Abstract
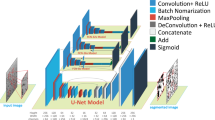
With the advancement of electron microscopy, industrial microscale objects are analyzed through image-based characterization. However, the automated and objective assessment of a vast number of images required for quality control is limited by the incomplete segmentation of individual objects in the image. In this study, the scanning electron microscope images of powder grains are selected as target images representing industrial microscale objects. A deep neural network based on the U-Net is developed and trained by manually labeled ground truth. Although the U-Net is a basic network originally devised for biomaterials, the network in this study achieves approximately 90% accuracy and outperforms conventional thresholding methods. However, the boundaries distinguishing individual are not completely classified. The inference results are further processed with morphological operations and watershed algorithms to quantitatively measure grain shapes. Discrepancies in shape parameters between ground truth and network prediction are also discussed.
Graphic Abstract











Similar content being viewed by others
References
Bradley D, Roth G (2007) Adaptive thresholding using the integral image. J Graph Tools 12:13–21. https://doi.org/10.1080/2151237X.2007.10129236
Csurka G, Larlus D, Perronnin F (2013) What is a good evaluation measure for semantic segmentation? In: Proceedings of the British Machine Vision Conference 2013. British Machine Vision Association, pp 32.1–32.11
Dumoulin V, Visin F (2016) A guide to convolution arithmetic for deep learning. Arxiv preprint arXiv: 1603.07285
Eppel S (2017) Hierarchical semantic segmentation using modular convolutional neural networks
Fabijańska A (2018) Segmentation of corneal endothelium images using a U-Net-based convolutional neural network. Artif Intell Med 88:1–13. https://doi.org/10.1016/j.artmed.2018.04.004
Foracchia M, Ruggeri A (2007) Corneal endothelium cell field analysis by means of interacting Bayesian shape models. In: 2007 29th Annual International Conference of the IEEE Engineering in Medicine and Biology Society. IEEE, pp 6035–6038
Gonzalez RC, Woods RE (Richard E, Eddins SL (2009) Digital Image Processing using MATLAB®. Gatesmark Publishing
Ji X, Vedaldi A, Henriques J (2019) Invariant Information Clustering for Unsupervised Image Classification and Segmentation. In: 2019 IEEE/CVF International Conference on Computer Vision (ICCV). IEEE, pp 9864–9873
Kingma DP, Ba J (2014) Adam: A Method for Stochastic Optimization
Kraus OZ, Ba JL, Frey BJ (2016) Classifying and segmenting microscopy images with deep multiple instance learning. Bioinformatics 32:i52–i59. https://doi.org/10.1093/bioinformatics/btw252
Krizhevsky A, Sutskever I, Hinton GE (2012) ImageNet classification with deep convolutional neural networks. In: Proceedings of the 25th International Conference on Neural Information Processing Systems. pp 1907–1105
Lateef F, Ruichek Y (2019) Survey on semantic segmentation using deep learning techniques. Neurocomputing 338:321–348. https://doi.org/10.1016/j.neucom.2019.02.003
Lee J, Kim H, Cho H et al (2019) Deep-learning-based label-free segmentation of cell nuclei in time-lapse refractive index tomograms. IEEE Access 7:83449–83460. https://doi.org/10.1109/ACCESS.2019.2924255
Liang Z, Nie Z, An A et al (2019) A particle shape extraction and evaluation method using a deep convolutional neural network and digital image processing. Powder Technol 353:156–170. https://doi.org/10.1016/j.powtec.2019.05.025
Long J, Shelhamer E, Darrell T (2015) Fully convolutional networks for semantic segmentation. In: 2015 IEEE Conference on Computer Vision and Pattern Recognition (CVPR). IEEE, pp 3431–3440
Maurer CR, Qi R, Raghavan V (2003) A linear time algorithm for computing exact Euclidean distance transforms of binary images in arbitrary dimensions. IEEE Trans Pattern Anal Mach Intell 25:265–270. https://doi.org/10.1109/TPAMI.2003.1177156
Odena A, Dumoulin V, Olah C (2016) Deconvolution and Checkerboard Artifacts. Distill. https://doi.org/10.23915/distill.00003
Oktay AB, Gurses A (2019) Automatic Detection, Localization and Segmentation of Nano-Particles with Deep Learning in Microscopy Images. Micron. doi: https://doi.org/10.1016/j.micron.2019.02.009
Olson E (2011) Particle shape factors and their use in image analysis part 1: Theory. J GXP Compliance 15:85–96
Otsu N (1979) A Threshold Selection Method from Gray-Level Histograms. IEEE Trans Syst Man Cybern 9:62–66. https://doi.org/10.1109/TSMC.1979.4310076
Raj PM, Cannon WR (1999) 2-D particle shape averaging and comparison using Fourier descriptors. Powder Technol 104:180–189. https://doi.org/10.1016/S0032-5910(99)00046-7
Ronneberger O, Fischer P, Brox T (2015) U-Net: Convolutional Networks for Biomedical Image Segmentation. In: Lecture Notes in Computer Science (including subseries Lecture Notes in Artificial Intelligence and Lecture Notes in Bioinformatics). Springer Verlag, pp 234–241
Sanchez-Marin FJ (1999) Automatic segmentation of contours of corneal cells. Comput Biol Med 29:243–258. https://doi.org/10.1016/S0010-4825(99)00010-4
Yuan W, Chin KS, Hua M et al (2016) Shape classification of wear particles by image boundary analysis using machine learning algorithms. Mech Syst Signal Process 72–73:346–358. https://doi.org/10.1016/j.ymssp.2015.10.013
Zheng J, Hryciw RD (2018) Identification and characterization of particle shapes from images of sand assemblies using pattern recognition. J Comput Civ Eng 32:04018016
Zheng J, Hryciw RD (2016) Segmentation of contacting soil particles in images by modified watershed analysis. Comput Geotech 73:142–152. https://doi.org/10.1016/j.compgeo.2015.11.025
Acknowledgement
This work was supported by the National Research Foundation of Korea (NRF) grant funded by the Korea government (MSIT) (No. 2020R1A5A8018822).
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Appendix
Appendix
This section briefly explains the shape indices discussed in this study. A more detailed explanation of these descriptors and additional indices are found in the work of Olson (Olson 2011).
The AR of an object is defined as the ratio of the maximum Feret diameter (Fmax) to the minimum Feret diameter (Fmin). The Feret diameter is the size of an object along a specified orientation. The longest distance between any two parallel tangent lines of the object boundary is Fmax; conversely, the shortest distance between any two parallel tangent lines on the object is Fmin.
The circularity (C) of an object represents the similarity of the object to a circle and is calculated as follows:
where A and P are the area and perimeter of the particle, respectively.
The convex hull of an object can be explained as the shape of the elastic and tense bounding string of an object. The perimeter (PC) and area (AC) of the convex hull are distinguished from those of the original object by subscript C.
The convexity (HP) of an object is defined as the ratio of PC to P. The maximum value of HP is unity and is achieved when the shape of the object is the same as that of the convex hull. From unity, HP decreases as the shape deviates from the convex hull and becomes more complex. The solidity (HA) of an object is an area-based measure of the extent that the shape of the object deviates from its convex hull; HA is defined as A divided by AC.
Rights and permissions
About this article
Cite this article
Kwon, D., Yeom, E. Shape evaluation of highly overlapped powder grains using U-Net-based deep learning segmentation network. J Vis 24, 931–942 (2021). https://doi.org/10.1007/s12650-021-00748-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12650-021-00748-0