Abstract
Purpose
This study aimed to explore novel tumor immune microenvironment (TIME)-associated biomarkers in prostate adenocarcinoma (PRAD).
Methods
PRAD RNA-sequencing data were obtained from UCSC Xena database as the training dataset. The ESTIMATE package was used to evaluate stromal, immune, and tumor purity scores. Differentially expressed genes (DEGs) related to TIME were screened using the immune and stromal scores. Gene functions were analyzed using DAVID. The LASSO method was performed to screen prognostic TIME-related genes. Kaplan–Meier curves were used to evaluate the prognosis of samples. The correlation between the screened genes and immune cell infiltration was explored using Tumor IMmune Estimation Resource. The GSE70768 dataset from the Gene Expression Omnibus was used to validate the expression of the screened genes.
Results
The ESTIMATE results revealed that high immune, stromal, and ESTIMATE scores and low tumor purity had better prognoses. Function analysis indicated that DEGs are involved in the cytokine–cytokine receptor interaction signaling pathway. In TIME-related DEGs, METTL7B, HOXB8, and TREM1 were closely related to the prognosis. Samples with low expression levels of METTL7B, HOXB8, and TREM1 had better survival times. Similarly, both the validation dataset and qRT-PCR suggested that METTL7B, HOXB8, and TREM1 were significantly decreased. The three genes showed a positive correlation with immune infiltration.
Conclusions
This study identified three TIME-related genes, namely, METTL7B, HOXB8, and TREM1, which correlated with the prognosis of patients with PRAD. Targeting the TIME-related genes might have important clinical implications when making decisions for immunotherapy in PRAD.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Prostate adenocarcinoma (PRAD) remains the leading cause of cancer related mortality among men in the United States [1]. Despite advances in clinical care, mortality rates remain high, indicating a need for better understanding of the factors influencing PRAD prognosis and treatment response [2, 3].
Recent evidence suggests that PRAD prognosis is heavily dependent on the tumor microenvironment [4]. The tumor immune microenvironment (TIME), comprised of extracellular matrix, stromal cells, and other tumor associated cells, can modulate the tumor’s response to therapy and can influence the progression of the tumor [5]. TIME has been reported to profoundly influence the growth and metastasis of cancer. It affects prognosis, tumor growth, and treatment response through a variety of mechanisms [6]. The composition of the tumor microenvironment can affect prognosis by influencing the proliferation, metastasis, and drug resistance of cancer cells [7]. Factors like the abundance of immune cells, angiogenesis (new blood vessel formation), and cytokines (types of proteins) released by the environment can either inhibit or promote the growth of cancer cells [8]. These signals can also have a large impact on how a patient responds to treatment; for example, certain treatments may be targeted more directly at immunoprivileged environments, and certain drugs tested in preclinical trials may not reach the tumor due to an immunosuppressive microenvironment [9]. In terms of tumor growth, TIME affect the rate of metastasis, and provide signals for cell growth, movement, and survival [10]. TIME could also be a source of genetic mutations, which can lead to the selection of drug resistant tumor cells [11]. Additionally, the presence of certain immune cells can influence the growth and progression of tumors [12]. Finally, the microenvironment can also influence response to treatments. Factors like hypoxia (low oxygen) or the immune cell composition can affect the effectiveness of certain therapies [13]. Additionally, certain signaling pathways in the microenvironment can be targeted by drugs to suppress tumor growth and improve outcomes [14]. Overall, the tumor microenvironment is an essential component of tumor biology and can affect the prognosis, growth, and response to treatments of cancer.
In this study, we downloaded the PRAD RNA-sequencing (RNA-seq) data from the UCSC Xena database. Then, the ESTIMATE method was employed to analyze the immune and stromal scores and tumor purity of PRAD samples. In addition, we analyzed the correlation between TIME and clinical information, from which we obtained the novel TIME-related prognostic genes for PRAD.
2 Materials and methods
2.1 Data processing
RNA-seq data of PRAD were obtained from UCSC Xena (https://xenabrowser.net/datapages/) according to Illumina HiSeq 2000 RNA-seq platform. The data included 551 samples of which 546 samples (494 tumor samples and 52 controls) with clinical information were used as the training dataset. Meanwhile, the GSE70768 dataset was obtained from NCBI Gene Expression Omnibus (https://www.ncbi.nlm.nih.gov/) database, including 199 samples (25 tumor samples and 74 controls), which was regarded as the validation dataset [15].
2.2 Evaluation of immune and stromal scores of samples using ESTIMATE
To calculate the immune, stromal, tumor purity, and ESTIMATE scores of samples in the UCSC dataset, the ESTIMATE package in R3.6.1 (http://127.0.0.1:29606/library/estimate/html/estimateScore.html) [16] was used. The samples were divided into high or low- scores groups according to the median value of immune, stromal, tumor purity, and ESTIMATE scores, respectively. Then, by combining with clinical information of the samples (including overall survival [OS], recurrence-free survival (RFS, and disease-free survival [DFS]), the Kaplan–Meier (KM) of survival package (version 2.41-1, http://bioconductor.org/packages/survivalr/) [17] in R3.6.1 was employed to assess the prognostic difference in different levels of TIME scores.
2.3 Screening of TIME-related genes
According to the immune and stromal scores, samples were separated into two groups including high and low stromal scores and high and low immune scores. Then, the limma package in R3.6.1 (version 3.34.7, https://bioconductor.org/packages/release/bioc/html/limma.html) [18] was employed to screen differentially expressed genes (DEGs) in the two groups with the threshold of the false discovery rate (FDR) of < 0.05 and |log2fold change (FC)|>0.5. The screened DEGs were visualized using heatmap constructed by pheatmap (https://cran.r-project.org/web/packages/pheatmap/index.html) in R3.6.1 [19, 20]. Then, the overlapped DEGs between the two groups were retained for the subsequent analysis. Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes analyses were performed on the overlapped DEGs via DAVIA (version 6.8, https://david.ncifcrf.gov/) [21, 22] with the threshold of FDR < 0.05.
2.4 Screening of TIME-related prognostic genes
The univariate Cox regression analysis of the survival package (version 2.41-1) [17] was conducted to obtain the TIME-related prognostic genes on the basis of the abovementioned TIME-related DEGs. Then, LASSO in LARS package (version 1.2, https://cran.r-project.org/web/packages/lars/index.html.) [23] was used to further obtain prognosis-related DEGs. Furthermore, the multivariate Cox regression analysis in R3.6.1 (version 2.41-1) [17] was used to screen independent TIME-related prognostic genes, and the results were visualized using forestplot (version 1.10, https://cran.r-project.org/web/packages/forestplot/index.html.) [24]. Then, the TIME-related genes were differentially expressed in the PRAD tumor and control samples from the UCSC dataset. Based on the DEG expression, the samples were separated into high and low expression groups. The Fisher precision test in R3.6.1 was used to analyze the difference between the two groups. Moreover, the significance of OS, RFS, and DFS in different groups was assessed using the KM curves [17]. The immunohistochemistry staining of TIME-related prognostic DEGs was validated using the Human Protein Atlas (HPA) (https://www.proteinatlas.org/) [25].
2.5 Correlation analysis between novel TIME-related DEGs and TIME
The TIME-related DEG expression level was extracted from the UCSC dataset, and cor function (https://www.rdocumentation.org/packages/stats/versions/3.6.1/topics/cor) was then performed to calculate the association between TIME-related DEGs and TIME scores. The results were represented using a correlation scatter plot.
Tumor IMmune Estimation Resource (https://cistrome.shinyapps.io/timer/) [26] was applied to analyze the association between TIME-related DEGs and six tumor-infiltrating immune cells (i.e., B cells, CD4 + T cells, CD8 + T cells, neutrophils, macrophages, and dendritic cells). Then, the correlation between TIME-related genes and infiltration of various immune cell subtypes was analyzed.
2.6 Validation of TIME-related DEGs
Initially, the expression levels of TIME-related DEGs were obtained from GSE70768 (validation dataset), and the difference in TIME-related DEGs between PRAD tumor samples and controls was then analyzed. Meanwhile, we collected 10 pairs PRAD tissues and adjacent tissues to validate the TIME-related DEGs. The expression of the TIME-related DEGs using quantitative real-time PCR (qRT-PCR) through the instruction of previous study. The results were calculated using 2−△△Ct method. Then, the samples were divided into high and low expression groups according to the clinical prognostic information of PRAD samples. KM curves were made to assess the significance of sample prognosis in the high and low expression groups.
3 Results
3.1 Evaluation of immune and stromal scores of samples using ESTIMATE
ESTIMATE results indicated that the ESTIMATE, immune, and stromal scores in PRAD tumors were significantly lower than those in controls; contrary results were obtained in tumor purity (Fig. 1A, Supplementary Table 1). Then, the samples were divided into high or low scores of ESTIMATE, immune, and stromal. As shown in Fig. 1B, high ESTIMATE, immune, and stromal scores correlated with good prognosis including OS, RFS, and DFS. By contrast, the higher tumor the purity score, the worse the patient’s prognosis.
Evaluation of immune and stromal scores of samples using ESTIMATE. A Stromal, immune, and ESTIMATE scores and purity in controls and PRAD tumor tissues. B KM curves for evaluating the OS, RFS, and DFS in different levels of stromal, immune, and ESTIMATE scores and purity. DFS disease-free survival, OS overall survival, PRAD prostate adenocarcinoma, RFS recurrence-free survival
3.2 Screening of TIME-associated genes
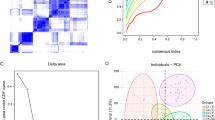
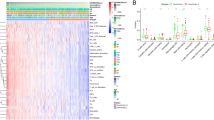
The samples were divided into high and low stromal scores and high and low immune scores. A total of 2017 and 1808 DEGs were screened from these two groups, which were visualized using heatmap and volcano map (Fig. 2A, B). Then, the overlapped DEGs between the stromal and immune groups were obtained; as a result, 16 downregulated and 1213 upregulated DEGs were screened (Fig. 2C).
The GO and KEGG analysis results indicated that the overlapped DEGs were involved in 136 biology processes (BPs), 17 cellular components, 31 molecular functions, and 19 KEGG pathways (Supplementary Table 2). The top 10 GO and KEGG are presented in Fig. 3, indicating that the overlapped DEGs significantly participated in immune response, plasma membrane, transmembrane signaling receptor activity, and cytokine–cytokine receptor interaction. Among the GO function and KEGG pathways, cytokine–cytokine receptor interaction was the most significant pathway that is separately displayed in Supplementary Fig. 1. Furthermore, the overlapped DEGs in this pathway were labeled, such as CCR8, CCL21, and IL2RA.
3.3 Screening of TIME-related prognostic genes
In total, 62 TIME-associated prognostic genes were screened from the overlapped TIME-related DEGs. Then, 14 optimized DEGs were obtained from the TIME-related prognostic genes using the LASSO method. Finally, three DEGs (METTL7B, HOXB8, and TREM1) were screened, which were significantly related to independent prognosis (Fig. 4A).
Screening of TIME-related prognostic genes. A LASSO parameter and multivariate Cox regression analysis forest display map. B KM curves for evaluating the importance of METTL7B, HOXB8, and TREM1 through OS, RFS, and DFS. DFS disease-free survival, KM Kaplan–Meier, OS overall survival, RFS recurrence-free survival
According to the expression levels of METTL7B, HOXB8, and TREM1, the samples in the UCSC dataset were divided into the high and low expression groups. The KM curves indicated that METTL7B, HOXB8, and TREM1 correlated with the OS, RFS, and DFS of the samples. Samples with low expression levels of METTL7B, HOXB8, and TREM1 had better prognoses (Fig. 4B). Moreover, the correlation between the gene level and clinicopathological information of PRAD is analyzed in Table 1. Notably, low expression levels of HOXB8 and TREM1 significantly correlated with a low tumor recurrence rate.
3.4 Correlation analysis of METTL7B, HOXB8, and TREM1 and TIME
To evaluate the association between METTL7B, HOXB8, and TREM1 and TIME, the cor function was performed to obtain the relationship between TIME scores and gene expression. A shown in Fig. 5A, the findings revealed that the expression levels of METTL7B, HOXB8, and TREM1 showed a significant positive correlation with the ESTIMATE, stromal, and immune scores and a negative correlation with tumor purity. Moreover, we analyzed the association between the expression of the three genes and immune infiltration level in PRAD. As shown in Fig. 5B, the expression of METTL7B significantly correlated with CD4 + T cells, neutrophils, and dendritic cells (R > 0.3). HOXB8 showed a significant correlation with CD4 + T cells and dendritic cells (R > 0.3). TREM1 was positively related to the infiltration levels of neutrophils and dendritic cells (R = 0.503; R = 0.426). METTL7B, HOXB8, and TREM1 were positively correlated with macrophages (R > 0.2).
3.5 Validation of METTL7B, HOXB8, and TREM1 in PRAD
To validate the importance of METTL7B, HOXB8, and TREM1 in PRAD, we obtained their expression levels from the GSE70768 dataset. Figure 6A–C shows that the expression levels of the three genes were consistent with the UCSC dataset. Moreover, KM curves revealed that low expression levels of METTL7B, HOXB8, and TREM1 correlated with a good prognosis, which was also consistent with the training dataset (Fig. 6D). Furthermore, we examined these genes between PRAD tissues and adjacent tissues, similar results that the expression of METTL7B, HOXB8, and TREM1 in PRAD was significantly decreased (P < 0.01, Fig. 6E).
Validation of METTL7B, HOXB8, and TREM1 in PRAD. Expression levels of METTL7B, HOXB8, and TREM1 in PRAD tumors and normal control samples from TCGA (A), GSE70768 (B) and GSE29079 (C). D. KM curves for evaluating the importance of METTL7B, HOXB8, and TREM1 through OS, RFS, and DFS. E. Expression levels of METTL7B, HOXB8, and TREM1 in PRAD tumors and normal control samples using qRT-PCR. DFS disease-free survival, KM Kaplan–Meier, OS overall survival, RFS recurrence-free survival, qRT-PCR quantitative real-time PCR
4 Discussion
Multiple factors, stages, and genes are involved in the occurrence and development of PRAD, where TIME is an important factor. Recently, immunotherapy was the novel treatment for PRAD tumors, whereas the clinical outcome was related to the characteristics of malignant tumors, such as hormone dependence, low tumor mutation load, and immunosuppressive microenvironment. Besides, previous studies have found that TIME correlated with the prognosis of patients with PRAD [27]. Thus, further exploration of TIME in PRAD was significant to help doctors make decisions about the treatment method and predict the prognosis for PRAD.
The results of this study revealed that the PRAD samples with high immune and stromal scores had a better prognosis, and those with high tumor purity had a worse prognosis. In addition, we collected the OS, RFS, and DFS of samples to examine the correlation in immune and stromal scores and survival time using KM curves. The results obtained were similar to the ESTIMATE results. These observations were consistent with the results of previous studies. For example, Chen et al. [28] suggested that stromal, immune, and ESTIMATE scores closely correlated with the OS of patients with PRAD. In addition, similar results were obtained in multiple cancer types, such as breast cancer, bladder cancer, and lung adenocarcinoma, indicating that the stromal, immune, and ESTIMATE scores and tumor purity in TIME played a significant role in immunotherapy [29,30,31]. Xiang et al. [32] indicated that the stromal, immune, and ESTIMATE scores and tumor purity in the microenvironment were associated with TIME. Thus, our study screened TIME-related DEGs by comparing stromal scores and immune scores. Subsequently, 1229 TIME-related DEGs were screened using the limma package. GO BP results indicated the involvement of DEGs in the immune response. Immune response regulated the development of PRAD tumors, which played an important role when making decisions for immunotherapy [33, 34]. The DEGs might be novel biomarkers for treating PRAD. In addition, KEGG results indicated that the DEGs were related to the cytokine–cytokine receptor interaction pathway. Previous studies have found that this pathway always involved some immune-related genes that are involved in different cancers, such as renal cell carcinoma and hepatocellular carcinoma [35, 36]. This pathway might be an important factor in the immunotherapy for PRAD. Further experiments must be performed to understand the mechanism of immunotherapy in PRAD.
Previous studies have indicated that the TIME was related to the prognosis of patients with PRAD. In this study, three prognostic DEGs, namely, METTL7B, HOXB8, and TREM1, were identified. The KM curves were used to evaluate the correlation between the three DEGs and patient prognosis, suggesting that patients with low expression levels of METTL7B, HOXB8, and TREM1 had good OS, RFS, and DFS. The results were validated using the external cohort, which obtained the same results as TCGA dataset. Our results were similar with the previous studies that have used external cohorts to validated the prognostic value of the biomarkers in PRAD [37,38,39]. The method further confirmed the results in this study. This novel TIME hub genes-related risk score model provides a new theoretical basis for the prognosis assessment of PRAD patients, which is expected to be further applied in the future clinical management. A prospective study of clinical cohorts recruiting PRAD patients in different stage will help validating this risk score model. The expression of METTL7B, HOXB8, and TREM1 was examined each month. Then, the follow-up was performed to observe the prognosis of PRAD patients. The KM curves and survival analysis will be carried out to the correlation between risk score model and prognosis. This study is expected to be conducted for 5 years or even longer to obtain good persuasiveness.
Meanwhile, significant differences in METTL7B, HOXB8, and TREM1 were found between the controls and tumor samples in both mRNA levels. The three genes might be novel TIME-related biomarkers for PRAD. METTL7B, an alkyl thiol methyltransferase, could metabolize hydrogen sulfide (H2S) [40]. H2S was found to participate in the epithelial–mesenchymal transition and tumor migration and invasion [41]. A recent study found that the expression of METTL7B positively correlated with immunosuppressive cells suggesting that it might play a significant role in modulating TIME [42]. Meanwhile, METTL7B expression was positively associated with CD4 + T cells and dendritic cells. All the results indicated that METTL7B could be used to predict the TIME in PRAD. Moreover, Redecke et al. [43] reported that HOXB8 transfected in mouse bone marrow cells with unlimited proliferative capacity that could enable investigations of immune cell differentiation and function. This study found that HOXB8 is closely correlated with CD4 + T cells. Besides, Zhao et al. [44] pointed out that high expression levels of TREM1 had improved the infiltration of regulatory T cells and reduced the infiltration of CD8 + T cells. Similarly, this study found that the expression of TREM1 could regulate the TIME, including neutrophils and dendritic cells. Previous studies have suggested that CD4 + T cells, CD8 + T cells, neutrophils, and dendritic cells are associated with the impairment of proliferation, cytokine production, and migratory capacities of effector T cells [45]. Besides, Meng et al. [46] firstly pointed out the infiltration of immunocytes among PRAD via the CIBERSORT algorithm. This study indicated that M2 macrophages was related to gene markers, whick could predict the prognosis of PRAD patients. These results were consistent with our study that we found that METTL7B, HOXB8, and TREM1 were positively correlated with M2 macrophages. Regulating the expression levels of METTL7B, HOXB8, and TREM1 may have remarkable clinical applications in enhancing immunotherapy. Immunotherapy has shown good prospects in treating cancer. We will continue to focus on the genes related to tumor microenvironment of PRAD in the future. Further exploration on genes related to tumor microenvironment will help treating patients with PRAD using immunotherapy as soon. Thus, more experiments such as mice experiments, molecular biology research and clinical test need be performed to validate these results in this study.
5 Conclusions
This study explored the expression levels of three TIME-related genes including METTL7B, HOXB8, and TREM1, which correlated with the prognosis of patients with PRAD. Moreover, targeting the TIME-related genes might have important clinical implications when making decisions for immunotherapy in PRAD.
Data Availability
The datasets generated and/or analysed during the current study are available in the UCSC Xena repository, (https://xenabrowser.net/datapages/?dataset=TCGA-PRAD.htseq_fpkm.tsv&host=https%3A%2F%2Fgdc.xenahubs.net&removeHub=https%3A%2F%2Fxena.treehouse.gi.ucsc.edu%3A443). GSE70768: https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE70768.
References
Rebbeck TR. Prostate cancer genetics: variation by race, ethnicity, and geography. Semin Radiat Oncol. 2017;27(1):3–10.
Kumar C, Bagga J, Chiliveru S, Kohli S, Bharadwaj A, Jain M, et al. Substantial remission of prostate adenocarcinoma with dendritic cell therapy APCEDEN(®) in combination with chemotherapy. Future Sci OA. 2019;5(10):Fso435.
Yunger S, Bar El A, Zeltzer LA, Fridman E, Raviv G, Laufer M, et al. Tumor-infiltrating lymphocytes from human prostate tumors reveal anti-tumor reactivity and potential for adoptive cell therapy. Oncoimmunology. 2019;8(12): e1672494.
Xu Z, Chen S, Zhang Y, Liu R, Chen M. Roles of m5C RNA modification patterns in biochemical recurrence and tumor microenvironment characterization of prostate adenocarcinoma. Front Immunol. 2022;13: 869759.
Gu CY, Dai B, Zhu Y, Lin GW, Wang HK, Ye DW, et al. The novel transcriptomic signature of angiogenesis predicts clinical outcome, tumor microenvironment and treatment response for prostate adenocarcinoma. Mol Med. 2022;28(1):78.
Wang N, Zhu L, Wang L, Shen Z, Huang X. Identification of SHCBP1 as a potential biomarker involving diagnosis, prognosis, and tumor immune microenvironment across multiple cancers. Comput Struct Biotechnol J. 2022;20:3106–19.
Zhang H, Luo YB, Wu W, Zhang L, Wang Z, Dai Z, et al. The molecular feature of macrophages in Tumor immune microenvironment of glioma patients. Comput Struct Biotechnol J. 2021;19:4603–18.
Zhong T, Zhang W, Guo H, Pan X, Chen X, He Q, et al. The regulatory and modulatory roles of TRP family channels in malignant tumors and relevant therapeutic strategies. Acta Pharm Sinica B. 2022;12(4):1761–80.
Wu T, Wang W, Shi G, Hao M, Wang Y, Yao M, et al. Targeting HIC1/TGF-β axis-shaped prostate cancer microenvironment restrains its progression. Cell Death Dis. 2022;13(7):624.
Masetti M, Carriero R, Portale F, Marelli G, Morina N, Pandini M, et al. Lipid-loaded tumor-associated macrophages sustain tumor growth and invasiveness in Prostate cancer. J Exp Med. 2022;219(2):e20210564.
Wang W, Zhang J, Wang Y, Xu Y, Zhang S. Identifies microtubule-binding protein CSPP1 as a novel cancer biomarker associated with ferroptosis and tumor microenvironment. Comput Struct Biotechnol J. 2022;20:3322–35.
Zhao C, Xiong K, Ji Z, Liu F, Li X. The prognostic value and immunological role of STEAP1 in pan-cancer: a result of data-based analysis. Oxidative Med Cell Longevity. 2022;2022:8297011.
Yang L, Zhang Y, Wang Y, Jiang P, Liu F, Feng N. Ferredoxin 1 is a cuproptosis-key gene responsible for tumor immunity and drug sensitivity: a pan-cancer analysis. Front Pharmacol. 2022;13: 938134.
Boumahdi S, de Sauvage FJ. The great escape: tumour cell plasticity in resistance to targeted therapy. Nat Rev Drug Discovery. 2020;19(1):39–56.
Ross-Adams H, Lamb AD, Dunning MJ, Halim S, Lindberg J, Massie CM, et al. Integration of copy number and transcriptomics provides risk stratification in prostate cancer: a discovery and validation cohort study. EBio Med. 2015;2(9):1133–44.
Hu D, Zhou M, Zhu X. Deciphering immune-associated genes to predict survival in clear cell renal cell cancer. Biomed Res Int. 2019;2019:2506843.
Wang P, Wang Y, Hang B, Zou X, Mao JH. A novel gene expression-based prognostic scoring system to predict survival in gastric cancer. Oncotarget. 2016;7(34):55343–51.
Ritchie ME, Phipson B, Wu D, Hu Y, Law CW, Shi W, et al. Limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 2015;43(7):e47.
Wang L, Cao C, Ma Q, Zeng Q, Wang H, Cheng Z, et al. RNA-seq analyses of multiple meristems of soybean: novel and alternative transcripts, evolutionary and functional implications. BMC Plant Biol. 2014;14: 169.
Eisen MB, Spellman PT, Brown PO, Botstein D. Cluster analysis and display of genome-wide expression patterns. Proc Natl Acad Sci USA. 1998;95(25):14863–8.
Huang da W, Sherman BT, Lempicki RA. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat Protoc. 2009;4(1):44–57.
Huang da W, Sherman BT, Lempicki RA. Bioinformatics enrichment tools: paths toward the comprehensive functional analysis of large gene lists. Nucleic Acids Res. 2009;37(1):1–13.
Goeman JJ. L1 penalized estimation in the Cox proportional hazards model. Biometrical J Biometrische Z. 2010;52(1):70–84.
Liu XF, Gao ZM, Wang RY, Wang PL, Li K, Gao S. Comparison of Billroth I, Billroth II, and Roux-en-Y reconstructions after distal gastrectomy according to functional recovery: a meta-analysis. Eur Rev Med Pharmacol Sci. 2019;23(17):7532–42.
Miura K, Ishida K, Fujibuchi W, Ito A, Niikura H, Ogawa H, et al. Differentiating rectal carcinoma by an immunohistological analysis of carcinomas of pelvic organs based on the NCBI Literature Survey and the human protein Atlas database. Surg Today. 2012;42(6):515–25.
Li T, Fan J, Wang B, Traugh N, Chen Q, Liu JS, et al. TIMER: a web server for comprehensive analysis of tumor-infiltrating immune cells. Cancer Res. 2017;77(21):e108–10.
Andersen LB, Nørgaard M, Rasmussen M, Fredsøe J, Borre M, Ulhøi BP, et al. Immune cell analyses of the tumor microenvironment in prostate cancer highlight infiltrating regulatory T cells and macrophages as adverse prognostic factors. J Pathol. 2021;255(2):155–65.
Chen L, Zhang M, Zhou J, Zhang L, Liang C. Establishment of an age- and tumor microenvironment-related gene signature for survival prediction in Prostate cancer. Cancer Med. 2022. https://doi.org/10.1002/cam4.4776.
Xu M, Li Y, Li W, Zhao Q, Zhang Q, Le K, et al. Immune and stroma related genes in breast cancer: a comprehensive analysis of tumor microenvironment based on the cancer genome atlas (TCGA) database. Front Med. 2020;7: 64.
Samstein RM, Lee CH, Shoushtari AN, Hellmann MD, Shen R, Janjigian YY, et al. Tumor mutational load predicts survival after immunotherapy across multiple cancer types. Nat Genet. 2019;51(2):202–6.
Wu J, Li L, Zhang H, Zhao Y, Zhang H, Wu S, et al. A risk model developed based on tumor microenvironment predicts overall survival and associates with Tumor immunity of patients with lung adenocarcinoma. Oncogene. 2021;40(26):4413–24.
Xiang S, Li J, Shen J, Zhao Y, Wu X, Li M, et al. Identification of prognostic genes in the tumor microenvironment of hepatocellular carcinoma. Front Immunol. 2021;12: 653836.
Zhang P, An Z, Sun C, Xu Y, Zhang Z. FLG gene mutation up-regulates the abnormal Tumor immune response and promotes the progression of Prostate cancer. Curr Pharm Biotechnol. 2022;23:1658.
Liu Z, Zhong J, Zeng J, Duan X, Lu J, Sun X, et al. Characterization of the m6A-associated tumor immune microenvironment in prostate cancer to aid immunotherapy. Front Immunol. 2021;12: 735170.
Tao H, Li Z, Mei Y, Li X, Lou H, Dong L, et al. Integrative bioinformatics analysis of a prognostic index and immunotherapeutic targets in renal cell carcinoma. Int Immunopharmacol. 2020;87: 106832.
Zhu D, Wu ZH, Xu L, Yang DL. Single sample scoring of hepatocellular carcinoma: a study based on data mining. Int J ImmunoPathol Pharmacol. 2021;35:20587384211018388.
Meng J, Guan Y, Wang B, Chen L, Chen J, Zhang M, et al. Risk subtyping and prognostic assessment of prostate cancer based on consensus genes. Commun Biol. 2022;5(1):233.
Gao L, Meng J, Zhang Y, Gu J, Han Z, Wang X, et al. Development and validation of a six-RNA binding proteins prognostic signature and candidate drugs for prostate cancer. Genomics. 2020;112(6):4980–92.
Meng J, Lu X, Zhou Y, Zhang M, Gao L, Gao S, et al. Characterization of the prognostic values and response to immunotherapy/chemotherapy of Krüppel-like factors in Prostate cancer. J Cell Mol Med. 2020;24(10):5797–810.
Maldonato BJ, Russell DA, Totah RA. Human METTL7B is an alkyl thiol methyltransferase that metabolizes hydrogen sulfide and captopril. Sci Rep. 2021;11(1):4857.
Wang M, Yan J, Cao X, Hua P, Li Z. Hydrogen sulfide modulates epithelial-mesenchymal transition and angiogenesis in non-small cell lung cancer via HIF-1α activation. Biochem Pharmacol. 2020;172: 113775.
Chen X, Li C, Li Y, Wu S, Liu W, Lin T, et al. Characterization of METTL7B to evaluate TME and predict prognosis by integrative analysis of multi-omics data in glioma. Front Mol Biosci. 2021;8: 727481.
Redecke V, Wu R, Zhou J, Finkelstein D, Chaturvedi V, High AA, et al. Hematopoietic progenitor cell lines with myeloid and lymphoid potential. Nat Methods. 2013;10(8):795–803.
Zhao Y, Zhang C, Zhu Y, Ding X, Zhou Y, Lv H, et al. TREM1 fosters an immunosuppressive tumor microenvironment in papillary thyroid cancer. Endocr Relat Cancer. 2022;29(2):71–86.
Haque S, Morris JC. Transforming growth factor-β: a therapeutic target for cancer. Hum Vaccines Immunother. 2017;13(8):1741–50.
Meng J, Liu Y, Guan S, Fan S, Zhou J, Zhang M, et al. The establishment of immune infiltration based novel recurrence predicting nomogram in prostate cancer. Cancer Med. 2019;8(11):5202–13.
Acknowledgements
not applicable.
Funding
This research did not receive any specific grant from funding agencies in the public, commercial, or not-for-profit sectors.
Author information
Authors and Affiliations
Contributions
SZ: Conceptualization; Formal analysis; Writing—original draft; Writing—review & editing. JG: Data curation; Investigation; Writing—review & editing.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zong, S., Gao, J. Identifying the tumor immune microenvironment-associated prognostic genes for prostate cancer. Discov Onc 15, 42 (2024). https://doi.org/10.1007/s12672-023-00856-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s12672-023-00856-3