Abstract
Background
Acute Lymphoblastic Leukemia (ALL) is a neoplasm of the hematopoietic system characterized by a clonal expansion of abnormal lymphocyte precursor cells. ALL is the most common form of cancer in children, but despite advances in treatment, it can still be fatal. Ethnic differences influence survival rates, and genomic ancestry plays an important role, especially in mixed-race populations such as Latin America. This study aims to analyze the influence of genomic ancestry on toxicity in children with ALL in the Amazon region.
Methods
The study included 171 patients (protocol number 119,649/2012—Ethics Committee) with ALL treated at a pediatric treatment center in Belém do Pará, in the Brazilian Amazon. The patients were submitted to the BFM protocol of induction therapy for ALL. Toxicity was assessed based on laboratory tests and adverse events, classified according to the CTC-NCI guide. Genomic ancestry was determined using autosomal informative markers.
Results
The majority of children (94.74%) developed some type of toxicity during treatment, 87.04% of which were severe. Infectious toxicity was the most common, present in 84.8% of cases, 77.24% of which were severe. Amerindian ancestry showed an association with the risk of severe general toxicity and severe infectious toxicity, with a contribution of 35.0% demonstrating a significant increase in risk. In addition, post-induction refractoriness and relapse were also associated with an increased risk of death.
Conclusion
This study highlights the influence of Amerindian genomic ancestry on response to therapy and toxicity in children with ALL in the Amazon region. Understanding these associations can contribute to personalizing treatment and improving clinical outcomes.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Acute Lymphoblastic Leukemia (ALL) is a hematopoietic neoplasm, defined as a clonal expansion of abnormal precursor lymphocytes (B and/or T cells), with complex and multifactorial events leading to malignant transformation, influenced by various factors [1, 2]. ALL is the most common form of cancer in children worldwide, and while therapeutic alternatives exist, it is still clinically manageable. However, ALL can lead to a fatal relapse, resulting in early mortality [3, 4].
Advancements in chemotherapy and knowledge about diagnostic alternatives for ALL have emerged in recent decades. The possibility of new drugs and therapeutic targets has led to the introduction of cellular therapies and the implementation of precision medicine to identify genomic markers that assist in treatment guidance and prognosis, significantly improving the survival rate of ALL. It has reached a cure and recovery rate of 90% in children living in developed countries, a reality not experienced in Latin American or low-income countries, where outcomes for children and adults are considerably worse than those in developed countries [4,5,6,7,8].
Ethnic differences in survival rates after childhood ALL vary worldwide [9,10,11,12,13,14], with Hispanic-origin children exhibiting poorer survival rates. Interestingly, Hispanic patients treated in high-income countries showed lower survival rates, highlighting ancestral genomic bias as a contributing factor to these outcomes. Particularly for the Latin American population, a highly heterogeneous group with genetic heritage contributions from European, African, and Native American ancestry [4, 15,16,17].
Taking these ancestral influences into account, studies conducted with more recent data in populations with a high Amazonian Amerindian ancestry have shown that 87% of ALL patients treated with the European Berlin-Frankfurt-Münster (BFM) Group protocol experienced grade 3 and 4 toxicities [16], a higher frequency than that found in other global populations subjected to the same protocol (26%) [18]. In addition to toxicity from standard ALL treatment, other studies in populations with high Native American ancestry showed a high risk of ALL recurrence [15].
Therefore, the different responses to medications related to ethnic differences can be partially explained by variations in the frequencies of important functional gene variants that make up the pharmacokinetic and pharmacodynamic pathways of chemotherapy agents, and these changes may be directly linked to ancestral genomic heritage. Considering these pieces of evidence in the literature, the present study aims to analyze the influence of genomic ancestry on the development of toxicity in pediatric patients diagnosed with ALL in the Amazon region.
2 Methods
2.1 Ethical considerations
The protocol used in this study was approved by the Research Ethics Committee of the Institute of Health Sciences of the Federal University of Pará, under protocol number 119,649/2012. Individuals under 18 years of age and their guardians were duly informed about the research. Volunteers agreed to participate in the study by signing the Informed Consent Form, and their guardians also signed the Informed Consent Form. Our methodology has already been described by Da Silva et al. [19]
2.2 Study populations
This is a prospective study that included 171 patients newly diagnosed with ALL by immunophenotyping and treated at a reference center for pediatric oncology (Otávio Lobo Hospital) in Belém do Pará, in the Brazilian Amazon. The included patients were between 1 and 18 years old, had no recurrence, comorbidities, or other types of cancer, and had morphological, immunophenotypic, and, when available, molecular diagnoses. We excluded patients with Down's syndrome, sickle cell anemia, HIV, tuberculosis, hepatitis B or C, who had previously undergone radiotherapy or chemotherapy for any type of cancer and who had previously undergone hematopoietic stem cell transplantation.
2.3 LLA induction therapy protocol
The included patients underwent induction therapy for ALL with the BFM 2009 protocol. During the induction therapy, which lasts a total of 64 days, patients receive phase 1 corticosteroids for 36 days; four doses of vincristine; 2 doses of doxorubicin (in low and intermediate-risk patients) or 4 doses of doxorubicin (in high-risk patients); and l-asparaginase (8 doses). During the second phase of induction with cyclophosphamide (two doses), patients received 16 doses of cytarabine and 28 consecutive days of mercaptopurine. Supplementary Table 1 summarizes the details of the treatment protocol used during this study.
2.4 Toxicity evaluation and classification
Laboratory tests, including transaminases and complete blood counts, were recorded in a table at three time points during the induction phase: on the first day of treatment before using any medication, before the second phase of induction before using mercaptopurine but with patients already sensitized to other drugs, and after the second phase of induction. Adverse events such as anorexia, colitis, diarrhea, dyspepsia, mucositis, nausea, vomiting, neutropenia, and documented or undocumented infection were recorded, and numerical stratification was applied to the events according to the CTC-NCI (Common Toxicity Criteria of the US National Cancer Institute). After collecting information, adverse effects were classified by grade, considering CTC-NCI: mild/moderate (grades 0, 1, and 2) and severe (grades 3 and 4).
2.5 DNA extraction and quantification
DNA was extracted using the conventional phenol–chloroform method according to Sambrook. Samples were quantified using the NanoDrop ND-1000 equipment (Thermo Scientific NanoDrop Products, Wilmington, DE, USA).
2.6 Genomic ancestry
The analysis was performed using a panel of 61 informative autosomal ancestry markers according to [23]. Two multiplex PCRs were performed, followed by electrophoresis on the ABI Prism 3130 sequencer (Applied Biosystems, Life Technologies, Carlsbad, CA, USA) and analysis using the GeneMapper ID v.3.2 software (Applied Biosystems, Life Technologies, Carlsbad, CA, USA). Individual proportions of European, African, and Amerindian genetic ancestry were estimated using Structure software v.2.3.3.
2.7 Statistical analysis
A descriptive analysis of the data related to the sample characterization was performed. Quantitative variables were first subjected to the Kolmogorov–Smirnov test to analyze the normality distribution. The protocol for analysis of this study has been reported by Da Silva et al. [19].
For comparative analysis between the study groups regarding sample characterization variables, the chi-square test was applied for categorical variables, and the Mann–Whitney test was used for continuous variables. To analyze the association of variables with the risk of overall severe toxicity, severe infectious toxicity, and the occurrence of deaths in the treatment of children with ALL, simple logistic regression was performed.
All statistical analyses were conducted using the SPSS 20.0 (IBM, Stanford University, USA) statistical package, respecting the significance level of 5% (p ≤ 0.05).
3 Results
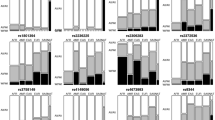
In the present study, it can be observed that 94.74% (95% CI 91.2–97.7) of children with ALL developed some type of toxicity during treatment, among these, 87.04% (95% CI 82.1–92.0) presented severe toxicity. Among the toxicities, the most prevalent was infectious toxicity, present in 84.8% (95% CI 78.9–90.1) of cases, and of these, 77.24% (95% CI 70.3–84.8) developed severe infectious toxicity, as shown in Table 1. Furthermore, it can be observed that the children with ALL included had predominantly European genomic ancestry (47.5%), followed by Amerindian (32.6%) and African (19.5%) (Table 1 and Fig. 1).
When assessing the risk factors associated with the occurrence of severe general toxicity, the contribution of Amerindian genomic ancestry was greater among children who developed severe general toxicity when compared to those who did not (OR 28.076; 95% CI 1.007–782.92). In addition, Amerindian ancestry was significantly associated with an increased risk of severe general toxicity (OR: 28.076; p-value: 0.049), as shown in Table 2 and Fig. 2.
The influence of the distribution of Amerindian ancestry on the susceptibility to developing severe general toxicity in children with ALL was assessed using a logistic regression model by contribution ranges (Fig. 2). The analysis showed that Amerindian ancestry in the range between 22.5% and 40.0% showed an increased risk for this event. However, only a contribution of 35.0% showed a significant increase in risk (OR:2.795; 95% CI 1.113–10.077; p-value: 0.035). The other genomic ancestries showed no significant associations.
In assessing the risk factors associated with the occurrence of severe infectious toxicity, it can be seen that the contribution of Amerindian genomic ancestry was greater among children who developed severe infectious toxicity when compared to those who did not. In addition, Amerindian ancestry was significantly associated with an increased risk of severe infectious toxicity (OR: 13.689; 95% CI 1.050–178.65; p = 0.046), as can be seen in Table 3 and Fig. 3.
The influence of the distribution of Amerindian ancestry on susceptibility to the development of severe infectious toxicity in children with ALL was assessed using a logistic regression model by contribution ranges (Fig. 3). The analysis showed that Amerindian ancestry in the range between 22.5% and 40.0% contribuition of Amerindian genetic profile showed an increased risk of this event. However, only a contribution of 35.0% showed a significant increase in risk (OR: 2.281; 95% CI 1.182–4.732; p-value: 0.020). The other genomic ancestries showed no significant associations.
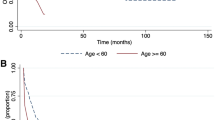
When evaluating the risk factors associated with the occurrence of deaths, it can be seen that post-induction refractoriness was higher among patients who died, significantly increasing the risk of death (p < 0.001; OR: 19.198). The same was observed for relapse, with a higher prevalence among children with ALL who died, significantly increasing the risk of death (p < 0.001; OR: 28.420). In addition, the occurrence of severe general toxicity (p-value: 0.012; OR: 3.421) and severe infectious toxicity (p-value: 0.002; OR: 2.933) were also associated with an increased risk of death among children undergoing treatment for ALL, as detailed in Table 2 in the Supplementary Material.
4 Discussion
Genomic studies are still very rare in populations of Amerindian origin and populations mixed with them. Different studies in the world literature have shown a worse prognosis for Acute Lymphoid Leukemia and high toxicities associated with standard therapy—BFM protocol—for the disease in populations with high Amerindian ancestry [8].
Therefore, it is important to highlight that racial and ethnic differences are significant in the prognosis of patients, since children of populations with Amerindian ancestry have, in addition to a worse treatment outcome, a higher incidence of ALL [20,21,22].
The results of this study shows that the population in the Brazilian Amazon, characterized by a high degree of miscegenation with Amerindians (around 30%), has proportionally high general (94.7%) and severe (87.04%) toxicities [23]. These data show significant differences from the worldwide data on the standard protocol used in ALL therapy.
In our patient sample undergoing therapy for ALL in the Brazilian Amazon, it was possible to identify approximately 10 indigenous individuals with lethal toxicities to the treatment. Therefore, based on this impactful result, the study's objective is to investigate the correlation between the genomic ancestry of these populations and the respective toxicity data.
The results based on the average ancestry in the patient group show similarity to the results obtained from the same population in northern Brazil [23]. Specifically, in Indigenous genomic ancestry, we observed results of 30,6 contribuition of Amerindian genetic profile in the population comprising the patient sample and 30.95% in the group of patients with ALL. When we separately analyzed individuals with severe and non-severe toxicity, we found a significant difference in our results for Amerindian ancestry. This result reinforces studies that suggest Amerindian ancestry may be a risk factor for toxicity and a worse prognosis [24,25,26,27].
When correlating ancestries with toxicities, we found interesting results that indicate severe and infectious toxicity were positively associated with Amerindian ancestry starting from 35%. When quantifying the influence of Amerindian ancestry in the study, it becomes apparent that patients with 35% or more contribuition of Amerindian genetic profile exhibited a significantly elevated risk for the development of toxicity. Therefore, we can assert that, concerning ALL patients, individuals with ancestry above 35% Amerindian have a higher risk of experiencing toxicity with the use of standard ALL therapy. This result may be supported by other studies in the global literature that highlight this elevated risk in individuals of mixed Amerindian heritage and traditional Amazonian indigenous populations [21,22,23,24].
Our hypothesis regarding these results leads us to the idea that there is limited research on the genomics of Amerindians, particularly Amerindian pharmacogenetics and mixed populations. Thus, the correlation of Amerindian ancestry with the severe toxicities observed may potentially result from specific gene variants within this ethnic group found in mixed populations. Other studies investigating the exomes of indigenous people have already demonstrated the existence of novel variants in other genes important for pharmacogenetics, which further supports our hypothesis [29,30,31].
Numerous studies have illustrated that Native American populations exhibit a distinct pattern of variability within the immune system. With regard to our finding of a correlation between Amerindian ancestry and severe infectious toxicity, we can explain it by the fact that Amazonian Amerindians have, throughout their generations, intermarried and have undergone drastic population reductions, which has a direct impact on the diversity of the immune response [26,27,28].
The Brazilian population is the largest admixture in the world, and this has a direct impact on the variation of important genetic variants related to pharmacogenomics. In this way, the international protocols that recommend the use of drugs do not consider the fluctuation of these variants in the Brazilian population, or even the presence of new variants that are specific to Brazilian admixture populations. Knowledge of these very poorly studied mixed-race populations in Latin America could directly lead to the reformulation of new precision medicine protocols for therapies that can be used in these populations, taking into account, in this case, the fluctuation in the allele frequencies of important variants already identified by international agencies and new variants that have not yet been identified.
5 Conclusion
Our work reinforces the fact that mixed-race and indigenous Amerindian populations have a higher risk of toxicity and a worse prognosis for treatment with the protocols used for ALL therapy. Therefore, the data presented is of great value for understanding the variability of response, and may provide new insights for the investigation and genomic analysis of genetic variants that may be directly related to greater severity of therapy for ALL.
Data availability
All data generated or analysed during this study are included in this published article.
References
Karol SE, Yang JJ. Pharmacogenomics and ALL treatment: How to optimize therapy. Semin Hematol. 2020;57:130–6. https://doi.org/10.1053/j.seminhematol.2020.10.001.
Aslam S, Ameer S, Shabana NA, Ahmed M. Pharmacogenetics of induction therapy-related toxicities in childhood acute lymphoblastic leukemia patients treated with UKALL 2003 protocol. Sci Rep. 2021;11:23757. https://doi.org/10.1038/s41598-021-03208-9.
Inaba H, Greaves M, Mullighan CG. Acute lymphoblastic leukaemia. Lancet. 2013;381:1943–55. https://doi.org/10.1016/S0140-6736(12)62187-4.
Lee SHR, Antillon-Klussmann F, Pei D, et al. Association of genetic ancestry with the molecular subtypes and prognosis of childhood acute lymphoblastic leukemia. JAMA Oncol. 2022;8:354–63. https://doi.org/10.1001/jamaoncol.2021.6826.
Bhatia S, Sather HN, Heerema NA, et al. Racial and ethnic differences in survival of children with acute lymphoblastic leukemia. Blood. 2002;100:1957–64. https://doi.org/10.1182/blood-2002-02-0395.
Gupta S, Dai Y, Chen Z, et al. Racial and ethnic disparities in childhood and young adult acute lymphocytic leukaemia: secondary analyses of eight Children’s Oncology Group cohort trials. Lancet Haematol. 2023;10:e129–41. https://doi.org/10.1016/S2352-3026(22)00371-4.
Eche IJ, Aronowitz T. A literature review of racial disparities in overall survival of black children with acute lymphoblastic leukemia compared with white children with acute lymphoblastic leukemia. J Pediatr Oncol Nurs. 2020;37:180–94. https://doi.org/10.1177/1043454220907547.
Quiroz E, Aldoss I, Pullarkat V, et al. The emerging story of acute lymphoblastic leukemia among the Latin American population - biological and clinical implications. Blood Rev. 2019;33:98–105. https://doi.org/10.1016/j.blre.2018.08.002.
Suarez-Kurtz G, Paula DP, Struchiner CJ. Pharmacogenomic implications of population admixture: Brazil as a model case. Pharmacogenomics. 2014;15:209–19. https://doi.org/10.2217/pgs.13.238.
da Silva ME, de Moraes FCA, de Nazaré C-P, et al. Influence of genetic variations in miRNA and genes encoding proteins in the miRNA synthesis complex on toxicity of the treatment of pediatric B-Cell ALL in the Brazilian Amazon. Int J Mol Sci. 2023;24:4431. https://doi.org/10.3390/ijms24054431.
Schmiegelow K, Attarbaschi A, Barzilai S, et al. Consensus definitions of 14 severe acute toxic effects for childhood lymphoblastic leukaemia treatment: a Delphi consensus. Lancet Oncol. 2016;17:e231–9. https://doi.org/10.1016/S1470-2045(16)30035-3.
JPM | Free Full-Text | The Genomic Profile Associated with Risk of Severe Forms of COVID-19 in Amazonian Native American Populations. https://www.mdpi.com/2075-4426/12/4/554. Accessed 29 Sep 2023
Oliveira MLG, Castelli EC, Veiga-Castelli LC, et al. Genetic diversity of the LILRB1 and LILRB2 coding regions in an admixed Brazilian population sample. HLA. 2022;100:325–48. https://doi.org/10.1111/tan.14725.
de Fernandes SSM, Leitão LPC, de Cohen-Paes AN, et al. The role of SLC22A1 and genomic ancestry on toxicity during treatment in children with acute lymphoblastic leukemia of the Amazon Region. Genes. 2022;13:610. https://doi.org/10.3390/genes13040610.
Yang JJ, Cheng C, Devidas M, et al. Ancestry and pharmacogenomics of relapse in acute lymphoblastic leukemia. Nat Genet. 2011;43:237–41. https://doi.org/10.1038/ng.763.
Yao S, Zhu Q, Cole PD, et al. Genetic ancestry and skeletal toxicities among childhood acute lymphoblastic leukemia patients in the DFCI 05–001 cohort. Blood Adv. 2021;5:451–8. https://doi.org/10.1182/bloodadvances.2020003060.
Shoag JM, Barredo JC, Lossos IS, Pinheiro PS. Acute lymphoblastic leukemia mortality in Hispanic Americans. Leuk Lymphoma. 2020;61:2674–81. https://doi.org/10.1080/10428194.2020.1779260.
Stary J, Zimmermann M, Campbell M, et al. Intensive chemotherapy for childhood acute lymphoblastic leukemia: results of the randomized intercontinental trial ALL IC-BFM 2002. J Clin Oncol. 2014;32:174–84. https://doi.org/10.1200/JCO.2013.48.6522.
da Silva EM, Fernandes MR, de Carvalho DC, et al. Effect of genetic ancestry to the risk of susceptibility to gastric cancer in a mixed population of the Brazilian Amazon. BMC Res Notes. 2017;10:646. https://doi.org/10.1186/s13104-017-2963-4.
Zapata-Tarrés M, Balandrán JC, Rivera-Luna R, Pelayo R. Childhood acute leukemias in developing nations: successes and challenges. Curr Oncol Rep. 2021;23:56. https://doi.org/10.1007/s11912-021-01043-9.
Rabin KR. Genetic ancestry and childhood acute lymphoblastic leukemia subtypes and outcomes in the genomic era. JAMA Oncol. 2022;8:342–3. https://doi.org/10.1001/jamaoncol.2021.6785.
Walsh KM, de Smith AJ, Welch TC, et al. Genomic ancestry and somatic alterations correlate with age at diagnosis in Hispanic children with B-cell acute lymphoblastic leukemia. Am J Hematol. 2014;89:721–5. https://doi.org/10.1002/ajh.23727.
Santos NPC, Ribeiro-Rodrigues EM, Ribeiro-dos-Santos ÂKC, et al. Assessing individual interethnic admixture and population substructure using a 48–insertion-deletion (INSEL) ancestry-informative marker (AIM) panel. Hum Mutat. 2010;31:184–90. https://doi.org/10.1002/humu.21159.
Association of Genetic Ancestry With the Molecular Subtypes and Prognosis of Childhood Acute Lymphoblastic Leukemia | Genetics and Genomics | JAMA Oncology | JAMA Network. https://jamanetwork.com/journals/jamaoncology/fullarticle/2788576. Accessed 29 Sep 2023
Wanderley A, vieira, Santos NP, Khayat A salim, et al. Risk of infectious toxicity associated with polymorphism in GSTP1 in children with acute lymphoblastic leukemia in a population of Amazonia. JCO. 2014;32:e21006–e21006. https://doi.org/10.1200/jco.2014.32.15_suppl.e21006.
Leitão LPC, de Carvalho DC, Rodrigues JCG, et al. Identification of genomic variants associated with the risk of acute lymphoblastic leukemia in native Americans from Brazilian Amazonia. J Pers Med. 2022;12:856. https://doi.org/10.3390/jpm12060856.
Zawitkowska J, Lejman M, Zaucha-Prażmo A, et al. Grade 3 and 4 toxicity profiles during therapy of childhood acute lymphoblastic leukemia. In Vivo. 2019;33:1333–9. https://doi.org/10.21873/invivo.11608.
Cohen-Paes A, de Alcântara AL, de Souza ME, et al. Characterization of DNA polymerase genes in Amazonian Amerindian populations. Genes. 2023;14:53. https://doi.org/10.3390/genes14010053.
de Cohen-Paes AN, de Alcântara AL, Moreira FC, et al. Molecular epidemiology in amerindians of the brazilian amazon reveals new genetic variants in DNA repair genes. Genes. 2022;13:1869. https://doi.org/10.3390/genes13101869.
Rodrigues JCG, Fernandes MR, Ribeiro-dos-Santos AM, et al. Pharmacogenomic profile of Amazonian Amerindians. J Pers Med. 2022;12:952. https://doi.org/10.3390/jpm12060952.
Rodrigues JCG, de Souza TP, Pastana LF, et al. Identification of NUDT15 gene variants in Amazonian Amerindians and admixed individuals from northern Brazil. PLoS ONE. 2020;15:e0231651. https://doi.org/10.1371/journal.pone.0231651.
Acknowledgements
We thank the Federal University of Pará (UFPA); the Center for Research in Oncology (NPO/UFPA), the Graduate Program in Genetics and Molecular Biology (PPGM/UFPA) and the Laboratory of Human and Medical Genetics (LGHM/UFPA). We also thank the contributions and ethical evaluation of indigenous rights of Josicleyce Pereira.
Author information
Authors and Affiliations
Contributions
A.V.W., idealization, contextualization, investigation and preparation of the original writing; R.R, statistical analysis; F.C.A.M., writing-review and editing; G.G.d.C.N, writing-review and editing; G.G.d.C.N, translation and preparation of the original script; F.C.A.M., adjustments to the rules and preparation of the original script; A.T.M.T, original writing preparation; L.d.C.P, visualization; L.P.C.L, original writing preparation; E.E.B.P., original writing preparation; B.C.M.K., methodology; A.K.R.d.S., methodology and research; R.M.R.B., financial acquisition, writing proofreading and editing; M.B.O, methodology and software; A.S.K., data curation and original writing preparation; P.P.d.A., proofreading and editing of writing; S.E.B.d.S., conceptualization, financial acquisition, proofreading, and writing editing; M.R.F., preparation of original writing, revision and editing of writing; N.P.C.d.S., conceptualization, revision and editing of writing.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
This study was conducted according to the guidelines of the Declaration of Helsinki and approved by the National Committee for Ethics in Research (CONEP), and the Research Ethics Committee of the Ethics Committee of the Institute of Health Sciences of the Federal University of Pará with the opinion 119,649/2012.
Competing interests
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
12672_2024_1014_MOESM1_ESM.docx
Supplementary material 1: Table S1. online provides the Analysis of the Prevalence of Toxicities by Treatment Phase in Children with Acute Lymphoblastic Leukemia in the Amazon Region. GTI Gastrointestinal. Table S2. online provides the Analysis of factors associated with mortality in children with Acute Lymphoblastic Leukemia in the Amazon Region. Figure S1. online provides the Genetic Ancestry and Susceptibility to Severe General Toxicity in Children with Acute Lymphoblastic Leukemia in the Amazon Region. Case-Toxicity; Control; without toxicity. Figure S2. online provides the . Genetic Ancestry and Susceptibility to Severe Infectious Toxicity in Children with Acute Lymphoid Leukemia in the Amazon Region.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Wanderley, A.V., de Moraes, F.C.A., da Costa Nunes, G.G. et al. Effect of American genomic ancestry on severe toxicities in children with acute lymphoblastic leukemia in the Amazon region. Discov Onc 15, 171 (2024). https://doi.org/10.1007/s12672-024-01014-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s12672-024-01014-z