Abstract
Performance status (PS) is widely used as an assessment of general condition in patients before performing comprehensive genomic profiling (CGP). However, PS scoring is dependent on each physician, and there is no objective and universal indicator to identify appropriate patients for CGP. Overall, 263 patients were scored using the modified Glasgow prognostic score (mGPS) from 0 to 2 based on the combination of serum albumin and c-reactive protein (CRP): 0, albumin ≥ 3.5 g/dl and CRP ≤ 0.5 mg/dl; 1, albumin < 3.5 g/dl or CRP > 0.5 mg/dl; and 2, albumin < 3.5 g/dl and CRP > 0.5 mg/dl. Overall survival was compared between mGPS 0–1 and mGPS 2 groups. The prognosis of patients with PS 0–1 and mGPS 2 was also evaluated. Thirty-nine patients (14.8%) were mGPS 2. Patients with mGPS 2 had significant shorter survival (14.7 months vs 4.6 months, p < 0.01). Twenty-eight patients were PS 0–1 and mGPS 2, and their survival was also short (5.6 months). Evaluation of mGPS is a simple and useful method for identifying patients with adequate prognosis using CGP.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Next-generation sequencing (NGS) is a remarkable technology for detecting genomic alterations in DNA and RNA sequences [1], and enables the high burden of genomic analysis to be undertaken in a single sequence run at low cost [2]. As technology advances, comprehensive genomic profiling (CGP) has been widely used in oncology practice. The primary goal of patients undergoing CGP is to have rapid access to molecular targeted therapy. Patients must participate in clinical trials to gain access to these drugs because it is often not covered by insurance. However, drug accessibility in clinical practice is very low [3,4,5,6,7,8]. The most common reason for not receiving targeted therapy was that by the time the CGP results were explained to the patients, their performance status (PS) had already declined and they were considered to have a poor prognosis [6]. One of the reasons leading to this situation is the use of PS for the assessment of general condition at the time of performing CGP. PS is a powerful prognostic marker and is used by many physicians to identify appropriate patients for performing CGP in cancer care. However, PS is dependent on the individual physician’s judgement. Neeman et al. reported that physicians' PS ratings do not seem to predict important outcomes [9]. Additionally, blind enforcement of CGP has a negative impact on the healthcare economy [2]. Hence, objective indicators are needed when implementing CGP.
Inflammation-based scores, such as GPS or modified GPS (mGPS), have been reported as good predictive indicators in cancer patients. GPS is a complex indicator that uses C-reactive protein (CRP) and serum albumin. CRP is an acute phase protein whose expression is increased by interleukin-6 (IL-6) activity [10]. IL-6 represents the degree of inflammation in cancer tissue, and IL-6 elevation leads to increased CRP and decreased serum albumin. Thus, GPS is indirect indicator of cancer cachexia and an independent prognostic marker regardless of disease stage [11]. Because various types of cancer patients in various situations receive CGP, it is considered to be meaningful to evaluate GPS in these patients. Additionally, GPS is an objective indicator, unlike PS. Hence, we evaluated whether GPS is useful compared with PS in patients receiving CGP.
2 Methods
2.1 Patients
We retrospectively analyzed collected data from consecutive patients who underwent CGP using the FoundationOne CDx Cancer Genomic Profile (Cambridge, MA, USA) or the OncoGuide NCC oncopanel system (Tokyo, Japan) at the Cancer Institute Hospital of Japanese Foundation for Cancer Research (JFCR) between November 2019 and July 2021 [12, 13]. Patient data comprised age, sex, Eastern Cooperative Oncology Group PS (ECOG PS) [14], serum albumin, and CRP. Laboratory data were used within 2 weeks before or after CGP. In the present study, GPS or mGPS was determined according to best predictive value of serum CRP using receiver operating characteristic (ROC) curves [15]. On the basis of this analysis, the cut-off value for CRP was calculated to be 0.45 mg/dl (sensitivity 69.1%, specificity 59.8%). Hence, in this study, mGPS was used because of its closer cut-off value. mGPS was scored 0 to 2 on the basis of the combination of serum albumin and CRP: 0, albumin ≥ 3.5 g/dl and CRP ≤ 0.5 mg/dl; 1, albumin < 3.5 g/dl or CRP > 0.5 mg/dl; and 2, albumin < 3.5 g/dl and CRP > 0.5 mg/dl [15].
2.2 Statistical analysis
We used chi-squared test and Mann–Whitney U test to compare patients with mGPS 0–1 and mGPS 2. Overall survival (OS) was estimated using the Kaplan–Meier method and the log-rank test. Data were censored on 31 December 2021. Patients who were lost to follow-up were censored at the date of last contact or follow-up. OS was calculated from the date of performing CGP to the date of death from any cause. We performed univariate and multivariate analyses to estimate factors potentially prognostic for OS by calculating hazard ratios using the Cox proportional hazards model. The level of significance was set at p < 0.05 for univariate and multivariate analyses and was two-sided. All analyses were performed using EZR (www.r-project.org) [16].
2.3 Ethics approval and consent to participate
All procedures were performed in accordance with the ethical standards of the responsible committee on human experimentation (institutional and national) and with the Helsinki Declaration of 1964 and later versions. This observational study was approved by the institutional review board of the JFCR (approval number 2021-GA-1075). Patient consent for this study was in the form of an opt-out form.
3 Results
3.1 Patients’ characteristics
In total, 341 patients were included in this study. Of these, 78 patients were excluded because mGPS could not be scored due to a lack of laboratory data. The remaining 263 patients were enrolled in this study. The median age was 58 years (range, 18–84 years). The median observation time was 8.4 months (range, 0.1–24.7 months). At the time of censoring, 127 patients (48.2%) had died from the primary disease. The median survival time was 11.8 months (range, 0.1–22.2 months). The patients were classified into two groups according to mGPS score (mGPS 0–1 and mGPS 2). Thirty-nine patients (14.8%) were mGPS 2. The characteristics of the 263 patients are shown in Table 1. Patients with mGPS 2 were older and had worse PS. The median turnaround time was 1.4 months (range, 0.8–3.3 months). No patients underwent liquid-based CGP.
3.2 Short-term survival
Thirteen patients (4.9%) had already died before the CGP results were explained. Nine of the 13 patients were mGPS 2. It is noteworthy that approximately a quarter of patients in the mGPS 2 group had already died.
3.3 Long-term survival
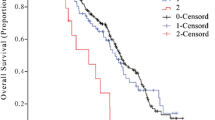
Figure 1 shows overall survival. Patients in the mGPS 2 group had significantly shorter survival (14.7 months vs 4.6 months, p < 0.01).
3.4 Risk factor of prognosis
Univariate and multivariate Cox proportional hazard analyses indicated that PS2–3 and mGPS 2 were significant poor prognostic factors (Table 2). PS and mGPS did not have multicollinearity (VIF: 1.0).
3.5 Correlation between PS and mGPS
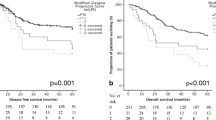
Two hundred thirty-five patients were PS 0–1. Of those, 28 patients (11.9%) were mGPS 2. Therefore, we classified patients into four groups to evaluate the utility of mGPS: a) PS 0–1 and mGPS 0–1; b) PS 0–1 and mGPS 2; c) PS 2–3 and mGPS 0–1; and d) PS 2–3 and mGPS 2. Figure 2 shows the overall survival of these four groups. Only patients with group a had good survival (median, 16.3 months). Other 3 groups had poor survival (median, b; 5.0 months, c; 4.9 months and d; 2.5 months). There was significant difference between group a and group c (p < 0.01). Particularly noteworthy is that group b had short survival, similar to group d (p = 0.16).
4 Discussion
This is the first study to evaluate overall survival after CGP using mGPS. Our study showed that patients with mGPS 2 had a poor prognosis. More than 10% of patients were judged as PS 0–1 by their physicians, but they were in fact mGPS 2. Notably, these patients’ prognoses were very short. These results indicate that PS is insufficient to identify appropriate patients with adequate prognosis for performing CGP. Physicians tend to assign a good PS score to their patients. According to Neeman et al., physicians frequently encountered their patients during their treatment span, and their assessment of patients may have been affected by their long-term familiarity [9]. In fact, it has been reported that the longer the relationship with the patient, the less accurate the prognostic prediction [17]. Another explanation is that physicians are hesitant to assign a low PS to a patient because the PS directly affects the treatment plan. Patients also attempt to hide their symptoms, such as chemotherapy adverse events, for fear of not being able to receive treatment. Patients who are undergoing CGP may have a long-term relationship with their physician. Hence, this is why mGPS is useful and is a good predictor of prognosis because of its objective nature.
There are several combined systematic inflammation and nutritional biomarkers associated with host–tumor interactions. GPS (mGPS) is a widely used classification that was proposed by McMillan and developed by Toyama[11, 15]. The amount of IL-6 in the circulating blood reflects the cancer state and when IL-6 is elevated, CRP is increased and albumin is decreased. Thus, it is logical that GPS is a surrogate of prognosis. Many reports focus on the relationship between GPS (mGPS) and prognosis [18,19,20,21,22,23]. Recently, the utility of the CRP–albumin ratio (CAR) has been reported [24,25,26]. Neutrophil–lymphocyte ratio (NLR) and platelet–lymphocyte ratio (PLR) are classical nutritional assessments using blood cell components [27]. Low nutritional status lowers lymphocyte counts and the systemic inflammatory response increases neutrophil and platelet counts. Many reports have investigated the relationship between NLR or PLR and prognosis [28,29,30,31,32]. However, the cut-off value is not fixed. Prognostic nutritional index (PNI) contains several indexes, among which Onodera’s PNI is widely used [33]. Some studies that have investigated the relationship between PNI and prognosis [34, 35] have reported that the cut-off value is not fixed. Thus, there are several biomarkers, but only GPS (mGPS) has a cut-off value. It is considered important for eliminating differences between physicians or facilities. Hence, GPS (mGPS) was used as an indicator in this study.
The turnaround time (TAT) of this study was 1.4 months. Considering the TAT, the prognosis of patients with GPS 2 is too short to enable participation in clinical trials. CGP should be considered before mGPS scores worsen. Several reports have investigated the advantages and disadvantages of CGP. An advantage is its potential to improve prognosis by leading to targeted therapy [36]. Targeted therapies have been reported to be cost-effective compared with standard care because CGP has the potential to reduce the use of inappropriate drugs [37,38,39]. Conversely, a disadvantage is its low contribution to treatment plans despite its high cost [40]. It is important to confer a benefit to patients. In any case, it would be best to conduct CGP at a time that would benefit the patient.
This study had several limitations. First, this was a single-institution retrospective study with potential selection bias and a short follow-up time. Second, treatment after CGP was not taken into consideration. Patients in the mGPS 0–1 group may have a better prognosis because of the continued treatment or because they are likely to have access to targeted therapies. Third, this cohort included subgroup which the subjective PS was better than mGPS in predicting survival. PS and mGPS were independent prognostic factors in this analysis. It is unclear whether PS or mGPS is a better indicator, however, mGPS would be at least meaningful complement when physicians assign a good PS.
In future work, it would be important to extend this analysis to show the relationship between the PS/mGPS combination and the CGP results since identification of pro-inflammatory mutations is of considerable interest to the scientific and medical community.
5 Conclusion
Our study demonstrated that evaluation of mGPS may help physicians to identify appropriate patients with an adequate prognosis to receive CGP.
Data availability
The datasets generated during and/or analyzed during the current study are available from the corresponding author on reasonable request.
References
Marino P, Touzani R, Perrier L, Rouleau E, Kossi DS, et al. Cost of cancer diagnosis using next-generation sequencing targeted gene panels in routine practice: a nationwide French study. Eur J Hum Genet. 2018;26:314–23. https://doi.org/10.1038/s41431-017-0081-3.
van Nimwegen KJ, van Soest RA, Veltman JA, Nelen MR, van der Wilt GJ, et al. Is the $1000 genome as near as we think? a cost analysis of next-generation sequencing. Clin Chem. 2016;62:1458–64. https://doi.org/10.1373/clinchem.2016.258632.
Singh AP, Shum E, Rajdev L, Cheng H, Goel S, et al. Impact and diagnostic gaps of comprehensive genomic profiling in real-world clinical practice. Cancers. 2020;12:1156. https://doi.org/10.3390/cancers12051156.
Sunami K, Ichikawa H, Kubo T, Kato M, Fujiwara Y, et al. Feasibility and utility of a panel testing for 114 cancer-associated genes in a clinical setting: a hospital-based study. Cancer Sci. 2019;110:1480–90. https://doi.org/10.1111/cas.13969.
Nesline MK, Depietro P, Dy GK, Early A, Papanicolau-Sengos A, et al. Oncologist uptake of comprehensive genomic profile guided targeted therapy. Oncotarget. 2019;10:4616–29. https://doi.org/10.18632/oncotarget.27047.
Hilal T, Nakazawa M, Hodskins J, Villano JL, Mathew A, et al. Comprehensive genomic profiling in routine clinical practice leads to a low rate of benefit from genotype-directed therapy. BMC Cancer. 2017;17:602. https://doi.org/10.1186/s12885-017-3587-8.
Hirshfield KM, Tolkunov D, Zhong H, Ali SM, Stein MN, et al. Clinical actionability of comprehensive genomic profiling for management of rare or refractory cancers. Oncologist. 2016;21:1315–25. https://doi.org/10.1634/theoncologist.2016-0049.
Schwaederle M, Daniels GA, Piccioni DE, Fanta PT, Schwab RB, et al. On the road to precision cancer medicine: analysis of genomic biomarker actionability in 439 patients. Mol Cancer Ther. 2015;14:1488–94. https://doi.org/10.1158/1535-7163.MCT-14-1061.
Neeman E, Gresham G, Ovasapians N, Hendifar A, Tuli R, et al. Comparing physician and nurse eastern cooperative oncology group performance status (ECOG-PS) ratings as predictors of clinical outcomes in patients with cancer. Oncologist. 2019;24:e1460-6. https://doi.org/10.1634/theoncologist.2018-0882.
Wakuda R, Miki C, Kusunoki M. Autoreactivity against interleukin 6 as a risk factor in elderly patients with colorectal carcinoma. Arch Surg. 2001;136:1274–9. https://doi.org/10.1001/archsurg.136.11.1274.
McMillan DC. Systemic inflammation, nutritional status and survival in patients with cancer. Curr Opin Clin Nutr Metab Care. 2009;12:223–6. https://doi.org/10.1097/MCO.0b013e32832a7902.
Ebi H, Bando H. Precision oncology and the universal health coverage system in Japan. JCO Precis Oncol. 2019;3:1–12. https://doi.org/10.1200/po.19.00291.
Chugai Pharmaceutical Co., Ltd. Chugai Launches Genomic Mutation Analysis Program, FoundationOne CDx Cancer Genomic Profile. 2019. https://www.chugai-pharm.co.jp/english/news/detail/20190603150001_625.html. Accessed 1 Apr 2022.
Oken MM, Creech RH, Tormey DC, Horton J, Davis TE, et al. Toxicity and response criteria of the eastern cooperative oncology group. Am J Clin Oncol. 1982;5:649–55.
Toiyama Y, Miki C, Inoue Y, Tanaka K, Mohri Y, et al. Evaluation of an inflammation-based prognostic score for the identification of patients requiring postoperative adjuvant chemotherapy for stage II colorectal cancer. Exp Ther Med. 2011;2:95–101. https://doi.org/10.3892/etm.2010.175.
Kanda Y. Investigation of the freely available easy-to-use software ‘EZR’ for medical statistics. Bone Marrow Transplant. 2013;48:452–8. https://doi.org/10.1038/bmt.2012.244.
Christakis NA, Lamont EB. Extent and determinants of error in doctors’ prognoses in terminally ill patients: prospective cohort study. BMJ. 2000;320:469–72. https://doi.org/10.1136/bmj.320.7233.469.
Ishizuka M, Kubota K, Kita J, Shimoda M, Kato M, et al. Impact of an inflammation-based prognostic system on patients undergoing surgery for hepatocellular carcinoma: a retrospective study of 398 Japanese patients. Am J Surg. 2012;203:101–6. https://doi.org/10.1016/j.amjsurg.2010.09.030.
Jamieson NB, Mohamed M, Oien KA, Foulis AK, Dickson EJ, et al. The relationship between tumor inflammatory cell infiltrate and outcome in patients with pancreatic ductal adenocarcinoma. Ann Surg Oncol. 2012;19:3581–90. https://doi.org/10.1245/s10434-012-2370-y.
La Torre M, Nigri G, Cavallini M, Mercantini P, Ziparo V, et al. The glasgow prognostic score as a predictor of survival in patients with potentially resectable pancreatic adenocarcinoma. Ann Surg Oncol. 2012;19:2917–23. https://doi.org/10.1245/s10434-012-2348-9.
Roxburgh CS, Platt JJ, Leitch EF, Kinsella J, Horgan PG, et al. Relationship between preoperative comorbidity, systemic inflammatory response, and survival in patients undergoing curative resection for colorectal cancer. Ann Surg Oncol. 2011;18:997–1005. https://doi.org/10.1245/s10434-010-1410-8.
Jiang X, Hiki N, Nunobe S, Kumagai K, Kubota T, et al. Prognostic importance of the inflammation-based Glasgow prognostic score in patients with gastric cancer. Br J Cancer. 2012;107:275–9. https://doi.org/10.1038/bjc.2012.262.
Dutta S, Crumley AB, Fullarton GM, Horgan PG, McMillan DC. Comparison of the prognostic value of tumour- and patient-related factors in patients undergoing potentially curative resection of oesophageal cancer. World J Surg. 2011;35:1861–6. https://doi.org/10.1007/s00268-011-1130-7.
Wu M, Guo J, Guo L, Zuo Q. The C-reactive protein/albumin ratio predicts overall survival of patients with advanced pancreatic cancer. Tumour Biol. 2016;37:12525–33. https://doi.org/10.1007/s13277-016-5122-y.
Ishizuka M, Nagata H, Takagi K, Iwasaki Y, Shibuya N, et al. Clinical significance of the C-reactive protein to albumin ratio for survival after surgery for colorectal cancer. Ann Surg Oncol. 2016;23:900–7. https://doi.org/10.1245/s10434-015-4948-7.
Liu X, Sun X, Liu J, Kong P, Chen S, et al. Preoperative C-reactive protein/albumin ratio predicts prognosis of patients after curative resection for gastric cancer. Transl Oncol. 2015;8:339–45. https://doi.org/10.1016/j.tranon.2015.06.006.
Miller CL. Immunological assays as measurements of nutritional status: a review. JPEN J Parenter Enteral Nutr. 1978;2:554–66. https://doi.org/10.1177/014860717800200406.
Wang DS, Luo HY, Qiu MZ, Wang ZQ, Zhang DS, et al. Comparison of the prognostic values of various inflammation based factors in patients with pancreatic cancer. Med Oncol. 2012;29:3092–100. https://doi.org/10.1007/s12032-012-0226-8.
Hung HY, Chen JS, Yeh CY, Changchien CR, Tang R, et al. Effect of preoperative neutrophil-lymphocyte ratio on the surgical outcomes of stage II colon cancer patients who do not receive adjuvant chemotherapy. Int J Colorectal Dis. 2011;26:1059–65. https://doi.org/10.1007/s00384-011-1192-x.
Shimada H, Takiguchi N, Kainuma O, Soda H, Ikeda A, et al. High preoperative neutrophil-lymphocyte ratio predicts poor survival in patients with gastric cancer. Gastric Cancer. 2010;13:170–6. https://doi.org/10.1007/s10120-010-0554-3.
Kinoshita A, Onoda H, Imai N, Iwaku A, Oishi M, et al. Comparison of the prognostic value of inflammation-based prognostic scores in patients with hepatocellular carcinoma. Br J Cancer. 2012;107:988–93. https://doi.org/10.1038/bjc.2012.354.
Kwon HC, Kim SH, Oh SY, Lee S, Lee JH, et al. Clinical significance of preoperative neutrophil-lymphocyte versus platelet-lymphocyte ratio in patients with operable colorectal cancer. Biomarkers. 2012;17:216–22. https://doi.org/10.3109/1354750X.2012.656705.
Onodera T, Goseki N, Kosaki G. Prognostic nutritional index in gastrointestinal surgery of malnourished cancer patients. Nihon Geka Gakkai Zasshi. 1984;85:1001–5.
Kanda M, Mizuno A, Tanaka C, Kobayashi D, Fujiwara M, et al. Nutritional predictors for postoperative short-term and long-term outcomes of patients with gastric cancer. Medicine. 2016;95: e3781. https://doi.org/10.1097/MD.0000000000003781.
Kanda M, Fujii T, Kodera Y, Nagai S, Takeda S, et al. Nutritional predictors of postoperative outcome in pancreatic cancer. Br J Surg. 2011;98:268–74. https://doi.org/10.1002/bjs.7305.
Sholl LM, Do K, Shivdasani P, Cerami E, Dubuc AM, Kuo FC, et al. Institutional implementation of clinical tumor profiling on an unselected cancer population. JCI Insight. 2016;1: e87062. https://doi.org/10.1172/jci.insight.87062.
Kasztura M, Richard A, Bempong N-E, Loncar D, Flahault A. Cost-effectiveness of precision medicine: a scoping review. Int J Public Health. 2019;64:1261–71. https://doi.org/10.1007/s00038-019-01298-x.
Berm EJJ, Looff MD, Wilffert B, Boersma C, Annemans L, et al. Economic evaluations of pharmacogenetic and pharmacogenomic screening tests: a systematic review second update of the literature. PLOS ONE. 2016;11:e0146262. https://doi.org/10.1371/journal.pone.0146262.
Akhmetov I, Bubnov RV. Assessing value of innovative molecular diagnostic tests in the concept of predictive, preventive, and personalized medicine. EPMA J. 2015. https://doi.org/10.1186/s13167-015-0041-3.
Chawla A, Janku F, Wheler JJ, Miller VA, Ryan J, et al. Estimated cost of anticancer therapy directed by comprehensive genomic profiling in a single-center study. JCO Precis Oncol. 2018. https://doi.org/10.1200/PO.18.00074.
Acknowledgements
We thank the medical staff of the Department of Genomic Medicine, The Cancer Institute Hospital of Japanese Foundation for Cancer Research, for their support during this study. We also thank H. Nikki March, PhD, from Edanz (https://jp.edanz.com/ac) for editing a draft of this manuscript.
Funding
This research received no specific grant from any funding agency in the public, commercial, or not-for-profit sectors.
Author information
Authors and Affiliations
Contributions
Conception and design: NH. Development of methodology: NH and AO. Data acquisition: NH, IP, MY, XW, and MF. Analysis and interpretation of data: NH. Draft writing, review, and/or revision of the manuscript: NH, IF, AO, and ST. Final approval of the manuscript: all authors. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interest
All the authors, except one, report that they have no conflict of interest to disclose. ST reports grants and personal fees from ONO Pharmaceutical Co., Ltd., Bristol-Myers Squibb, MSD, AstraZeneca, Chugai, and BAYER.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Hayashi, N., Fukada, I., Ohmoto, A. et al. Evaluation of an inflammation-based score for identification of appropriate patients for comprehensive genomic profiling. Discov Onc 13, 109 (2022). https://doi.org/10.1007/s12672-022-00574-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s12672-022-00574-2