Abstract
Purpose
Breath profiling has gained importance in recent years as it is a non-invasive technique to identify biomarkers for various diseases. Breath profiling of abnormal liver function in individuals for identifying potential biomarkers in exhaled breath could be a useful diagnostic tool. The objective of this study was to identify potential biomarkers in exhaled breath that remain stable and consistent during different physiological states, including rest and brief workouts, intending to develop a non-invasive diagnostic tool for detecting abnormal liver function.
Method
Our study employed a gas chromatography and mass-spectrometer quantified dataset for analysis. Machine learning techniques, including feature selection and model training, were used to rank and evaluate potential biomarkers' contributions to the model's performance. Statistical methods were applied to filter significant and consistent biomarkers. The final selected biomarkers were iterated for all possible combinations using machine learning algorithms to determine their accuracy range. Furthermore, classification models were used to evaluate the performance metrics of the biomarkers and compare models.
Result
The final selected biomarkers, including 2-Myristynoyl Pantetheine, Pterin-6 Carboxylic Acid, Methyl Mercaptan, N-Acetyl Cysteine, and Butyric Acid, exhibited stable levels in exhaled breath during different physiological states. They showed high accuracy and precision in detecting abnormal liver function. Our machine learning models achieved an accuracy rate ranging from 0.7 to 0.95 in all conditions, with precision, recall, prediction probability, and a 95% confidence interval ranging from 0.84 to 0.94, using various combinations of these biomarkers.
Conclusion
Our statistical and machine learning analysis identified significant and potential biomarkers that contribute to the detection of abnormal liver function. These biomarkers were consistent across different physiological states of the body in both patient and healthy groups. The use of breath samples and feature selection machine learning methods proved to be an accurate and reliable approach for identifying these biomarkers. Our findings provide valuable insights for future research in this field and can inform the development of non-invasive and cost-effective diagnostic tests for liver disease.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Liver disease is a major cause of death globally, responsible for approximately 2 million deaths each year [1]. Prevention measures, including reducing alcohol consumption, promoting healthy lifestyles, and vaccinating against viral hepatitis, are important in reducing the incidence of liver disease and related deaths [2]. Efforts are underway to address the global burden of liver disease by increasing awareness, improving access to screening and treatment, and developing new therapies [3, 4]. Breath biomarkers have the potential to revolutionize healthcare by providing a non-invasive and convenient method for diagnosing and monitoring a wide range of health conditions [5]. These biomarkers are identified by analyzing the volatile organic compounds (VOCs) in a person's exhaled breath, which can provide valuable information about their health status. One of the main advantages of using breath biomarkers is that it is a simple and painless procedure that can be easily repeated over time. Breath biomarkers have shown promise in diagnosing and monitoring a range of diseases [6], including cancer [7], diabetes, infectious diseases and COVID-19 [8,9,10,11].
Machine learning has the potential to revolutionize healthcare by improving patient outcomes, reducing costs, and advancing medical research [12]. It is being used for predictive analytics to identify risk factors and predict the likelihood of disease onset or progression, improve the accuracy of medical diagnoses, and drug discovery and development [13]. It is also being used for personalized medicine to identify the most effective treatments for individual patients, which can improve treatment outcomes and reduce the risk of adverse reactions [14, 15].
Previous research has shown that breath biomarkers can accurately diagnose liver fibrosis using a single VOC or a panel of VOCs [16, 17]. The breath test was found to be comparable in accuracy to traditional blood tests, but less invasive and more convenient for patients [10, 18]. To achieve maximum diagnostic accuracy, multiple biomarkers need to be used as individual VOC biomarkers linked to the cellular state may not differentiate between causative agents or symptoms. The liver is responsible for metabolism, and the liver disease affects multiple VOCs because it alters various metabolic pathways. These pathways are related to some VOCs of one or more functional groups [9, 19].
It has been seen that the variability in laboratory test results can lead to confusion in diagnosis, treatment, and disease monitoring. This variability is caused by pre-analytical variation, biological variation, and analytical variation. To avoid misinterpretation of results, it is essential to consider an individual's overall health status and medical history when interpreting laboratory test results [20]. A study is mentioned that highlights the importance of stable biomarkers for predicting schizophrenia in the human connectome [21].
The objective of this investigation is to identify a new panel of stable potential biomarkers using statistical and machine learning techniques that can accurately detect abnormal liver function, intending to improve diagnostic and prognostic tools for liver disease. Breath samples were collected from liver patients and healthy volunteers at different physiological states. Samples were collected at various physiological states to identify biomarkers that remain consistent across different conditions. By analyzing the samples collected at rest, after exercise, and during recovery, the study aimed to identify biomarkers that exhibit stability and reliability irrespective of the individual's physiological state. This approach enables the identification of robust biomarkers that can be consistently used for diagnostic or monitoring purposes. The samples were analyzed and quantified to obtain the existing compound names and their relative concentration. To obtain a stable and potential panel of biomarkers multiple strategies were adopted. At first, the common compounds among different physiological states were shortlisted. Then the common compounds were ranked based on the contribution to predicting the samples belong to the either patient or healthy group. After that a significance test was conducted to verify the statistical significance of the ranked best compounds. Finally, the compound which is commonly ranked best and stable or consistent is chosen in the panel of potential and stable biomarkers. The selected biomarkers are validated to determine their accuracy and reliability. The development of accurate and reliable diagnostic tests for the liver disease could improve patient outcomes and provide clinicians with a powerful tool for monitoring disease progression and developing personalized treatment plans.
2 Method
2.1 Experimental setup
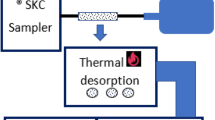
To identify common and consistent biomarkers for detecting abnormal liver function, breath samples were collected three times from two groups of study subjects consisting of 30 liver patients and 33 healthy individuals, recruited and tested at National Taiwan University Hospital, Taipei, Taiwan (REC: 201912138RINB). The Research Ethics Committee B of National Taiwan University Hospital, Taipei, Taiwan granted approval for the breath test protocol and research method. All methods were performed in accordance with the relevant guidelines and regulations to ensure ethical conduct of the research study. The healthy volunteers in the study were recruited voluntarily, and their participation involved conducting liver function tests at NTU Hospital. A preliminary liver donor criterion determines healthy participants based on blood test results, age, and BMI. In a recent study of 473 liver donors over ten years revealed that healthy participants are typically under 30 years old [22]. This age group is chosen to avoid older individuals with other health conditions that could affect the study's findings. On the other hand, the patients included in the study were hospitalized at NTU Hospital for diagnosis and treatment purposes of liver illness. This ensured that the participants in both the healthy and diseased groups underwent evaluation and monitoring within the controlled hospital setting, minimizing potential confounding effects related to environmental exposure in exhaled breath samples. In the supplementary file Figs. S1, S2 and S3 represents the distribution of the study-subjects Child-Pugh’s score, AST to PLT ratio (APRI) and model of end-stage liver disease (MELD) score, respectively. A significant portion of the patient group, specifically over 20 out of the total 30, exhibit low scores in the Child-Pugh, APRI, and MELD assessments. This pattern suggests a prevalence of initial stage patients within this subset, indicating that a substantial proportion of these individuals are in the early stages of their respective conditions. Exclusion criteria included patients with lung disease, those who were advised bed rest, and those who did not fit within the age range of 20 to 70 years. Study participants provided informed consent and performed staircase walking with a minimum heart rate of 100 bpm for two minutes. Breath samples were collected using the experimental setup shown in Fig. 1, with three bags of samples collected before exercise, after exercise, and after 15 min of rest. The collected exhaled breath from the bags is sampled into desorption tubes (Carbotrap, Perkin Elmer ®). The flow rate of a pump was adjusted to collect one liter of sample from the bag to the tube. Likely three tubes containing samples from three different bags are fitted to the automatic thermal desorption (ATD) unit, Turbomatrix Perkin Elmer ®. The sample from the tube was heated and collected in a trap unit. Then it is transferred into the gas chromatography (GC) (Claurus 680 Perkin Elmer ®) column and the output is detected using the mass spectrometer (MS). A detail regarding, the breath collection protocol and GC-MS setup were discussed in length in this article [17].
2.2 Data collection
Here we used the PerkinElmer TurboMass software® which is a data acquisition and analysis software for mass spectrometry used to analyze and process mass spectral data. The software is designed to work with PerkinElmer's mass spectrometry systems and can perform a range of functions such as data acquisition, instrument control, and data processing. It has a quantitative analysis method that can be linked with the National Institute of Standards and Technology (NIST) database. This feature allows users to generate quantitative reports for their samples based on the comparison of the sample spectrum with the NIST database. Then use the external calibration method, where a calibration curve is generated using a standard compound or compounds, and then the response values of the analytes in your sample are used to determine their concentrations. A four-point calibration is prepared using acetone as standard and the response value of various compounds is quantified to a relative concentration. This approach can be useful for a limited number of samples to quantify and quantify a variety of compounds in your sample simultaneously [23, 24]. All the concentration and compound names from the three bags are separated as three datasets with each containing data from both groups. The dataset has a positive class label for patients and a negative class label for healthy samples which makes it a binary classification problem. Every compound is treated as a feature and discussed in further sections.
2.3 Feature selection process
At first, the compounds which are found in at least 50% of the total number of samples are selected, this is a common approach in statistical analysis to ensure that only the most prevalent and consistent features are considered for further analysis. This is a type of frequency-based feature selection, which involves selecting features (in this case, compounds) that occur commonly found across the samples [25]. By adopting this method, noise and irrelevant features can be eliminated, and the most informative features can be identified for further analysis. This helps to avoid background noise, and residuals and increase the reliability of the results. A conceptual flowchart (Fig. 2) describes the feature selection process to select the final set of potential features or biomarkers to predict the samples belonging to the liver patient group.
The features which are common in the three datasets are considered for further steps. These features were then subjected to a recursive feature elimination (RFE) technique to rank the features and assess their impact on model accuracy [26]. RFE is a powerful method for identifying the most important features in a dataset for a given problem. Using a decision tree (100 estimators chosen), RFE works by repeatedly training a model on subsets of features and eliminating the least important ones until a desired number of features is reached. The algorithm ranks the features based on their contribution to model performance and evaluates various combinations of ranked features to identify the best ones. Additionally, fivefold cross-validation with 3 times repetition is conducted to ensure the analysis is robust and reliable. RFE with a decision tree estimator can improve model performance while minimizing overfitting. The model and data analysis methods were constructed using Python programming language, with Jupyter Notebook serving as the integrated development environment for the development process.
Then a reasonable number of features are selected with the possible maximum cross-validated accuracy achieved. The features are then subjected to a statistical test to verify the significant difference in each feature between two groups in an individual dataset. In order to assess the normality of the data distribution, the Shapiro-Wilk test was conducted. For group-wise significance analysis, non-parametric tests, specifically the Mann-Whitney U test, were employed. This test was chosen due to its robustness against non-normality and its suitability for comparing two independent groups. Furthermore, to explore the significance of features across different conditions of sample collection, the Kruskal-Wallis test was employed. This non-parametric test was utilized to evaluate potential differences among multiple independent groups, taking into account the non-normal distribution of the data [27].
The features which are selected using RFE and the features that showed a significant difference between the two groups (p < 0.01) and were present in all the datasets were selected. The common sort listed features are selected to find the consistent biomarkers. Again the Kruskall-Wallis test (p > 0.5) is adapted to verify the means of a compound is not significantly different in the three datasets. A Kruskall-Wallis test with a null hypothesis that the means of all the groups are equal if the Kruskall-Wallis test does not reject the null hypothesis, suggests that the means of all the groups are similar. In the end, the features which are ranked contributing (using RFE), significant between groups (p < 0.01), and common and consistent among datasets (p > 0.5) are the final features in the features set.
All possible combinations of input features are enumerated then, each subset of features trains a decision tree model. The accuracy of the model is evaluated using cross-validation with repeated stratification. This gives a detailed idea about the various combinations of best features selected using RFE techniques and the accuracy of various feature combinations.
2.4 Machine learning model
Additionally, the other two models, a simple Naïve-Bayes (NB) classifier, and a Random Forest (RF) classifier were trained and tested to gain insights into the strengths of the features selected and to validate the results of the analysis.
2.4.1 Naive bayes classifier
Naive Bayes is a popular machine learning algorithm based on the Bayes theorem. It predicts categorical data using probability theory and is efficient in handling high-dimensional data. The algorithm estimates the probability distribution of each input feature given the target class, assuming each feature is independent. When presented with new data, it calculates the posterior probability of each class and predicts the class with the highest probability as the output [28].
2.4.2 Random forest classifier
Random Forest is a powerful ensemble machine-learning algorithm that creates multiple decision trees by randomly selecting subsets of data and features. The trees are built using recursive partitioning and each independently predicts the class of a new data point. The final prediction is made by aggregating the predictions of all trees using a majority voting scheme, reducing the effects of individual incorrect predictions and leading to more accurate overall performance [29, 30]. The hyperparameters are kept at default value with 500 estimators.
2.5 Performance metrics
There are five important parameters for model evaluation accuracy, precision, recall (sensitivity), F1-score and specificity [31]. To evaluate model performance, accuracy is suitable for even datasets, but when dealing with uneven classes, F1-score is the better option. Precision indicates how well a model predicts a class label, while recall measures misclassified labels for a class. Depending on the type of misclassification, either recall or specificity is a better metric for evaluation.
To further assess the reliability of the best-performed model, a bootstrap and confidence interval method was applied to find a 95% confidence interval that covered the true skill of the model [32]. Finally, a classification report, Receiver Operating Characteristics (ROC), Precision-Recall (PR) Curve and their area under the curve (AUC) provide a comprehensive overview of the model's performance [33]. The three datasets are analyzed parallelly as described to verify the results obtained from the sample at different times are close enough.
3 Results
3.1 Feature selection result
The recruited study-subjects health status was confirmed at the hospital by conducting a blood test. The liver function test parameters data are shown in Table 1.
The quantified VOC concentration forms the dataset, in total, there are three datasets obtained from the collected 3 bags of all study subjects. A total of 36 compounds in Bag-1, 35 compounds in Bag-2 and 29 compounds are found in half of the samples. The 15 compounds which are common in the three datasets are considered for further analysis. Then three conditions are applied. First, in all datasets, the 15 common compounds are ranked using the RFE method. Then, the significance test is performed using Mann Whitney test and the compounds are marked with p < 0.01. At last the compounds which are contributing based on RFE analysis, significant features and their mean values are close (not significant) in all bags and are treated as stable VOCs forming the final panel of biomarkers. In the supplementary Table S1 describe the names of the common compounds, rank of the compounds measured at different physiological state and the significance test result. The following figure explains the RFE method result.
The box and whisker plot shown in Fig. 3 explains the effect of feature elimination on the model accuracy. In each features combination the cross-validation accuracy results for the possible combinations from a maximum of 15 to a minimum of 2 are shown. At every combination, the least ranked feature is removed from the combination. Overall, 6 to 2 combinations give the best combination. Those features are significantly different in both groups and all the datasets' mean values are close. All the features are ranked based on performance. A combination of three to five features was found to be contributing maximum accuracy (mean and medium), significant in discriminating the class in each bag, and common and consistent in three datasets with a not significant difference in the mean value.
Box plot showing the classification accuracy of decision tree models with varying numbers of selected features using RFE on bag 1 data a, bag 2 data b and bag 3 data c. The green diamond indicates the mean value, the red box represents the interquartile range (IQR), the whiskers extend to the lowest and highest values within 1.5 times the IQR, and the circles represent outliers
The Fig. 4 box and whisker plots provide valuable insights into the concentration of these compounds in liver patients and healthy individuals, as well as the variation in concentration across different sample bags. The p-value of their significance based on two said groups of study subjects are shown using the standard ‘*’ system [34]. Then all the possible combinations with three to five features are iterated, trained and tested with a decision tree model with 100 estimators and all the accuracies obtained are shown in Table 2.
The figure presents 15 box and whisker plots representing the concentration of five different compounds in three different sample bags (Bag-1, Bag-2, Bag-3) for both liver patients (red boxes) and healthy individuals (blue boxes). The compounds and their concentrations in parts per billion (ppb) are 2-Myristynoyl Pantetheine, N_Acetyl Cystine, Pterin-6 Carboxylic Acid, Butanoic Acid, and Methyl Mercaptan. The p-values are represented by asterisks, with *p < 0.05, **p < 0.01, and ***p < 0.001
Table 2 presents the results of different feature combinations for five compounds across three sample bags. The "Range of accuracy for all combinations" column shows the accuracy range achieved for all possible feature combinations. The "Best accuracy (combinations)" column shows the highest accuracy achieved, along with the corresponding feature combinations used. The "Accuracy with selected five features" column displays the accuracy obtained by using all five features together. These findings offer valuable insights into the effectiveness of various feature combinations for predicting the accuracy of the five compounds in different sample bags. Notably, any three combinations of the selected features achieved an accuracy of approximately a minimum of 0.77 for a decision tree model.
3.2 Classification model result
The five selected features are further used with the RF classifier and NB classifier for training and testing. The RF classifier with 500 estimators and NB classifier fitted with 70% of the training data. Then the trained models are tested with 30% of the data. The predicted class and real class are used to obtain the classification reports. Table 3 shows the classification reports from the RF classifier and NB classifier.
Table 3 shows the performance metrics of two different classifiers on three different bags of data (Bag-1, Bag-2, and Bag-3) for a binary classification problem. The metrics are presented separately for healthy and patient classes. Overall, the RF classifier performed better than the NB classifier in most cases. In Bag-1 and Bag-2, the precision, recall, F1 score, and accuracy of the RF classifier were consistently higher than those of the NB classifier for both healthy and patient classes. In Bag-3, however, both classifiers' performances close in terms of precision, recall, F1 score, and accuracy both classes.
The results obtained from bootstrapping and computing 95% confidence intervals for the classification model on the three datasets indicate the range of accuracies that can be expected when using the model on similar datasets. For the first dataset (Bag-1), the 95% confidence interval was 94.7%. This means that, with 95% confidence, the true accuracy of the model on similar datasets will be 94.7%. For the second dataset (Bag-2), the 95% confidence interval was 89.2%. This is a very narrow range and suggests that the model is highly accurate on this type of data. For the third dataset (Bag-3), the 95% confidence interval was 89.5%. For the NB model, the 95% confidence interval was 84.2% for all datasets. The ROC and PRC graphs of three datasets for the RF model and NB model are shown in Figs. 5 and 6.
Performance evaluation of the RF classifier on three independent test datasets (Bag 1, Bag 2, and Bag 3) using ROC and PR curves. a ROC curves for the RF classifier on Bag 1, Bag 2, and Bag 3 are shown, with the AUC indicated for each dataset. b PR curves for the RF classifier on Bag 1, Bag 2, and Bag 3 are shown, with the AUC indicated for each dataset
Performance evaluation of the NB classifier on three independent test datasets (Bag 1, Bag 2, and Bag 3) using ROC and PR curves. a ROC curves for the NB classifier on Bag 1, Bag 2, and Bag 3 are shown, with the AUC indicated for each dataset. b PR curves for the NB classifier on Bag 1, Bag 2, and Bag 3 are shown, with the AUC indicated for each dataset
4 Discussion
The concept of stability or consistency is relevant to the study of breath biomarkers in general, as stable biomarkers are necessary to ensure accurate and reliable diagnosis and monitoring of diseases. In this study, the breath samples are collected at three different physiological states to identify potential biomarkers for liver dysfunction. The common biomarkers are identified in all three samples and the RFE algorithm ranked them based on their ability to predict liver dysfunction. The significance of the biomarkers between healthy and patient classes was determined by conducting an Mann Whitney test. The stability and consistency of the biomarkers by checking their means are not significantly different in all sample bags. The findings suggest that this approach provides a unique and effective means of identifying stable biomarkers for liver dysfunction and may have broader implications for the development of biomarker-based diagnostic tools.
The RFE algorithm works by repeatedly training a model using subsets of the 15 common features and eliminating the least important features at each iteration. Figure 3(a–c) explains about a combination of three features gives the highest mean and median accuracy for all datasets. Selecting the top three ranked compounds across all datasets results in a list of five compounds (Table 2). The selected compounds are significant between the healthy and patent group. Their mean values are not significantly different among all datasets which are tested by considering.
Many compounds among the 15 common compounds are found significant and have potential as biomarkers but are not consistently found significant or stable in all cases. The Mann Whitney test qualified some common compounds are Acetone, Alkane, Toluene, Isopropyl alcohol, Ethyl acetate and Furan. Some abundant biomarkers are more in concentration are fluctuates with heart rate and the biomarkers with less concentration are not commonly found in all bags. Some biomarkers are not only potential biomarkers for liver disease but also for other diseases as described in other studies such as Acetone [16], Acetaldehyde, Alkanes [35], Toluene, Furan, Dimethyl Sulphide, and Terpenes [36].
The stable biomarkers that do not change with physical activity or heart rate were chosen as the final panel of biomarkers, and their possible source of origin, pathway, and relation to body metabolism are discussed from the available supporting literature. The name of the compounds listed in the panel of biomarkers are: 1) N- Acetyl Cystine, 2) 2-Myristynayl Pantetheine, 3) Pterin-6 Carboxylic Acid, 4) Butanoic Acid, and 5) Methyl Mercaptan.
N-Acetyl cysteine, a precursor for glutathione synthesis [37], can reduce inflammation, lower liver enzymes and risk of alcohol-induced liver damage [38, 39], and may benefit those with inadequate production or genetic variations affecting its metabolism [40, 41]. The patient group consistently shows lower levels of N-Acetyl Cystine across all bags compared to the healthy group may be because of ill-liver function. The patient group has relatively consistent low mean values of Myristoyl pantetheine across all three bags, while the healthy group displays consistently high levels in all bags. It is a derivative of coenzyme A (CoA) synthesized in the liver from pantothenic acid and cysteine [42, 43], plays a role in various metabolic pathways as a component of CoA and in its myristoylated form [44,45,46,47]. The results dictate that liver dysfunction may dysregulate the concentration of Myristoyl pantetheine. The patient group displays higher mean values of Pterin-6 Carboxylic Acid than the healthy group across all bags, while the healthy group shows consistently low levels in all bags. Pterin-6 Carboxylic Acid, a metabolite of tetrahydrobiopterin, which is involved in producing nitric oxide [48], may serve as a biomarker for the presence of tumours and cancer [49,50,51], though its association with liver disease is not well established, and altered liver function may affect its concentration in breath samples. The patient group has consistently low levels of butyric acid, while the healthy group has consistently high levels in all bags. It is suggested that the disruption of the gut microbiome in liver disease may play a role, leading to a decrease in butyrate-producing bacteria in the gut and hence lower levels of butyric acid. Additionally, impaired liver function may affect the metabolism and absorption of butyric acid in the body, contributing to lower levels in liver disease sample [52, 53]. The patient group has consistently higher levels of Methyl Mercaptan than the healthy group across all three bags. Methyl Mercaptan is a compound produced by gut bacteria when breaking down certain amino acids. The liver normally processes it, but if liver function is poor, it can build up and harm health. A study looked at mercaptans' role in inducing coma in liver disease or methanethiol gas exposure [54]. In cirrhosis patients have more mercaptans in their breath than healthy individuals, suggesting a connection between sulfur-containing amino acids and mercaptan production in liver disease [55].
The results of the three datasets with the selected biomarkers as features are quite similar, with high recall, F1 score, accuracy, and precision obtained for both classes. The balanced result indicates that the model is equally good at predicting both healthy and patient classes. The results obtained from all datasets are similar and the performance of the two models is also similar. A higher ROC AUC (more than 0.9 for all observations) results suggest that the model performed well in distinguishing between the positive and negative classes, with Bag2 having the highest AUC value of 0.989. The PRC AUC of the classification models is more than 0.9 for all datasets and the model’s combination. These results suggest that the model performed well in identifying positive examples with high precision, with Bag2 having the highest AUC value and F1 score. The results obtained from bootstrapping and computing 95% confidence intervals for the classification model on the three datasets indicate the range of accuracies that can be expected when using the model on similar datasets. For all the datasets and RF model, the 95% confidence interval is a minimum of 89% to 94% and for the NB model, the 95% confidence interval is 84%. This suggests that the model is likely to perform well on this type of data.
The five biomarkers have successfully met the requirements for RFE test qualification. Three conditions were applied to different classes, and statistical tests were run to determine their significance. These biomarkers also showed stability under various conditions, which is significant. The significance of the five biomarkers has been thoroughly investigated, both individually and in relation to possible combinations. The above data analysis procedure and result discussed shows the model potential to successfully monitoring the possibility of liver disease. This study highlights the findings of potential, stable and consistent biomarkers at different stage. The findings were further supported by use machine learning model to use them to predict liver function.
5 Conclusion
In conclusion, our study successfully identified 2-Myristynoyl Pantetheine, Pterin-6 Carboxylic Acid, Methyl Mercaptan, N-Acetyl Cystine, and Butanoic Acid as stable biomarkers found in breath profiling that has the potential to detect abnormal liver function. Our extensive GC-MS data analysis, statistical analysis, and feature selection technique enabled us to rank and select the most significant and consistent biomarkers. We iterated the final selected biomarkers for all combinations and identified the range of accuracy. Our results showed that the model test accuracy for various possible combinations of biomarkers ranged from 0.7 to 0.9 in all conditions. Moreover, the precision, recall, prediction probability, and 95% confidence interval ranged from 0.89 to 0.94 in all conditions. Our findings pave the way for future research in this field and provide a non-invasive approach to detecting potential biomarkers for various diseases.
Availability of data and materials
The datasets used and/or during the current study are available from the corresponding author on reasonable request.
Abbreviations
- VOC:
-
Volatile organic compound
- ATD:
-
Automatic thermal desorption
- GC:
-
Gas chromatography
- MS:
-
Mass spectrometer
- NIST:
-
National Institute of Standard and Technology
- RFE:
-
Recursive feature elimination
- NB:
-
Naïve Bayes
- RF:
-
Random Forest
- ROC:
-
Recursive operating characteristics
- PR:
-
Precision-Recall
- AUC:
-
Area under the curve
- IQR:
-
Interquartile range
- Ppb:
-
Parts per billion
- CoA:
-
Coenzyme A
References
Polaris T, Hcv O. Global prevalence and genotype distribution of hepatitis C virus infection in 2015: a modelling study. 2015;161–76.
Akinyemiju T, Abera S, Ahmed M, Alam N, Alemayohu MA, Allen C, et al. The Burden of Primary Liver Cancer and Underlying Etiologies From 1990 to 2015 at the Global, Regional, and National Level: Results From the Global Burden of Disease Study 2015. JAMA Oncol. 2017;3(12):1683–91.
Global hepatitis report. 2017. WHO. https://www.who.int/publications/i/item/global-hepatitis-report-2017.
Lavanchy D. The global burden of hepatitis C. Liver Int Off J Int Assoc Study Liver. 2009;29(Suppl 1):74–81.
Dragonieri S, Schot R, Mertens BJA, Le Cessie S, Gauw SA, Spanevello A, et al. An electronic nose in the discrimination of patients with asthma and controls. J Allergy Clin Immunol. 2007;120(4):856–62.
Moura PC, Raposo M, Vassilenko V. Breath volatile organic compounds (VOCs) as biomarkers for the diagnosis of pathological conditions: A review. Biomed J. 2023;46(4):100623.
Herman-Saffar O, Boger Z, Libson S, Lieberman D, Gonen R, Zeiri Y. Early non-invasive detection of breast cancer using exhaled breath and urine analysis. Comput Biol Med. 2018;96:227–32.
Filipiak W, Beer R, Sponring A, Filipiak A, Ager C, Schiefecker A, et al. Breath analysis for in vivo detection of pathogens related to ventilator-associated pneumonia in intensive care patients: a prospective pilot study. J Breath Res. 2015;9(1):16004.
De Lacy CB, Amann A, Al-Kateb H, Flynn C, Filipiak W, Khalid T, et al. A review of the volatiles from the healthy human body. J Breath Res. 2014;8(1):14001.
Pereira J, Porto-Figueira P, Cavaco C, Taunk K, Rapole S, Dhakne R, et al. Breath analysis as a potential and non-invasive frontier in disease diagnosis: an overview. Metabolites. 2015;5(1):3–55.
Chen Z, Li M, Wang R, Sun W, Liu J, Li H, et al. Diagnosis of COVID-19 via acoustic analysis and artificial intelligence by monitoring breath sounds on smartphones. J Biomed Inform. 2022;130:104078.
Aliper A, Plis S, Artemov A, Ulloa A, Mamoshina P, Zhavoronkov A. Deep Learning Applications for Predicting Pharmacological Properties of Drugs and Drug Repurposing Using Transcriptomic Data. Mol Pharm. 2016;13(7):2524–30.
Saria S, Goldenberg A. Subtyping: What It is and Its Role in Precision Medicine. IEEE Intell Syst. 2015;30(4):70–5.
Kim JH, Choi A, Kim MJ, Hyun H, Kim S, Chang H-J. Development of a machine-learning algorithm to predict in-hospital cardiac arrest for emergency department patients using a nationwide database. Sci Rep. 2022;12(1):21797.
McKinney SM, Sieniek M, Godbole V, Godwin J, Antropova N, Ashrafian H, et al. International evaluation of an AI system for breast cancer screening. Nature. 2020;577(7788):89–94.
Alkhouri N, Cikach F, Eng K, Moses J, Patel N, Yan C, et al. Analysis of breath volatile organic compounds as a noninvasive tool to diagnose nonalcoholic fatty liver disease in children. Eur J Gastroenterol Hepatol. 2014;26(1):82–7.
Patnaik RK, Lin Y-C, Agarwal A, Ho M-C, Yeh JA. A pilot study for the prediction of liver function related scores using breath biomarkers and machine learning. Sci Rep. 2022;12(1):2032.
De Vincentis A, Vespasiani-Gentilucci U, Sabatini A, Antonelli-Incalzi RPA. Exhaled breath analysis in hepatology: State-of-the-art and perspectives. World J Gastroenterol. 2019;25(30):4043–50.
Smoleńska Ż, Zdrojewski Z. Metabolomics and its potential in diagnosis, prognosis and treatment of rheumatic diseases. Reumatologia. 2015;53(3):152–6.
Pradhan S, Gautam K, Pant V. Variation in Laboratory Reports: Causes other than Laboratory Error. JNMA J Nepal Med Assoc. 2022;60(246):222–4.
Gutiérrez-Gómez L, Vohryzek J, Chiêm B, Baumann PS, Conus P, Cuenod K Do, et al. Stable biomarker identification for predicting schizophrenia in the human connectome. NeuroImage Clin. 2020;27:102316.
Ho CM, Huang YM, Hu RH, Wu YM, Ho MC, Lee PH. Revisiting donor risk over two decades of single-center experience: More attention on the impact of overweight. Asian J Surg. 2019;42(1):172–9.
Chu S, Haffner GD, Letcher RJ. Simultaneous determination of tetrabromobisphenol A, tetrachlorobisphenol A, bisphenol A and other halogenated analogues in sediment and sludge by high performance liquid chromatography-electrospray tandem mass spectrometry. J Chromatogr A. 2005;1097(1):25–32.
Busardò FP, Carlier J, Giorgetti R, Tagliabracci A, Pacifici R, Gottardi M, et al. Ultra-high-performance liquid chromatography-tandem mass spectrometry assay for quantifying fentanyl and 22 analogs and metabolites in whole blood, urine, and hair. Front Chem. 2019;7:1–13.
Christopher D. Manning, Prabhakar Raghavan and Hinrich Schütze, Introduction to Information Retrieval, Cambridge University Press. 2008. https://nlp.stanford.edu/IR-book/html/htmledition/frequency-based-feature-selection-1.html.
Kuhn M, Johnson K. Applied Predictive Modeling. Springer New York; 2013. https://books.google.com.tw/books?id=xYRDAAAAQBAJ.
Girden ER. ANOVA: Repeated Measures,. SAGE Publications; 1992. (ANOVA: Repeated Measures). https://books.google.com.tw/books?id=JomGKpjnfPcC.
Feng P-M, Ding H, Chen W, Lin H. Naïve Bayes classifier with feature selection to identify phage virion proteins. Comput Math Methods Med. 2013;2013:530696.
VanderPlas J. Python Data Science Handbook: Essential Tools for Working with Data. O’Reilly Media 2016. https://books.google.com.tw/books?id=6omNDQAAQBAJ.
Deist TM, Dankers FJWM, Valdes G, Wijsman R, Hsu I-C, Oberije C, et al. Machine learning algorithms for outcome prediction in (chemo)radiotherapy: An empirical comparison of classifiers. Med Phys. 2018;45(7):3449–59.
Mantas J, Hasman A, Househ MS. The Importance of Health Informatics in Public Health during a Pandemic. IOS Press; 2020. (Studies in Health Technology and Informatics). https://books.google.com.tw/books?id=GpP-DwAAQBAJ.
Clark JS. Model Assessment and Selection. Models for Ecol Data. 2020;143–160.
Ma Y, He H. Imbalanced Learning: Foundations, Algorithms, and Applications. Wiley; 2013. https://books.google.com.tw/books?id=CVHx-Gp9jzUC.
Al-Khelaifi F, Diboun I, Donati F, Botrè F, Alsayrafi M, Georgakopoulos C, et al. A pilot study comparing the metabolic profiles of elite-level athletes from different sporting disciplines. Sport Med - Open. 2018;4(1).
Aghdassi E, Allard JP. Breath alkanes as a marker of oxidative stress in different clinical conditions. Free Radic Biol Med. 2000;28(6):880–6.
Koureas M, Kirgou P, Amoutzias G, Hadjichristodoulou C, Gourgoulianis K, Tsakalof A. Target Analysis of Volatile Organic Compounds in Exhaled Breath for Lung Cancer Discrimination from Other Pulmonary Diseases and Healthy Persons. Metabolites. 2020;10(8).
Lu SC. Glutathione synthesis. Biochim Biophys Acta. 2013;1830(5):3143–53.
Dludla P V, Nkambule BB, Mazibuko-Mbeje SE, Nyambuya TM, Marcheggiani F, Cirilli I, et al. N-Acetyl Cysteine Targets Hepatic Lipid Accumulation to Curb Oxidative Stress and Inflammation in NAFLD: A Comprehensive Analysis of the Literature. Antioxidants (Basel, Switzerland). 2020;9(12).
Baniasadi S, Eftekhari P, Tabarsi P, Fahimi F, Raoufy MR, Masjedi MR, et al. Protective effect of N-acetylcysteine on antituberculosis drug-induced hepatotoxicity. Eur J Gastroenterol Hepatol. 2010;22(10):1235–8.
De Andrade KQ, Moura FA, dos Santos JM, de Araújo ORP, de Farias Santos JC, Goulart MOF. Oxidative Stress and Inflammation in Hepatic Diseases: Therapeutic Possibilities of N-Acetylcysteine. Int J Mol Sci. 2015;16(12):30269–308.
Gao B, Jeong W-I, Tian Z. Liver: An organ with predominant innate immunity. Hepatology. 2008;47(2):729–36.
Berg JM, Stryer L, Tymoczko JL, Gatto GJ. Biochemistry. Macmillan Learning; 2015. https://books.google.com.tw/books?id=5bjzrQEACAAJ.
Lehninger AL, Cox MM, Nelson DL, Lehninger Principles of Biochemistry. W. H. Freeman; 2005. https://books.google.com.tw/books?id=7chAN0UY0LYC.
Leonardi R, Jackowski S. Biosynthesis of Pantothenic Acid and Coenzyme A. EcoSal Plus. 2007;2(2).
Theodoulou FL, Sibon OCM, Jackowski S, Gout I. Coenzyme A and its derivatives: renaissance of a textbook classic. Biochem Soc Trans. 2014;42(4):1025–32.
Wagner GR, Payne RM. Widespread and enzyme-independent Nε-acetylation and Nε-succinylation of proteins in the chemical conditions of the mitochondrial matrix. J Biol Chem. 2013;288(40):29036–45.
Leonardi R, Zhang Y-M, Rock CO, Jackowski S. Coenzyme A: back in action. Prog Lipid Res. 2005;44(2–3):125–53.
Buglak AA, Kapitonova MA, Vechtomova YL, Telegina TA. Insights into Molecular Structure of Pterins Suitable for Biomedical Applications. Int J Mol Sci. 2022;23(23).
Kośliński P, Pluskota R, Mądra-Gackowska K, Gackowski M, Markuszewski MJ, Kędziora-Kornatowska K, et al. Comparison of Pteridine Normalization Methods in Urine for Detection of Bladder Cancer. Diagnostics. 2020;10(9).
Zvarik M, Martinicky D, Hunakova L, Sikurova L. Differences in pteridine urinary levels in patients with malignant and benign ovarian tumors in comparison with healthy individuals. J Photochem Photobiol B Biol. 2015;153:191–7.
Zeitler HJ, Andondonskaja-Renz B. Evaluation of pteridines in patients with different tumors. Cancer Detect Prev. 1987;10(1–2):71–9.
Zhao Z-H, Wang Z-X, Zhou D, Han Y, Ma F, Hu Z, et al. Sodium Butyrate Supplementation Inhibits Hepatic Steatosis by Stimulating Liver Kinase B1 and Insulin-Induced Gene. Cell Mol Gastroenterol Hepatol. 2021;12(3):857–71.
Zheng Z, Wang B. The Gut-Liver Axis in Health and Disease: The Role of Gut Microbiota-Derived Signals in Liver Injury and Regeneration. Front Immunol. 2021;12:775526.
Al Mardini H, Bartlett K, Record CO. Blood and brain concentrations of mercaptans in hepatic and methanethiol induced coma. Gut. 1984;25(3):284–90.
Chen S, Zieve L, Mahadevan V. Mercaptans and dimethyl sulfide in the breath of patients with cirrhosis of the liver. Effect of feeding methionine. J Lab Clin Med. 1970;75(4):628–35.
Acknowledgements
We thank members of Dr. Ming Chih Ho laboratory for helping us to carry out the test and patient recruitment. We also like to acknowledge Dr. Chen kuo Chiang for his valuable suggestion about data analysis.
Funding
This work is supported by the Ministry of Science and Technology, Taiwan research project funding (Project Grant: MOST 110-2221-E-007-068-MY3).
Author information
Authors and Affiliations
Contributions
Rakesh Kumar Patnaik did the study design, test the samples, and analysed the data. Yu-Chen Lin help to do equipment setup and sample collection. Ming-Chih Ho verified the study design, look after the data collection and confirm the health status of study participants. J. Andrew Yeh conceived, guided, and analysed the results. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Ethics approval and consent to participate and publication
We have obtained ethics approval and signed consent to participate and publish from all individuals involved in the research project. All study subjects' blood tests were conducted at National Taiwan University Hospital, Taipei, Taiwan. The Research Ethics Committee B passed this study design, reference REC: 201912138RINB.
Competing interests
The authors declare that they have no competing interests.
Disclaimer
All authors agree with the content of the article and approve of its submission to the journal. The material contained in the manuscript is a part of a study which is explore a new insight and is not being concurrently submitted elsewhere. The experiments reported in the article were undertaken in compliance with the current laws of the country in which the experiments were performed.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Patnaik, R., Lin, YC., Ho, M. et al. Selection of consistent breath biomarkers of abnormal liver function using feature selection: a pilot study. Health Technol. 13, 957–969 (2023). https://doi.org/10.1007/s12553-023-00787-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12553-023-00787-7