Abstract
Competition for resources is a common microbial interaction in the gut microbiome. Inulin is a well-studied prebiotic dietary fiber that profoundly shapes gut microbiome composition. Several community members and some probiotics, such as Lacticaseibacillus paracasei, deploy multiple molecular strategies to access fructans. In this work, we screened bacterial interactions during inulin utilization in representative gut microbes. Unidirectional and bidirectional assays were used to evaluate the effects of microbial interactions and global proteomic changes on inulin utilization. Unidirectional assays showed the total or partial consumption of inulin by many gut microbes. Partial consumption was associated with cross-feeding of fructose or short oligosaccharides. However, bidirectional assays showed strong competition from L. paracasei M38 against other gut microbes, reducing the growth and quantity of proteins found in the latter. L. paracasei dominated and outcompeted other inulin utilizers, such as Ligilactobacillus ruminis PT16, Bifidobacterium longum PT4, and Bacteroides fragilis HM714. The importance of strain-specific characteristics of L. paracasei, such as its high fitness for inulin consumption, allows it to be favored for bacterial competence. Proteomic studies indicated an increase in inulin-degrading enzymes in co-cultures, such as β-fructosidase, 6-phosphofructokinase, the PTS D-fructose system, and ABC transporters. These results reveal that intestinal metabolic interactions are strain-dependent and might result in cross-feeding or competition depending on total or partial consumption of inulin. Partial degradation of inulin by certain bacteria favors coexistence. However, when L. paracasei M38 totally degrades the fiber, this does not happen. The synergy of this prebiotic with L. paracasei M38 could determine the predominance in the host as a potential probiotic.
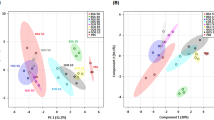
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The gut microbiome comprises the collective genome of microbes that inhabit the gut, including bacteria, archaea, viruses, and fungi [1]. These microorganisms can provide nutrients and vitamins to the host and protect against colonization by pathogenic microorganisms [2]. The dietary intake of specific non-digestible carbohydrates is increasingly seen as a highly effective approach to manipulating the composition and activities of the human gut microbiota to benefit health [3]. Dietary fibers are complex carbohydrate polymers found in fruits, vegetables, legumes, seeds, and cereals, which endogenous human enzymes cannot hydrolyze. However, the intestinal microbiome can selectively metabolize them through anaerobic fermentation [4, 5].
Some dietary fibers may act as prebiotics, enhancing the proliferation of beneficial microbes in the gut and host health [6]. Its consumption is associated with antidiabetic and antihypertensive properties [7]. The most common prebiotics contained in foods are fructooligosaccharides (FOS) and inulin [6]. Inulin is a fructan polysaccharide (polymer of fructose chains) linked by β-2,1 bonds (between 2 and 60 units) with glucose at its end [8, 9]. Inulin stimulates the growth of bifidobacteria and lactobacilli, which are beneficial for health [10]. It is found in roots and vegetables such as onions, artichokes, and chicory and can be fermented by several bacterial genera, such as Lactobacillus [11], Bifidobacterium [12, 13], and Bacteroides [14]. Multiple studies have shown the benefits of inulin consumption in stimulating the growth of health-promoting species that produce short-chain fatty acids (SCFAs) [15]. Inulin promotes increased intestinal calcium absorption, colonic pH regulation, gastrointestinal transit [16], improves blood lipid profiles, relieving constipation [17, 18] and protecting intestinal barrier function by restoring the microbiome [15].
Enzymes that metabolize inulin belong to the GH32 and GH91 glycosyl hydrolase families. These families include enzymes, such as inulinase, invertase, and levanase. Inulinases act precisely on the β-2,1-linkages of inulin, producing fructose and FOS [19]. These enzymes are classified as exo and endoinulinases [20]. Exoinulinase (fructan β-fructosidase) degrades inulin from its non-reducing terminal end to release the fructose units. Lacticaseibacillus paracasei has an inulin gene cluster that includes fructose-PTS system proteins (FosA, FosB, FosC, and FosD) and an extracellular β-fructosidase (FosE) [21]. In contrast, endoinulinase disrupts the internal bonds of inulin to produce smaller FOS [22, 23]. Intracellular β-fructofuranosidases capable of fermenting inulin have been found in L. paracasei BGP1 [24], Bifidobacterium adolescentis [19], and Bifidobacterium longum [25]. When inulin is degraded into FOS, it is transported by the ATP-binding cassette (ABC transport) identified in B. longum NCC2705. The fructan metabolic pathway derives fructose from the bifid shunt pathway in these bacteria [26]. This pathway converts these monosaccharides into fructose-6-phosphate, intermediates of the hexose fermentation pathway. In Lactobacillus, the pathway proposed by Buntin et al., (2017) [27] for inulin metabolization includes the degradation by β-fructosidase and the entry of FOS (by ABC transporter) or fructose (by PTS transporter) to the cell. Fructose is then catabolized by 1-phosphofructokinase and 6-phosphofructokinase to synthesize β-D-fructose-1,6-bisphosphate. The latter can also be synthesized from FOS by sucrose-6-phosphate hydrolase, fructokinase, and 1-phosphofructokinase. Fructose-bisphosphate aldolase class II converts β-D-fructose-1,6-bisphosphate to glyceraldehyde-3-phosphate, a compound metabolized in glycolysis.
Bacterial interactions are fundamental in shaping the gut microbiome. They can occur between microorganisms of the same species or between different species, genera, families, and domains [1, 28]. Interactions can be positive (cooperation) or negative (competition) [29]. In this context, microbial life strategies can influence the outcomes of interactions [30] and determine various consequences for microbial fitness, population dynamics, and functional capabilities within the microbiome [31]. However, competition is estimated to be prevalent among many bacterial species, but few cooperative interactions [32, 33]. Competition can be passive or active. During passive competition, strain can negatively affect the other through resource competition. In active competition (for resources or space), strains inhibit and kill each other through direct interference [34], bacteriocins, or the production of toxic waste products [35]. Species that compete for similar resources produce antimicrobial compounds for adaptive advantages [36]. For example, Escherichia coli K-12, Lactobacillus johnsonii NCC533, and B. longum NCC2705 [37] in the gut of gnotobiotic mice showed that the addition of E. coli Nissle 1917 led to the elimination of L. johnsonii and E. coli K-12, whereas B. longum only decreased its population. On the other hand, Bifidobacterium animalis BB04 produces the bacteriocin bifidocin A and acts against E. coli, Listeria monocytogenes, and Staphylococcus aureus [38].
Competition is a phenomenon that can help to understand how bacteria adapt to adverse conditions. The combined synergistic effect of inulin and potential probiotics needs further investigation to understand the impact of dietary fiber on gut bacteria. It is essential to unveil bacteria-bacteria interactions [3, 39]. In this study, we used proteomics on inulin bidirectional cultures to determine the resource competence interactions between L. paracasei M38 and other bacteria of different phyla (Ligilactobacillus ruminis PT16, B. longum PT4, and Bacteroides fragilis HM714). Proteomics is a robust platform with great potential for studying antagonistic mechanisms between bacteria, such as pathogen inhibition by Lactobacillus [40]. Therefore, studying gut microbiome commensal bacteria used in this investigation could reveal different affinities for inulin metabolism. This knowledge could contribute to a better understanding of the competitive mechanisms of paired bacterial interactions at the gut level in the presence of this dietary fiber.
Materials and Methods
Strains and Culture Medium
Among the 16 strains used in this work (Supplementary file Table S1) were B. fragilis HM714 (Bf, BEI Resources) and strains isolated from fecal samples of Chilean young adults such as L. paracasei M38 (Lp), L. ruminis PT16 (Lr), B. longum PT4 (Bl1), B. longum PT8 (Bl2), and B. longum PT33 (Bl3) [41]. The base culture medium used in this work was modified ZMB (mZMB) [42], which is a complex medium of known composition (minerals, ions, and vitamins) and is adequate for the growth of anaerobic bacteria. All the substrates used in modified mZMB, such as L-cysteine (cys; Sigma-Aldrich, St. Louis, MO, USA), inulin (Piping Rock, Ronkonkoma, NY, USA), or lactose (Sigma-Aldrich), were sterilized by filtration through 0.22 µm filters (Jet Biofil, China). Clostridium-reinforced medium (RCM; Becton, Dickinson, Franklin Lakes, NJ, USA) and Man-Rogosa-Sharpe (MRS; Difco Laboratories, Detroit, MI, USA) were autoclaved for 15 min at 121 °C. Bacterial growths in an anaerobic jar (Anaerocult; Merck, Darmstadt, Germany) with anaerobic packs (Gaspak EM; Becton, Dickinson, Franklin Lakes, NJ, USA) were reactivated from a -80 °C stock in a 1 mL (inoculum 8% v/v) of its respective culture medium (Supplementary file Table S1), for 48 h at 37 °C. Next, cultures were centrifuged (5000 × g), washed with pre-reduced mZMB medium, and the pellets were dissolved in mZMB medium supplemented with 1% (w/v) inulin (mZMB IN) or 2% (w/v) lactose (mZMB LAC) as a positive control, as appropriate.
Genomic Search of Inulin-Degrading Enzymes
Microbial strains were sequenced by MicrobesNG (Birmingham, UK) using Illumina MiSeq. B. longum PT4 (JAPJDT000000000) and B. longum PT8 (JARCPQ000000000) genome sequences were deposited at NCBI. L. paracasei M38 genome (PRJNA861286) was previously deposited by Torres-Miranda et al. (2022) [21]. B. fragilis HM714 genome was obtained from the Integrated Microbial Genome Database (IMG) [43]. Finally, we individually searched for enzymes of interest related to inulin degradation by Uniprot or literature [27, 44, 45]. Specifically, we searched for enzymes linked to the catabolism of inulin, FOS, and fructose and their transports (ABC and PTS transporters). These include β-fructofuranosidase and 6-phosphofructokinase, enzymes involved in the degradation of fructans.
Monoculture Growths
Each strain was reactivated and inoculated (8% v/v) following the methodology described by Hirmas et al. (2022) [46], with some modifications. Assays were performed in triplicate to evaluate the consumption of mZMB IN or mZMB LAC. Growth kinetics were performed in 96-well plates covered with a mineral oil layer, and the strains were cultured for 48 h at 37 °C in an anaerobic chamber (Sheldon Manufacturing INC, Bactronez-2 Anaerobic Chamber Workstation, Cornelius, OR, USA). The optical density (OD) at 600 nm was measured in a Tecan F50 spectrophotometer (Tecan Trading AG, Infinite F50, Männedorf, Switzerland) every 30 min with shaking every 5 s before measuring.
Unidirectional Growths
Unidirectional cultures (using a bacterial supernatant of 24 h for culturing another bacterium) were performed as described above with some modifications. Primary degrader strains (Lp, Bl1, Bl2, Bl3, and Bf) were cultured in mZMB IN under anaerobic conditions for 24 h at 37 °C in 5 mL tubes (8% v/v). The culture was then centrifuged at 10,000 × g for 1 min, and the supernatant was sterilized using a 0.22 μm filter. Each bacterium (secondary degrader) grew in the supernatant of the primary degrader for 48 h, as previously indicated. Growths were performed in triplicates and represented as ΔOD = ODfinal – ODinitial.
Bidirectional Growths
Bidirectional assays correspond to two bacteria grown simultaneously and separated by a membrane. The strains selected for this assay were those that best degraded inulin in monoculture. The considered pairs were (insert/well): Lr/Lp; Bl1/Lp; Bf/Lp. Both monocultures and bicultures were analyzed. The bacterium with the best growth on inulin (Lp) was plated in the bottom well in 1 mL of mZMB IN. The other bacteria of the evaluated pair (Lr, Bl1, Bf) were grown in the upper insert, which contained 250 μL of mZMB IN. The procedure was performed as described by Hirmas et al. (2022) [46]. Briefly, bacteria were first reactivated in RCM (Bf) or MRS (Lp, Lr, and Bl1) for 48 h, and centrifuged at 10,000 × g for 1 min, then washed with the mZMB (without carbon source). Bacteria were then inoculated at 8% v/v in mZMB IN or mZMB LAC (basal treatment), as appropriate, onto Transwell plates (JetBiofil, China). The plates were incubated in an anaerobic jar using anaerobic packs at 37 °C for 48 h. At the end of the experiment, the contents of the Transwell plates were transferred into a 96-well plate, and OD 600 (at 0 h and 48 h) was measured using a Tecan Infinite M200 Pro plate reader (Tecan Trading AG, Grödig, Austria). Finally, the culture was centrifuged at 10,000 × g for 1 min, and the supernatant and precipitate were stored at − 80 °C until further use.
Carbohydrate Consumption
Thin-layer chromatography (TLC) was performed as described by Hirmas et al. (2022) [46]. TLC was performed on F-60 silica plates (Merck, Germany) using 1-butanol/ethanol/ethanol/water 10:8:5 v/v as a run buffer and 1% orcinol in 10% H2SO4 in ethanol as the developer reagent. Two μL were taken from each sample. The chromatogram was developed after two runs and the sample was allowed to dry. The silica gel was heated at 100 °C until the bands were visually detectable. The carbohydrate consumption was evaluated in unidirectional (Lp, Bl1, Bl2, Bl3, and Bf supernatants) and bidirectional assays (“Lr vs. Lp,” “Bl1 vs. Lp,” and “Bf vs. Lp” supernatants).
SCFA Quantification
SCFAs were quantified in selected supernatants from bidirectional and unidirectional assays at the end of the experiment (48 h) using a Lachrom liquid chromatograph (Merck-Hitachi) equipped with a UV detector at 210 nm. The Aminex HPX-87H ion exclusion column (300 mm, 7.8 mm; Bio-Rad) was operated with five mM H2SO4 at a flow rate of 0.45 mL/min at 35 °C for 35 min. Acetic, butyric, lactic, propionic, and succinic acid standards of known concentrations were used for column calibration (Sigma-Aldrich, St. Louis, MO, USA). Thirty microliters of the sample were injected and ran in duplicate. Data analysis was performed using Multi-HSM Manager software (Hitachi).
Label-Free Comparative Proteomics
Pellets were obtained from bidirectional bacterial assays, and both monocultures and bicultures were analyzed (n = 4 biological replicates). Samples were lyophilized from 1.5 mL tubes in a 2.5 L lyophilizer at a temperature of − 50 °C (Labconco, USA) and stored at − 80 °C until further use. Extraction and proteomic analyses were performed following the methodology described by Caballero et al. (2022) [47]. Data were obtained from a Top15 method for MS/MS scans [48]. The label-free quantitative (LFQ) algorithm was used to normalize spectral intensities and calculate relative protein abundance [49], using MaxQuant software (v.1.6.15.9; https://www.maxquant.org/) [50]. Carbamidomethylation of cysteines was set as a fixed modification, whereas methionine oxidation and N-terminal acetylation were set as variable modifications. Maximum peptide/protein false discovery rates (FDR) were set at 1 % as the maximum compared to a reverse database. Perseus software (v.1.6.14.0) was applied for data organization and statistical analysis [51]. A t-test for quantitative analysis was used to compare the different batches with the control batch. Statistical differences were set at p < 0.05. A protein database of Lp, Lr, Bl1, and Bf from Uniprot (https://www.uniprot.org/) was used to perform the search. Qualitative analysis was performed by detecting proteins in at least two replicates from the same batch but none from the compared batch. ClueGO software [52] was used for gene ontology enrichment analysis [53]. To define term-term interrelationships and functional groups based on shared genes between terms, the Kappa score was set to 0.4. A minimum of three GO terms and 4% of covered genes were set to be selected. The p value was corrected using the Bonferroni downward step and was established as p ≤ 0.05 [52]. Fold change with respect to lactose was expressed as ΔlcLog2. When the protein was only found in one condition, the label-free quantitative (LFQ) intensity was expressed as Log2. Heatmaps containing those discriminant proteins with biological relevance were elaborated in R studio 4.2.2.
Statistical Analysis
Multiple comparison ANOVA was performed for studies in SCFAs (2-factor ANOVA, Tukey’s test). In unidirectional assays, bacterial SCFAs in the supernatant were compared with the basal state, and SCFAs in bidirectional assays were compared biculture with monoculture. OD 600 of bidirectional assays (1-factor ANOVA) was obtained using GraphPad Prism 7 software. Statistical significance was set at p ≤ 0.05.
Results
Monoculture Assays and Enzyme Search
Figure 1 shows the growth of different gut microbes included in this work (Supplementary file Table S1), using inulin as the sole carbon source. All bacteria grew on this substrate with different vigorousness except Bifidobacterium breve I1 (Bb1), Bifidobacterium bifidum JCM-1254 (Bb2), B. adolescentis D3 (Ba), and B. longum SC664 (Bl5). The strains that grew best were Lp (ΔOD = 0.89), followed by Bf (ΔOD = 0.60), Bl1 (ΔOD = 0.55), Bl2 (ΔOD = 0.45), and Bl3 (ΔOD = 0.33). The results showed that inulin was widely consumed by different strains but differed in the degree of utilization according to their growth.
Bacterial growth curves of strains grown on inulin. IN: mZMB supplemented with inulin. LAC: mZMB supplemented with lactose. A The bacteria Phocaeicola dorei 5_1_36/D4 (Pd), Bacteroides thetaiotaomicron VPI-5482 (Bt1), and B. thetaiotaomicron HM23 (Bt2). B The bacteria B. fragilis HM714 (Bf), Bacteroides ovatus HM222 (Bo), and Phocaeicola vulgatus S1 (Pv). C The bacteria L. paracasei M38 (Lp), L. ruminis PT16 (Lr), and B. longum PT4 (Bl1). D The bacteria B. longum PT8 (Bl2), B. longum PT33 (Bl3), and B. longum PT7 (Bl4)
Table 1 shows the enzymes found in the bacteria that presented the highest growth on inulin (Lp, Bf, Bl1, and Bl2), which could be related to the fermentation of this dietary fiber. All the bacteria analyzed contained ABC transporters or PTS systems (necessary to transport the released sugars). Lp, Bl1, and Bl2 encoded sugar metabolism enzymes like sucrose-6-phosphate hydrolase and phosphofructokinase. Lp and Bf had a β-fructofuranosidase in their genome, an enzyme essential in inulin catabolism. The details of the enzymes found in the Lp genome were analyzed by Torres-Miranda et al. [21].
Unidirectional Assays
The supernatants used as initial substrates were from the strains that grew best on inulin (Lp > Bf > Bl1 > Bl2 > Bl3). Table 2 summarizes the growth observed. Lp showed the highest growth in all spent supernatants (ΔOD of 0.99, 0.55, 0.76, 0.81 in Bl1, Bl2, Bl3, and Bf supernatants, respectively). On the other hand, the supernatant of Lp allowed little growth of the other bacteria. The bacteria that grew in the Bl1 supernatant were Bacteroides strains. In the supernatant of Bl2 and Bl3, this behavior was also observed. Finally, in the supernatant of Bf (Table 2), there was not much bacterial growth (except Lp).
The TLC results of supernatants from unidirectional assays (Fig. 2) showed consumption mainly of monosaccharides or oligosaccharides derived from inulin. However, Lp was an exception, as it always consumed all fractions of inulin in the supernatants evaluated (Bl1, Bl2, Bl3, and Bf supernatants). In the supernatant of the Lp (Fig. 2A) other bacteria had no consumption, except for Bl1 (Fig. 2B), which consumed fructose. In the other supernatants, inulin metabolization preferences and partial degradation by Bacteroides and Bifidobacterium were observed in relation to oligofructose size. Specifically, the evaluated strains partially consumed small oligosaccharides in the Bl2 supernatant (Fig. 2A, B) and Bf supernatant (Fig. 2A–C). In the supernatant of Bl3, Bf consumed approximately half of the inulin (Fig. 2C), whereas Bl1 only consumed fructose (Fig. 2B). Fructose consumption was mainly observed in the Bl1 supernatant (Fig. 2D). On the other hand, among the SCFAs measured in unidirectional assays, acetate and lactate production predominated (Supplementary file Fig. S1).
TLC of strains of interest using Lp (A, B), Bl1 (D), Bl2 (A, B), Bl3 (B, C), Bf (A, B, C) supernatants previously grown for 24 h on mZMB inulin 1%. SUP: Supernatant. C-: Initial supernatant (negative control). C- IN: mZMB inulin without bacterial growth. The red box indicates the fractions of inulin consumed
Bidirectional Assays
Later, we evaluated how a bacterium with high inulin consumption capacity (Lp) can impact the growth of other bacteria of different genera (Lactobacillus, Bifidobacterium, and Bacteroides) that also degraded inulin. Three bidirectional interactions were analyzed (Lr/Lp; Bl1/Lp; Bf/Lp). In the “Lr vs. Lp” growth (Fig. 3A), Lr decreased by 64%, and Lp increased by 30%, both in bicultures. A similar trend was observed in both “Bl1 vs. Lp” (Fig. 3B) and “Bf vs. Lp” (Fig. 3C) when grown in bicultures (with respect to monoculture), where Bl1 and Bf decreased (reduced by 79% and 61%, respectively), and Lp increased its growth (174%, and 31%, respectively). These results indicate that Lp resource competition dominated the three bacterial interactions evaluated.
Bidirectional bacterial growth on inulin after 48 h. A Bidirectional assay of Lr and Lp. B Bidirectional assay of Bl1 and Lp. C Bidirectional assay of Bf and Lp. BID: Strain grown in a bidirectional assay, C+: Strain grew in monoculture. Lr: L. ruminis PT16. Bl1: B. longum PT4. Bf: B. fragilis HM714. Lp: L. paracasei M38. The monoculture was compared with the biculture in each strain. ANOVA statistical analysis was performed. **p < 0.01, ***p < 0.001, ****p < 0.0001
Figure 4A shows the inulin consumption in the three metabolic interactions evaluated. Lr, Bl1, and Bf showed the same trend (Fig. 4A), where monocultures partially metabolized inulin at 24 h, and degradation was almost complete at 48 h. However, in the presence of Lp, inulin was almost completely utilized at 24 h because there was only an inulin remnant at the base of the TLC. And at 48 h, the inulin consumption was complete. As for Lp (Fig. 4B), only one oligosaccharide stayed at 24 h in the monoculture. At 48 h, the total substrate used was. In the bicultures of Lp with different bacteria, the total catabolism of inulin was observed (both at 24 h and 48 h).
TLC of bidirectional assays of interest. A TLC of insert supernatants of Lr, Bl1, and Bf at 24–48 h. B TLC of well supernatants of Lp at 24–48 h. C+: Monoculture assay. BID, bidirectional assay; C- mZMB IN, mZMB inulin without bacterial growth; Bi1, biculture, in the presence of Lr; Bi2, biculture, in the presence of Bl1; Bi3, biculture, in the presence of Bf; Lr, L. ruminis PT16; Bl1, B. longum PT4; Bf, B. fragilis HM714; Lp, L. paracasei M38
Figure 5 shows the SCFAs produced in bidirectional assays, where the SCFAs were compared with their monocultures. These results indicated the absence of butyrate. Propionate and succinate were mainly detected when Bf was grown in a monoculture (Fig. 5C) but not in biculture. On the other hand, less acetate was detected in Lp supernatants in the presence of the three bacteria evaluated, since in monoculture it produced 33.7 mmol/L, and in biculture, it produced 20.6 mmol/L (in presence of Lr), 18.5 mmol/L (in presence of Bl1), and 26.9 mmol/L (in presence of Bf). In addition, acetate also declined in supernatants of other bacteria in bicultures, specifically decreased in Lr 20.5% (Fig. 5A), in Bl1 64.2% (Fig. 5B), and in Bf 45.4% (Fig. 5C). Finally, the amount of lactate remained little changed in the supernatant of Lp, but its concentration increased in the bicultures of other bacteria evaluated (Lr, Bl1, and Bf). However, this was probably due to the diffusion of this SCFA through the membrane, which corresponds to the lactate produced in high amounts by Lp.
Quantification of short-chain fatty acids in bidirectional assays. ANOVA statistical analysis was performed. **p < 0.01, ***p < 0.001, ****p < 0.0001. IN, mZMB supplemented with inulin; C+, monoculture assay; BID, bidirectional assay; Bi1, biculture, in the presence of Lr; Bi2, biculture, in the presence of Bl1; Bi3, biculture, in the presence of Bf; Bi4, biculture, in the presence of Lp; Lr, L. ruminis PT16; Bl1, B. longum PT4; Bf, B. fragilis HM714; Lp, L. paracasei M38
Label-Free Comparative Proteomics in Inulin
Proteomics used in this study was based on comparing cultures on mZMB IN with the basal state (mZMB LAC). Table 3 shows the four bacteria that were evaluated under different conditions. L. paracasei M38 (monoculture and bicultures in bidirectional assays, in the presence of Lr, Bl1, and Bf). In addition, proteomes from monocultures and bicultures in bidirectional assays (in the presence of Lp) were analyzed in L. ruminis PT16, B. longum PT4, and B. fragilis HM714 strains.
The proteomic assay showed that the diversity of metabolic pathways was higher in Lp monoculture (Supplementary file Fig. S2A) compared with bicultures. Specifically, this could be related to the increase in its OD shown in previous experiments. In the presence of Bf (Supplementary file Fig. S2D), Lp increased pathways suggesting an accelerated sugar consumption (carbohydrate derivate metabolic process and carbohydrate transport). This correlates with the increment in OD in co-culture. In general, bacteria competing with Lp decreased the number of metabolic pathways (Supplementary file Fig. S2). Carbohydrate metabolism from bacteria competing with Lp was negatively affected, which correlates with the low growth observed for these bacteria (Lr, Bl1, and Bf) in previous experiments (Fig. 3).
Figure 6 shows the number of identified proteins that increased or decreased in quantity when comparing proteomes of bacteria grown in inulin to growth in lactose. Lp in biculture displayed a slight increase of total proteins in inulin (147 in the presence of Lr, 124 in the presence of Bl1, and 193 in the presence of Bf), with respect to monoculture (117). The greatest increment in these proteins was 65% when Lp grew in the presence of Bf. In contrast, bacterial strains in the presence of Lp decreased the proteins higher relative quantity in inulin concerning monoculture. Specifically, biculture proteins from Lr (106), Bl1 (117), or Bf (139), with respect to their monocultures (200, 119, 332, respectively). Bf was the bacterium that reduced most of the total protein in inulin (13%) in bicultures.
Heat map (hieratical clustering) based in the number of proteins in different conditions found in bacterial interactions when compared with the basal treatment, lactose. The x-axis labels indicate those conditions. For proteins found in higher or lower abundance, p value < 0.05. For proteins only found in one treatment, they were found in at least two biological replicates and not found in any of the replicates of the counterpart. Mo, monoculture; Bi1, biculture, in the presence of L. ruminis PT16; Bi2, biculture, in the presence of B. longum PT4; Bi3, biculture, in the presence of B. fragilis HM714; Bi4, biculture, in the presence of L. paracasei M38
Proteins Found in L. paracasei M38, When Grown on Inulin
In the three bicultures, Lp increased the proteins found in higher quantities when inulin was consumed, with respect to the monoculture (Fig. 7). Generally, these proteins were mainly related to sugar metabolism, such as glycosyltransferases or ABC transporters that can transport FOS into the cellular interior, and 6-phosphofructokinase (ΔlcLog2 0.19 in PI, ΔlcLog2 0.40 in PRI, ΔlcLog2 0.30 in PLI, and ΔlcLog2 0.57 in PFI), which participates in the phosphorylation processes of inulin degradation intermediates. A phosphotransferase system for fructose (ΔlcLog2 4.46 in PI, ΔlcLog2 3.70 in PRI, ΔlcLog2 3.62 in PLI, and ΔlcLog2 4.33 in PFI) and β-fructosidase/levanase/invertase (ΔlcLog2 3.72 in PI, ΔlcLog2 4.41 in PRI, ΔlcLog2 4.77 in PLI, and ΔlcLog2 4.02 in PFI) were found. They belong to the operon inulin-degrading fosRABCDXE, and β-fructosidase was higher in biculture than in monoculture, mainly increasing in the presence of Bl1, probably due to its strong competition for the substrate. In addition, 50S ribosomal protein L18 increased in the presence of Bl1, indicating accelerated growth. Proteins involved in cell proliferation, such as ribonuclease and cell wall-associated hydrolase, or conjugation proteins, such as the type IV secretion system (T4SS), were found only in some bicultures (Fig. 7). Other proteins found in greater numbers in Lp in bicultures were related to bacterial growth, such as DNA helicase and DNA polymerase III, which that are essential for DNA replication. In addition, were found protein RecA, which has DNA repair functions, DNA topoisomerase 4 that relaxes supercoiled DNA before replication, and cell division protein FtsA. Furthermore, in bicultures increased the acetate kinase (related to acetate production) and sortase (a surface protein). Sortase only increased its relative quantity in the presence of Bl1 (ΔlcLog2 0.45 in PI, ΔlcLog2 0.66 in PLI). Ribonuclease R, which is involved in RNA metabolism, increased in Lp in the presence of Lr and Bl1. Finally, enolase was only found in Lp when grown with Lr and Bl1. Among the proteins only found in Lp when grew on inulin (not found on lactose), relaxase and T4SS were only found in bicultures (concerning monoculture), in the presence of Lr, or Bl1 (Supplementary file Fig. S3).
Heat map of proteins increased in abundance when compared with the basal medium, lactose (ΔlcLog2) in L. paracasei M38 in the four conditions analyzed. For proteins only found in one treatment, they were found in at least two biological replicates and not found in any of the replicates of the counterpart. Mo, monoculture; Bi1, biculture, in presence of L. ruminis PT16; Bi2, biculture, in presence of B. longum PT4; Bi3, biculture, in presence of B. fragilis HM714
Proteins Found in L. ruminis PT16, B. longum PT4, and B. fragilis HM714, When Grown on Inulin
In general, in the evaluated strains (Lr, Bl1, and Bf), the proteins of high relative quantity in inulin (Fig. 8) were reduced or not detected in the presence of Lp (bicultures) with respect to monoculture. Specifically, only in monocultures were found certain enzymes related to sugar consumption (not found in biculture probably due to competition with Lp). For example, ABC transporters (found in Lr, Bl1, and Bf), and proteins of glucose degradation such as β-glucosidase (found in Lr), oligo-1,6-glucosidase (found in Bl1), and exported β-glucosidase (found in Bf). Furthermore, proteins with several essential functions were found only in the monocultures. Some were helicase (found in Lr, Bl1, and Bf), repair protein RecF (found in Bf), DNA polymerase III (found in Lr), and DNA primase (found in Bl1) related to DNA replication. Flagellar proteins (found in Lr) were associated with bacterial movement, and succinate-CoA ligase (found in Bf) was related to succinate production.
Heat map of proteins increased in abundance in L. ruminis PT16 (A), or B. longum PT4 (B), or B. fragilis HM714 (C) when compared with the basal medium, lactose (ΔlcLog2), in two conditions analyzed. For proteins only found in one treatment, they were found in at least two biological replicates and not found in any of the replicates of the counterpart. Mo, monoculture; Bi4, biculture, in presence of L. paracasei M38
On the other hand, among the proteins found in both monocultures and bicultures, a general decrease in fold change was observed in the presence of Lp. In Lr (Fig. 8A), among these proteins decreased in bicultures were of sugar transport (PTS family fructose, PTS system sucrose, and ABC transporter) or proteins of inulin degradation, such as fructokinase (ΔlcLog2 3.67 in RI, ΔlcLog2 2.21 in RPI), and sucrose-6-phosphate hydrolase (ΔlcLog2 5.95 in RI, ΔlcLog2 5.45 in RPI). In Bl1 (Fig. 8B), the proteins found were related to sugar transport, such as ABC transport and glucose phosphotransferase (both decreased in biculture). In addition, fructose-6-phosphate phosphoketolase (ΔlcLog2 1.92 in LI, ΔlcLog2 1.61 in LPI) was found, an enzyme relevant in bifid shunt metabolism [55]. In Bf (Fig. 8C), were found lipoproteins (decreased in biculture) and proteins that linked to DNA replication, such as DNA helicase (ΔlcLog2 0.29 in FI, ΔlcLog2 -2.45 in FPI) and DNA ligase (ΔlcLog2 0.62 in FI, ΔlcLog2 -0.95 in FPI).
Interestingly, in Bf, several enzymes related to sugar metabolism were found, but they were not directly involved in the degradation of inulin. Proteins involved in inulin degradation, such as levanase were found only in the monocultures (Supplementary file Fig. S4C). Finally, proteins were generally only found in bacteria when they grew on inulin (not found on lactose). More proteins were found in the monoculture than in the bicultures (Supplementary file Fig. S4). In summary, the effect of Lp, when interacting with the strains evaluated (Lr, Bl1, and Bf), manifested in the reduction of proteins relevant to the growth of the latter.
Discussion
Plants rich in inulin lead to beneficial modifications in the composition and function of the intestinal microbiome [56]. Their effects on the human microbiome have been extensively studied, focusing on cooperative interactions with health-relevant bacteria. However, interactions between bacteria can be dominated by competition and depend on the substrate degradation capacity [57]. Single cultures showed vast consumption for the most part, except for four bifidobacteria (Bb1, Bb2, Ba, and Bl5). Although these bacteria did not grow on inulin, studies show that these species can consume fructans [58,59,60]. However, inulin size preference and consumption rate are strain specific [61, 62]. On the other hand, previous studies have shown that Lactobacillus, Bifidobacterium, and Bacteroides can grow on inulin [63, 64]. The growth of Lp was remarkable because it consumed inulin quickly and completely. This coincides with L. paracasei W20 [45].
In unidirectional assays (growth in supernatants), it was observed that Lp always consumed the substrate and showed the highest growth in the supernatants, repeating the behavior seen in monocultures. The TLCs (Fig. 2) supported these data because of the total sugar consumption of the supernatants by Lp. The other strains generally consumed the FOS remaining from the initial substrate (except for Lp, where Bl1 consumed fructose), which can be easily assimilated due to their small size. B. longum can use β-(2,1)-fructans, growing better with short-chain FOS than long-chain inulin [25].
In bidirectional assays (Fig. 3), bacteria showed competition interaction in the three pairs evaluated (Lp with Lr, Bl1, or Bf), where Lp was always favored, probably due to the dominance of substrate consumption [65]. Although L. paracasei has been shown to have beneficial effects on other members of the intestinal microbiota [17, 64] and has been reported to allow growth on inulin of Lactobacillus salivarius W57 by cross-feeding. However, competing strains may be distantly related species or, conversely, differ only in a single mutation, depending on whether they overlap in resource use [34]. In addition, bacteria with similar nutritional requirements compete to acquire nutrients that are depleted in the environment [66]. The interaction most affected by competition was "Bl1 vs. Lp", where Bl1 reduced its growth more than the other strains, and Lp showed the opposite effect (174% increased with respect to monoculture). This may be due to the competition for fructose (observed in unidirectional TLC). The preference for this sugar has already been reported in the proteome of B. longum NCC2705 [67].
Interestingly, Lactobacillus is found in low amounts in the intestinal microbiome [68] but can alter the gut microbiome population [69]. Therefore, Lp reduced the growth of other strains. The highly competitive and nutrient-limited intestinal environment may be reflected in the consumption of inulin [70]. For example, L. paracasei populations reduced the survival of L. monocytogenes, Salmonella enterica subsp. enterica, and E. coli on inulin of artichokes foods [71]. The consumption of inulin was shown by TLC (Fig. 4) in Lr, Bl1, and Bf, where in bicultures, they showed accelerated degradation (compared to monoculture) due to Lp [45]. On the other hand, in bicultures, Lp (in the three conditions), Bl1, and Bf reduced acetate concentration (Fig. 5), which can be consumed as a carbon source [64] or decreased by low bacterial growth (of Bl1, or Bf). Furthermore, lactate increased in Lr, Bl1, and Bf due to the presence of Lp [72]. This change affects pH, an essential factor governing competition between bacterial species [73]. Finally, Bf in biculture did not produce succinate, prioritizing the use of the carbon source for primary metabolic pathways [74].
Proteins Found in Lp
Bacteria in the presence of inulin increased the relative abundance of carbohydrate metabolism pathways [75]. In general, Lp increased the relative quantity of proteins found in inulin bicultures compared to monocultures (Fig. 7). As for proteins, Lp in biculture increased the ABC transporter, which is used in Lactobacillus to transport inulin or FOS to the cells [76, 77], and is degraded by cytoplasmic β-fructosidase [78]. In bicultures, increased 6-phosphofructokinase catalyzes the phosphorylation of D-fructose 6-phosphate to fructose 1,6-bisphosphate during inulin degradation [27]. In addition, proteins of the operon fosRABCDXE were found [21], phosphotransferase system fructose (PTS transport), and β-fructosidase/levanase/invertase (FosE), which hydrolyze the terminal non-reducing β-D-fructofuranoside residues in β-D-fructofuranosides.
In the presence of Bl1, Lp increased its β-fructosidase, which correlates with the best growth rates in previous trials (Fig. 3B). L. paracasei 1195 degrades FOS (DP < 10) extracellularly through β-fructosidase anchored in the cell wall. Each PTS transporter takes the released fructose and sucrose into the cells [79]. Proteomic analyses of Lactobacillus plantarum on inulin revealed an increase of β-fructosidase in monocultures [27], and L. paracasei consumed short-chain inulin using an exoinulinase enzyme (GH32) [45]. Furthermore, L. paracasei 1195 contains a cell wall-anchored β-fructosidase that degrades fructan outside the cell [27, 79].
As for proteins related to possible advantages in competition, was a sortase in PLI, which increased (regarding monoculture), functioning as an adhesin [80], increasing the chances of colonization [66]. Enolase was only found in Lp when grew with Lr and Bl1. This enzyme can also exhibit lyase activity [81]. Finally, the enzymes were found only in bicultures and not in lactose (Supplementary file Fig. S3), such as relaxase, binds to DNA and directs it to the recipient cells [82]. This can be complemented by T4SS (found only in the presence of Lr or Bl1). T4SS is used for genetic exchange in conjugation in Lactobacillus [83] and translocation of effectors with consequent impacts on genome plasticity [84, 85].
In summary, proteomic evidence showed that in bicultures, there was an increase in critical proteins for inulin degradation, bacterial growth (replicative DNA helicase and protein RecA), and phenotypic characteristics, which confer adaptive advantages to Lp when competing with other strains. These results suggest strong synergy between Lp and inulin. This performance has been shown in L. paracasei BGP1, together with inulin, contributing to the extension of the shelf life of foods, possibly due to competition or the production of antimicrobial compounds [24]. In addition, The symbiosis of L. paracasei I321 and inulin allowed a complete inhibition of Salmonella by antibacterial secretion and competitive adhesion [70].
Proteins Found in Lr, Bl1, and Bf
In general, in all strains (Lr, Bl1, and Bf), certain proteins in the bicultures were not found or decreased with respect to those in the monocultures (Fig. 8). Because Lp had a higher prevalence of fermenting inulin [11]. ABC transporters and helicases were not detected in the presence of Lp. Only in the monoculture of Lr were flagellar proteins found that can provide motility in competition. This affects the ability of some bacteria to compete, whereas other bacteria use active locomotion to avoid competition [66]. Furthermore, succinate-CoA ligase was found only in FI and was correlated with succinate reduction in bicultures (Fig. 5C). This effect was contrary to cross-feeding, where the proteome of B. fragilis has been observed after consuming bifidobacterial EPS and activating the succinate pathway [86]. As for proteins in all bicultures, enzymes decreased related to inulin degradation or DNA replication, with respect to the monoculture (Fig. 8).
Interestingly, only in FPI was found transporter efflux component protein associated with antimicrobial resistance [87], probably because of Lp. In summary, all bacteria were negatively affected by Lp. However, the inhibitory effect on gram-positive bacteria (Lr, Bl1) was mainly based on reducing their ability to metabolize inulin. While in Bf, it primarily affected their growth, thereby affecting DNA replication. The inhibition of their growth by Lp drastically affected the production of many important enzymes, such as levanase (Supplementary file Fig. S4). Furthermore, it is known that Bacteroides spp. grows less when inulin is present at an acidic pH because it is an essential factor of competition between bacteria [73]. In this case, lactate was produced by Lp. Inulin regulates the gut microbiome composition and promotes the proliferation of beneficial bacteria [3]. But, when a competitive strain dominates the community, it extinguishes the weaker strain [66, 88].
Conclusions
This work demonstrated how intestinal bacteria could modify their growth, proteomes, and sugar consumption when interacting. Unidirectional assays showed partial degradation of inulin by certain bacteria. It favors the coexistence of other microorganisms, which consume oligosaccharides. However, bidirectional assays showed that competition is preferred when a bacterium completely degrades the prebiotic substrate. In this context, L. paracasei M38, when interacting with different commensal bacteria (L. ruminis PT16, B. longum PT4, and B. fragilis HM714), it competed for the inulin (carbon source) and modified its proteome. The antagonistic effects favored L. paracasei M38, which increased the abundance of relevant enzymes in inulin catabolism, such as β-fructosidase, and sugar transporters. These proteins gave an adaptive advantage for inulin consumption over other bacteria evaluated. These latter reduced the number of proteins crucial for their development, leading to their poor growth. The synergy of inulin and L. paracasei M38 allows enhancing this bacterium to search for probiotic characteristics that displace the harmful host bacteria by competitive inhibition or other mechanisms.
Data Availability
The datasets are available from the corresponding author on reasonable request.
References
Berg G, Rybakova D, Fischer D et al (2020) Microbiome definition re-visited: old concepts and new challenges. Microbiome 8:103. https://doi.org/10.1186/s40168-020-00875-0
Sekirov I, Russell SL, Antunes LCM, Finlay BB (2010) Gut microbiota in health and disease. Physiol Rev 90:859–904. https://doi.org/10.1152/physrev.00045.2009
Tawfick MM, Xie H, Zhao C et al (2022) Inulin fructans in diet: role in gut homeostasis, immunity, health outcomes and potential therapeutics. Int J Biol Macromol 208:948–961. https://doi.org/10.1016/j.ijbiomac.2022.03.218
Wang M, Wichienchot S, He X et al (2019) In vitro colonic fermentation of dietary fibers: fermentation rate, short-chain fatty acid production and changes in microbiota. Trends Food Sci Technol 88:1–9. https://doi.org/10.1016/j.tifs.2019.03.005
Zhang F, Fan D, Huang J, Zuo T (2022) The gut microbiome: linking dietary fiber to inflammatory diseases. Med Microecol 14:100070. https://doi.org/10.1016/j.medmic.2022.100070
Abreu CDD, Caldas BV, Ribeiro GHM et al (2022) Inulin prebiotic dietary supplementation improves metabolic parameters by reducing the Toll-like receptor 4 transmembrane protein gene and interleukin 6 expression in adipose tissue. PharmaNutrition 22:100316. https://doi.org/10.1016/j.phanu.2022.100316
Rosa MC, Carmo MRS, Balthazar CF et al (2021) Dairy products with prebiotics: an overview of the health benefits, technological and sensory properties. Int Dairy J 117:105009. https://doi.org/10.1016/j.idairyj.2021.105009
López-Molina D, Chazarra S, How CW et al (2015) Cinnamate of inulin as a vehicle for delivery of colonic drugs. Int J Pharm 479:96–102. https://doi.org/10.1016/j.ijpharm.2014.12.064
Strieder MM, Arruda HS, Pastore GM, Silva EK (2023) Inulin-type dietary fiber stability after combined thermal, mechanical, and chemical stresses related to ultrasound processing of prebiotic apple beverage. Food Hydrocoll 139:108489. https://doi.org/10.1016/j.foodhyd.2023.108489
Akram W, Garud N, Joshi R (2019) Role of inulin as prebiotics on inflammatory bowel disease. Drug Discov Ther 13:1–8. https://doi.org/10.5582/ddt.2019.01000
Zhu Y, Liu J, Lopez JM, Mills DA (2020) Inulin fermentation by lactobacilli and bifidobacteria from dairy calves. Appl Environ Microbiol 87. https://doi.org/10.1128/AEM.01738-20
Vandeputte D, Falony G, Vieira-Silva S et al (2017) Prebiotic inulin-type fructans induce specific changes in the human gut microbiota. Gut 66:1968–1974. https://doi.org/10.1136/gutjnl-2016-313271
Watson AW, Houghton D, Avery PJ et al (2019) Changes in stool frequency following chicory inulin consumption, and effects on stool consistency, quality of life and composition of gut microbiota. Food Hydrocoll 96:688–698. https://doi.org/10.1016/j.foodhyd.2019.06.006
Sonnenburg ED, Zheng H, Joglekar P et al (2010) Specificity of polysaccharide use in intestinal Bacteroides species determines diet-induced microbiota alterations. Cell 141:1241–1252. https://doi.org/10.1016/j.cell.2010.05.005
Zou J, Chassaing B, Singh V et al (2018) Fiber-mediated nourishment of gut microbiota protects against diet-induced obesity by restoring IL-22-mediated colonic health. Cell Host Microbe 23:41-53.e4. https://doi.org/10.1016/j.chom.2017.11.003
Markowiak P, Slizewska K (2017) Effects of probiotics, prebiotics, and synbiotics on human health. Nutrients. https://doi.org/10.3390/nu9091021
De Bellis P, Sisto A, Lavermicocca P (2021) Probiotic bacteria and plant-based matrices: an association with improved health-promoting features. J Funct Foods 87:104821. https://doi.org/10.1016/j.jff.2021.104821
Ahmed W, Rashid S (2019) Functional and therapeutic potential of inulin: a comprehensive review. Crit Rev Food Sci Nutr 59:1–13. https://doi.org/10.1080/10408398.2017.1355775
Sarup R, Chauhan K, Kennedy JF (2017) A panorama of bacterial inulinases : production, purification, characterization and industrial applications. Int J Biol Macromol 96:312–322. https://doi.org/10.1016/j.ijbiomac.2016.12.004
Kango N, Jain SC (2011) Production and properties of microbial inulinases : production and properties of microbial inulinases : Recent 5436. https://doi.org/10.1080/08905436.2011.590763
Torres-Miranda A, Melis-Arcos F, Garrido D (2022) Characterization and identification of probiotic features in Lacticaseibacillus Paracasei using a comparative genomic analysis approach. Probiotics Antimicrob Proteins 14:1211–1224. https://doi.org/10.1007/s12602-022-09999-1
Singh P, Gill PK (2006) Production of inulinases : recent advances. 44:151–162
Chi Z, Chi Z, Zhang T, Liu G (2009) Inulinase-expressing microorganisms and applications of inulinases. 211–220. https://doi.org/10.1007/s00253-008-1827-1
Rubel IA, Iraporda C, Manrique GD, Genovese DB (2022) Jerusalem Artichoke (Helianthus tuberosus L.) inulin as a suitable bioactive ingredient to incorporate into spreadable ricotta cheese for the delivery of probiotic. Bioact Carbohydrates Diet Fibre 28:100325. https://doi.org/10.1016/j.bcdf.2022.100325
Valdés-Varela L, Ruas-Madiedo P, Gueimonde M (2017) In vitro fermentation of different fructo-oligosaccharides by Bifidobacterium strains for the selection of synbiotic combinations. Int J Food Microbiol 242:19–23. https://doi.org/10.1016/j.ijfoodmicro.2016.11.011
Wei X, Guo Y, Shao C et al (2012) Fructose uptake in Bifidobacterium longum NCC2705 is mediated by an ATP-binding cassette transporter. J Biol Chem 287:357–367. https://doi.org/10.1074/jbc.M111.266213
Buntin N, Hongpattarakere T, Ritari J et al (2017) An inducible operon is involved in inulin utilization in Lactobacillus plantarum strains, as revealed by comparative proteogenomics and metabolic profiling. Appl Environ Microbiol 83. https://doi.org/10.1128/AEM.02402-16
Faust K, Raes J (2012) Microbial interactions: from networks to models. Nat Rev Microbiol 10:538–550. https://doi.org/10.1038/nrmicro2832
Saa P, Urrutia A, Silva-Andrade C et al (2022) Modeling approaches for probing cross-feeding interactions in the human gut microbiome. Comput Struct Biotechnol J 20:79–89. https://doi.org/10.1016/j.csbj.2021.12.006
Ho A, Lonardo DP Di, Bodelier PLE (2017) Revisiting life strategy concepts in environmental microbial ecology. FEMS Microbiol Ecol fix006. https://doi.org/10.1093/femsec/fix006
Banerjee S, Schlaeppi K, van der Heijden MGA (2018) Keystone taxa as drivers of microbiome structure and functioning. Nat Rev Microbiol 16:567–576. https://doi.org/10.1038/s41579-018-0024-1
Freilich S, Zarecki R, Eilam O et al (2011) Competitive and cooperative metabolic interactions in bacterial communities. Nat Commun 2:589. https://doi.org/10.1038/ncomms1597
O’Brien EJ, Monk JM, Palsson BO (2015) Using genome-scale models to predict biological capabilities. Cell 161:971–987. https://doi.org/10.1016/j.cell.2015.05.019
Ghoul M, Mitri S (2016) The ecology and evolution of microbial competition. Trends Microbiol 24:833–845. https://doi.org/10.1016/j.tim.2016.06.011
Venturelli OS, Carr AC, Fisher G et al (2018) Deciphering microbial interactions in synthetic human gut microbiome communities. Mol Syst Biol 14:e8157. https://doi.org/10.15252/msb.20178157
Menon R, Ramanan V, Korolev KS (2018) Interactions between species introduce spurious associations in microbiome studies. PLoS Comput Biol 14(1): e1005939. https://doi.org/10.1371/journal.pcbi.1005939
Denou E, Rezzonico E, Panoff J-M et al (2009) A mesocosm of Lactobacillus johnsonii, Bifidobacterium longum, and Escherichia coli in the mouse gut. DNA Cell Biol 28:413–422. https://doi.org/10.1089/dna.2009.0873
Liu G, Song Z, Yang X et al (2016) Antibacterial mechanism of bifidocin A, a novel broad-spectrum bacteriocin produced by Bifidobacterium animalis BB04. Food Control 62:309–316. https://doi.org/10.1016/j.foodcont.2015.10.033
Patnode ML, Beller ZW, Han ND et al (2019) Interspecies Competition Impacts Targeted Manipulation of Human Gut Bacteria by Fiber-Derived Glycans. Cell 179:59-73.e13. https://doi.org/10.1016/j.cell.2019.08.011
Orihuel A, Terán L, Renaut J et al (2018) Differential proteomic analysis of lactic acid bacteria—Escherichia coli O157:H7 interaction and its contribution to bioprotection strategies in meat. Front Microbiol 9. https://doi.org/10.3389/fmicb.2018.01083
Thomson P, Santibañez R, Aguirre C et al (2019) Short-term impact of sucralose consumption on the metabolic response and gut microbiome of healthy adults. Br J Nutr 122:856–862. https://doi.org/10.1017/S0007114519001570
Pinto F, Medina DA, Pérez-Correa JR, Garrido D (2017) Modeling metabolic interactions in a consortium of the infant gut microbiome. Front Microbiol 8. https://doi.org/10.3389/fmicb.2017.02507
Chen I-MA, Chu K, Palaniappan K et al (2022) The IMG/M data management and analysis system vol 7: content updates and new features. Nucleic Acids Res. https://doi.org/10.1093/nar/gkac976
Petrova P, Petrov K (2017) Prebiotic–probiotic relationship: the genetic fundamentals of polysaccharides conversion by Bifidobacterium and Lactobacillus genera. In: Food Bioconversion. Elsevier, pp 237–278
Boger MCL, Lammerts van Bueren A, Dijkhuizen L (2018) Cross-feeding among probiotic bacterial strains on prebiotic inulin involves the extracellular exo -inulinase of Lactobacillus paracasei strain W20. Appl Environ Microbiol 84. https://doi.org/10.1128/AEM.01539-18
Hirmas B, Gasaly N, Orellana G et al (2022) Metabolic modeling and bidirectional culturing of two gut microbes reveal cross-feeding interactions and protective effects on intestinal cells. mSystems 7. https://doi.org/10.1128/msystems.00646-22
Caballero V, Estévez M, Tomás-Barberán FA et al (2022) Biodegradation of punicalagin into ellagic acid by selected probiotic bacteria: a study of the underlying mechanisms by MS-based proteomics. J Agric Food Chem. https://doi.org/10.1021/acs.jafc.2c06585
Delgado J, Owens RA, Doyle S et al (2017) Quantitative proteomics reveals new insights into calcium-mediated resistance mechanisms in Aspergillus flavus against the antifungal protein PgAFP in cheese. Food Microbiol 66:1–10. https://doi.org/10.1016/j.fm.2017.03.015
Luber CA, Cox J, Lauterbach H et al (2010) Quantitative proteomics reveals subset-specific viral recognition in dendritic cells. Immunity 32:279–289. https://doi.org/10.1016/j.immuni.2010.01.013
Cox J, Mann M (2008) MaxQuant enables high peptide identification rates, individualized p.p.b.-range mass accuracies and proteome-wide protein quantification. Nat Biotechnol. https://doi.org/10.1038/nbt.1511
Álvarez M, Delgado J, Núñez F et al (2021) Proteomic analyses reveal mechanisms of action of biocontrol agents on ochratoxin A repression in Penicillium nordicum. Food Control 129:108232. https://doi.org/10.1016/j.foodcont.2021.108232
Bindea G, Mlecnik B, Hackl H et al (2009) ClueGO: a Cytoscape plug-in to decipher functionally grouped gene ontology and pathway annotation networks. Bioinformatics 25:1091–1093. https://doi.org/10.1093/bioinformatics/btp101
Álvarez M, Delgado J, Núñez F et al (2022) Proteomic approach to unveil the ochratoxin A repression by Debaryomyces hansenii and rosemary on Penicillium nordicum during dry-cured fermented sausages ripening. Food Control 137:108695. https://doi.org/10.1016/j.foodcont.2021.108695
Rawi MH, Zaman SA, Pa’ee KF et al (2020) Prebiotics metabolism by gut-isolated probiotics. J Food Sci Technol 57:2786–2799. https://doi.org/10.1007/s13197-020-04244-5
Egan M, Van Sinderen D (2018) Carbohydrate metabolism in bifidobacteria. In: The bifidobacteria and related organisms. Elsevier, pp 145–164
Hiel S, Bindels LB, Pachikian BD et al (2019) Effects of a diet based on inulin-rich vegetables on gut health and nutritional behavior in healthy humans. Am J Clin Nutr 109:1683–1695. https://doi.org/10.1093/ajcn/nqz001
Moens F, Weckx S, De Vuyst L (2016) Bifidobacterial inulin-type fructan degradation capacity determines cross-feeding interactions between bifidobacteria and Faecalibacterium prausnitzii. Int J Food Microbiol 231:76–85. https://doi.org/10.1016/j.ijfoodmicro.2016.05.015
Pohlentz JC, Gallala N, Kosciow K, Hövels M (2022) Growth behavior of probiotic microorganisms on levan- and inulin-based fructans. J Funct Foods 99:105343. https://doi.org/10.1016/j.jff.2022.105343
Elaheh M, Ali MS, Elnaz M, Ladan N (2016) Prebiotic effect of Jerusalem artichoke (Helianthus tuberosus) fructans on the growth performance of Bifidobacterium bifidum and Escherichia coli. Asian Pacific J Trop Dis 6:385–389. https://doi.org/10.1016/S2222-1808(15)61053-2
Rattanakiat S, Pulbutr P, Khunawattanakul W et al (2020) Prebiotic activity of polysaccharides extracted from Jerusalem artichoke tuber and development of prebiotic granules. Pharmacogn J 12:1402–1411. https://doi.org/10.5530/pj.2020.12.194
Falony G, Lazidou K, Verschaeren A et al (2009) In vitro kinetic analysis of fermentation of prebiotic inulin-type fructans by Bifidobacterium species reveals four different phenotypes. Appl Environ Microbiol 75:454–461. https://doi.org/10.1128/AEM.01488-08
Selak M, Rivière A, Moens F et al (2016) Inulin-type fructan fermentation by bifidobacteria depends on the strain rather than the species and region in the human intestine. Appl Microbiol Biotechnol 100:4097–4107. https://doi.org/10.1007/s00253-016-7351-9
Feng J, Qian Y, Zhou Z et al (2022) Polysaccharide utilization loci in Bacteroides determine population fitness and community-level interactions. Cell Host Microbe 30:200-215.e12. https://doi.org/10.1016/j.chom.2021.12.006
Moens F, Verce M, De Vuyst L (2017) Lactate- and acetate-based cross-feeding interactions between selected strains of lactobacilli, bifidobacteria and colon bacteria in the presence of inulin-type fructans. Int J Food Microbiol 241:225–236. https://doi.org/10.1016/j.ijfoodmicro.2016.10.019
Hindson J (2019) Bacteroides compete for fibre-derived glycans. Nat Rev Gastroenterol Hepatol 16:706–707. https://doi.org/10.1038/s41575-019-0223-x
Kumar N, Chaturvedi S, Khurana SMP (2022) Survival and thriving behavior of bacteria in microbial jungle. In: Microbial Resource Technologies for Sustainable Development. Elsevier, pp 1–21
Liu D, Wang S, Xu B et al (2011) Proteomics analysis of Bifidobacterium longum NCC2705 growing on glucose, fructose, mannose, xylose, ribose, and galactose. Proteomics 11:2628–2638. https://doi.org/10.1002/pmic.201100035
Azad MAK, Sarker M, Li T, Yin J (2018) Probiotic species in the modulation of gut microbiota: an overview. Biomed Res Int 2018. https://doi.org/10.1155/2018/9478630
Lye H-S, Kato T, Low W-Y et al (2017) Lactobacillus fermentum FTDC 8312 combats hypercholesterolemia via alteration of gut microbiota. J Biotechnol 262:75–83. https://doi.org/10.1016/j.jbiotec.2017.09.007
Kanjan P, Hongpattarakere T (2017) Prebiotic efficacy and mechanism of inulin combined with inulin-degrading Lactobacillus paracasei I321 in competition with Salmonella. Carbohydr Polym 169:236–244. https://doi.org/10.1016/j.carbpol.2017.03.072
Valerio F, Lonigro SL, Di Biase M et al (2013) Bioprotection of ready-to-eat probiotic artichokes processed with Lactobacillus paracasei LMGP22043 against foodborne pathogens. J Food Sci 78:M1757–M1763. https://doi.org/10.1111/1750-3841.12282
Chen L, Shen Y, Wang C et al (2019) Megasphaera elsdenii lactate degradation pattern shifts in rumen acidosis models. Front Microbiol 10. https://doi.org/10.3389/fmicb.2019.00162
Chung WSF, Walker AW, Louis P et al (2016) Modulation of the human gut microbiota by dietary fibres occurs at the species level. BMC Biol 14:3. https://doi.org/10.1186/s12915-015-0224-3
Seyedsayamdost MR (2019) Toward a global picture of bacterial secondary metabolism. J Ind Microbiol Biotechnol 46:301–311. https://doi.org/10.1007/s10295-019-02136-y
Chen N, Liu Y, Wei S et al (2023) Dynamic changes of inulin utilization associated with longitudinal development of gut microbiota. Int J Biol Macromol 229:952–963. https://doi.org/10.1016/j.ijbiomac.2022.12.318
Tsujikawa Y, Ishikawa S, Sakane I et al (2021) Identification of genes encoding a novel ABC transporter in Lactobacillus delbrueckii for inulin polymers uptake. Sci Rep 11:16007. https://doi.org/10.1038/s41598-021-95356-1
Barrangou R, Altermann E, Hutkins R et al (2003) Functional and comparative genomic analyses of an operon involved in fructooligosaccharide utilization by Lactobacillus acidophilus. Proc Natl Acad Sci 100:8957–8962. https://doi.org/10.1073/pnas.1332765100
Park J-H, Song W-S, Lee J et al (2022) An integrative multiomics approach to characterize prebiotic inulin effects on Faecalibacterium prausnitzii. Front Bioeng Biotechnol 10. https://doi.org/10.3389/fbioe.2022.825399
Goh YJ, Lee J-H, Hutkins RW (2007) Functional analysis of the fructooligosaccharide utilization operon in Lactobacillus paracasei 1195. Appl Environ Microbiol 73:5716–5724. https://doi.org/10.1128/AEM.00805-07
Vélez MP, De Keersmaecker SCJ, Vanderleyden J (2007) Adherence factors of Lactobacillus in the human gastrointestinal tract. FEMS Microbiol Lett 276:140–148. https://doi.org/10.1111/j.1574-6968.2007.00908.x
Ferreira TG, dos Trindade CN, R, Bell P et al (2018) Identification of the alpha-enolase P46 in the extracellular membrane vesicles of Bacteroides fragilis. Mem Inst Oswaldo Cruz 113:178–184. https://doi.org/10.1590/0074-02760170340
Guzmán-Herrador DL, Llosa M (2019) The secret life of conjugative relaxases. Plasmid 104:102415. https://doi.org/10.1016/j.plasmid.2019.102415
Wuyts S, Allonsius CN, Wittouck S et al (2019) Comparative genome analysis of Lactobacillus mudanjiangensis, an understudied member of the Lactobacillus plantarum group. Microb Genomics 5. https://doi.org/10.1099/mgen.0.000286
Wallden K, Rivera-Calzada A, Waksman G (2010) Microreview: Type IV secretion systems: versatility and diversity in function. Cell Microbiol 12:1203–1212. https://doi.org/10.1111/j.1462-5822.2010.01499.x
Bi D, Liu L, Tai C et al (2013) SecReT4: a web-based bacterial type IV secretion system resource. Nucleic Acids Res 41:D660–D665. https://doi.org/10.1093/nar/gks1248
Rios-Covian D, Sánchez B, Salazar N et al (2015) Different metabolic features of Bacteroides fragilis growing in the presence of glucose and exopolysaccharides of bifidobacteria. Front Microbiol 6. https://doi.org/10.3389/fmicb.2015.00825
Dong N, Zeng Y, Wang Y et al (2022) Distribution and spread of the mobilised RND efflux pump gene cluster tmexCD-toprJ in clinical Gram-negative bacteria: a molecular epidemiological study. The Lancet Microbe 3:e846–e856. https://doi.org/10.1016/S2666-5247(22)00221-X
Wei S, Wang C, Zhang Q et al (2022) Dynamics of microbial communities during inulin fermentation associated with the temporal response in SCFA production. Carbohydr Polym 298:120057. https://doi.org/10.1016/j.carbpol.2022.120057
Funding
Open Access funding provided thanks to the CRUE-CSIC agreement with Springer Nature. This work was supported by the “Agencia Nacional de Investigacion y Desarrollo de Chile” (ANID) regular Fondecyt project 1190074, ANID Fondequip EQM190070, and ANID Scholarship 21190742. Q-Exactive mass spectrometer to proteomic research was supported by the Spanish Ministerio de Economía y Competitividad (Ref. UNEX-AE-3394).
Author information
Authors and Affiliations
Contributions
MVS: conceptualization, data curation, formal analysis, investigation, methodology, validation, visualization, writing—original draft, and writing—review and editing. ECC: data curation. JD: data curation, methodology, resources, supervision, writing—review and editing. SRM: methodology, resources, supervision, writing—review and editing. DG: conceptualization, methodology, resources, supervision, writing—review and editing.
Corresponding authors
Ethics declarations
Conflict of Interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Vega-Sagardía, M., Cabezón, E.C., Delgado, J. et al. Screening Microbial Interactions During Inulin Utilization Reveals Strong Competition and Proteomic Changes in Lacticaseibacillus paracasei M38. Probiotics & Antimicro. Prot. 16, 993–1011 (2024). https://doi.org/10.1007/s12602-023-10083-5
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12602-023-10083-5