Abstract
The proposition of a post-antimicrobial era is all the more realistic with the continued rise of antimicrobial resistance. The development of new antimicrobials is failing to counter the ever-increasing rates of bacterial antimicrobial resistance. This necessitates novel antimicrobials and drug targets. The bacterial cell membrane is an essential and highly conserved cellular component in bacteria and acts as the primary barrier for entry of antimicrobials into the cell. Although previously under-exploited as an antimicrobial target, the bacterial cell membrane is attractive for the development of novel antimicrobials due to its importance in pathogen viability. Bacterial cell membranes are diverse assemblies of macromolecules built around a central lipid bilayer core. This lipid bilayer governs the overall membrane biophysical properties and function of its membrane-embedded proteins. This mini-review will outline the mechanisms by which the bacterial membrane causes and controls resistance, with a focus on alterations in the membrane lipid composition, chemical modification of constituent lipids, and the efflux of antimicrobials by membrane-embedded efflux systems. Thorough insight into the interplay between membrane-active antimicrobials and lipid-mediated resistance is needed to enable the rational development of new antimicrobials. In particular, the union of computational approaches and experimental techniques for the development of innovative and efficacious membrane-active antimicrobials is explored.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Antimicrobial resistance (AMR) is one of the foremost threats facing global public health. A 2016 review on antimicrobial resistance gave a conservative estimate of 700,000 deaths caused by AMR annually (O’Neil Jim 2016). As a result of the continuing rise of AMR infections in conjunction with limited advances in the development of novel antimicrobials, the number of AMR-related deaths is predicted to increase to an alarming 10 million annually by 2050 (O’Neil Jim 2016). Traditional antimicrobial therapeutics generally function either by altering bacterial cell wall synthesis (β-lactams, glycopeptides), inhibiting protein or nucleic acid synthesis (macrolides, quinolones and tetracyclines), or interfering with metabolic pathways (sulfonamides) (Reygaert 2018; Streicher 2021). Treatment strategies often use a combination of antimicrobials to simultaneously target multiple biochemical sites. Resistance to a single antimicrobial can arise through a myriad of point mutations in different genes, whilst resistance to combination therapies commonly occurs when several mutations are acquired which reduce or eliminate susceptibility to multiple antimicrobials, making the pathogen multidrug resistant (MDR) (Alekshun and Levy 2007). The ongoing acquisition of AMR against most currently available antimicrobials requires new and innovative solutions. The bacterial cell membrane is the primary barrier for antimicrobial entry to the cell and is a critical mediator of antimicrobial resistance and pathogen survival. As such, the bacterial cell membrane is an attractive target for the development of new antimicrobials (Moellering 2011; Hurdle et al. 2011; Mingeot-Leclercq and Décout 2016; Mehta et al. 2021).
In this mini-review, we outline the lipid-mediated mechanisms by which the bacterial cell membrane causes and controls resistance, particularly focusing on alterations of membrane lipid composition, the chemical modification of membrane lipids and the role of multidrug efflux systems. We also highlight the need to gain a comprehensive understanding of the interplay between lipid-mediated AMR and the mechanisms of antimicrobial action and efflux to guide the rational development of new and effective membrane-active antimicrobials.
The bacterial cell membrane and membrane-active antimicrobials
Bacterial membranes are dynamic and heterogenous assemblies of macromolecules that contain lipids, proteins and glycans, and act as the primary physical barrier for the cell entry of antimicrobial agents (Strahl and Errington 2017; May and Grabowicz 2018; Willdigg and Helmann 2021). Consequently, membrane-active antimicrobial agents which target the bacterial membrane, either by disrupting the functional integrity of the bacterial membrane itself or modulating the function of essential membrane-associated proteins, can greatly hinder bacterial viability (Zasloff 2002; Fjell et al. 2012; Mingeot-Leclercq and Décout 2016). Although there is widespread resistance to many current classes of antimicrobials, the highly conserved and essential nature of the bacterial membrane would suggest a reduced potential for bacteria to acquire resistance to membrane-active antimicrobials (Hurdle et al. 2011; Fjell et al. 2012; Spohn et al. 2019). Furthermore, as the bacterial membrane and its associated components act as the primary barriers for antimicrobial entry, targeting membrane function and integrity can allow for increased sensitivity to other co-administrated antimicrobials (Mingeot-Leclercq and Décout 2016; Pizzolato-Cezar et al. 2019). One such example is the synergy between azithromycin and the membrane-active antimicrobial peptides, LL-37 and colistin in the treatment of several MDR Gram-negative pathogenic bacteria (Lin et al. 2015).
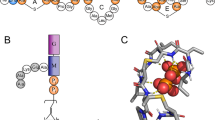
The different cellular architectures of Gram-positive and Gram-negative bacteria are critical to the functional mechanism of membrane-active antimicrobials (Fig. 1A and B). Gram-negative bacteria contain two membranes; an inner cytoplasmic membrane and an outer membrane, separated by a thin peptidoglycan layer. The inner membrane of Gram-negative bacteria is composed of lipids and integral membrane proteins, whilst the outer membrane also contains lipoproteins, porins and lipopolysaccharides (LPS), found primarily in the extracellular leaflet (Silhavy et al. 2010). In comparison, Gram-positive bacteria possess a single cytoplasmic membrane and a thicker peptidoglycan cell wall. The membrane of Gram-positive bacteria is comprised of phospholipids, integral and associated membrane proteins, and lipoteichoic acid (LTA) components that anchor the membrane to the cell wall (Silhavy et al. 2010). The additional barrier presented by the outer membrane in Gram-negative bacteria makes them a particularly challenging antimicrobial target and has contributed to the rise of numerous strains of MDR Gram-negative pathogens (Brown and Wright 2016). Indeed, nine out of the twelve bacterial pathogens listed by the WHO on their AMR priority list are Gram-negative (WHO 2017).
Lipid-mediated mechanisms of AMR. (A) Diagrammatic representation of the Gram-negative bacterial cell envelope and (B) the Gram-positive bacterial cell envelope. (C) Chemical modifications of Lipid A and (D) lysylation of PG provide antimicrobial resistance. (E) Lipid acyl tail remodelling, such as increases in overall membrane unsaturation levels lead to changes in membrane biophysical properties, and control antimicrobial susceptibility. (F) The transport of lipids between membranes and across membrane leaflets by lipid transport systems, including the Mla and Lpt complexes, governs membrane-antimicrobial interactions. (G) Membrane-embedded drug efflux pumps can actively efflux a range of antimicrobials and are regulated by the membrane lipid environment. Figure created with BioRender.com
Within bacterial membranes, the distribution and chemical composition of membrane lipids is highly varied across different species, and even different bacterial strains (Sohlenkamp and Geiger 2015; López-Lara and Geiger 2017). The precise lipid composition of a given membrane has clear implications for the modulation of membrane biophysical properties and the function of membrane-embedded proteins, which in turn governs the activity of membrane-active antimicrobials (Harayama and Riezman 2018; Lee et al. 2019b). Mammalian cell membranes are largely composed of phosphatidylcholine (PC) lipids and cholesterol, whilst bacterial membranes are rich in zwitterionic phosphatidylethanolamine (PE) lipids, anionic phosphatidylglycerol (PG) lipids and poly-anionic cardiolipin (CL). Within these major lipid classes, there is considerable species- and environment-dependent diversity in acyl chain lengths, degree of saturation, and the incorporation of branched chain, cyclopropane-containing, or ω-alicyclic fatty acyl chains (Oshima and Ariga 1975; Sohlenkamp and Geiger 2015; López-Lara and Geiger 2017). Additionally, there are a number of other unique lipid components associated with bacterial membranes. These include endotoxic LPS, composed of Lipid A and the polysaccharide O-antigen; Lipid II, a precursor for bacterial cell wall synthesis; and LTA, which anchors the cell wall of Gram-positive bacteria to the membrane (Epand and Epand 2009a; Silhavy et al. 2010). Notably, the differential distribution of lipid species between bacterial and mammalian membranes, as well as between bacterial species can be used to design membrane-active antimicrobials with highly specific bacterial selectivity (Dias and Rauter 2019).
Bacterial alterations to membrane lipid composition—the phantom menace
Both Gram-negative and Gram-positive pathogens can alter their membrane composition to adapt to their biological niche (Adams et al. 2021) or evade antimicrobials and host immune mechanisms (Epand and Epand 2009b; Hewelt-Belka et al. 2016; Hines et al. 2017; Sperandeo et al. 2019). These changes in their membranes are induced via transcriptional control of lipid synthesis, or through alterations to the active transport of lipids between membrane leaflets, and also between the inner and outer membranes of Gram-negative bacteria. Consequently, resistant bacterial strains often possess functional mutations in the proteins or regulatory elements controlling membrane composition, lipid synthesis, and lipid transport systems (Hachmann et al. 2009, 2011; Olaitan et al. 2014; May and Grabowicz 2018; Jiang et al. 2019a). As a result, lipid composition-dependent changes in membrane biophysical properties, such as surface charge, membrane thickness, membrane fluidity, and modulation of curvature are heavily implicated in antimicrobial resistance (Epand et al. 2016). In addition, changes in the surface display of signalling or structural lipids (Soto et al. 2019; Giordano et al. 2020), and lipid modifications that change the propensity of the membrane to form segregated domains (Epand and Epand 2009b) have also been highlighted as key determinants of AMR phenotypes.
Surface charge modification
Membrane surface charge plays a major role in controlling the susceptibility of bacteria to membrane-active antimicrobials (Table 1). As the major lipid species in bacterial membranes are zwitterionic PE, anionic PG and poly-anionic CL, the relative concentration of anionic PG and CL in the bacterial membrane imparts an overall negative surface charge. This provides an electrostatic basis for interactions with charged membrane-active antimicrobials, particularly cationic antimicrobial peptides (CAMPs). Consequently, changes in the relative ratios of these lipid classes alters the overall surface charge of the membrane and governs the strength of membrane interactions with antimicrobial agents. CAMP-resistant bacterial isolates have been observed to carry functional mutations in the pgsA (PG) or cls2 (CL) lipid synthesis enzymes, resulting in a net reduction in the overall negative charge of the membrane surface (Hachmann et al. 2011; Davlieva et al. 2013; Hines et al. 2017; Jiang et al. 2019a). This reduces the propensity of CAMPs to aggregate on the membrane surface, providing resistance against these antimicrobials.
Lipid chemical modifications
Another mechanism bacteria employ to reduce membrane surface charge is the chemical modification of constituent lipids (Fig. 1C and D, Table 1). Gram-negative pathogens readily modify Lipid A through a wide variety of enzymatic mechanisms, including dephosphorylation (Ingram et al. 2010; Cullen et al. 2015; Zhao et al. 2019), acylation (Guo et al. 1998), diacylation (Kawasaki et al. 2004), addition of various sugars and decoration with PE (Liu et al. 2017), glycine (Henderson et al. 2017), or aminoarabinose (Zhou et al. 2001; Raetz et al. 2007; Olaitan et al. 2014; Sperandeo et al. 2019). These Lipid A modifications also affect other membrane biophysical properties, altering LPS fluidity, packing, and interactions with divalent cations (Wu et al. 2013; Rice and Wereszczynski 2018). These properties are key in modulating interactions with antimicrobial agents, enabling AMR acquisition against outer membrane antimicrobials, such as the CAMP polymyxin (Olaitan et al. 2014). Gram-positive bacteria also possess methods for enzymatic modification of lipids to modulate membrane surface charge. For example, they can reduce CAMP susceptibility by increasing production of cationic lysyl-PG, which lessens interactions with cationic antimicrobials by increasing electrostatic repulsion (Willdigg and Helmann 2021). In particular, the MprF system, which catalyses the aminoacyl-tRNA-dependent lysylation of PG to lysyl-PG, also acts as a flippase (Ernst and Peschel 2011), translocating lysl-PG from the inner to the outer leaflet, which reduces the overall negative membrane surface charge and provides protection against CAMPs (Andrä et al. 2011; Mishra et al. 2011; Kumariya et al. 2015; Ernst et al. 2018; Sabat et al. 2018; Ernst and Peschel 2019). In addition to lysl-PG formation, resistance phenotypes can also exhibit the aminoacyl-tRNA dependent formation of cationic arginyl-PG or zwitterionic alanyl-PG, highlighting the range of chemical modification of membrane lipids (Sohlenkamp et al. 2007; Klein et al. 2009; Roy and Ibba 2009). Gram-positive bacteria also possess a range of other charge modification systems, including the Dlt system which modifies poly-anionic lipoteichoic acids (LTAs) through the transfer a D-alanine moiety, reducing the overall membrane surface charge and providing protection against CAMPs (McBride and Sonenshein 2011; Sabat et al. 2018).
Acyl tail remodelling
Lipid acyl tail remodelling (Fig. 1E, Table 1) impacts membrane thickness and membrane fluidity and has been implicated in resistance to antimicrobials (Aricha et al. 2004; Maria-Neto et al. 2015; Kumariya et al. 2015). Collectively, bacterial acyl tail remodelling includes changes in unsaturated fatty acid content and speciation, acyl tail length and the relative proportions of acyl tails that incorporate cyclopropane or branched lipid tails (Sohlenkamp and Geiger 2015; López-Lara and Geiger 2017). Experimental studies of lipid bilayers indicate longer, more saturated, and minimally branched lipid tails promote thick, ordered, compact membranes with slowly diffusing lipids (Filippov et al. 2007; Kučerka et al. 2011; Poger et al. 2014; Levental et al. 2016; Marquardt et al. 2016). This is important to consider as acyl tail induced changes to membrane fluidity and ordering have been implicated in resistance to AMPs. For example, increased unsaturation can inhibit the assembly of daptomycin oligomers, modulating daptomycin pore formation (Taylor et al. 2017; Beriashvili et al. 2018). Consequently, increased unsaturated fatty acid content has been observed in resistance to daptomycin and other lipophilic peptide antimicrobials, including colistin (Saito et al. 2014; Harp et al. 2016; Tao et al. 2021). Increases in unsaturation and concomitant decreases in cyclopropane containing lipids that provide resistance to apidaecin 1b or naringenin have been shown to alter membrane fluidity (Wang et al. 2018; Schmidt et al. 2018). In addition to changing membrane biophysical properties, acyl tail remodelling can occur as a secondary effect from changes to enzymatic pathways that provide resistance. For example, mutations in pgsA, resulting in decreased PG content, can also result in upstream accumulation of fatty acids and subsequent remodelling of bacterial lipid fatty acid profiles (Hines et al. 2017). The acyl tail profile of some pathogens is also influenced by the availability of exogenous fatty acids, such as host-derived long-chain polyunsaturated fatty acids which have antimicrobial properties (Yao and Rock 2015; Churchward et al. 2018; Adams et al. 2021; Kengmo Tchoupa et al. 2021). Acyl-tail remodelling induced by exogenous fatty acids has been associated with reduced fitness and increased antimicrobial susceptibility in several pathogens (Kengmo Tchoupa et al. 2021), raising the possibility of combining membrane-active antimicrobial therapy with exogenous fatty acids for increased efficacy.
Lipid domains and curvature
Some membrane-active antimicrobials target or have their activity modulated by segregated membrane lipid domains (Table 1). The formation of segregated lipid domains is influenced by bacterial membrane composition and biophysical properties, showing dependencies on headgroup content, intrinsic lipid curvature and relative tail order (Epand and Epand 2009b). In bacteria, curvature driven segregation of CL and PE domains at the cell poles is thought to play key roles in polar protein localisation (Matsumoto et al. 2006; Mileykovskaya and Dowhan 2009; Renner and Weibel 2011; Beltrán-Heredia et al. 2019). Some amphiphilic aminoglycosides target CL rich domains, including those at the poles, resulting in domain disassembly and disrupting overall cellular function (El Khoury et al. 2017; Swain et al. 2018). Interactions of antimicrobials with CL rich domains are proposed to be driven by a combination of high negative charge density, high intrinsic curvature and the unique domain segregation preferences of CL (El Khoury et al. 2017; Swain et al. 2018). Increases in glycolipid content are also observed in some antimicrobial resistant strains (Hines et al. 2017). In experimental studies of model lipid bilayers, increases in glycolipid content results in the formation of ordered domains with high structural integrity (Levental et al. 2020) which may play a role in modulating antimicrobial mediated membrane disruption. In addition to targeting pre-existing membrane domains, the mechanism of action of several antimicrobials involves either the induction of altered lateral lipid domains to destabilise the membrane, or the formation of domains that disrupt protein localisation and function (Epand and Epand 2009a; Scheinpflug et al. 2017; Su et al. 2020).
Lipid transport systems—a further menace
The transport of lipids between membranes or across membrane leaflets can mediate changes in membrane structure and biophysical properties that limit the interaction of antimicrobials with the membrane, affecting antimicrobial susceptibility (Fig. 1F) (May and Grabowicz 2018; Bogdanov et al. 2020; Paulowski et al. 2020). In the outer membrane of Gram-negative bacteria, a number of regulatory systems help maintain the highly asymmetric distribution of lipopolysaccharides between the inner and outer leaflet, required for both membrane integrity and resistance to antimicrobials (May and Grabowicz 2018). In the Gram-negative outer membrane, the phospholipase PldA plays an integral role in membrane homeostasis and membrane asymmetry, by directly degrading phospholipids mis-localised to the outer leaflet of the outer membrane (May and Silhavy 2018). Between the Gram-negative inner and outer membranes, the Mla and Lpt transport systems are integral to the transport of lipids and lipopolysaccharides, respectively (Malinverni and Silhavy 2009; Sperandeo et al. 2017). These systems are critical for bacterial fitness and antimicrobial resistance, and mutations in these systems are present in AMP resistant strains (Lewis et al. 2009; Spohn et al. 2019). Other membrane homeostasis pathways involve the Lipid A palmitoyltransferase, PagP, which transfers a palmitoyl chain from mis-localised lipids to Lipid A (Guo et al. 1998; Bishop 2005; Boll et al. 2015). As previously noted, the MprF system is key in regulating lysyl-PG translocation between the inner and outer leaflets of the Gram-positive membrane (Ernst and Peschel 2011). All of these mechanisms function to preserve the lipid-dependent biophysical properties of bacterial membranes that allow them to function as a key protective barrier. As such, these protein systems are possible targets for future antimicrobial development.
Membrane efflux pumps—the bacterial empire strikes back
Another pertinent mechanism of membrane-mediated AMR is the active efflux of compounds from the bacterial cell membrane by membrane-embedded transport proteins that act as drug efflux pumps (Fig. 1G) (Henderson et al. 2021). Bacteria can upregulate the expression of drug efflux pumps to efflux a range of substrates, including antimicrobial peptides, lipids, and other antimicrobials (Du et al. 2018). Drug efflux pumps are not only involved in the active extrusion of lipid substrates, but their functioning is also linked to their membrane lipid microenvironment (Fig. 1G inset) (Corradi et al. 2019; Stieger et al. 2021). There are seven key drug efflux superfamilies present in bacteria: the ABC (ATP binding cassette), RND (resistance-nodulation-cell-division), MFS (major facilitator superfamily), MATE (multidrug and toxic compound extrusion), DMT (drug/metabolite transporter), PACE (proteobacterial antimicrobial compound efflux), and AbgT (p-aminobenzoyl-glutamate transporter) families (Henderson et al. 2021). While many of these transporters are found in both Gram-positive and Gram-negative bacteria, Gram-negative bacteria also contain tripartite efflux systems, in which a RND, ABC or MFS inner membrane transporter is coupled to an outer membrane porin or channel via a periplasmic coupling protein. These tripartite efflux systems are a powerful first-defence mechanism for efficient antimicrobial efflux across both the inner and outer membrane of Gram-negative bacteria (Henderson et al. 2021).
Although antimicrobials have only been in widespread use since the 1940s, the ubiquitous distribution of drug efflux pumps across bacterial species indicates they have underlying physiological roles in addition to antimicrobial efflux. Many of these efflux pumps are also associated with membrane homeostasis, or act as virulence factors by facilitating bacterial colonization through the efflux of xenobiotics such as host-derived hormones, signaling molecules or fatty acids (Henderson et al. 2021) (Table 2). In Escherichia coli, the ABC multidrug efflux pump MsbA plays a critical role in the transport of Lipid A and LPS for assembly in the cell envelope (Mi et al. 2017; Voss and Stephen Trent 2018). MsbA also effluxes the antimicrobial ethidium (Singh et al. 2016), and the drug daunorubicin (Siarheyeva and Sharom 2009). Investigations into MsbA-substrate interactions show daunorubicin binding decreases the binding affinity of Lipid A to MsbA (Siarheyeva and Sharom 2009). This suggests that in the presence of drugs and antimicrobials, the affinity of MsbA for its natural lipid substrates is decreased, instead facilitating the preferential efflux of antimicrobials.
ABC efflux pumps
There is mounting evidence that the function and modulation of many membrane proteins, including drug efflux pumps, is influenced by the local membrane environment and its biophysical properties (Corradi et al. 2019) (Table 2). A key example of this is the archetypal eukaryotic multidrug efflux pump, P-glycoprotein (P-gp, ABCB1), a member of the ABC transporter superfamily. P-gp function is increased in the presence of membrane cholesterol and modulated by the overall ordering and phase of the membrane (Rothnie et al. 2001; Sharom 2014). This previous work on eukaryotic drug transporters has provided evidence for a conserved functional mechanism of lipid modulation of bacterial ABC multidrug efflux pumps (Neumann et al. 2017). For example, mass spectrometry analysis of the ABC multidrug efflux pump TmrAB, from Thermus thermophilus, has shown that the integrity of the lipid annulus and the high-affinity binding of annular PG lipids to TmrAB is essential for both the structural integrity of the transporter and its ability to hydrolyse ATP to power antimicrobial efflux (Bechara et al. 2015).
RND efflux pumps
Bacterial RND multidrug efflux pumps actively efflux a range of membrane associated substrates (Table 2). For example, in the human pathogens Neisseria gonorrhoeae and Neisseria meningitidis, the RND efflux pump, MtrD, is essential for both bacterial virulence and the efflux of a range of host-derived, lipid-based antimicrobials including bile salts, progesterone and fatty acids, as well as other pharmaceutical antimicrobials (Shafer et al. 1998; Warner et al. 2008), and the structurally diverse antimicrobial peptides LL-37, protegrin-1 and polymyxin B (Tzeng et al. 2005). Recent simulation studies of MtrD have demonstrated that the binding of the antimicrobial hormone progesterone to MtrD induces allosteric couplings that govern efflux (Fairweather et al. 2021), and that these allosteric couplings can be deregulated by mutations that impact substrate uptake and the orientation of MtrD within the membrane (Chitsaz et al. 2021). Additionally, the Klebsiella pneumoniae AcrAB RND multidrug efflux pump is implicated in resistance to a number of antimicrobial peptides, including polymyxin B (Padilla et al. 2010).
The function of RND efflux pumps is also modulated by their membrane lipid environment. Recent studies linking the membrane composition and AMR in Acinetobacter baumannii have revealed an important link between membrane homeostasis, antimicrobial susceptibility and RND efflux pump function (Jiang et al. 2019b; Zang et al. 2021). Modifications of the membrane lipid composition due to incorporation of host-derived polyunsaturated fatty acids in bacterial lipid synthesis gave decreased resistance to antimicrobials. Analysis of A. baumannii efflux systems indicated that AdeABC-mediated efflux was impacted by this change in membrane lipid composition, due to a reduction in membrane ordering that abrogated the integrity of the protein–protein interface between AdeB subunits, disrupting AdeB functioning (Zang et al. 2021). Lipid modulation of RND transporter functioning has also been demonstrated for the E. coli AcrB RND transporter, whose interaction with a small regulatory protein AcrZ was modulated by the presence of CL, thus resulting in increased sensitivity to chloramphenicol in the absence of CL (Du et al. 2020).
Other efflux pump families
Lipid modulation of other efflux pump families has also been noted (Table 2). The MFS efflux pump LmrP from Lactococcus lactis undergoes proton-dependent conformational transitions during antimicrobial efflux. These conformational changes are sensitive to the presence of PE lipids and CL; however, differences in acyl chain length do not impact conformational switching (Martens et al. 2016). Likewise, the Pyrococcus furiosus ion-coupled MATE antimicrobial antiporter, pfMATE, is also strictly lipid dependent: the protein is only functional when reconstituted in a lipid environment (not in detergent), highlighting functional dependence on the presence of lipids (Zakrzewska et al. 2019; Jagessar et al. 2020). Additionally, the assembly of an E. coli ion-coupled small multidrug resistance (SMR) transporter EmrE, responsible for resistance to aromatic cationic compounds, is dependent on its lipid environment (Schuldiner 2009; Nathoo et al. 2013; Dutta et al. 2014).
Collectively, these examples highlight the growing body of evidence demonstrating that the lipid environment is integral to the function and modulation of many multidrug efflux pumps that are critical to the development of AMR. The importance of this must be considered to gain a holistic understanding of the impact of the membrane in modulating multidrug resistance.
Understanding the role of lipid-mediated resistance for antimicrobial development—a new hope
The development of membrane-active antimicrobials is a multi-faceted area of research. Extensive links between the efficacy of membrane-active antimicrobials and lipid-mediated AMR means that successful antimicrobial development must be founded on a detailed understanding of membrane dynamics, membrane biophysical properties and the functional mechanisms of transporters involved in AMR. This is best achieved using a combination of experimental techniques and computational approaches (Fig. 2A) (Fjell et al. 2012; Lopes et al. 2017; Li et al. 2017). Although, experimental methods, such as nuclear magnetic resonance, X-ray crystallography, circular dichroism and cryo-EM provide valuable structural information about bacterial cell membranes and membrane proteins, they are often limited by either temporal or spatial resolution. In silico approaches, such as molecular dynamics simulations, allow for high resolution characterisation of the functional dynamics and interactions between antimicrobials and their membrane targets (Berglund et al. 2015; Ulmschneider and Ulmschneider 2018; Palmer et al. 2021). Thus, iterative combinations of molecular simulations and experimental methodologies will allow for the streamlined, innovative, and rational design of novel antimicrobials (Li et al. 2015; Chen et al. 2019).
Guiding the development of novel membrane-active antimicrobials. (A) Iterative cycles of in vivo and in vitro experiments, combined with experimental biophysical techniques, such as NMR and X-ray crystallography, and computational approaches allows for the streamlined and rational design of novel antimicrobials. (B) Modified AMPs, such as lipoglycopeptides with added lipophilic groups allow for enhanced membrane interactions. (C) The inhibition of drug efflux systems and (D) lipid transport systems are emerging membrane-associated drug targets. (E) Host-derived fatty acids can have antimicrobial activity against membranes via changes to membrane biophysical properties. (F) Non-peptide cationic antimicrobials, such as metal nanoparticles and steroid-based antimicrobials can interact with negatively charged bacterial membranes to exert their antimicrobial properties. Figure created with BioRender.com
Antimicrobial peptides
The continued exploration into naturally occurring antimicrobial peptides (AMPs) from the innate host immune response has created a vast natural library of membrane-active compounds (Ashby et al. 2017). Whilst AMPs generally carry a net positive charge, their large variations in size (between ~ 9 and 100 amino acids) and structural diversity (they can be linear or cyclical and adopt α-helical, β-sheet or mixed structures) (Shai 2002) makes AMPs and AMP-based structural modifications a promising future direction for expanding the current range of antimicrobials. Additionally, AMPs can act as conjunct therapeutics to restore antimicrobial sensitivity in resistant bacteria (Lin et al. 2015; Pizzolato-Cezar et al. 2019). Growing computational and experimental insight into the interactions between AMPs and the membrane has already allowed for the development of modified AMPs with improved action (Fig. 2B). For example, lipoglycopeptides such as telavancin, oritavancin and dalbavancin are derivatives of earlier generation glycopeptides, with the addition of lipophilic alkyl and aryl groups to the sugar subunit. These lipophilic moieties act as membrane anchoring groups, enhancing interactions with the hydrophobic lipid tails, resulting in membrane permeabilization and the loss of membrane integrity (Blaskovich et al. 2018).
Efflux pump inhibitors
Membrane-active peptides are also inhibitors of some multidrug efflux pumps, opening avenues for the inhibition of efflux pump systems via the co-administration of known or engineered peptides (Fig. 2C). This strategy has been effective against the RND efflux pump AcrB in E. coli (Jesin et al. 2020), and against an SMR efflux protein of Pseudomonas aeruginosa, PAsmr, inhibiting antimicrobial efflux (Mitchell et al. 2019). These and other efflux pump inhibitors (EPIs), including PAβN, pyridopyrimidines, quinoline derivatives, arylpiperidines and arylpiperazines have been identified as having the potential to abrogate resistance in a number of problematic pathogens (Kabra et al. 2019). Continued exploration of the peptide EPI sequence space, coupled with improved knowledge of the functional mechanisms of drug efflux pumps, and their relationship with membrane composition and membrane biophysical properties will better enable the development of highly efficacious and specific membrane-active AMPs for resistant bacterial pathogens.
Targeting lipid synthesis and transport
Another area for the development of membrane-active agents includes the targeting of proteins involved in lipid synthesis and membrane homeostasis (Fig. 2D). Potential protein targets include the condensing enzymes of the fatty acid biosynthesis cycle (FabH, FabB, FabF); the sn-glycerol-3-phosphate acyltransferase (PlsB) involved in linking fatty acids to the phospholipid glycerol-3-phosphate backbone; or members of the Lpt complex involved in transport of LPS to the extracellular leaflet of the membrane (Heath et al. 2001; Heath and Rock 2004; Choi and Lee 2019; Walker and Black 2021). For example, the outer membrane assembly inhibitor, murepavadin, which is based on the protegrin-1 AMP, is a likely inhibitor of members of the Lpt complex (Lehman and Grabowicz 2019). Further exploration of lipid synthesis and transport inhibitors is needed to better target this under-exploited avenue for membrane-active antimicrobial development.
Host-derived fatty acids
Host-derived fatty-acids, including lauric and sapienic acid, can also act as membrane-active antimicrobials (Fig. 2E) (Thormar and Hilmarsson 2007; Fischer 2020). However, the mechanism of action of these fatty acids is highly varied, and includes the disruption of oxidative phosphorylation via binding to membrane embedded proteins, alteration of membrane biophysical properties and permeability, and the inhibition of enzymes involved in fatty acid synthesis pathways (Desbois and Smith 2010). Detailed computational and experimental investigation of the mechanisms by which host-derived antimicrobial fatty-acids disrupt the bacterial membrane may enable the renewal of existing therapeutics through co-administration of host-derived membrane-active fatty acids.
Non-peptide cationic antimicrobials
Due to the high concentration of negatively charged lipids in bacterial membranes, there is increasing interest in the development of non-peptide cationic antimicrobial agents that use a combination of electrostatics and lipophilicity to bind to and disrupt bacterial membranes (Fig. 2F). One such application is the design of membrane-disrupting metal nanoparticles and metal ion clusters which increase membrane permeability and promote oxidation of bacterial membrane lipids (Godoy-Gallardo et al. 2021). Cationic steroid antimicrobials such as ceragenins also utilise this mechanism. Ceragenins are synthetic sterol-based compounds decorated with amino acids or other chemical groups to aid lipid partitioning. These cationic compounds cause membrane permeabilization and depolarization. Importantly, the efficacy of ceragenins is impacted by membrane PE and anionic lipid content (Epand et al. 2007, 2010). As with AMPs, understanding of the biophysical mechanisms by which non-peptide cationic antimicrobials disrupt the bacterial membrane will aid in their development and enhance their effectiveness.
Computational approaches
Molecular dynamics simulations are poised to play an essential role in elucidating the membrane and transporter dynamics critical to lipid-based AMR. Importantly, bacterial lipid composition can now be examined using high resolution mass-spectrometry techniques allowing unprecedented insight into the lipid composition of bacterial membranes and responses to antimicrobials (Rustam and Reid 2018; Appala et al. 2020). When combined with recent advances in cryo-EM structural elucidation, structure-based drug design, as well as improved computational hardware and algorithms, these advances allow for the development of detailed molecular models that can reveal the mechanisms that underpin lipid-mediated AMR in bacteria. Molecular simulation of bacterial membranes, and biological membranes in general, has advanced significantly over the past 20 years, with larger, more complex and more realistic simulations becoming feasible (Marrink et al. 2019; Wilson et al. 2020b; Im and Khalid 2020). Advances in forcefields, particularly the CHARMM atomistic forcefield and MARTINI coarse-grained forcefield (Marrink et al. 2004, 2007; Wu et al. 2014; Lee et al. 2019a), coupled with increasing hardware capabilities and improvements in simulation tooling, has enabled simulations of membranes approaching realistic chemical diversity in constituent protein and lipid components (Ingólfsson et al. 2014, 2017; Reddy et al. 2015; Wilson et al. 2020a, 2021).
For example, large scale coarse-grained simulations have identified the preferential interaction of some AMPs with ordered domains in phase separated membranes (Su et al. 2020). Molecular dynamics simulations have also been used to show the preference of AMPs to bind in regions of high membrane curvature, as well as the curvature-dependence of AMP secondary structure (Chen and Mark 2011). Simulation work on the mechanisms governing lipid modification induced reduction in AMP binding has examined the role of lysl-PG in AMP surface binding and membrane disruption in a model bacterial membrane (Simcock et al. 2021). Molecular insights into transporter regulation by lipid components and membrane biophysical properties derived from simulations is gaining momentum (Corradi et al. 2018, 2019; Corey et al. 2021) and has recently been applied to AdeB and AdeJ transporter regulation in A. baumannii (Zang et al. 2021). Advanced simulations of Gram-negative membranes now contain the inner and outer membrane, LPS and peptidoglycan layers, and multiple copies of inner and outer membrane proteins, enabling even more detailed insights into the molecular mechanisms of lipid-based resistance (Im and Khalid 2020).
Conclusion
Lipid-mediated antimicrobial resistance is a multi-faceted phenomenon with inter-dependencies arising not only from the lipid membrane composition, but also from bacterial metabolic pathways, sequestration of lipids from the host environment, enzymatic modifications of lipid targets and the activity of drug efflux pumps. These interact synergistically to alter antimicrobial targets, decrease the specificity of membrane-active antimicrobials and actively efflux antimicrobial agents, increasing the inherent AMR of bacteria. As a result of rapidly escalating rates of AMR, there is continued interest in the development of membrane-active antimicrobials that may restore efficacy of now redundant antimicrobials or present new avenues for antimicrobial therapy. When combined with the improvements in the resolution of experimental methods to determine membrane lipid composition, membrane protein structural elucidation and advances in molecular simulation, the stage is set to develop more effective approaches to overcome lipid-mediated antimicrobial resistance.
Data availability
Not applicable.
Code availability
Not applicable.
References
Adams FG, Trappetti C, Waters JK et al (2021) To make or take: Bacterial lipid homeostasis during infection. Mbio 12:e0092821. https://doi.org/10.1128/mBio.00928-21
Alekshun MN, Levy SB (2007) Molecular Mechanisms of Antibacterial Multidrug Resistance. Cell 128:1037–1050. https://doi.org/10.1016/j.cell.2007.03.004
Andrä J, Goldmann T, Ernst CM et al (2011) Multiple Peptide Resistance Factor (MprF)-mediated Resistance of Staphylococcus aureus against Antimicrobial Peptides Coincides with a Modulated Peptide Interaction with Artificial Membranes Comprising Lysyl-Phosphatidylglycerol. J Biol Chem 286:18692–18700. https://doi.org/10.1074/JBC.M111.226886
Appala K, Bimpeh K, Freeman C, Hines KM (2020) Recent applications of mass spectrometry in bacterial lipidomics Anal Bioanal Chem 1–9https://doi.org/10.1007/s00216-020-02541-8
Aricha B, Fishov I, Cohen Z et al (2004) Differences in membrane fluidity and fatty acid composition between phenotypic variants of Streptococcus pneumoniae. J Bacteriol 186:4638–4644. https://doi.org/10.1128/JB.186.14.4638-4644.2004
Ashby M, Petkova A, Gani J et al (2017) Use of Peptide Libraries for Identification and Optimization of Novel Antimicrobial Peptides. Curr Top Med Chem 17:537–553. https://doi.org/10.2174/1568026616666160713125555
Bechara C, Nöll A, Morgner N et al (2015) (2015) A subset of annular lipids is linked to the flippase activity of an ABC transporter. Nat Chem 73(7):255–262. https://doi.org/10.1038/nchem.2172
Beltrán-Heredia E, Tsai FC, Salinas-Almaguer S et al (2019) Membrane curvature induces cardiolipin sorting. Commun Biol 2:1–7. https://doi.org/10.1038/s42003-019-0471-x
Berglund NA, Piggot TJ, Jefferies D et al (2015) Interaction of the Antimicrobial Peptide Polymyxin B1 with Both Membranes of E. coli: A Molecular Dynamics Study. PLOS Comput Biol 11:e1004180. https://doi.org/10.1371/JOURNAL.PCBI.1004180
Beriashvili D, Taylor R, Kralt B et al (2018) Mechanistic studies on the effect of membrane lipid acyl chain composition on daptomycin pore formation. Chem Phys Lipids 216:73–79. https://doi.org/10.1016/J.CHEMPHYSLIP.2018.09.015
Bishop RE (2005) The lipid A palmitoyltransferase PagP: molecular mechanisms and role in bacterial pathogenesis. Mol Microbiol 57:900–912. https://doi.org/10.1111/J.1365-2958.2005.04711.X
Blaskovich MAT, Hansford KA, Butler MS et al (2018) Developments in Glycopeptide Antibiotics. ACS Infect Dis 4:715–735. https://doi.org/10.1021/acsinfecdis.7b00258
Bogdanov M, Pyrshev K, Yesylevskyy S et al (2020) Phospholipid distribution in the cytoplasmic membrane of Gram-negative bacteria is highly asymmetric, dynamic, and cell shape-dependent. Sci Adv 6:eaaz6333. https://doi.org/10.1126/SCIADV.AAZ6333
Boll JM, Tucker AT, Klein DR et al (2015) Reinforcing lipid A acylation on the cell surface of acinetobacter baumannii promotes cationic antimicrobial peptide resistance and desiccation survival. Mbio 6:1–11. https://doi.org/10.1128/mBio.00478-15
Brown ED, Wright GD (2016) Antibacterial drug discovery in the resistance era. Nature 529:336–343. https://doi.org/10.1038/nature17042
Chen CH, Starr CG, Troendle E et al (2019) Simulation-Guided Rational de Novo Design of a Small Pore-Forming Antimicrobial Peptide. J Am Chem Soc 141:4839–4848. https://doi.org/10.1021/jacs.8b11939
Chen R, Mark AE (2011) The effect of membrane curvature on the conformation of antimicrobial peptides: implications for binding and the mechanism of action. Eur Biophys J 40:545–553. https://doi.org/10.1007/s00249-011-0677-4
Chitsaz M, Gupta V, Harris B et al (2021) A Unique Sequence Is Essential for Efficient Multidrug Efflux Function of the MtrD Protein of Neisseria gonorrhoeae. Mbio 12:e01675-e1721. https://doi.org/10.1128/mbio.01675-21
Choi U, Lee CR (2019) Antimicrobial agents that inhibit the outer membrane assembly machines of gram-negative bacteria. J Microbiol Biotechnol 29:1–10. https://doi.org/10.4014/jmb.1804.03051
Churchward CP, Alany RG, Snyder LAS (2018) Alternative antimicrobials: the properties of fatty acids and monoglycerides. Crit Rev Microbiol 44:561–570. https://doi.org/10.1080/1040841X.2018.1467875
Corey RA, Sansom MSP, Stansfeld PJ (2021) Identification and Characterization of Specific Protein-Lipid Interactions Using Molecular Simulation. Methods in Molecular Biology. Humana, New York, NY, pp 121–139. https://doi.org/10.1007/978-1-0716-1468-6_8
Corradi V, Mendez-Villuendas E, Ingólfsson HI et al (2018) Lipid-protein interactions are unique fingerprints for membrane proteins. ACS Cent Sci 4:709–717. https://doi.org/10.1021/acscentsci.8b00143
Corradi V, Sejdiu BI, Mesa-Galloso H et al (2019) Emerging Diversity in Lipid-Protein Interactions. Chem Rev 119:5775–5848. https://doi.org/10.1021/acs.chemrev.8b00451
Cullen TW, Schofield WB, Barry NA et al (2015) Antimicrobial peptide resistance mediates resilience of prominent gut commensals during inflammation. Science 80(347):170–175. https://doi.org/10.1126/SCIENCE.1260580
Davlieva M, Zhang W, Arias CA, Shamoo Y (2013) Biochemical characterization of cardiolipin synthase mutations associated with daptomycin resistance in enterococci. Antimicrob Agents Chemother 57:289–296. https://doi.org/10.1128/AAC.01743-12
Desbois AP, Smith VJ (2010) Antibacterial free fatty acids: activities, mechanisms of action and biotechnological potential. Appl Microbiol Biotechnol 85:1629–1642. https://doi.org/10.1007/s00253-009-2355-3
Dias C, Rauter AP (2019) Membrane-targeting antibiotics: Recent developments outside the peptide space. Future Med Chem 11:211–228. https://doi.org/10.4155/fmc-2018-0254
Du D, Neuberger A, Orr MW et al (2020) Interactions of a Bacterial RND Transporter with a Transmembrane Small Protein in a Lipid Environment. Structure 28:625-634.e6. https://doi.org/10.1016/J.STR.2020.03.013
Du D, Wang-Kan X, Neuberger A et al (2018) (2018) Multidrug efflux pumps: structure, function and regulation. Nat Rev Microbiol 169(16):523–539. https://doi.org/10.1038/s41579-018-0048-6
Dutta S, Morrison EA, Henzler-Wildman KA (2014) EmrE dimerization depends on membrane environment. Biochim Biophys Acta - Biomembr 1838:1817–1822. https://doi.org/10.1016/j.bbamem.2014.03.013
El Khoury M, Swain J, Sautrey G et al (2017) Targeting bacterial cardiolipin enriched microdomains: An antimicrobial strategy used by amphiphilic aminoglycoside antibiotics. Sci Rep 7:10697. https://doi.org/10.1038/s41598-017-10543-3
Epand RF, Pollard JE, Wright JO et al (2010) Depolarization, bacterial membrane composition, and the antimicrobial action of ceragenins. Antimicrob Agents Chemother 54:3708–3713. https://doi.org/10.1128/AAC.00380-10
Epand RF, Savage PB, Epand RM (2007) Bacterial lipid composition and the antimicrobial efficacy of cationic steroid compounds (Ceragenins). Biochim Biophys Acta - Biomembr 1768:2500–2509. https://doi.org/10.1016/j.bbamem.2007.05.023
Epand RM, Epand RF (2009a) Lipid domains in bacterial membranes and the action of antimicrobial agents. Biochim Biophys Acta - Biomembr 1788:289–294. https://doi.org/10.1016/j.bbamem.2008.08.023
Epand RM, Epand RF (2009b) Domains in bacterial membranes and the action of antimicrobial agents. Mol Biosyst 5:580–587. https://doi.org/10.1039/B900278M
Epand RM, Walker C, Epand RF, Magarvey N (2016) Molecular mechanisms of membrane targeting antibiotics. Biochim Biophys Acta 1858:980–987. https://doi.org/10.1016/J.BBAMEM.2015.10.018
Ernst CM, Peschel A (2011) Broad-spectrum antimicrobial peptide resistance by MprF-mediated aminoacylation and flipping of phospholipids. Mol Microbiol 80:290–299. https://doi.org/10.1111/j.1365-2958.2011.07576.x
Ernst CM, Peschel A (2019) MprF-mediated daptomycin resistance. Int J Med Microbiol 309:359–363. https://doi.org/10.1016/J.IJMM.2019.05.010
Ernst CM, Slavetinsky CJ, Kuhn S et al (2018) Gain-of-function mutations in the phospholipid flippase mprf confer specific daptomycin resistance. Mbio 9:1–12. https://doi.org/10.1128/MBIO.01659-18
Fairweather SJ, Gupta V, Chitsaz M et al (2021) Coordination of Substrate Binding and Protonation in the N. gonorrhoeae MtrD Efflux Pump Controls the Functionally Rotating Transport Mechanism. ACS Infect Dis 7:1833–1847. https://doi.org/10.1021/ACSINFECDIS.1C00149
Filippov A, Orädd G, Lindblom G (2007) Domain formation in model membranes studied by Pulsed-Field Gradient-NMR: The role of lipid polyunsaturation. Biophys J 93:3182–3190. https://doi.org/10.1529/BIOPHYSJ.107.111534
Fischer CL (2020) Antimicrobial activity of host-derived lipids Antibiotics 9https://doi.org/10.3390/antibiotics9020075
Fjell CD, Hiss JA, Hancock REW, Schneider G (2012) Designing antimicrobial peptides: Form follows function. Nat Rev Drug Discov 11:37–51. https://doi.org/10.1038/nrd3591
Giordano NP, Cian MB, Dalebroux ZD (2020) Outer membrane lipid secretion and the innate immune response to gram-negative bacteria Infect Immun 88https://doi.org/10.1128/IAI.00920-19
Godoy-Gallardo M, Eckhard U, Delgado LM et al (2021) Antibacterial approaches in tissue engineering using metal ions and nanoparticles: From mechanisms to applications. Bioact Mater 6:4470–4490. https://doi.org/10.1016/J.BIOACTMAT.2021.04.033
Guo L, Lim KB, Poduje CM et al (1998) Lipid A acylation and bacterial resistance against vertebrate antimicrobial peptides. Cell 95:189–198. https://doi.org/10.1016/S0092-8674(00)81750-X
Hachmann A-B, Sevim E, Gaballa A et al (2011) Reduction in Membrane Phosphatidylglycerol Content Leads to Daptomycin Resistance in Bacillus subtilis. Antimicrob Agents Chemother 55:4326. https://doi.org/10.1128/AAC.01819-10
Hachmann AB, Angert ER, Helmann JD (2009) Genetic analysis of factors affecting susceptibility of bacillus subtilis to daptomycin. Antimicrob Agents Chemother 53:1598–1609. https://doi.org/10.1128/AAC.01329-08
Harayama T, Riezman H (2018) Understanding the diversity of membrane lipid composition. Nat Rev Mol Cell Biol 19:281–296. https://doi.org/10.1038/nrm.2017.138
Harp JR, Saito HE, Bourdon AK et al (2016) Exogenous fatty acids protect Enterococcus faecalis from daptomycin-induced membrane stress independently of the response regulator LiaR. Appl Environ Microbiol 82:4410–4420. https://doi.org/10.1128/AEM.00933-16
Heath RJ, Rock CO (2004) Fatty acid biosynthesis as a target for novel antibacterials. Curr Opin Investig Drugs 5:146
Heath RJ, White SW, Rock CO (2001) Lipid biosynthesis as a target for antibacterial agents. Prog Lipid Res 40:467–497. https://doi.org/10.1016/S0163-7827(01)00012-1
Henderson JC, Herrera CM, Trent MS (2017) AlmG, responsible for polymyxin resistance in pandemic Vibrio cholerae, is a glycyltransferase distantly related to lipid A late acyltransferases. J Biol Chem 292:21205–21215. https://doi.org/10.1074/jbc.RA117.000131
Henderson PJF, Maher C, Elbourne LDH et al (2021) Physiological Functions of Bacterial “Multidrug” Efflux Pumps. Chem Rev 121:5417–5478. https://doi.org/10.1021/ACS.CHEMREV.0C01226
Hewelt-Belka W, Nakonieczna J, Belka M et al (2016) Untargeted Lipidomics Reveals Differences in the Lipid Pattern among Clinical Isolates of Staphylococcus aureus Resistant and Sensitive to Antibiotics. J Proteome Res 15:914–922. https://doi.org/10.1021/acs.jproteome.5b00915
Hines KM, Waalkes A, Penewit K, et al (2017) Characterization of the Mechanisms of Daptomycin Resistance among Gram-Positive Bacterial Pathogens by Multidimensional Lipidomics. mSphere 2 https://doi.org/10.1128/msphere.00492-17
Hurdle JG, O’Neill AJ, Chopra I, Lee RE (2011) Targeting bacterial membrane function: an underexploited mechanism for treating persistent infections. Nat Rev Microbiol 9:62–75. https://doi.org/10.1038/NRMICRO2474
Im W, Khalid S (2020) Molecular simulations of Gram-negative bacterial membranes come of age. Annu Rev Phys Chem 71:171–188. https://doi.org/10.1146/annurev-physchem-103019-033434
Ingólfsson HI, Carpenter TS, Bhatia H et al (2017) Computational lipidomics of the neuronal plasma membrane. Biophys J 113:2271–2280. https://doi.org/10.1016/j.bpj.2017.10.017
Ingólfsson HI, Melo MN, Van Eerden FJ et al (2014) Lipid organization of the plasma membrane. J Am Chem Soc 136:14554–14559. https://doi.org/10.1021/ja507832e
Ingram BO, Sohlenkamp C, Geiger O, Raetz CRH (2010) Altered lipid A structures and polymyxin hypersensitivity of Rhizobium etli mutants lacking the LpxE and LpxF phosphatases. Biochim Biophys Acta - Mol Cell Biol Lipids 1801:593–604. https://doi.org/10.1016/j.bbalip.2010.02.001
Jagessar KL, Claxton DP, Stein RA, Mchaourab HS (2020) Sequence and structural determinants of ligand-dependent alternating access of a MATE transporter. Proc Natl Acad Sci 117:4732–4740. https://doi.org/10.1073/PNAS.1917139117
Jesin JA, Stone TA, Mitchell CJ et al (2020) Peptide-Based Approach to Inhibition of the Multidrug Resistance Efflux Pump AcrB. Biochemistry 59:3973–3981. https://doi.org/10.1021/acs.biochem.0c00417
Jiang JH, Bhuiyan MS, Shen HH et al (2019a) Antibiotic resistance and host immune evasion in Staphylococcus aureus mediated by a metabolic adaptation. Proc Natl Acad Sci U S A 116:3722–3727. https://doi.org/10.1073/pnas.1812066116
Jiang JH, Hassan KA, Begg SL, et al (2019b) Identification of novel Acinetobacter baumannii host fatty acid stress adaptation strategies Mbio 10https://doi.org/10.1128/mBio.02056-18
Kabra R, Chauhan N, Kumar A et al (2019) Efflux pumps and antimicrobial resistance: Paradoxical components in systems genomics. Prog Biophys Mol Biol 141:15–24. https://doi.org/10.1016/j.pbiomolbio.2018.07.008
Kawasaki K, Ernst RK, Miller SI (2004) 3-O-Deacylation of Lipid A by PagL, a PhoP/PhoQ-regulated Deacylase of Salmonella typhimurium, Modulates Signaling through Toll-like Receptor 4. J Biol Chem 279:20044–20048. https://doi.org/10.1074/JBC.M401275200
Kengmo Tchoupa A, Eijkelkamp BA, Peschel A (2021) Bacterial adaptation strategies to host-derived fatty acids. Trends Microbiol. https://doi.org/10.1016/j.tim.2021.06.002
Klein S, Lorenzo C, Hoffmann S et al (2009) Adaptation of Pseudomonas aeruginosa to various conditions includes tRNA-dependent formation of alanyl-phosphatidylglycerol. Mol Microbiol 71:551–565. https://doi.org/10.1111/J.1365-2958.2008.06562.X
Kučerka N, Nieh MP, Katsaras J (2011) Fluid phase lipid areas and bilayer thicknesses of commonly used phosphatidylcholines as a function of temperature. Biochim Biophys Acta - Biomembr 1808:2761–2771. https://doi.org/10.1016/j.bbamem.2011.07.022
Kumariya R, Sood SK, Rajput YS, et al (2015) Increased membrane surface positive charge and altered membrane fluidity leads to cationic antimicrobial peptide resistance in Enterococcus faecalis. 1848:1367–1375. https://doi.org/10.1016/j.bbamem.2015.03.007
Lee J, Patel DS, Ståhle J et al (2019a) CHARMM-GUI Membrane Builder for Complex Biological Membrane Simulations with Glycolipids and Lipoglycans. J Chem Theory Comput 15:775–786. https://doi.org/10.1021/acs.jctc.8b01066
Lee TH, Hofferek V, Separovic F et al (2019b) The role of bacterial lipid diversity and membrane properties in modulating antimicrobial peptide activity and drug resistance. Curr Opin Chem Biol 52:85–92. https://doi.org/10.1016/j.cbpa.2019.05.025
Lehman KM, Grabowicz M (2019) Countering Gram-Negative Antibiotic Resistance: Recent Progress in Disrupting the Outer Membrane with Novel Therapeutics Antibiotics 8https://doi.org/10.3390/antibiotics8040163
Levental I, Levental KR, Heberle FA (2020) Lipid Rafts: Controversies Resolved, Mysteries Remain. Trends Cell Biol 30:341–353. https://doi.org/10.1016/j.tcb.2020.01.009
Levental KR, Lorent JH, Lin X et al (2016) Polyunsaturated lipids regulate membrane domain stability by tuning membrane order. Biophys J 110:1800–1810. https://doi.org/10.1016/j.bpj.2016.03.012
Lewis LA, Choudhury B, Balthazar JT et al (2009) Phosphoethanolamine substitution of lipid A and resistance of Neisseria gonorrhoeae to cationic antimicrobial peptides and complement-mediated killing by normal human serum. Infect Immun 77:1112–1120. https://doi.org/10.1128/IAI.01280-08
Li J, Koh JJ, Liu S et al (2017) Membrane active antimicrobial peptides: Translating mechanistic insights to design. Front Neurosci 11:73. https://doi.org/10.3389/fnins.2017.00073
Li J, Liu S, Koh J-J et al (2015) A novel fragment based strategy for membrane active antimicrobials against MRSA. Biochim Biophys Acta - Biomembr 1848:1023–1031. https://doi.org/10.1016/j.bbamem.2015.01.001
Lin L, Nonejuie P, Munguia J et al (2015) Azithromycin Synergizes with Cationic Antimicrobial Peptides to Exert Bactericidal and Therapeutic Activity Against Highly Multidrug-Resistant Gram-Negative Bacterial Pathogens. EBioMedicine 2:690–698. https://doi.org/10.1016/J.EBIOM.2015.05.021
Liu YY, Chandler CE, Leung LM, et al (2017) Structural modification of lipopolysaccharide conferred by mcr-1 in gram-negative ESKAPE pathogens Antimicrob Agents Chemother 61https://doi.org/10.1128/AAC.00580-17
Lopes D, Jakobtorweihen S, Nunes C et al (2017) Shedding light on the puzzle of drug-membrane interactions: Experimental techniques and molecular dynamics simulations. Prog Lipid Res 65:24–44. https://doi.org/10.1016/j.plipres.2016.12.001
López-Lara IM, Geiger O (2017) Bacterial lipid diversity. Biochim Biophys Acta - Mol Cell Biol Lipids 1862:1287–1299. https://doi.org/10.1016/J.BBALIP.2016.10.007
Malinverni JC, Silhavy TJ (2009) An ABC transport system that maintains lipid asymmetry in the Gram-negative outer membrane. Proc Natl Acad Sci 106:8009–8014. https://doi.org/10.1073/PNAS.0903229106
Maria-Neto S, de Almeida KC, Macedo MLR, Franco OL (2015) Understanding bacterial resistance to antimicrobial peptides: From the surface to deep inside. Biochim Biophys Acta - Biomembr 1848:3078–3088. https://doi.org/10.1016/j.bbamem.2015.02.017
Marquardt D, Heberle FA, Greathouse DV et al (2016) Lipid bilayer thickness determines cholesterols location in model membranes. Soft Matter 12:9417–9428. https://doi.org/10.1039/c6sm01777k
Marrink SJ, Corradi V, Souza PCTT et al (2019) Computational modeling of realistic cell membranes. Chem Rev 119:6184–6226. https://doi.org/10.1021/acs.chemrev.8b00460
Marrink SJ, De Vries AH, Mark AE (2004) Coarse grained model for semiquantitative lipid simulations. J Phys Chem B 108:750–760. https://doi.org/10.1021/jp036508g
Marrink SJ, Risselada HJ, Yefimov S et al (2007) The MARTINI force field: Coarse grained model for biomolecular simulations. J Phys Chem B 111:7812–7824. https://doi.org/10.1021/jp071097f
Martens C, Stein RA, Masureel M et al (2016) (2016) Lipids modulate the conformational dynamics of a secondary multidrug transporter. Nat Struct Mol Biol 238(23):744–751. https://doi.org/10.1038/nsmb.3262
Matsumoto K, Kusaka J, Nishibori A, Hara H (2006) Lipid domains in bacterial membranes. Mol Microbiol 61:1110–1117. https://doi.org/10.1111/J.1365-2958.2006.05317.X
May KL, Grabowicz M (2018) The bacterial outer membrane is an evolving antibiotic barrier. Proc Natl Acad Sci 115:8852–8854. https://doi.org/10.1073/PNAS.1812779115
May KL, Silhavy TJ (2018) The Escherichia coli phospholipase PldA regulates outer membrane homeostasis via lipid signaling Mbio 9https://doi.org/10.1128/MBIO.00379-18
McBride SM, Sonenshein AL (2011) The dlt operon confers resistance to cationic antimicrobial peptides in Clostridium difficile. Microbiology 157:1457. https://doi.org/10.1099/MIC.0.045997-0
Mehta D, Saini V, Aggarwal B, et al (2021) Unlocking the bacterial membrane as a therapeutic target for next-generation antimicrobial amphiphiles Mol Aspects Med 100999https://doi.org/10.1016/j.mam.2021.100999
Mi W, Li Y, Yoon SH et al (2017) (2017) Structural basis of MsbA-mediated lipopolysaccharide transport. Nat 5497671(549):233–237. https://doi.org/10.1038/nature23649
Mileykovskaya E, Dowhan W (2009) Cardiolipin membrane domains in prokaryotes and eukaryotes. Biochim Biophys Acta - Biomembr 1788:2084–2091. https://doi.org/10.1016/J.BBAMEM.2009.04.003
Mingeot-Leclercq M-P, Décout J-L (2016) Bacterial lipid membranes as promising targets to fight antimicrobial resistance, molecular foundations and illustration through the renewal of aminoglycoside antibiotics and emergence of amphiphilic aminoglycosides. Medchemcomm 7:586–611. https://doi.org/10.1039/C5MD00503E
Mishra NN, McKinnell J, Yeaman MR et al (2011) In Vitro cross-resistance to daptomycin and host defense cationic antimicrobial peptides in clinical methicillin-resistant Staphylococcus aureus isolates. Antimicrob Agents Chemother 55:4012–4018. https://doi.org/10.1128/AAC.00223-11
Mitchell CJ, Stone TA, Debera CM (2019) Peptide-Based Efflux Pump Inhibitors of the Small Multidrug Resistance Protein from Pseudomonas aeruginosa. Antimicrob Agents Chemother 63:e00730-e819. https://doi.org/10.1128/AAC.00730-19
Moellering RC (2011) Discovering new antimicrobial agents. Int J Antimicrob Agents 37:2–9. https://doi.org/10.1016/j.ijantimicag.2010.08.018
Nathoo S, Litzenberger JK, Bay DC et al (2013) Visualizing a multidrug resistance protein, EmrE, with major bacterial lipids using Brewster angle microscopy. Chem Phys Lipids 167–168:33–42. https://doi.org/10.1016/j.chemphyslip.2013.01.007
Neumann J, Rose-Sperling D, Hellmich UA (2017) Diverse relations between ABC transporters and lipids: An overview. Biochim Biophys Acta - Biomembr 1859:605–618. https://doi.org/10.1016/J.BBAMEM.2016.09.023
O’Neil Jim (2016) Tackling Drug-Resistance Infections Globally: Final Report and Recommendations.
Olaitan AO, Morand S, Rolain J-M (2014) Mechanisms of polymyxin resistance: acquired and intrinsic resistance in bacteria Front Microbiol 5https://doi.org/10.3389/FMICB.2014.00643
Oshima M, Ariga T (1975) Omega-cyclohexyl fatty acids in acidophilic thermophilic bacteria. Studies on their presence, structure, and biosynthesis using precursors labeled with stable isotopes and radioisotopes. J Biol Chem 250:6963–6968
Padilla E, Llobet E, Doménech-Sánchez A et al (2010) Klebsiella pneumoniae AcrAB Efflux Pump Contributes to Antimicrobial Resistance and Virulence. Antimicrob Agents Chemother 54:177–183. https://doi.org/10.1128/AAC.00715-09
Palmer N, Maasch JRMA, Torres MDT, De La Fuente-Nunez C (2021) Molecular dynamics for antimicrobial peptide discovery. Infect Immun 89:e00703-e720. https://doi.org/10.1128/IAI.00703-20
Paulowski L, Donoghue A, Nehls C et al (2020) The Beauty of Asymmetric Membranes: Reconstitution of the Outer Membrane of Gram-Negative Bacteria. Front Cell Dev Biol 0:586. https://doi.org/10.3389/FCELL.2020.00586
Pizzolato-Cezar LR, Okuda-Shinagawa NM, Machini MT (2019) Combinatory Therapy Antimicrobial Peptide-Antibiotic to Minimize the Ongoing Rise of Resistance. Front Microbiol 10:1703. https://doi.org/10.3389/FMICB.2019.01703
Poger D, Caron B, Mark AE (2014) Effect of Methyl-Branched Fatty Acids on the Structure of Lipid Bilayers. J Phys Chem B 118:13838–13848. https://doi.org/10.1021/JP503910R
Raetz CRH, Reynolds CM, Trent MS, Bishop RE (2007) Lipid A modification systems in gram-negative bacteria. Annu Rev Biochem 76:295–329. https://doi.org/10.1146/annurev.biochem.76.010307.145803
Reddy T, Shorthouse D, Parton DL et al (2015) Nothing to Sneeze At: A Dynamic and Integrative Computational Model of an Influenza A Virion. Structure 23:584–597. https://doi.org/10.1016/j.str.2014.12.019
Renner LD, Weibel DB (2011) Cardiolipin microdomains localize to negatively curved regions of Escherichia coli membranes. Proc Natl Acad Sci 108:6264–6269. https://doi.org/10.1073/PNAS.1015757108
Reygaert WC (2018) An overview of the antimicrobial resistance mechanisms of bacteria. AIMS Microbiol 4:482. https://doi.org/10.3934/MICROBIOL.2018.3.482
Rice A, Wereszczynski J (2018) Atomistic Scale Effects of Lipopolysaccharide Modifications on Bacterial Outer Membrane Defenses. Biophys J 114:1389–1399. https://doi.org/10.1016/J.BPJ.2018.02.006
Rothnie A, Theron D, Soceneantu L et al (2001) The importance of cholesterol in maintenance of P-glycoprotein activity and its membrane perturbing influence. Eur Biophys J 30:430–442. https://doi.org/10.1007/s002490100156
Roy H, Ibba M (2009) Broad Range Amino Acid Specificity of RNA-dependent Lipid Remodeling by Multiple Peptide Resistance Factors. J Biol Chem 284:29677–29683. https://doi.org/10.1074/JBC.M109.046367
Rustam YH, Reid GE (2018) Analytical Challenges and Recent Advances in Mass Spectrometry Based Lipidomics. Anal Chem 90:374–397. https://doi.org/10.1021/acs.analchem.7b04836
Sabat AJ, Tinelli M, Grundmann H et al (2018) Daptomycin Resistant Staphylococcus aureus Clinical Strain With Novel Non-synonymous Mutations in the mprF and vraS Genes: A New Insight Into Daptomycin Resistance. Front Microbiol 6:2705. https://doi.org/10.3389/FMICB.2018.02705
Saito HE, Harp JR, Fozo EM (2014) Incorporation of exogenous fatty acids protects Enterococcus faecalis from membrane-damaging agents. Appl Environ Microbiol 80:6527 LP – 6538. https://doi.org/10.1128/AEM.02044-14
Scheinpflug K, Wenzel M, Krylova O, et al (2017) Antimicrobial peptide cWFW kills by combining lipid phase separation with autolysis Sci Rep 7https://doi.org/10.1038/srep44332
Schmidt R, Yonghong D, Hoffmann R (2018) Phospholipid composition of the outer membrane of Escherichia coli influences its susceptibility against antimicrobial peptide apidaecin 1b. Diagn Microbiol Infect Dis 90:316–323. https://doi.org/10.1016/j.diagmicrobio.2017.11.008
Schuldiner S (2009) EmrE, a model for studying evolution and mechanism of ion-coupled transporters. Biochim Biophys Acta - Proteins Proteomics 1794:748–762. https://doi.org/10.1016/j.bbapap.2008.12.018
Shafer WM, Qu XD, Waring AJ, Lehrer RI (1998) Modulation of Neisseria gonorrhoeae susceptibility to vertebrate antibacterial peptides due to a member of the resistance/nodulation/division efflux pump family. Proc Natl Acad Sci U S A 95:1829–1833. https://doi.org/10.1073/pnas.95.4.1829
Shai Y (2002) Mode of action of membrane active antimicrobial peptides. Biopolym - Pept Sci Sect 66:236–248. https://doi.org/10.1002/bip.10260
Sharom FJ (2014) Complex Interplay between the P-Glycoprotein Multidrug Efflux Pump and the Membrane: Its Role in Modulating Protein Function. Front Oncol 0:41. https://doi.org/10.3389/FONC.2014.00041
Siarheyeva A, Sharom FJ (2009) The ABC transporter MsbA interacts with lipid A and amphipathic drugs at different sites. Biochem J 419:317–328. https://doi.org/10.1042/BJ20081364
Silhavy TJ, Kahne D, Walker S (2010) The Bacterial Cell Envelope Cold Spring Harb Perspect Biol 2https://doi.org/10.1101/CSHPERSPECT.A000414
Simcock PW, Bublitz M, Cipcigan F et al (2021) Membrane Binding of Antimicrobial Peptides Is Modulated by Lipid Charge Modification. J Chem Theory Comput 17:1218–1228. https://doi.org/10.1021/ACS.JCTC.0C01025
Singh H, Velamakanni S, Deery MJ et al (2016) (2016) ATP-dependent substrate transport by the ABC transporter MsbA is proton-coupled. Nat Commun 71(7):1–11. https://doi.org/10.1038/ncomms12387
Sohlenkamp C, Galindo-Lagunas KA, Guan Z et al (2007) The lipid lysyl-phosphatidylglycerol is present in membranes of Rhizobium tropici CIAT899 and confers increased resistance to polymyxin B under acidic growth conditions. Mol Plant-Microbe Interact 20:1421–1430. https://doi.org/10.1094/MPMI-20-11-1421
Sohlenkamp C, Geiger O (2015) Bacterial membrane lipids: Diversity in structures and pathways. FEMS Microbiol Rev 40:133–159. https://doi.org/10.1093/femsre/fuv008
Soto MJ, Calatrava-Morales N, López-Lara IM (2019) Functional Roles of Non-membrane Lipids in Bacterial Signaling Biog Fat Acids, Lipids Membr 273–289https://doi.org/10.1007/978-3-319-50430-8_16
Sperandeo P, Martorana AM, Polissi A (2017) The lipopolysaccharide transport (Lpt) machinery: A nonconventional transporter for lipopolysaccharide assembly at the outer membrane of Gram-negative bacteria. J Biol Chem 292:17981. https://doi.org/10.1074/JBC.R117.802512
Sperandeo P, Polissi A, De Fabiani E (2019) Fat Matters for Bugs: How Lipids and Lipid Modifications Make the Difference in Bacterial Life. Eur J Lipid Sci Technol 121:1900204. https://doi.org/10.1002/ejlt.201900204
Spohn R, Daruka L, Lázár V et al (2019) Integrated evolutionary analysis reveals antimicrobial peptides with limited resistance. Nat Commun 10:1–13. https://doi.org/10.1038/s41467-019-12364-6
Stieger B, Steiger J, Locher KP (2021) Membrane lipids and transporter function. Biochim Biophys Acta - Mol Basis Dis 1867:166079. https://doi.org/10.1016/j.bbadis.2021.166079
Strahl H, Errington J (2017) Bacterial Membranes: Structure, Domains, and Function. Annu Rev Microbiol 71:519–538. https://doi.org/10.1146/annurev-micro-102215-095630
Streicher LM (2021) Exploring the future of infectious disease treatment in a post-antibiotic era: A comparative review of alternative therapeutics. J Glob Antimicrob Resist 24:285–295. https://doi.org/10.1016/J.JGAR.2020.12.025
Su J, Marrink SJ, Melo MN (2020) Localization Preference of Antimicrobial Peptides on Liquid-Disordered Membrane Domains. Front Cell Dev Biol 8:350. https://doi.org/10.3389/fcell.2020.00350
Swain J, El KM, Kempf J et al (2018) Effect of cardiolipin on the antimicrobial activity of a new amphiphilic aminoglycoside derivative on Pseudomonas aeruginosa. PLoS One 13:e0201752. https://doi.org/10.1371/JOURNAL.PONE.0201752
Tao Y, Acket S, Beaumont E, et al (2021) Colistin treatment affects lipid composition of Acinetobacter baumannii Antibiotics 10https://doi.org/10.3390/ANTIBIOTICS10050528
Taylor R, Beriashvili D, Taylor S, Palmer M (2017) Daptomycin Pore Formation Is Restricted by Lipid Acyl Chain Composition. ACS Infect Dis 3:797–801. https://doi.org/10.1021/acsinfecdis.7b00138
Thormar H, Hilmarsson H (2007) The role of microbicidal lipids in host defense against pathogens and their potential as therapeutic agents. Chem Phys Lipids 150:1–11. https://doi.org/10.1016/j.chemphyslip.2007.06.220
Tzeng YL, Ambrose KD, Zughaier S et al (2005) Cationic antimicrobial peptide resistance in Neisseria meningitidis. J Bacteriol 187:5387–5396. https://doi.org/10.1128/JB.187.15.5387-5396.2005
Ulmschneider JP, Ulmschneider MB (2018) Molecular Dynamics Simulations Are Redefining Our View of Peptides Interacting with Biological Membranes. Acc Chem Res 51:1106–1116. https://doi.org/10.1021/ACS.ACCOUNTS.7B00613
Voss BJ, Stephen Trent M (2018) LPS Transport: Flipping Out over MsbA. Curr Biol 28:R30–R33. https://doi.org/10.1016/j.cub.2017.10.067
Walker SS, Black TA (2021) Are outer-membrane targets the solution for MDR Gram-negative bacteria? Drug Discov Today. https://doi.org/10.1016/J.DRUDIS.2021.03.027
Wang LH, Zeng XA, Wang MS et al (2018) Modification of membrane properties and fatty acids biosynthesis-related genes in Escherichia coli and Staphylococcus aureus: Implications for the antibacterial mechanism of naringenin. Biochim Biophys Acta - Biomembr 1860:481–490. https://doi.org/10.1016/j.bbamem.2017.11.007
Warner DM, Shafer WM, Jerse AE (2008) Clinically relevant mutations that cause derepression of the Neisseria gonorrhoeae MtrC-MtrD-MtrE Efflux pump system confer different levels of antimicrobial resistance and in vivo fitness. Mol Microbiol 70:462–478. https://doi.org/10.1111/j.1365-2958.2008.06424.x
WHO (2017) Global Priority List of Antibiotic-resistant Bacteria to Guide Research, Discovery, and Development of New Antibiotics
Willdigg JR, Helmann JD (2021) Mini Review: Bacterial Membrane Composition and Its Modulation in Response to Stress. Front Mol Biosci 0:338. https://doi.org/10.3389/FMOLB.2021.634438
Wilson KA, Fairweather SJ, MacDermott-Opeskin HI et al (2021) The role of plasmalogens, Forssman lipids, and sphingolipid hydroxylation in modulating the biophysical properties of the epithelial plasma membrane. J Chem Phys 154:95101. https://doi.org/10.1063/5.0040887
Wilson KA, MacDermott-Opeskin HI, Riley E et al (2020a) Understanding the link between lipid diversity and the biophysical properties of the neuronal plasma membrane. Biochemistry 59:3010–3018. https://doi.org/10.1021/acs.biochem.0c00524
Wilson KA, Wang L, MacDermott-Opeskin H, O’Mara ML (2020b) The Fats of Life: Using computational chemistry to characterise the eukaryotic cell membrane. Aust J Chem 73:85–95. https://doi.org/10.1071/CH19353
Wu EL, Cheng X, Jo S et al (2014) CHARMM-GUI Membrane Builder toward realistic biological membrane simulations. J Comput Chem 35:1997–2004. https://doi.org/10.1002/jcc.23702
Wu EL, Engström O, Jo S et al (2013) Molecular dynamics and NMR spectroscopy studies of E. coli lipopolysaccharide structure and dynamics. Biophys J 105:1444–1455. https://doi.org/10.1016/j.bpj.2013.08.002
Yao J, Rock CO (2015) How bacterial pathogens eat host lipids: Implications for the development of fatty acid synthesis therapeutics. J Biol Chem 290:5940–5946. https://doi.org/10.1074/jbc.R114.636241
Zakrzewska S, Mehdipour AR, Malviya VN et al (2019) Inward-facing conformation of a multidrug resistance MATE family transporter. Proc Natl Acad Sci 116:12275–12284. https://doi.org/10.1073/PNAS.1904210116
Zang M, MacDermott-Opeskin H, Adams FG, et al (2021) The membrane composition defines the spatial organization and function of a major Acinetobacter baumannii drug efflux system Mbio 12https://doi.org/10.1128/mbio.01070-21
Zasloff M (2002) Antimicrobial peptides of multicellular organisms. Nature 415:389–395. https://doi.org/10.1038/415389a
Zhao J, An J, Hwang D, et al (2019) The lipid A 1-phosphatase, LpxE, functionally connects multiple layers of bacterial envelope biogenesis Mbio 10https://doi.org/10.1128/MBIO.00886-19
Zhou Z, Ribeiro AA, Lin S et al (2001) Lipid A modifications in polymyxin-resistant Salmonella typhimurium: PmrA-dependent 4-amino-4-deoxy-L-arabinose, and phosphoethanolamine incorporation. J Biol Chem 276:43111–43121. https://doi.org/10.1074/jbc.M106960200
Acknowledgements
The authors would like to their friends and family for support during the COVID-19 pandemic.
Funding
Open Access funding enabled and organized by CAUL and its Member Institutions. This work was supported by the National Health and Medical Research Council Project Grant APP1140554.
Author information
Authors and Affiliations
Contributions
H.I.M.O, V.G and M.L.O conducted the literature review and wrote the manuscript.
Corresponding author
Ethics declarations
Not applicable.
Ethics approval
Not applicable.
Consent to participate
Not applicable.
Consent for publication
All authors have given final approval for this version of the manuscript.
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Vrinda Gupta and Hugo I. MacDermott-Opeskin are equally contributing authors.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
MacDermott-Opeskin, H.I., Gupta, V. & O’Mara, M.L. Lipid-mediated antimicrobial resistance: a phantom menace or a new hope?. Biophys Rev 14, 145–162 (2022). https://doi.org/10.1007/s12551-021-00912-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12551-021-00912-8