Abstract
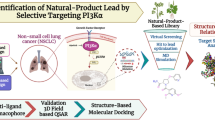
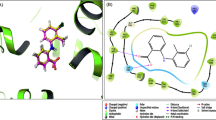
The rate-limiting enzyme cyclooxygenase-2 (COX-2) is considered as an insightful prognostic target for non-small cell lung cancer (NSCLC) therapy. Now, administration and prolonged utilization of selective COX-2 inhibitors (COXIBs) towards moderating the NSCLC has been associated with different side effects. In the present study, we focused on the structure-based drug repositioning approaches for predicting therapeutic potential de novo candidates for human COX-2. Due to discrepancies in the eminence of x-ray diffraction structures, creates a big barrier in drug discovery approach. Hence, the adaptable COX-2 structure was investigated using multi-template modeling method. Next, a dataset of twenty-six celebrex-associated optimized scaffolds were screened from ZINC database. Comparative docking approaches were then utilized to identify five compounds as best binders to the active site of COX-2 structures and strongly agree with enormous experimental consequences. MD simulations of regarded protein–ligand complexes reveals that lead molecules were stabilized dynamically in inside the cyclooxygenase site by forming potential salt bridges with Tyr348, Tyr385 and Ser530 residues. These significant results revealed that, identified druggables could prevent the tyrosyl radicals and prostaglandin production that reduces NSCLC progression. Furthermore, pharmacokinetics assets of respected ligands were analyzed, which incorporates similarity ensemble approach, druglikeness and ADMET properties. Finally, the identified novel candidates could serve as COX-2 inhibitors for NSCLC therapy, and coxibs are the best choices for designing new scaffolds to treat cyclooxygenases regard disorders.








Similar content being viewed by others
References
Molina JR, Yang P, Cassivi SD, Schild SE, Adjei AA (2008) Non-small cell lung cancer: epidemiology, risk factors, treatment, and survivorship. Mayo Clin Proc 83(5):584–594. https://doi.org/10.4065/83.5.584
Okimoto RA, Bivona TG (2014) Recent advances in personalized lung cancer medicine. Per Med 11(3):309–321
Cooper WA, Lam DC, O’Toole SA, Minna JD (2013) Molecular biology of lung cancer. J Thorac Dis 5:S479–S490. https://doi.org/10.3978/j.issn.2072-1439
Mascaux C, Martin B, Paesmans M, Berghmans T, Dusart M, Haller A, Lothaire P, Meert AP, Lafitte JJ, Sculier JP (2006) Has Cox-2 a prognostic role in non-small-cell lung cancer? A systematic review of the literature with meta-analysis of the survival results. Br J Cancer 95(2):139–145
Vane JR, Bakhle YS, Botting RM (1998) Cyclooxygenases 1 and 2. Annu Rev Pharmacol Toxicol 38:97–120. https://doi.org/10.1146/annurev.pharmtox.38.1.97
Claria J (2003) Cyclooxygenase-2 biology. Curr Pharm Des 9(27):2177–2190
Kurumbail RG, Kiefer JR, Marnett LJ (2001) Cyclooxygenase enzymes: catalysis and inhibition. Curr Opin Struct Biol 11(6):752–760
Lipsky PE (1999) Role of cyclooxygenase-1 and -2 in health and disease. Am J Orthop 3:8–12
Smith WL, Langenbach R (2001) Why there are two cyclooxygenase isozymes. J Clin Invest 107(12):1491–1495. https://doi.org/10.1172/JCI13271
Crofford LJ (1997) COX-1 and COX-2 tissue expression: implications and predictions. J Rheumatol Suppl 49:15–19
Hida T, Kozaki K, Muramatsu H, Masuda A, Shimizu S, Mitsudomi T, Sugiura T, Ogawa M, Takahashi T (2000) Cyclooxygenase-2 inhibitor induces apoptosis and enhances cytotoxicity of various anticancer agents in non-small cell lung cancer cell lines. Clin Cancer Res 6(5):2006–2011
Nie D, Honn KV (2002) Cyclooxygenase, lipoxygenase and tumor angiogenesis. Cell Mol Life Sci 59:799–807
Castelao JE, Bart RD, DiPerna CA, Sievers EM, Bremner RM (2003) Lung cancer and cyclooxygenase-2. Ann Thorac Surg 76(4):1327–1335
Schuller HM, Tithof PK, Williams M, Plummer H (1999) The tobacco-specific carcinogen 4-(methylnitrosamino)-1-(3-pyridyl)-1-butanone is a beta-adrenergic agonist and stimulates DNA synthesis in lung adenocarcinoma via beta-adrenergic receptor-mediated release of arachidonic acid. Cancer Res 59(18):4510–4515
Schuller HM, Plummer HK, Bochsler PN, Dudric P, Bell JL, Harris RE (2001) Co-expression of beta-adrenergic receptors and cyclooxygenase-2 in pulmonary adenocarcinoma. Int J Oncol 19(3):445–449
Harris RE, Beebe-Donk J, Alshafie GA (2007) Reduced risk of human lung cancer by selective cyclooxygenase 2 (COX-2) blockade: results of a case control study. Int J Biol Sci 3(5):328–334
Rao P, Knaus EE (2008) Evolution of nonsteroidal anti-inflammatory drugs (NSAIDs): cyclooxygenase (COX) inhibition and beyond. J Pharm Pharm Sci 11(2):81s–110s
Schreinemachers DM, Everson RB (1994) Aspirin use and lung, colon and breast cancer incidence in a prospective study. Epidemiology 5:138–146
Thun MJ, Henley SJ, Patrono C (2002) Nonsteroidal anti-inflammatory drugs as anticancer agents: mechanistic, pharmacologic, and clinical issues. J Natl Cancer Inst 94(4):252–266
Zarghi A, Arfaei S (2011) Selective COX-2 Inhibitors: a review of their structure-activity relationships. Iran J Pharm Res 10(4):655–683
Bresalier RS, Sandler RS, Quan H, Bolognese JA, Oxenius B, Horgan K, Lines C, Riddell R, Morton D, Lanas A, Konstam MA, Baron JA, Adenomatous Polyp Prevention on Vioxx (APPROVe) Trial Investigators (2005) Cardiovascular events associated with rofecoxib in a colorectal adenoma chemoprevention trial. N Engl J Med 352:1092–1102
Nussmeier NA, Whelton AA, Brown MT, Langford RM, Hoeft A, Parlow JL, Boyce SW, Verburg KM (2005) Complications of the COX-2 inhibitors parecoxib and valdecoxib after cardiac surgery. N Engl J Med 352:1081–1091
Giercksky KE, Huseby G, Rugstad HE (1989) Epidemiology of NSAID-related gastrointestinal side effects. Scand J Gastroenterol Suppl 163:3–8
Lenzer J (2005) FDA advisers warn: COX 2 inhibitors increase risk of heart attack and stroke. BMJ 330(7489):440
Sun SX, Lee KY, Bertram CT, Goldstein JL (2007) Withdrawal of COX-2 selective inhibitors rofecoxib and valdecoxib: impact on NSAID and gastroprotective drug prescribing and utilization. Curr Med Res Opin 8:1859–1866
Ostermeier C, Michel H (1997) Crystallization of membrane proteins. Curr Opin Struct Biol 5:697–701
Lucido MJ, Orlando BJ, Vecchio AJ, Malkowski MG (2016) Crystal structure of aspirin-acetylated human cyclooxygenase-2: insight into the formation of products with reversed stereochemistry. Biochemistry 55(8):1226–1238
Orlando BJ, Malkowski MG (2016) Substrate-selective inhibition of cyclooxygenase-2 by fenamic acid derivatives is dependent on peroxide tone. J Biol Chem 291(29):15069–15081
Sali A, Blundell TL (1993) Comparative protein modelling by satisfaction of spatial restraints. J Mol Biol 234(3):779–815
Guex N, Peitsch MC (1997) SWISS-MODEL and the Swiss-PdbViewer: an environment for comparative protein modeling. Electrophoresis 15:2714–2723
Laskowski RA, MacArthur MW, Moss DS, Thornton JM (1993) PROCHECK: a program to check the stereochemical quality of protein structures. J Appl Crystallogr 26:283–291
Luthy R, Bowie JU, Eisenberg D (1992) Assessment of protein models with three-dimensional profiles. Nature 356(6364):83–85
Colovos C, Yeates TO (1993) Verification of protein structures: patterns of non-bonded atomic interactions. Protein Sci 9:1511–1519
Wiederstein M, Sippl MJ (2007) ProSA-web: interactive web service for the recognition of errors in three-dimensional structures of proteins. Nucleic Acids Res 35:W407–W410
Irwin JJ, Sterling T, Mysinger MM, Bolstad ES, Coleman RG (2012) ZINC: a free tool to discover chemistry for biology. J Chem Inf Model 52(7):1757–1768. https://doi.org/10.1021/ci3001277
Chitrala KN, Yeguvapalli S (2014) Computational Prediction and analysis of breast cancer targets for 6-Methyl-1, 3, 8-trichlorodibenzofuran. PLoS One 9(11):e109185. https://doi.org/10.1371/journal.pone.0109185
Zoete V, Cuendet MA, Grosdidier A, Michielin O (2011) SwissParam: a fast force field generation tool for small organic molecules. J Comput Chem 32(11):2359–2368
Dallakyan S, Olson AJ (2015) Small-molecule library screening by docking with PyRx. Methods Mol Biol 1263:243–250. https://doi.org/10.1007/978-1-4939-2269-7_19
Keiser MJ, Roth BL, Armbruster BN, Ernsberger P, Irwin JJ, Shoichet BK (2007) Relating protein pharmacology by ligand chemistry. Nat Biotechnol 2:197–206
Nelson MT, Humphrey W, Gursoy A et al (1996) NAMD: a parallel, object-oriented molecular dynamics program. Int J Supercomput 10(4):251–268
Humphrey W, Dalke A, Schulten K (1996) VMD: visual molecular dynamics. J Mol Graph 14(1):33–38
Tanner DE, Phillips JC, Schulten K (2012) GPU/CPU algorithm for generalized Born/solvent-accessible surface area implicit solvent calculations. J Chem Theor Comp 8:2521–2530
Aqvist J, Medina C (1994) A new method for predicting binding affinity in computer-aided drug design. Protein Eng 7:385–391
Feller SE, Zhang Y, Pastor RW, Brooks BR (1995) Constant pressure molecular dynamics simulation: the Langevin piston method. J Chem Phys 103:4613–4621
Daina A, Michielin O, Zoete V (2014) iLOGP: a simple, robust, and efficient description of n-octanol/water partition coefficient for drug design using the GB/SA approach. J Chem Inf Model 54(12):3284–3301
Sliwoski G, Kothiwale S, Meiler J Jr, Lowe EW (2013) Computational methods in drug discovery. Pharmacol Rev 66(1):334–395
Dong GQ, Fan H, Schneidman-Duhovny D, Webb B, Sali A (2013) Optimized atomic statistical potentials: assessment of protein interfaces and loops. Bioinformatics 29(24):3158–3166
Yokouchi H, Kanazawa K (2015) Revisiting the role of COX-2 inhibitor for non-small cell lung cancer. Transl Lung Cancer Res 5:660–664
Penning TD, Talley JJ, Bertenshaw SR, Carter JS, Collins PW, Docter S, Graneto MJ, Lee LF, Malecha JW, Miyashiro JM, Rogers RS, Rogier DJ, Yu SS, Anderson GD, Burton EG, Cogburn JN, Gregory SA, Koboldt CM, Perkins WE, Seibert K, Veenhuizen AW, Zhang YY, Isakson PC (1997) Synthesis and biological evaluation of the 1,5-diarylpyrazole class of cyclooxygenase-2 inhibitors: identification of 4-[5-(4-methylphenyl)-3-(trifluoromethyl)-1H-pyrazol-1-yl]benzenesulfonamide (SC-58635, celecoxib). J Med Chem 40(9):1347–1365
Kufareva I, Abagyan R (2012) Methods of protein structure comparison. Methods Mol Biol 857:231–257
Acknowledgements
B. Uma Devi [F1-17.1/2012-13/RGNF-2012-13-SC-AND-32761/(SA-III/WEBSITE)] and K. Kranthi Kumar [F1-17.1/2013-14/RGNF-2013-14-SC-AND-40670/(SAIII/Website)] grateful to University Grants Commission (UGC) New Delhi, India for awarding research association as Rajiv Gandhi National Fellowship (RGNF) to carry out this work. Y. Suneetha is thankful for Human Resource Development for Health Research, New Delhi (F.No. V.25011/542-HRD/2016-HR) and extremely grateful to the Prof. P. Sreedhara Reddy, Department of Physics, Sri Venkateswara University, Tirupati for providing computational facilities.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
The authors announce that they have no conflict of interest about this work.
Additional information
Uma Devi Bommu and Kranthi Kumar Konidala contributed equally.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Bommu, U.D., Konidala, K.K., Pamanji, R. et al. Structural Probing, Screening and Structure-Based Drug Repositioning Insights into the Identification of Potential Cox-2 Inhibitors from Selective Coxibs. Interdiscip Sci Comput Life Sci 11, 153–169 (2019). https://doi.org/10.1007/s12539-017-0244-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12539-017-0244-5