Abstract
Purpose
Machine learning models are used to develop and improve various disease prediction systems. Ensemble learning is a machine learning technique that combines many classifiers to increase performance by making more accurate predictions than a single classifier. Although several researchers have employed ensemble techniques for disease prediction, a comprehensive comparative study of these techniques still needs to be provided.
Methods
Using 16 disease datasets from Kaggle and the UCI Machine Learning Repository, this study compares the performance of 15 variants of ensemble techniques for disease prediction. The comparison was performed using six performance measures: accuracy, precision, recall, F1 score, AUC (Area Under the receiver operating characteristics Curve) and AUPRC (Area Under the Precision-Recall Curve).
Results
Stacking variant of Multi-level stacking showed superior disease prediction performance compared with other bagging and boosting variants, followed by another stacking variant (Classical stacking). Overall, stacking outperformed bagging and boosting for disease prediction. Logit Boost showed the worst performance.
Conclusion
The findings of this study can help researchers select an appropriate ensemble approach for future studies focusing on accurate disease prediction.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Disease diagnosis is a critical step in treating and managing various medical conditions. However, it can be challenging due to the complexity and variability of symptoms and signs. Correct disease diagnosis is essential for effective intervention and patient care [1]. Many scientists have developed machine learning algorithms that accurately identify a broad spectrum of diseases [2,3,4,5]. These algorithms can create disease prediction models, enabling early detection and intervention, which are crucial in reducing disease-related mortality [6]. As a result, most medical scientists are drawn to emerging machine learning-based predictive model technologies for disease prediction.
Diabetes, skin disease, kidney disease, liver disease and heart disease are all major chronic diseases that substantially impact health and, if left untreated, can lead to death [7]. Therefore, accurate disease prediction becomes vital in improving patient care and minimising the burden of these chronic conditions. By identifying hidden patterns and relationships in vast healthcare databases, machine learning techniques can assist healthcare professionals in making informed decisions and delivering timely interventions [8]. Ensemble learning is a machine learning technique that aims to improve prediction performance by combining forecasts from several models [1]. Ensemble models reduce the generalisation error in the forecast. The ensemble method reduces model prediction error when the fundamental models are diverse and independent [9].
Bagging, also known as bootstrap aggregating, reduces overfitting and variance by combining predictions from multiple models trained on different subsets of the data [10]. On the other hand, boosting adjusts the weights of misclassified samples iteratively, focusing on difficult-to-classify instances and improving the accuracy of the overall ensemble model. Stacking combines the predictions of multiple models using a meta-learner, which can outperform individual models and other ensemble techniques in various applications [11]. While researchers have used machine learning algorithms extensively for disease prediction, there is a lack of comprehensive studies comparing the performance measures of ensemble learning techniques, such as bagging, boosting, and stacking and their variants, against different significant chronic disease datasets.
A comparative analysis of different ensemble techniques and their variants for disease prediction is crucial in understanding the strengths and limitations of these ensemble approaches. It can help researchers identify the most effective methods for disease prediction [12]. Researchers compared supervised [13, 14] and unsupervised [15] machine learning algorithms for disease prediction. Mahajan et al. [16] conducted a literature review on applying ensemble approaches for disease prediction. However, no study in the current literature compares and contrasts ensemble approaches using multiple datasets. Therefore, the primary objective of this study is to uncover critical trends in disease prediction models based on ensemble learning techniques, specifically bagging, boosting and stacking, and their variants, using performance measures such as accuracy, precision, recall and F1 score. By comparing and evaluating these approaches across various chronic disease datasets, this research provides insights into the effectiveness of different ensemble learning methods for disease prediction.
The datasets used in this study encompass major chronic diseases, including diabetes, chronic kidney disease, liver disease, heart disease, and skin cancer. These diseases were selected due to their prevalence and impact on health outcomes. This study will conduct a comprehensive performance analysis of various ensemble learning techniques by implementing and conducting experiments on 16 machine learning datasets obtained from reputable sources, including Kaggle and the UCI Machine Learning Repository.
For analyses, we considered 15 ensemble algorithms: classical bagging, decision tree, random forest (RF), extra trees, dagging, random subspace, classical boosting, AdaBoost, CatBoost, XGBoost, LightGBM, Logit Boost, Classical stacking, Two-level stacking and Multi-level stacking for disease prediction. Table 1 provides the basic idea, pros and cons for each of these 15 ensemble variants.
2 Materials and methods
2.1 Data source
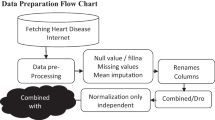
This study examines 16 datasets from Kaggle and the UCI Machine Learning Repository that are associated with five primary chronic diseases: heart disease, renal disease, liver disease, diabetes and skin cancer. The details of all 16 datasets are provided in Table 2. This table details each dataset’s source, number of attributes, total instances, and positive and negative instances. Of these 16 datasets, four are for heart disease, three for liver disease, four for diabetes, three for chronic kidney disease, and two for skin cancer. Data cleaning and preprocessing were performed before conducting the analysis to ensure the quality and integrity of the data. Normalisation was a critical step followed in this procedure since it kept all the data on the same scale and improved the accuracy of the results. While building the model, hyperparameter tuning was performed for all the classifiers to attain better performance.
2.2 Relative performance index
The relative performance index (RPI) is an assessor that collects data results of any performance measure and produces a comparative result for the final assessment [43]. For a given set of performance values, the RPI value is calculated by summing up the difference between each data instance and the minimum value of that dataset. A higher RPI value for an algorithm indicates its superior predictive power compared with other candidates and vice versa [44]. RPI is useful for researchers and practitioners looking to optimise their models for specific datasets. By analysing different variants and calculating their RPI values, it is possible to identify which ones are most effective for a given task or application, improving the overall quality of the data analysis and decision-making processes. This is the formula for RPI:
where, \({a}_{i}^{*}\) is the minimum value of the list, \({a}_{i}\) is the value for the variant under consideration for dataset \(i\), and \(d\) is the number of the datasets in the analyses.
2.3 Performance measures
2.3.1 Confusion matrix
A confusion matrix is a method for measuring performance used in statistics and machine learning to evaluate the precision of a classification model [8]. In a confusion matrix, columns correspond to the anticipated class labels, and rows correspond to the true class labels. A confusion matrix is made up of four basic parts (Fig. 1): (a) true positive (TP) is the number of instances that have been correctly predicted as positive from the positive class; (b) true negative (TN) is the number of instances that have been correctly predicted as negative from the negative class; (c) false positive (FP) is the number of instances that have been incorrectly predicted as positive from the negative class; and (d) false negative (FN) is the number of instances that have been incorrectly predicted as negative from the positive class.
Four performance measures considered in this study (i.e., accuracy, precision, recall, and F1 score) are calculated using these confusion matrix values [45]. These metrics can be calculated using the formulas mentioned below:
This study also considered two other commonly used performance measures. They are AUC (Area Under the receiver operating characteristics Curve) and AUPRC (Area Under the Precision-Recall Curve). AUC focuses on the trade-off between the true-positive rate (sensitivity) and the false-positive rate, making it appropriate for well-balanced datasets with equally distributed positive and negative examples [42]. AUPRC, on the other hand, focuses more on the precision-recall trade-off, making it appropriate for imbalanced datasets with few positive cases [43].
2.4 Experiment setup
The experimental setting for using ensemble approaches to improve binary classification task performance is described in this section. We specifically concentrated on ensemble techniques that use different basic classifiers and hyperparameter tuning techniques for bagging, boosting, and stacking. The intention was to show how various ensemble approaches can be used to increase prediction accuracy. The bagging ensemble methodology combines predictions from several base classifiers to increase performance. The process entails loading a dataset, labelling categorical features, dividing the dataset into test, validation, and training sets, and then instantiating a bagging classifier using a selected base estimator. Libraries must also be imported. GridSearchCV is used for hyperparameter tuning, and the model with the highest accuracy is chosen as the best performer. A thorough classification report is produced after the model has been trained and assessed. A similar procedure was followed for boosting and its variants. Fivefold cross-validation is used to optimise the hyperparameters of each method, improving model performance and offering a thorough evaluation of classification abilities. Both two-level and multi-level stacking are part of the experimental setting for stacking ensemble approaches. Based on the number of levels in the stacking classifier, base classifiers are trained. The predict_proba method is used to produce first-level predictions. GridSearchCV is used once again for hyperparameter tuning. The metamodel produces final predictions, and a classification report is included in the performance evaluation.
3 Results
3.1 Accuracy comparison
The accuracy outcomes of the ensemble algorithms and their variants are shown in Table 3 against all datasets considered in this study. A bold number in a cell indicates that the corresponding algorithm (column title) showed the best accuracy performance against the given dataset (row title). Interestingly, datasets D7 and D12 revealed 100% accuracy for all classification algorithms. The last row shows the number of times each algorithm revealed the best performance. Classical stacking (9) has been found to offer the best performance at most times, followed by multi-level stacking (8). Classical boosting and Logit boost performed worst against the same criteria, each revealing the best performance only three times.
Table 4 summarises the outcomes from Table 3 for the three basic ensemble approaches. In doing so, we considered all variants for a basic ensemble technique. For example, we considered all six variants while checking whether bagging produces the best result. If any of them has the best accuracy, we increase the count for the bagging technique. An “x” in a cell designates the ensemble technique that produced the best results for that dataset. For datasets D7, D11, D12, D14, and D15, all three approaches or their variants have shown the best accuracy performance. Again, stacking (14) was the best-performing method, as revealed in the last column.
Apart from the datasets showing the best accuracy for each ensemble technique (i.e., D7, D11, D12, D14 and D15), bagging showed the best accuracy only once (D13), and boosting showed three times (D1, D6 and D8). On the other hand, stacking performed the best nine times (D2-D5, D8-D10, D13 and D16). From this data analysis perspective, it is again stacking that performed best for disease prediction.
3.2 Precision comparison
Table 5 displays the results of precision scores for different ensemble techniques and their variants across disease datasets. All 15 ensemble classifiers considered in this study showed a 100% precision score for datasets D7 and D12. Datasets D12, D13, D15, and D16 consistently performed, giving a precision score of > 90% against each classifier. Regarding how many times a variant reveals the best precision performance (last row of Table 5), Classical Stacking (9) ranked first, followed by Two-level and Multi-level Stacking, each showing the best performance eight times. Classical boosting and logit boost were positioned the lowest in this regard, delivering the best performance four times each. Like the accuracy measure, Classical Boosting and Logit Boosting showed the worst outcome regarding the number of times revealing the best performance. They showed the best performance only four times, much lower than that of classical stacking, which showed the best performance the most times (9).
When variants converged to their corresponding parent ensemble approaches in terms of the number of times revealing the best precision performance, stacking appeared to be the best. The results are presented in Table 6. Stacking showed the best performance 14 times out of 16 datasets, followed by boosting (9) and bagging (8). All variants showed the best precision performance for datasets D7, D8, D12 and D14-D16. For the remaining ten datasets (D1-D6, D9-D11 and D13), stacking achieved the best precision eight times, followed by boosting (3) and bagging (2).
3.3 Recall comparison
For accuracy and precision, the variants of the stacking technique showed the best and second-best performance. Recall outcomes make an exception in this regard – there is a tie for the second-best recall score between random subspace and classical stacking. Each showed the best performance seven times, according to the last row of Table 7. Dataset D12 revealed 100% recall against all ensemble variants. Logit Boost led to the best performance minimum number of times (3) among all variants.
For the three parent ensemble approaches, there is a three-way tie for the best-performing score against datasets D7, D12, D14 and D15, according to Table 8. Stacking scored the best 12 times, followed by boosting (9) and bagging (7). For datasets D3, D7 and D12-D15, stacking showed a 100% recall score.
3.4 F1 score comparison
We observed a similar trend in the F1 score as what we observed for accuracy and precision. Stacking variants outperformed other candidate variants, as detailed in Table 9. Multi-level stacking appeared nine times as the best performer, followed by classical stacking (8) and two-level stacking (8). Datasets D7 and D12 showed a 100% F1 score for all variants. D16 showed the same F1 score (94%) for all variants. Classical boosting appeared minimum times (3) as the best performer.
At the meta-level (i.e., basic ensemble approaches), stacking showed the best F1 score performance 13 times, followed by boosting (10) and bagging (7), according to Table 10. For datasets D7, D12 and D14-D16, all the classifiers have shown the same F1 score.
3.5 AUC comparison
Like in accuracy, precision and F1 score, stacking variants outperformed other candidate variants for AUC, as detailed in Table 11. Multi-level stacking appeared nine times as the best performer, followed by classical stacking (7) and two-level stacking (7). Dataset D15 showed a 100% AUC value for all variants. D16 showed the same AUC score (89%) for all variants. Logit Boost appeared minimum times (3) as the best performer.
According to Table 12, at the meta-level (i.e., basic ensemble approaches), stacking showed the best AUC performance 13 times, followed by boosting (11) and bagging (7). For datasets D2, D7, D12 and D15-D16, all the classifiers showed the same AUC value.
3.6 AUPRC comparison
Multi-level stacking and classical stacking tied in the number of their appearance as the best peformer (8), according to Table 13. Decision tree, XGBoost and two-level stacking appeared six times each as the best performer. Like in AUC, dataset D15 showed a 100% AUPRC score for all variants. Dataset D12 showed the same AUPRC score (98%) for all variants. Classical Boosting and Logit Boost appeared minimum times (3) as the best performer.
According to Table 14, at the meta-level (i.e., basic ensemble approaches), stacking showed the best AUPRC performance 14 times, followed by boosting (9) and bagging (7). For datasets D7-D8, D12-D13 and D15-D16, all the classifiers showed the same AUPRC value.
3.7 Comparing RPI score
Using the results from Table 3, 5, 7, 9, 11 and 13 for 16 datasets, we calculated the RPI score for all performance measures against each variant. Table 15 presents the corresponding RPI score results. Classical stacking showed the highest RPI score for accuracy (11.31%), precision (16.81%) and recall (21.50%) measures. Multi-level stacking showed the highest RPI scores for AUC (9.56%) and AUPRC (12.69%). For the F1 score, Classical Boosting had the highest RPI score (7.06%).
3.8 Comparison of best count statistics
The last rows of Table 3, 5, 7, 9, 11 and 13 show the number of times each variant performed best against accuracy, precision, recall, F1 score, AUC and AUPRC, respectively. Table 16 summarises these six rows to reveal the number of times each variant performed best against all six measures. Stacking variants of multi-level stacking topped the list by appearing 50 times as the best-performing variant. This value is significantly higher than other list values \((p\le 0.02)\) according to the ‘inverse normal distribution’ test for a single value. The second highest value was revealed by another stacking variant of classical stacking (48), which is also significantly higher than other remaining values \((p\le 0.04)\). The Logit Boost variant appeared the minimum times (20) as the best performer in this table.
4 Discussion
The ensemble approach, which combines multiple prediction models, proves effective in disease prediction by reducing errors and improving the quality of forecasts. In this study, we evaluated the performance of 15 ensemble techniques, including bagging, boosting, and stacking, using 16 datasets containing information about various diseases. To ensure the reliability of our findings, we rigorously examined how well these ensemble methods performed based on different measures, such as accuracy, precision, recall, and F1 score. We also proceeded to preprocess the data, ensuring it was clean and standardised for accurate predictions.
Our analysis uncovered some interesting trends. For instance, we observed that decision trees performed less effectively in recall and F1 score than other ensemble methods, but bagging demonstrated substantial accuracy and precision. Classical boosting and logit boost performed relatively poorly among the boosting algorithms. However, stacking outperformed other methods, with classical and multi-level stacking exhibiting remarkable results. The repeated success of stacking indicates its reliability and effectiveness as an ensemble method for disease prediction, consistently surpassing other strategies. These findings suggest that stacking could have a meaningful impact on global healthcare by improving disease prediction and management. In addition to these results, our evaluation provides insights into the advantages and limitations of ensemble methods. We observed that ensemble approaches, especially stacking, improve accuracy by reducing outliers. The consistent performance of stacking across diverse datasets highlights its potential as a reliable approach for disease prediction.
Although this research considered 16 benchmark research datasets from two highly regarded open-access data repositories, these datasets may not capture the full complexity and variability of other real-world health data, such as the data from clinical settings. Moreover, most of these datasets are highly balanced. Clinical settings often encounter imbalanced data [44]. This limitation of our study opens a new research scope for the future, establishing research collaborations with healthcare providers that have access to such data to validate the findings of this study. Integrating any computation models, including the best one this study observed (i.e., classical and multi-level stacking), with the present healthcare environments is challenging, primarily due to computational complexity [45] and cost-effectiveness [46]. While this study focused on the robustness of the theoretical findings, it is crucial to research the potentiality of their real-world applications. Numerous studies [e.g., 47] highlighted the importance of adopting advanced technologies and computational models appropriately in healthcare settings. Our research echoed this importance once again. Overfitting could be another limitation of this study. Despite their effectiveness, ensemble approaches are sometimes prone to overfitting [48], especially when working with intricate models like stacking or boosting. Our consideration of GridSearchCV for hyperparameter tuning that maximises performance while minimising overfitting and cross-validation helps reduce the negative impact of this overfitting issue.
Our findings could potentially add new thoughts to improving ensemble model performance. These findings offer several directions for future research. Comparative analyses can help determine which ensemble strategy is most suitable for healthcare scenarios. Fine-tuning the methods and optimising individual algorithms can further enhance prediction accuracy. Furthermore, exploring specialised feature engineering techniques for specific domains may improve the predictive power of ensemble models. Real-world validation is essential to test their performance in healthcare settings to ensure the practical application of ensemble models. When using ensemble models for illness prediction, ethical considerations are critical. Future studies should focus on protecting privacy, minimising discrimination based on projected health effects, and ensuring responsible and equitable use. Integrating ensemble models with existing medical technologies holds promise for improving disease prediction accuracy and usefulness, ultimately benefiting patients and healthcare providers. Finally, our study highlights the strengths and potential of ensemble methods in disease prediction, with stacking emerging as a standout performer. Our recommendations for future research encompass comparative analysis, algorithm refinement, interpretability, validation, ethical considerations, and seamless integration with other healthcare technologies. These research avenues promise to advance ensemble approaches for disease prediction, leading to more accurate predictions and improved healthcare outcomes.
5 Conclusion
In this research, we evaluated the performance of various algorithms and their variations in the context of disease prediction through ensemble techniques. The findings consistently favoured the stacking technique over other ensemble strategies, revealing its effectiveness in accurately predicting diseases across diverse datasets. Notably, stacking achieved 100% accuracy on some datasets, highlighting its potential as a robust and reliable ensemble method. While bagging classifiers such as Dagging, Random Forest, Extra Trees, and Random Subspace demonstrated strong performance within the bagging ensemble models, stacking outperformed individual techniques such as CatBoost, XGBoost, and LightGBM in the boosting category. Classical boosting and LogitBoost emerged as the weakest classifiers among the various ensemble approaches assessed.
These results provide valuable guidance for selecting the most suitable algorithm for disease prediction. Notably, stacking, particularly the classical stacking and multi-level stacking algorithms, emerged as the most reliable and precise ensemble methods, outperforming other approaches across all performance metrics, showing the advantage of combining the strengths of multiple models and reducing bias and variance in predictions. The implications of these findings are significant for the field of disease prediction, as they enable healthcare professionals to enhance the accuracy of disease prediction models, potentially leading to earlier diagnosis, expedited treatment, and improved patient outcomes. Further research is warranted to explore aspects such as interpretability, optimisation, ethical considerations, and the integration of ensemble models with other medical technologies. Addressing these aspects can advance the field, resulting in more accurate and reliable predictive models for disease prediction.
Data availability
All data used in this study are publicly available in the Kaggle and UCI Machine Learning Repository.
Change history
05 April 2024
Springer Nature’s version of this paper was updated: Table 4 has been revised.
References
Mienye ID, Sun Y. A survey of ensemble learning: concepts, algorithms, applications, and prospects. IEEE Access. 2022;10:99129–49.
Ramesh D, Katheria YS. Ensemble method based predictive model for analyzing disease datasets: a predictive analysis approach. Health Technol. 2019;9:533–45.
Lu H, Uddin S. Embedding-based link predictions to explore latent comorbidity of chronic diseases. Health Inform Sci Syst. 2022;11(1):2.
Uddin S, Wang S, Lu H, Khan A, Hajati F, Khushi M. Comorbidity and multimorbidity prediction of major chronic diseases using machine learning and network analytics. Expert Syst Appl. 2022;205: 117761.
Hossain ME, Khan A, Uddin S. Understanding the comorbidity of multiple chronic diseases using a network approach. In Proc Austral Comput Sci Week Multiconference. 2019;1–7.
Nikookar E, Naderi E. Hybrid ensemble framework for heart disease detection and prediction. Int J Adv Comput Sci Appl. 2018;9(5):243–8.
Igodan EC, Thompson AF-B, Obe O, Owolafe O. Erythemato squamous disease prediction using ensemble multi-feature selection approach. Int J Comput Sci Inf Secur. 2022;20:95–106.
Alqahtani A, Alsubai S, Sha M, Vilcekova L, Javed T. Cardiovascular disease detection using ensemble learning. Comput Intell Neurosci. 2022;2022:9.
Ishaq A, Sadiq S, Umer M, Ullah S, Mirjalili S, Rupapara V, Nappi M. Improving the prediction of heart failure patients’ survival using SMOTE and effective data mining techniques. IEEE Access. 2021;9:39707–16.
Chaurasia V, Pandey MK, Pal S. Chronic kidney disease: a prediction and comparison of ensemble and basic classifiers performance. Human-Intelligent Syst Integr. 2022;4(1–2):1–10.
Zubair Hasan K, Hasan Z. Performance evaluation of ensemble-based machine learning techniques for prediction of chronic kidney disease. In: Emerging Research in Computing, Information, Communication and Applications: ERCICA 2018, vol. 1. Springer; 2019. pp. 415–26.
Yariyan P, Janizadeh S, Van Phong T, Nguyen HD, Costache R, Van Le H, Pham BT, Pradhan B, Tiefenbacher JP. Improvement of best first decision trees using bagging and dagging ensembles for flood probability mapping. Water Resour Manage. 2020;34:3037–53.
Uddin S, Khan A, Hossain ME, Moni MA. Comparing different supervised machine learning algorithms for disease prediction. BMC Med Inf Decis Mak. 2019;19(1):1–16.
Uddin S, Haque I, Lu H, Moni MA, Gide E. Comparative performance analysis of K-nearest neighbour (KNN) algorithm and its different variants for disease prediction. Sci Rep. 2022;12(1):1–11.
Lu H, Uddin S. Unsupervised machine learning for disease prediction: a comparative performance analysis using multiple datasets. Health Technol. 2024;14(1):141–54.
Mahajan P, Uddin S, Hajati F, Moni MA. Ensemble learning for disease prediction: a review. Healthcare. 2023;11(12):1808.
Kotsianti S, Kanellopoulos D. Combining bagging, boosting and dagging for classification problems. In Knowledge-Based Intelligent Information and Engineering Systems: 11th International Conference, KES 2007, XVII Italian Workshop on Neural Networks, Vietri sul Mare, Italy, September 12–14, 2007. Proceedings, Part II 11. 2007. Springer.
Basar MD, Akan A. Detection of chronic kidney disease by using ensemble classifiers. In 2017 10th International Conference on Electrical and Electronics Engineering (ELECO). IEEE; 2017. pp. 544–47.
Shorewala V. Early detection of coronary heart disease using ensemble techniques. Inf Med Unlocked. 2021;26:100655.
Qin Y, Wu J, Xiao W, Wang K, Huang A, Liu B, Yu J, Li C, Yu F, Ren Z. Machine learning models for data-driven prediction of diabetes by lifestyle type. Int J Environ Res Public Health. 2022;19(22):15027.
Nahar N, Ara F, Neloy MAI, Barua V, Hossain MS, Andersson K. A comparative analysis of the ensemble method for liver disease prediction. In 2019 2nd International Conference on Innovation in Engineering and Technology (ICIET). IEEE; 2019. pp. 1–6.
Singh V, Gourisaria MK, Das H. Performance analysis of machine learning algorithms for prediction of liver disease. In 2021 IEEE 4th International Conference on Computing, Power and Communication Technologies (GUCON). IEEE; 2021. pp. 1–7.
Liza FR, Samsuzzaman M, Azim R, Mahmud MZ, Bepery C, Masud MA, Taha B. An ensemble approach of supervised learning algorithms and artificial neural network for early prediction of diabetes. In 2021 3rd International Conference on Sustainable Technologies for Industry 4.0 (STI). IEEE; 2021. pp. 1–6.
Abdollahi J, Nouri-Moghaddam B. Hybrid stacked ensemble combined with genetic algorithms for diabetes prediction. Iran J Comput Sci. 2022;5:205–20.
Kuzhippallil MA, Joseph C, Kannan A. Comparative analysis of machine learning techniques for indian liver disease patients. In 2020 6th International Conference on Advanced Computing and Communication Systems (ICACCS). IEEE; 2020. pp. 778–82.
Alizadehsani R, Roshanzamir M, Abdar M, Beykikhoshk A, Khosravi A, Panahiazar M, Koohestani A, Khozeimeh F, Nahavandi S, Sarrafzadegan N. A database for using machine learning and data mining techniques for coronary artery disease diagnosis. Sci data. 2019;6(1):227.
Janosi A, Steinbrunn W, Pfisterer M, Detrano R. Heart disease UCI mach learn repository. 2020. https://doi.org/10.24432/C52P4X.
Lapp D. Heart disease dataset. 2019. https://www.kaggle.com/datasets/johnsmith88/heart-disease-dataset.
Chicco D, Jurman G. Machine learning can predict survival of patients with heart failure from serum creatinine and ejection fraction alone. BMC Med Inf Decis Mak. 2020;20(1):1–16.
Forsyth RS. Liver disorders data set. 1990. https://archive.ics.uci.edu/ml/datasets/Liver+Disorders.
Ramana BV. Indian liver patient dataset data set. 2012. https://archive.ics.uci.edu/ml/datasets/ILPD+%28Indian+Liver+Patient+Dataset%29.
Fedesoriano. COVID-19 effect on liver cancer prediction dataset. 2022. Available from: https://www.kaggle.com/datasets/fedesoriano/covid19-effect-on-liver-cancer-prediction-dataset.
Early stage diabetes risk prediction dataset. 2020. Available from: https://archive.ics.uci.edu/dataset/529/early+stage+diabetes+risk+prediction+dataset.
Mahgoub A. Diabetes prediction system with KNN algorithm. 2021. https://www.kaggle.com/abdallamahgoub/diabetes .
Tigga NP. Diabetes Dataset 2019. 2020. Available from: https://www.kaggle.com/datasets/tigganeha4/diabetes-dataset-2019.
Antal B, Hajdu A. An ensemble-based system for automatic screening of diabetic retinopathy. Knowl Based Syst. 2014;60:20–7.
Iqbal M. Chronic kidney disease dataset. 2017. https://www.kaggle.com/datasets/mansoordaku/ckdisease.
Pandit AK. Chronic kidney disease. 2020. Available from: https://www.kaggle.com/datasets/abhia1999/chronic-kidney-disease.
Ghadiya H. Kidney stone dataset. Available from: https://www.kaggle.com/datasets/harshghadiya/kidneystone.
Mader S, Skin Cancer MNIST. : HAM10000. 2018. Available from: https://www.kaggle.com/datasets/kmader/skin-cancer-mnist-ham10000.
Ilter N. Dermatology data set. 1998. https://archive.ics.uci.edu/ml/datasets/Dermatology.
de Hond AA, Steyerberg EW, van Calster B. Interpreting area under the receiver operating characteristic curve. Lancet Digit Health. 2022;4(12):e853-855.
Ozenne B, Subtil F, Maucort-Boulch D. The precision–recall curve overcame the optimism of the receiver operating characteristic curve in rare diseases. J Clin Epidemiol. 2015;68(8):855–9.
Tarekegn AN, Giacobini M, Michalak K. A review of methods for imbalanced multi-label classification. Pattern Recogn. 2021;118: 107965.
Chen P-T, Lin C-L, Wu W-N. Big data management in healthcare: adoption challenges and implications. Int J Inf Manag. 2020;53: 102078.
Lokkerbol J, Adema D, Cuijpers P, Reynolds CF III, Schulz R, Weehuizen R, Smit F. Improving the cost-effectiveness of a healthcare system for depressive disorders by implementing telemedicine: a health economic modeling study. Am J Geriatric Psychiatry. 2014;22(3):253–62.
Colicchio TK, Facelli JC, Del Fiol G, Scammon DL, Bowes WA III, Narus SP. Health information technology adoption: understanding research protocols and outcome measurements for IT interventions in health care. J Biomed Inform. 2016;63:33–44.
Grushka-Cockayne Y, Jose VRR, Lichtendahl Jr KC. Ensembles of overfit and overconfident forecasts. Manage Sci. 2017;63(4):1110–30.
Funding
Open Access funding enabled and organized by CAUL and its Member Institutions.
Author information
Authors and Affiliations
Contributions
PM: Analysis, Coding, Writing. SU: Conceptualisation, Supervision, Analysis and Writing. FH, MAM and EG: Writing.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Highlights:
• Our research examines 15 variations of ensemble approaches for disease prediction, providing useful insights into their performance.
• We evaluate the performance of these approaches using six commonly accepted performance measures: accuracy, precision, recall, F1-score, AUC (Area Under the receiver operating characteristics Curve) and AUPRC (Area Under the Precision-Recall Curve).
• Our findings show that stacking variants, notably classical and multi-level stacking, outperform other Bagging and boosting variations in disease prediction.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Mahajan, P., Uddin, S., Hajati, F. et al. A comparative evaluation of machine learning ensemble approaches for disease prediction using multiple datasets. Health Technol. 14, 597–613 (2024). https://doi.org/10.1007/s12553-024-00835-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12553-024-00835-w