening of CD8 T cell-related genes
Differentially expressed genes (DEGs) were identified via the "limma" package with FDR < 0.05 and a LogFoldChange > 1.5. These DEGs were intersected with CD8T cell module genes to obtain CD8T cell-related genes.
2.5 The risk stratification model was constructed via Cox-Lasso
Cox analysis was employed to screen CD8 T cell-related prognostic genes. Lasso regression analysis was utilized to determine the risk correlation coefficient (coef) of each identified prognostic gene. Through the risk formula to calculate risk score (risk score = ∑ (coefficient i × expression value of gene i)), the ICGC cohort served as an external test set for model validation.
2.6 Nomogram
Cox regression analysis was used to evaluate the prognostic independence of the risk score in HCC, and a prediction model was constructed combined with clinical factors. The prognostic value of the risk model was assessed using the calibration curve, ROC curve, and C-index.
2.7 GSVA and immune subtypes
C2.cp.kegg (v7.4) and Hallmark (v7.2) gene sets were used to perform Gene Set Enrichment Analysis (GSEA) and Gene Set Variation Analysis (GSVA) enrichment analysis of risk scores. The TCGA-LIHC expression matrix was computed using the "GSEABase" and "GSVA" R packages to derive enrichment values for the GSVA gene set in each sample. Subsequently, the Spearman method was employed to analyze the correlation between GSVA gene sets and risk genes/risk scores. Furthermore, we retrieved a list of immune checkpoint genes from relevant literature [11] and analyzed the correlation between these genes and risk scores. The TCGA-LIHC samples encompassed four distinct immune subtypes, namely wound healing (C1), INF-gamma dominant (C2), Inflammatory (C3), and Lymphocyte depleted (C4). Each immune subtype represented a specific immune microenvironment [12].
2.8 TME
We utilized the ssGSEA algorithm to evaluate the levels of infiltration of tumor-infiltrating immune cells (TIICs) and immune function. Additionally, The TCGA-LIHC samples were assessed using the “ESTIMATE” R package to derive ESTIMATE scores, Immune scores, Stromal scores, and Tumor purity [13]. Furthermore, the prediction of immunotherapy response in HCC patients was conducted using the Tumor Immune Dysfunction Exclusion (TIDE) and The Cancer Immunome Atlas (TCIA) databases.
2.9 Chemotherapy agents
The Drug Sensitivity reference matrix was obtained from the Genomics of Drug Sensitivity in Cancer (GDSC) database, and the IC50 values of chemotherapy agents were calculated using the "oncoPredict" R package.
2.10 qRT-PCR
The HCC cell lines L02, HuH7, Hep3B, and SK-Hep-1 were acquired from the Cell Resource Center in Beijing. These cells were cultured in DMEM medium (Gibco, USA) at 37 °C in a 5% CO2 atmosphere. We retrieved twelve paired specimens of post-hepatectomy primary HCC tissues and adjacent liver tissue from an ultra-cryogenic freezer maintained at − 80 °C in our laboratory. The conduct of this study was rigorously vetted and approved by the Medical Research Ethics Committee of our hospital, bearing the ethics batch number D2023-078, and adhered to the stringent ethical standards outlined in the 1975 Declaration of Helsinki. Total RNA was isolated from the cells and then reverse transcribed into cDNA using the cDNA Synthesis Kit (Vazyme Biotech Co., China). Quantitative PCR analysis was performed under the following reaction conditions: an initial denaturation step at 95 °C for 5 min, followed by 39 cycles of amplification at 95 °C for 10 s and annealing at 60 °C for 30 s in a PCR instrument (Bio-Rad CFX96, USA). The primer sequences for the genes are provided in Supplemental Table S1. The relative expression of the risk genes was determined using the 2−ΔΔCt method. The immunohistochemical images of risk gene protein expression in HCC patients’ clinical specimens were obtained from the Human Protein Atlas (HPA) database for this study.
2.11 Statistical analysis
The R (4.1.3) software was used for data analysis and graphical visualization. In this study, a significance level of P < 0.05 was considered statistically significant.
3 Results
3.1 Integration and clustering of HCC scRNA-seq
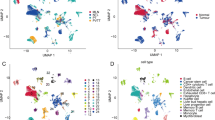
The study's design flowchart is depicted in Supplementary Fig. 1. GSE98638 represents a scRNA-seq dataset of T cells in HCC, encompassing 6 patients and 5063 cells. Utilizing the "Single" R package, cell subsets were annotated and visualized (Supplementary Fig. 2A). Subsequently, the scRNA-seq data underwent normalization and dimensionality reduction via PCA, UMAP, and t-SNE, leading to the identification of 15 distinct clusters (Supplementary Fig. 2B). A total of 4304 cluster markers were acquired, and a gene expression heat map for each cluster was generated (Supplementary Fig. 2C).
3.2 WGCNA
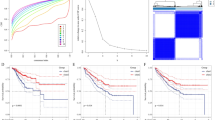
The content matrix of 22 TIICs in the TCGA-LIHC samples was calculated using the CIBERSORT algorithm. The WGCNA package was utilized for correlation analysis of cluster markers and the immune cell matrix, ensuring that each gene regulation relationship conformed to the scale-free distribution by achieving a scale-free topology fitting index of 0.9 (Fig. 1A, B). Cluster markers were grouped into different modules by creating the module clustering dendrogram (Fig. 1C, D). Subsequent analysis of correlation coefficients revealed that the red module showed the strongest association with CD8 T cells among all immune cell types (Fig. 1E).
Functional annotation of GO and enrichment analyses of KEGG were performed for genes related to CD8 T cells. The GO enrichment analysis revealed participation in functions such as T-cell activation, MHC II protein complex, and immune receptor activity (Supplementary Fig. 3A). The KEGG analysis identified genes associated with CD8 T cells participating in Th1 and Th2 cell differentiation and Th17 cell differentiation (Supplementary Fig. 3B).
3.3 Development and validation of prognosis models
Fifty-nine DEGs related to CD8 T cells were identified using a Venn diagram (Fig. 2A). Subsequently, Cox-Lasso regression was performed to select five prognostic genes (IKBKE, ATP1B3, MSC, ADA, BATF), which were used to develop a risk stratification model (Fig. 2B–D). The risk score was calculated as follows: Risk Score = IKBKEexp × 0.002064 + ATP1B3exp × 0.003343 + MSCexp × 0.002037 + ADAexp × 0.001229 + BATFexp × 0.002017. To assess the reliability of our risk model, we employed the TCGA-LIHC cohort as the training set and used the ICGC-LIHC cohort for external validation. The survival heat map indicates that as the risk score increased, a majority of patients experienced shorter survival durations (Fig. 3A, B). Kaplan–Meier analysis revealed a statistically significant decrease in overall survival among high-risk patients (Fig. 3C, D).
Risk scores of the TCGA training set and ICGC test set. A Survival state plots of risk scores for the TCGA training set. B Survival state plots of risk scores for the ICGC test set. C Kaplan–Meier survival curves of risk scores for the TCGA training set. D Kaplan–Meier survival curves of risk scores for the ICGC test set
The risk score exhibited significant correlations with grade, stage, and T stage (Fig. 4A, B). Cox regression analyses revealed that the risk score independently predicted the prognosis of HCC (Fig. 4C, D). A nomogram was constructed by incorporating both risk score and clinical factors (Fig. 4E). The calibration curve and ROC curve substantiated the excellent predictive value of the nomogram (Fig. 4F, G). Based on the ROC curve and C-index analysis of variables, it can be concluded that the risk score possesses superior predictive capability (Fig. 4H, I). Additionally, the Cox regression analysis conducted in the ICGC-LIHC external validation cohort confirmed that the risk score served as an independent prognostic factor (Supplementary Fig. 4A, B). By constructing a nomogram, calibration curve, ROC curve, and C-index, we observed that the risk score exhibited significant predictive value for the prognostic model in the ICGC external cohort (Supplementary Fig. 4C–G).
The model's predictive performance and clinical prognostic value. A, B The correlation between clinical factors and patients at high and low risk. C, D Univariate and multivariate Cox analysis of risk scores. E A nomogram was constructed based on clinical factors and risk scores. F The calibration curve of the nomogram. G The ROC curve of the nomogram. H The ROC curves of factors. I The C-index of factors
3.4 GSVA and immune checkpoints (ICPs)
GSEA analysis revealed the chemokine signaling pathway, ECM receptor interaction, and others in the high-risk group (Supplementary Fig. 5A). Conversely, the low-risk group exhibited significant enrichment in beta-alanine metabolism, retinol metabolism, and other pathways (Supplementary Fig. 5B). Additionally, GSVA analysis demonstrated a strong correlation between the high-risk group and pathways such as PI3K-AKT-mTOR signaling, Notch signaling, and IL6-JAK-STAT3 signaling (Fig. 5A). Furthermore, ICPs including PDCD1, CTLA4, LAG3, CD274, etc., were positively correlated with the risk score (Fig. 5B). Furthermore, notable differences were found in immune subtypes (C1, C2, C3, and C4) among the high and low-risk groups (Fig. 5C). These findings further support a relationship between high-risk scores and activation of immune processes.
3.5 TME
We conducted a thorough analysis of TIICs and immune function in HCC utilizing the ssGSEA algorithm. The findings indicated that in the high-risk group, the levels of CD8+ T cells and macrophage infiltration were elevated, and immune functions such as HLA and MHC-1 may be activated (Fig. 6A, B). Additionally, the high-risk group exhibited higher Estimate scores, Immune scores, and Stromal scores but lower Tumor purity compared to the low-risk group (Fig. 6C–F).
Tumor microenvironment. A The heatmap illustrates the correlation between risk score and TME. B Differences in immune cell infiltration levels between high and low risk groups. C-F Differences of ESTIMATE scores, Immune scores, Stromal scores, and Tumor purity in high-risk and low-risk groups. (*P < 0.05, **P < 0.01, ***P < 0.001)
3.6 Immunotherapy evaluation
We employed the TIDE database to examine tumor immune-related factors and observed significant correlations between Exclusion score, CAF, Merck18, MDSC, CD8, IFNG, and risk scores (Fig. 7A, B). Additionally, through the computation of IPS for HCC samples, we found that patients classified in the high-risk group and exhibiting positive CTLA-4 and PD-1 expressions displayed elevated IPS scores (Fig. 7C–F).
Differences in immune responses between high-risk and low-risk patients. A Correlation between risk score and various immune indicators in TIDE. B Differences in TIDE immune indicators between high and low-risk groups. C-F Differences in IPS between high and low-risk groups to PD1 and CTLA4 treatment
3.7 Chemotherapy drug sensitivity
Upon evaluating the IC50 values of chemotherapy drugs in HCC, the findings reveal that the low-risk group demonstrates lower IC50 values for cytarabine and axitinib (Fig. 8A, B), while the high-risk group exhibits lower IC50 values for dasatinib and gefitinib (Fig. 8C, D). A negative correlation between drug sensitivity and IC50 value was observed.
3.8 The expression of risk genes in HCC
The violin map and bubble plot depict the expression of risk genes (IKBKE, ATP1B3, MSC, ADA, and BATF) in distinct T cell clusters (Supplementary Fig. 6A, B). The t-SNE plot reveals that these risk genes exhibit high expression levels in T cells (Supplementary Fig. 6C). Additionally, the risk genes are up-regulated in HCC tissues (Fig. 9A–E), and individuals with elevated expression of risk genes tend to have a poor prognosis (Fig. 9F–J). Furthermore, the expression levels of five risk genes were higher in majority of HCC cell lines compared to L02 normal cells (Fig. 9K–O). Additionally, qRT-PCR analysis of 12 HCC samples validated that the expression levels of five risk genes were elevated in cancerous tissues compared to adjacent normal tissues (Fig. 9P–T). Immunohistochemical staining images of risk gene proteins in HCC tissues and normal liver tissues were acquired from the HPA database. The findings revealed elevated protein expression levels of IKBKE, ATP1B3, and ADA in HCC tissues compared to normal liver tissues, while the protein expression level of BATF was lower than that in normal liver tissues (Supplementary Fig. 7A–D).
The expression and prognosis of risk genes in HCC. A–E The expression levels of IKBKE, ATP1B3, MSC, ADA, and BATF in both normal liver and HCC tissues. F–J The Kaplan–Meier survival curves of IKBKE, ATP1B3, MSC, ADA, and BATF in HCC. K–O Relative expression levels of IKBKE, ATP1B3, MSC, ADA, and BATF in normal cell lines and HCC cell lines via qRT-PCR. P–T Relative expression levels of IKBKE, ATP1B3, MSC, ADA, and BATF were determined in 12 paired normal liver tissues and HCC tissues using qRT-PCR.*P < 0.05, **P < 0.01, ***P < 0.001
4 Discussion
More and more studies have demonstrated the influence of TME characteristics and components on tumor growth, metastasis, and treatment response [14]. Investigating the significance of immune cells in the TME and overcoming immune evasion and tolerance are crucial directions in tumor immunotherapy. CD8+ T cells play a pivotal role in antitumor immunity, carrying out essential functions including immune surveillance, antigen presentation and recognition, cytotoxicity, and establishment of immune memory [15, 16]. Considering the complex mechanisms of T-cell biology, identifying dependable biomarkers and investigating associated molecular mechanisms provide new perspectives for advancing novel approaches to cancer therapy.
In recent years, scRNA-seq has emerged as a powerful tool for accurately identifying cell types in tumor tissues and obtaining gene expression data at the single-cell level [17, 18]. The current research trend involves the integration of scRNA-seq and bulk RNA-seq to develop prognostic models. In this study, we have discovered 59 characteristic genes related to CD8 T cells and further identified five independent prognostic genes (IKBKE, ATP1B3, MSC, ADA, and BATF). IKBKE plays a crucial role in the regulation of inflammatory response and immune cells, and the pathogenesis of malignant tumors [19]. ATP1B3 serves as an independent prognostic indicator for HCC and is closely associated with immune cell infiltration and expression of immune factors in HCC [20]. MSC, functioning as a transcriptional repressor, plays a crucial role in the regulation of immune functions [21]. The overexpression of ADA enhances the proliferation and attenuates the exhaustion of CD19-specific and HER2-specific CAR T cells [22]. BATF plays a crucial role in T cell exhaustion and is indispensable for the initial stages of CD8+ T cell differentiation [23]. In this study, based on these five independent prognostic CD8 T cell-associated genes, we have developed a prognostic risk model for HCC. Both internal and external cohort tests have demonstrated the excellent predictive value of the risk stratification model. Additionally, a nomogram utilizing the risk score has demonstrated its efficacy in predicting the 1-year, 3-year, and 5-year survival rates of patients with HCC.
The TME exerts a pivotal influence on treatment response and clinical outcome. Immune checkpoints are essential in enabling tumor immune evasion [24]. In the past few years, significant advancements have been achieved in combining of immune checkpoint inhibitors (ICIs) with first-line drugs [25, 26]. This research unveiled that the high-risk group demonstrated comparatively elevated levels of infiltration by CD8+ T cells, macrophages, and HLA. High-risk patients also demonstrated elevated expression of immune-related markers such as CD274, CTLA-4, PDCD1, CD8, and IFNG. Currently, PD-1 and CTLA-4 inhibitors are widely used in solid tumor immunotherapy. Blocking PD-1 and PD-L1 can restore the function of effector CD8+ T cells [27], while anti-CTLA-4 therapy can increase the abundance of CD4+ T and CD8+ T cells in HCC patients [28, 29]. However, different patients exhibit different immune responses, making it particularly important to develop an individualized immunotherapy regimen for HCC patients with different prognostic stratification. The TCIA database enables the calculation of IPS from a tumor sample to predict the response to PD-1 and CTLA-4 inhibitors [30]. A higher IPS score indicates a better immune response. The present study revealed a significant association between positive expression of CTLA-4 and PD-1 and higher IPS in the high-risk patient group, indicating that inhibiting the PD-1 and CTLA-4 can enhance the abundance of CD8T cells within the TME, thereby further augmenting the efficacy of immunotherapy in HCC patients. Additionally, we have successfully identified chemotherapy agents that exhibit sensitivity towards high-risk patients through the utilization of pharmacogenomics.
Nevertheless, this study has certain limitations. The data were derived from public databases, and there was a shortage of prospective, multicenter clinical HCC sample cohorts to validate the risk stratification prediction model. The tumor microenvironment encompasses intricate interactions among diverse cell types. Our study solely focused on CD8T cells, which may not fully encompass the comprehensive role of the tumor microenvironment in tumorigenesis and development, thus presenting certain limitations. Moreover, the underlying mechanisms of risk prognostic genes have not been comprehensively investigated, and therefore warrant further exploration in future studies.
5 Conclusions
In summary, this study demonstrated variations in clinical outcomes and immune responses between low-risk and high-risk HCC patients through the integrated analysis of scRNA-seq and bulk RNA-seq data. We developed a risk stratification prognostic signature based on CD8 T cells, which accurately predicts the prognosis of HCC patients' outcomes and serves as a crucial reference for clinical treatment decision-making and drug selection.
Data availability
All data from this study were downloaded from online databases. The related data were downloaded from the GEO (https://www.ncbi.nlm.nih.gov/geo/), TCGA (https://dcc.icgc.org/), ICGC (https://dcc.icgc.org/projects-/LIRI-JP), HPA (https://www.proteinatlas.org/). Therefore, everyone can get the data online. If you have further inquiries, please contact the authors.
References
Sung H, Ferlay J, Siegel RL, Laversanne M, Soerjomataram I, Jemal A, et al. Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2021;71(3):209–49. https://doi.org/10.3322/caac.21660.
Kumari R, Sahu MK, Tripathy A, Uthansingh K, Behera M. Hepatocellular carcinoma treatment: hurdles, advances and prospects. Hepat Oncol. 2018;5(2): HEP08. https://doi.org/10.2217/hep-2018-0002.
Fu T, Dai L-J, Wu S-Y, Xiao Y, Ma D, Jiang Y-Z, et al. Spatial architecture of the immune microenvironment orchestrates tumor immunity and therapeutic response. J Hematol Oncol. 2021;14(1):98. https://doi.org/10.1186/s13045-021-01103-4.
Tang T, Huang X, Zhang G, Hong Z, Bai X, Liang T. Advantages of targeting the tumor immune microenvironment over blocking immune checkpoint in cancer immunotherapy. Signal Transduct Target Ther. 2021;6(1):72. https://doi.org/10.1038/s41392-020-00449-4.
Farhood B, Najafi M, Mortezaee K. CD8+ cytotoxic T lymphocytes in cancer immunotherapy: a review. J Cell Physiol. 2019;234(6):8509–21. https://doi.org/10.1002/jcp.27782.
Wang J, Li R, Cao Y, Gu Y, Fang H, Fei Y, et al. Intratumoral CXCR5+CD8+T associates with favorable clinical outcomes and immunogenic contexture in gastric cancer. Nat Commun. 2021;12(1):3080. https://doi.org/10.1038/s41467-021-23356-w.
Sanmamed MF, Nie X, Desai SS, Villaroel-Espindola F, Badri T, Zhao D, et al. A burned-out CD8+ T-cell subset expands in the tumor microenvironment and curbs cancer immunotherapy. Cancer Discov. 2021;11(7):1700–15. https://doi.org/10.1158/2159-8290.CD-20-0962.
Pan Q, Cheng Y, Cheng D. Identification of CD8+ T cell-related genes: correlations with immune phenotypes and outcomes of liver cancer. J Immunol Res. 2021;2021:9960905. https://doi.org/10.1155/2021/9960905.
Zhang Y, Wang D, Peng M, Tang L, Ouyang J, Xiong F, et al. Single-cell RNA sequencing in cancer research. J Exp Clin Cancer Res. 2021;40(1):81. https://doi.org/10.1186/s13046-021-01874-1.
Newman AM, Liu CL, Green MR, Gentles AJ, Feng W, Xu Y, et al. Robust enumeration of cell subsets from tissue expression profiles. Nat Methods. 2015;12(5):453–7. https://doi.org/10.1038/nmeth.3337.
Fang L, Yu W, Zhu P, Yu G, Ye B. TEDC2 correlated with prognosis and immune microenvironment in lung adenocarcinoma. Sci Rep. 2023;13(1):5006. https://doi.org/10.1038/s41598-023-32238-8.
Wang H, Cai H, Li L. Comprehensive analysis of m6A reader YTHDF2 prognosis, immune infiltration, and related regulatory networks in hepatocellular carcinoma. Heliyon. 2024;10(1): e23204. https://doi.org/10.1016/j.heliyon.2023.e23204.
Yoshihara K, Shahmoradgoli M, Martínez E, Vegesna R, Kim H, Torres-Garcia W, et al. Inferring tumour purity and stromal and immune cell admixture from expression data. Nat Commun. 2013;4:2612. https://doi.org/10.1038/ncomms3612.
Xiang X, Wang J, Lu D, Xu X. Targeting tumor-associated macrophages to synergize tumor immunotherapy. Signal Transduct Target Ther. 2021;6(1):75. https://doi.org/10.1038/s41392-021-00484-9.
Dolina JS, Van Braeckel-Budimir N, Thomas GD, Salek-Ardakani S. CD8+ T cell exhaustion in cancer. Front Immunol. 2021;12: 715234. https://doi.org/10.3389/fimmu.2021.715234.
He Q-F, Xu Y, Li J, Huang Z-M, Li X-H, Wang X. CD8+ T-cell exhaustion in cancer: mechanisms and new area for cancer immunotherapy. Brief Funct Genomics. 2019;18(2):999–106. https://doi.org/10.1093/bfgp/ely006.
Jovic D, Liang X, Zeng H, Lin L, Xu F, Luo Y. Single-cell RNA sequencing technologies and applications: a brief overview. Clin Transl Med. 2022;12(3): e694. https://doi.org/10.1002/ctm2.694.
Hwang B, Lee JH, Bang D. Single-cell RNA sequencing technologies and bioinformatics pipelines. Exp Mol Med. 2018;50(8):1–14. https://doi.org/10.1038/s12276-018-0071-8.
Yin M, Wang X, Lu J. Advances in IKBKE as a potential target for cancer therapy. Cancer Med. 2020;9(1):247–58. https://doi.org/10.1002/cam4.2678.
Lu S, Cai S, Peng X, Cheng R, Zhang Y. Integrative transcriptomic, proteomic and functional analysis reveals ATP1B3 as a diagnostic and potential therapeutic target in hepatocellular carcinoma. Front Immunol. 2021;12: 636614. https://doi.org/10.3389/fimmu.2021.636614.
Yu J, Liu Y, Zhang W, Yang X, Tang W, Liang H, et al. Musculin deficiency aggravates colonic injury and inflammation in mice with inflammatory Bowel disease. Inflammation. 2020;43(4):1455–63. https://doi.org/10.1007/s10753-020-01223-y.
Renauer P, Park JJ, Bai M, Acosta A, Lee W-H, Lin GH, et al. Immunogenetic metabolomics reveals key enzymes that modulate CAR T-cell metabolism and function. Cancer Immunol Res. 2023;11(8):1068–84. https://doi.org/10.1158/2326-6066.CIR-22-0565.
Sun C, Li D, Wang Z. BATF-mediated regulation of exhausted CD8+ T-cell responses and potential implications for chimeric antigen receptor-T therapy. Immunotherapy. 2024;16(5):331–40. https://doi.org/10.2217/imt-2023-0170.
Morad G, Helmink BA, Sharma P, Wargo JA. Hallmarks of response, resistance, and toxicity to immune checkpoint blockade. Cell. 2021;184(21):5309–37. https://doi.org/10.1016/j.cell.2021.09.020.
Plaz Torres MC, Lai Q, Piscaglia F, Caturelli E, Cabibbo G, Biasini E, et al. Treatment of hepatocellular carcinoma with immune checkpoint inhibitors and applicability of first-line atezolizumab/bevacizumab in a real-life setting. J Clin Med. 2021;10(15):3201. https://doi.org/10.3390/jcm10153201.
Yu Y, Chen K, Fan Y. Extensive-stage small-cell lung cancer: current management and future directions. Int J Cancer. 2023;152(11):2243–56. https://doi.org/10.1002/ijc.34346.
Ahn E, Araki K, Hashimoto M, Li W, Riley JL, Cheung J, et al. Role of PD-1 during effector CD8 T cell differentiation. Proc Natl Acad Sci USA. 2018;115(18):4749–54. https://doi.org/10.1073/pnas.1718217115.
Peggs KS, Quezada SA, Chambers CA, Korman AJ, Allison JP. Blockade of CTLA-4 on both effector and regulatory T cell compartments contributes to the antitumor activity of anti-CTLA-4 antibodies. J Exp Med. 2009;206(8):1717–25. https://doi.org/10.1084/jem.20082492.
Maker AV, Attia P, Rosenberg SA. Analysis of the cellular mechanism of antitumor responses and autoimmunity in patients treated with CTLA-4 blockade. J Immunol. 2005;175(11):7746–54.
Charoentong P, Finotello F, Angelova M, Mayer C, Efremova M, Rieder D, et al. Pan-cancer immunogenomic analyses reveal genotype-immunophenotype relationships and predictors of response to checkpoint blockade. Cell Rep. 2017;18(1):248–62. https://doi.org/10.1016/j.celrep.2016.12.019.
Acknowledgements
Thanks to all authors for their contributions to this manuscript.
Funding
The research was funded by The Project of Fundamental Research Funds for the Central Universities of Lanzhou University (lzujbky-2022-sp08); Medical Research Improvement Project of Lanzhou University (lzuyxcx-2022-154); Medical Innovation and Development Project of Lanzhou University (lzuyxcx-2022-141); Major Science and Technology Project of Gansu Province [20ZD7FA003, 22ZD6FA050, 22JR9KA002]; Project of Gansu Provincial Department of Education [2021jyjbgs-02]; Project of Gansu Provincial Development and Reform Commission [2020–2022]; Natural Science Foundation of Gansu Province (21JR1RA135, 23JRRA1001).
Author information
Authors and Affiliations
Contributions
All authors participated in this research, including conception and design (CHQ, YJW, and YML), data acquisition (CHQ, XY, FTT, and YML), data analysis and interpretation, material support, study supervision (FTT and YML), and drafting the article or critically revising.
Corresponding author
Ethics declarations
Ethical approval
The Institutional Research Ethics Committees of the Lanzhou University Second Hospital approved this study (protocol number: D2023-078).
Competing interests
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
12672_2024_1092_MOESM1_ESM.docx
Supplementary file1 (DOCX 13487 KB) Supplementary Table 1: The primer sequence of genes. Supplementary Fig. 1: The work flowchart of the study. Abbreviations: WGCNA: Weighted correlation network analysis, ICGC: International Cancer Genome Consortium, TME: tumor microenvironment, TIICs: Tumor-infiltrating immune cells. Supplementary Fig. 2: Integration and clustering of scRNA-Seq. A Annotation and visualization of cell subsets. B A total of 15 distinct clusters were identified via the t-SNE and UMAP algorithms. C Heatmap of cluster markers expression in each cluster. Supplementary Fig. 3: A GO enrichment analysis for CD8 T cell-related genes, including biological process (BP), cellular component (CC), and molecular function (MF). B KEGG enrichment analysis for CD8 T cell-related genes. Supplementary Fig. 4: The predictive value of the risk score in the ICGC external cohort model. A-B Univariate and multivariate Cox analysis of risk scores. C A nomogram was constructed based on risk scores. D The calibration curve of the nomogram. E The ROC curve of the nomogram. F The ROC curves of risk score. G The C-index of risk score. Supplementary Fig. 5: The KEGG signaling pathways enriched by the risk score. A High-risk score group. B Low-risk score group. Supplementary Fig. 6: Expression of risk genes in T cell clusters. A-B The violin map and bubble plot show the expression of risk genes in T cell clusters. C The t-SNE plot demonstrates the risk gene expression levels in T-cell clusters. Supplementary Fig. 7: IHC staining images of risk genes in HCC tissues and normal liver tissues were obtained from the HPA database. A IKBKE. B ATP1B3. C ADA. D BATF.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Qu, C., Yan, X., Wei, Y. et al. Establishment and validation of a novel CD8+ T cell-associated prognostic signature for predicting clinical outcomes and immunotherapy response in hepatocellular carcinoma via integrating single-cell RNA-seq and bulk RNA-seq. Discov Onc 15, 235 (2024). https://doi.org/10.1007/s12672-024-01092-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s12672-024-01092-z