Abstract
The recent discovery of Rhabdopleura in Singapore and the chance collecting of fresh material from northern New Zealand (Three Kings Shelf) provided an opportunity to sequence the specimens with an aim to determine their species identity. Phylogeny reconstructions of two new Rhabdopleura taxa based on ribosomal and mitochondrial genes suggest a different identity from known samples, including putative Rhabdopleura annulata, first described from the Three Kings Shelf but sequenced from the Great Barrier Reef. Pairwise distances between rhabdopleurids for the 16S rRNA locus were several magnitudes larger than that of 18S rRNA, and might potentially be a suitable barcoding gene once sufficient samples of conspecifics are collected to determine the barcoding gaps. Type material of R. annulata was re-examined, as well as Rhabdopleura material from eight other New Zealand localities from north of subtropical Raoul Island (Kermadec Ridge) at ~29° S to the subantarctic Campbell Plateau at ~49° S. Six morphological characters, four of them new, were applied to all samples. The findings from morphology suggest (1) the holotype and cotype of R. annulata might not be conspecific; (2) there are 3–4 variants (species?) on the Three Kings Shelf; and (3) there are 2–3 additional variants (species?) elsewhere in the New Zealand region.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The pterobranch hemichordate genus Rhabdopleura Allman, 1869, an extant graptolite taxon (Mitchell et al. 2013), is widely distributed in space and time. It is found at the present day in all major oceans, ranging across Arctic, tropical and sub-Antarctic latitudes and from the intertidal to c. 900 m (Tassia et al. 2016; Beli et al. 2018). Notwithstanding this distribution, only five named species are presently accepted out of the eight species described (Swalla and van der Land 2022). Fossil species nominally range from the Middle Cambrian to the Eocene (Chapman et al. 1995; Sennikov 2016) (but Mierzejewski (1986), Urbanek and Dilly (2000) and Mierzejewski and Kulick (2001) have interpreted the earliest true representatives of Rhabdopleura as Middle Jurassic in age). Sennikov (2016) recognizes eight fossil species. The paucity of fossil species is unsurprising, given that Rhabdopleura colonies not only lack a hard skeleton, but, unlike planktonic graptolites, are found on benthic hard substrata and typically not amenable to fossilization in fine sediments. On the other hand, the low number of described living species invites explanation, especially since their geographic and environmental distribution is substantial.
The low number of described species may be attributed to the lack of study of the range of morphological variation in nominal species and limited molecular sequencing to determine genetic diversity. Ironically, even though there are so few living species, some recent workers have noted the difficulty of determining the identity of newly discovered material in remote locations based on morphological criteria. Such was the case when Dilly and Ryland (1985) encountered an intertidal Rhabdopleura species in Fiji, commenting: “The claims for any of them to be separate species are somewhat tenuous, and depend, as always, on the estimation of what are differentiating characters.” The relative (or apparent) paucity of such characters led to early synonymy. For example, the type species, Rhabdopleura normani Allman, 1869, apparently widely distributed along western European coasts (Burdon-Jones 1954), was interpreted by Schepotieff (1909) to include four other eastern Atlantic species, namely Rhabdopleura mirabilis Sars, 1872, Rhabdopleura compacta Hincks, 1880, Rhabdopleura grimaldii Jullien, 1890 and Rhabdopleura manubialis Jullien and Calvet, 1903. In the same work, Schepotieff (1909) described a new species, Rhabdopleura striata, from Sri Lanka. Stebbing (1970a) subsequently clarified the separate status of Rhabdopleura compacta, removing it from synonymy with R. normani. The latter name has since been accorded to specimens from Bermuda (Dilly 1985), southern Argentina (López Gappa 1987) and, with less certainty, to the material from Fiji (Dilly and Ryland 1985). Authors agree that R. mirabilis is reliably synonymized but the status of R. grimaldii and R. manubialis, both from the Azores, is “uncertain/nomen dubium” (Swalla and van der Land 2022), underscoring the need for re-examination.
A similar example is that of Rhabdopleura annulata Norman, 1921. It was formally described from northern New Zealand, but was at the same time attributed by its author to unidentified Siboga Expedition material from Indonesia described by Harmer (1905). It was subsequently reported from southern Australia (Johnston 1937) (but Shepherd (1997) re-interpreted it as R. normani), Darwin, Northern Territory (Gordon 2009) and the Great Barrier Reef, Queensland (Ramírez-Guerrero et al. 2020). Only one other species has been described since Norman (1921), i.e. Rhabdopleura recondita Beli, Cameron & Piraino in Beli et al. 2018, a Mediterranean species hitherto overlooked because of its highly cryptic habit of growing in dead skeletal parts of several bryozoan species. Relatively few specimens of Rhabdopleura have been sequenced, including some that have not been taxonomically determined to species. The results suggest the possibility of cryptic species (Beli et al. 2018; Ramírez-Guerrero et al. 2020).
Recently, in the course of targeted collecting of bryozoans by five of the present authors for a global molecular phylogeny (Orr et al. 2022), the genus Rhabdopleura was newly discovered in Singapore (Jain et al. 2022). The sole specimen was not able to be determined to species on the basis of morphology and appeared to be unrelated to Rhabdopleura previously collected from Sulawesi and West Papua by the Siboga Expedition. The same targeted bryozoan collecting also yielded a sample of Rhabdopleura from the Three Kings Shelf, northern New Zealand, geographically close to the type locality of R. annulata. We sequenced both the Singaporean and Three Kings Shelf samples to determine their phylogenetic relationships.
At the same time, owing to the evident challenge of identifying Rhabdopleura on the basis of morphological criteria, the opportunity was taken to examine a collection of Rhabdopleura from several New Zealand localities, ranging from the Kermadec Ridge at ~28° S to the subantarctic Campbell Plateau at ~49° S. These were not amenable to molecular sequencing in the same manner as the Singapore and Three Kings Shelf samples, having either been first preserved in seawater-formalin, followed by isopropanol, or discovered on dried unpreserved dead molluscan shell (principally Pecten and glycymeridid bivalves). A goal of the comparative morphological study was to ascertain not only if the sequenced sample from the Three Kings Shelf matched R. annulata, the type material of which was newly re-examined, but also if there exist overlooked or undervalued morphological criteria that could be useful in highlighting within-species morphological variation or discriminating among putative other species.
The purpose of this paper, therefore, is as follows: (1) to reconstruct a phylogeny including the two new Rhabdopleura samples using genome skimming to recover mitochondrial and ribosomal genes, and (2) to discuss the most useful morphological criteria applicable to species discrimination in Rhabdopleura.
Materials and methods
Morphological study
The provenance of all specimens examined for this study is given in Table 1. Figure 1 plots localities for all the New Zealand samples, most of which are registered in the NIWA Invertebrate Collection of the National Institute of Water and Atmospheric Research, Wellington. Most specimens were examined by scanning electron microscopy (SEM). Selected type and other material were examined uncoated by SEM at the Natural History Museum, London (NHMUK), using a JEOL IT500 SEM, supplemented by light microscopy of slide material. Voucher specimens at the University of Oslo (NHMO) were imaged uncoated using a Hitachi TM4040PLUS Tabletop SEM. That prepared for SEM was air-dried from ethanol unless the specimen was already dry. Materials examined at NIWA were imaged using a Hitachi TM3000 Tabletop SEM and coated using gold-palladium in a sputter coater for 30 s to avoid charging.
Historically, erect and creeping tubes of Rhabdopleura have mostly been imaged and measured using transmitted light microscopy. We have found the clarity and magnification of SEM images to be highly optimal for providing metric data, measured using Fiji (ImageJ) software (Schindelin et al. 2012). Terminology for descriptions and measurements (Fig. 2) is based on that provided by Maletz et al. (2014). A variety of metric measures were applied to all specimens examined by us to determine which morphological characters might be more informative. We settled on four discriminating characters that provide unambiguous and easy-to-measure standard metrics using SEM (Table 2). They are as follows:
-
1)
External diameter of erect tubes between fusellar collars (Fig. 2c). Since tube diameter may vary from its base to its aperture, and sometimes just proximal to the aperture, we determined that measurements are best made in the distal half proximal to the aperture (where possible—sometimes a majority of tubes in a colony may be distally broken depending on method of collection, by hand if by diving or by dredge in deeper water, the latter being more damaging to the fragile colonies).
-
2)
Number of fusellar collars in 500 μm of erect tubes. As with character 1, we determined that these should be counted in the distal half of tubes proximal to the aperture where there seems to be more consistency in fusellus height (which can be shorter and compressed in the proximal quarter of erect tubes in some samples). This is a new character. Its value is in smoothing across the variability in fusellus height over a standard distance.
-
3)
Fusellus height (Fig. 2c). This is the distance between one fusellus collar and the next. As with characters 1 and 2, measurements are best made in the distal half of tubes below the aperture. In some colonies, individual fusellus height may be consistent, but mostly it can vary, so it is useful to make two measurements for each fusellus, one on both sides of the face that is presented in the SEM image.
-
4)
Point-to-point angles of divergence in zigzag sequences of fusellar sutures on creeping tubes (Fig. 2e). Inasmuch as some of the zigzag sutures may be curved, while others are straight, straight lines were drawn between points of intersection, regardless of curvature, to obtain measurable angles that can be compared across samples. (Curvature of zigzag suture components means that initial angles are small at points of intersection and then widen. Some measurements of initial angles were made but eventually deemed to be less informative, for comparative purposes, than point-to-point angles.) This is a new character.
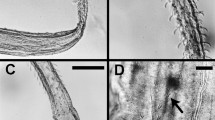
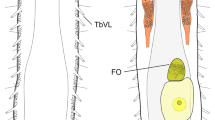
a–e Interpretative images illustrating morphological terms and measured characters. a Part of Rhabdopleura annulata Norman, 1921 cotype (NHMUK 1921.7.28.1) (but not actually this species) from Three Kings Shelf, New Zealand, showing an erect tube (et) and the broken base of an erect tube (bbet) on a creeping tube (ct); the top half of the figure is distal for both kinds of tubes, with the distal end of the creeping tube directed to the right; b light micrograph of a slide preparation of Rhabdopleura sp. (NHMUK 1928.3.5.3; Siboga Stn 163, West Papua), showing an erect tube with a contained zooid (z), with distal tentacles and proximal gut that is connected to a stolon (st); c distal end of an erect tube of Rhabdopleura sp. from Spirits Bay, New Zealand (NIWA 158157), showing a vertical series of ring-like fuselli (f), the distal rim of each having a curled fusellar collar (fc). Tube diameter (td) is measured between fusellar collars; fusellar height (fh) is the distance between the bases of a pair of fusellar collars; d line drawing of part of a creeping tube of Rhabdopleura sp. from SE Sulawesi, Indonesia (Siboga Stn 204), showing the raised central portion with a surface pattern of intersecting fusellar sutures and inside of which is a stolon (st; sometimes called a ‘black stolon’), flanked on either side by the flat basal layer (fbl) of the tube margins (from Harmer 1905, Fig. 2B); e part a creeping tube of Rhabdopleura sp. from Cavalli Seamount, New Zealand (NIWA 90267), with frontal fusellar sutures (fs, two arrowed). The white lines show the zigzag (zz) portion of the sutures (note the proximal-wards shift to the left), with zigzags rendered as straight lines between points of intersection, regardless of suture curvature, to obtain measured angles (one example shown). Scale bars: a, b, d 200 μm; c 100 μm; e 50 μm
Two other characters, both descriptive as used here, are included in Table 2. These are as follows:
-
5)
The width of the fusellar-suture zigzag sequence along uninterrupted straight sections of the creeping tube (Fig. 3). There are three main patterns. The zigzag sequence can cross the entire width of the creeping tube (e.g. Fig. 3a), occupy a narrow zone along the midline of the creeping tube (e.g. Fig. 3g–j), or be variably intermediate between these two extremes. There can be some variation within a single colony (e.g. Fig. 3g, h). This is a new character.
-
6)
Curvature of the zigzag components of the fusellar suture of the creeping tubes. They can be all or mostly curved (e.g. Fig. 3b), mostly straight (e.g. Fig. 3j), or variable within a zigzag sequence. This is a new character.
a–j Examples of zigzag patterns within the frontal fusellar sutures on creeping tubes of Rhabdopleura from most of the localities mentioned in the text and tables. The images are arranged in order of zigzag width from widest (a), through intermediate, to shortest (j). The top of each image is distal on the specimen (i.e. the growth direction of the creeping tube). Note variations from curved through straight for each zigzag component, and the shift of zigzag tips from the side to the middle of the creeping stolon or vice versa. All images pertain to specimens from the New Zealand region, apart from NHMO H2136 from Singapore. a Rhabdopleura annulata holotype NHMUK 21.7.28.4, Three Kings Shelf; b NIWA 158157, Spirits Bay; c NHMO H2136, Singapore; d R. annulata cotype NHMUK 21.7.28.1, Three Kings Shelf; e NIWA 90267, Cavalli Seamount; f NIWA 158156, Campbell Plateau; g, h NIWA 158517, Snares Plateau; i NHMO H2137, Three Kings Shelf; j NIWA 90266, Cavalli Seamount. Scale bars: a, b, d, e, g, h, j 200 μm; c, f, i 100 μm
DNA extraction, sequencing and genome skimming
Extraction of DNA and sequencing generally followed Orr et al. (2022). Briefly, samples were crushed with a pestle in a mixture of both the lysis buffer and proteinase K from DNeasy Blood and Tissue Kit (Qiagen). Subsequently, genomic DNA was extracted from digested samples following the manufacturer’s protocol using the aforementioned kit. Sequencing was performed at the Norwegian Sequencing Centre (Oslo, Norway) on an Illumina HiSeq 4000 lane (150×150 bp). Adapter removal and quality trimming on raw reads were performed using cutadapt v1.4.2 (Martin 2011). Assembly of trimmed reads was conducted using SPAdes v3.13.0 (Bankevich et al. 2012) under default settings. Identification of 18S rRNA genes was conducted with BLASTn by searching assembled sequences against Rhabdopleura sequences from GenBank (Table S1; see Ramírez-Guerrero et al. 2020). A single best hit was extracted for each sample sorted by highest bitscore, followed by e-value and then percentage identity. Overlapping regions with that of the subject were then extracted from the contig assembled to obtain the region of interest for downstream phylogenetic reconstruction.
Mitochondrial genomes were assembled for each sample using NOVOPlasty v4.3.1 (k-mer=33) (Dierckxsens et al. 2017), with the complete mitochondrial genome of congeneric R. compacta (FN908482; Perseke et al. 2011) as reference. Annotation of the mitochondrial genome was conducted first using MITOS2 (Donath et al. 2019), followed by NCBI’s ORFfinder (Wheeler et al. 2007) under translation table 24 (Rhabdopleuridae mitochondrial code) to refine the annotation for protein-coding genes (PCGs). Finally, we manually corrected PCG annotations with amino acid alignments of other hemichordate mitochondrial genomes. Alignment of sequences was done using MAFFT v7.427 under L-INS-i setting (Katoh and Standley 2013). Annotation of tRNA genes was determined by MITOS2 (Donath et al. 2019) and cross-checked using tRNAscan-SE (Chan and Lowe 2019).
Phylogenetic reconstructions and pairwise distance estimations
Two separate phylogenetic reconstructions were executed. The first was a mitochondrial genome phylogeny for hemichordates based on 13 protein-coding genes (PCGs), building upon the phylogeny in Li et al. (2019). The second was a rRNA phylogeny of Rhabdopleura based on Ramírez-Guerrero et al. (2020) (Table S2).
For the hemichordate mitochondrial genome phylogeny, PCG sequences were downloaded from both GenBank and Li et al. (2019). Sequences were translated using ExPASy (Gasteiger et al. 2003) and then aligned using MAFFT v7.427 under L-INS-i setting (Katoh and Standley 2013). Amino acid alignments were then back-translated using pal2nal v14 (Suyama et al. 2006) and trimmed using trimAl v1.4 with the heuristic algorithm activated (Capella-Gutiérrez et al. 2009). Evolutionary models for each gene were determined using ModelTest-NG (Darriba et al. 2020) under default settings. To ensure that the mitochondrial genome was properly assembled, a maximum likelihood phylogeny with models specified according to ModelTest-NG (Darriba et al. 2020) was reconstructed by RAxML-NG v0.8.1 (Kozlov et al. 2019) for each mitochondrial PCG with 10 random and 10 parsimony starting trees and 100 bootstrap pseudoreplicates. We expected that both samples sequenced here would be recovered as a monophyletic group with the other congeners.
After inspecting all gene trees and ensuring the mitochondrial genomes were accurately recovered, aligned DNA sequences from 13 PCGs from all taxa were concatenated into a single matrix. A maximum likelihood phylogeny for hemichordates was reconstructed using RAxML-NG v0.8.1 (Kozlov et al. 2019), with the matrix partitioned by gene and models specified as determined above. A total of 50 random and 50 parsimony trees were specified as starting trees, with 1000 bootstrap pseudoreplicates to estimate node support. Furthermore, Bayesian inference (BI) was performed using MrBayes v3.2.6 (Ronquist et al. 2012) on the concatenated data matrix partitioned by gene, and each partition model was estimated using ModelTest-NG (-T mrbayes) (Darriba et al. 2020). Two distinct runs over four Markov chains Monte Carlo, each with 12 million generation and one tree logged every 100 generations, were performed. Convergence was determined using Tracer v1.7 (Rambaut et al. 2018), which showed that the first 20,001 trees could be discarded as burn-in (average standard deviation of split frequencies = 0.000892).
For the rRNA phylogeny of Rhabdopleura, 16S and 18S rRNA genes were concatenated for analysis. Sequences were first aligned using MAFFT v7.427 under L-INS-i strategy (Katoh and Standley 2013). Subsequently, trimming and DNA evolution model selection, as well as maximum likelihood (ML) and Bayesian (BI) analyses, followed that of the hemichordate mitogenome phylogeny reconstruction.
Finally, uncorrected pairwise distances between Rhabdopleura sequences were separately computed for both 16S and 18S ribosomal genes from the trimmed alignment using MEGA X (Kumar et al. 2018) (Tables S1, S2).
Results
Mitochondrial genomes and gene order
A circularized mitochondrial genome was recovered for both individuals by NOVOPlasty v4.3.1 (Dierckxsens et al. 2017). The genome length of the New Zealand sample was 16,307 bp, and that of the Singapore sample was 16,500 bp. We recovered all 13 PCGs and both mitochondrial rRNA genes. Curiously, for both samples, tRNA-Arg remained unannotated, which was the only missing tRNA flagged by MITOS2 (Donath et al. 2019). In addition, the position for cytochrome c oxidase subunit II (COII) for both samples lay between ATPase 6 (ATP6) and NADH dehydrogenase subunit 6 (NAD6), whereas that in R. compacta was between cytochrome c oxidase subunit III and ATPase 8 (ATP8) (Fig. S1; Perseke et al. 2011).
Phylogeny reconstructions and pairwise distances
Rhabdopleura was recovered as a monophyletic clade in all reconstructions with maximum node support, and both samples in this study were nested within Rhabdopleura (Fig. 4). In both the concatenated rRNA and mitogenome trees, samples from Singapore and New Zealand were recovered as sister taxa. In the rRNA tree, weak support for this relationship was observed (bootstrap = 37, unresolved in Bayesian inference) (Fig. 4a). The phylogeny based on the mitochondrial PCGs performed better, with strong support for the close relationship between the two taxa (bootstrap = 73, posterior probability = 0.98), which were together sister to the remaining Rhabdopleura spp. (R. annulata and R. compacta) with maximal support (Fig. 4b).
Pairwise distances for both 16S and 18S genes suggested that the two samples from this study were more closely related to each other compared to other Rhabdopleura taxa (16S: 0.135 and 18S: 0.0058), but lower mean intraspecific distances were recorded for R. recondita (16S: 0.0226 ± SD 0.0194; 18S: 0.0022 ± SD 0.0012) (Tables S2 and S3). Overall, examination of mean interspecific distances (using a single representative of R. recondita GA2016) revealed that 16S rRNA distances (0.205 ± 0.0365) were several magnitudes more variable than 18S rRNA (0.0277 ± SD 0.0116) (Tables S2 and S3).
Morphological study
Descriptions are given below for the voucher material of the sequenced species from Singapore (NHMO H2136) and the Three Kings Shelf (NHMO H2137) at the University of Oslo. Partial descriptions only are given of re-examined type material of R. annulata, based on some new measurements of selected characters, in order to supplement the original description (Norman 1921). Following that, an overview is given of the other specimens examined for this comparative study, sourced from several other localities in the New Zealand region. Comparative metrics of selected characters from this range of specimens are provided in Table 2.
A. Description of sequenced species from Singapore and Three Kings Shelf
The descriptions below are based on small voucher specimens retained from fragments used for sequencing.
Rhabdopleura sp. from Pulau Tekukor, Singapore (Fig. 3c, 5a–d; Table 1)
Description of NHMO H2136 (University of Oslo): Colony on eroded coral surface, comprising adherent creeping tubes from which arise, at right angles, 31 erect tubes of varying length (most broken) up to 980 μm, the longest with 32 irregular full fusellae. Diameter of erect tubes 125–178 μm between fusellar collars (mean 148 μm), with tube openings subcircular to elongate-oval. Fusellus height variable (19–44 μm, mean 31 μm), fusellar collars irregular, projecting 5–13 μm (mean 8 μm) circumferentially from the tube. Creeping tubes comprising a generally parallel-sided convex median linear portion that is flanked by a flat basal layer, of varying width and unpatterned, on either side (see Fig. 2d). Convex part of creeping tube 144–263 μm (mean 212 μm). Surface fusellar pattern of creeping tubes variable; in places fusellar sutures are irregularly annular but they are more often oblique, forming a zigzag pattern in which the points of the zigzag are either at or very near the edges of the median linear part of the tube, or nearer its middle, forming parallel oblique strips in a criss-cross pattern (see Jain et al. 2022, Fig. 6e–g). Spacing between the latter parallel strips 42–106 μm (mean 75 μm). Zigzag angles (point-to-point angle of divergence) 28–44° (mean 36°). Width of stolon within creeping tube 19–57 μm (mean 35 μm).
-
Remarks
a–d Rhabdopleura from Singapore, voucher specimen from sample NHMO H2136, Pulau Tekukor. a Profile of the longest erect tube in the voucher specimen, attached at its base, showing c. 31 fuselli; b, c fusellar patterning on parts of creeping tubes, each immediately proximal to the base of an erect tube; d a pair of diverging stolons, both emerging from under the broken edge of a creeping tube at bottom of image; e, f Rhabdopleura from Three Kings Shelf, New Zealand, voucher specimen from NHMO H21378 sample; e profile of a broken erect tube, attached at its base, showing 16 fuselli; f fusellar suture pattern on frontal surface of part of a creeping tube a little distant from an erect tube. Scale bars: a 200 μm; b–f 100 μm
a, b Rhabdopleura annulata holotype (NHMUK 1921.7.28.4), Three Kings Shelf, New Zealand). a Profile of distal part of a broken erect tube showing fuselli and their collars; b light micrograph of a slide preparation from the holotype of a creeping tube showing the pattern of frontal fusellar sutures; note how the tips of the very wide zigzags extend onto the marginal basal layer; c Rhabdopleura annulata cotype (NHMUK 1921.7.28.1) (but not actually this species), Three Kings Shelf, New Zealand, showing the pattern of frontal fusellar sutures on the proximal part of the creeping tube in Fig. 2a; note that the tips of the zigzags are set back from the tube margins. Scale bars: a–c 200 μm
The voucher specimen was re-examined to obtain additional measurements (cf. Jain et al. 2022). The identity of the Singapore species remains in doubt. It is not identical to the holotype specimen of R. annulata, described from near the Three Kings Islands, New Zealand, at 549 m. Although the point-to-point angles of divergence of the zigzag sutures on the creeping tubes (Char 4) are practically identical, the number of fusellar collars in 500 μm (Char 2) is non-overlapping. Erect tubes of NHMO H2136 also have a smaller average diameter. Ramírez-Guerrero et al. (2020) compared their material from Heron Island, Great Barrier Reef, with Norman’s (1921) description of R. annulata and concluded that they had that species. Indeed, their figure 1C, which shows a wide zigzag suture on a creeping tube, is very close to that of Norman (1921, Fig. 3). Our phylogeny (Fig. 4a) shows that NHMO H2136 is not conspecific with R. annulata from Heron Island. On the other hand, it is closely related to a new specimen from The Three Kings Shelf, sequenced for this study (below).
Rhabdopleura sp. from Three Kings Shelf, New Zealand (Fig. 3i, 5e, f; Table 1)
Description of NHMO H2137 (University of Oslo): Colony on shelly surface, comprising adherent creeping tubes from which arise, at right angles, 8 erect tubes of varying length (some broken) up to 430 μm, the longest with 18 irregular full fusellae. Diameter of erect tubes 102–172 μm between fusellar collars (mean 146 μm), with tube openings subcircular to elongate-oval. Fusellus height irregular (18–33 μm, mean 26 μm). Fusellar collars irregular, projecting 7–16 μm (mean 10 μm) circumferentially from the tube. Creeping tubes of variable width, generally comprising a parallel-sided convex median linear portion flanked by smooth unpatterned lateral basal-wall extensions of varying width. Convex part of creeping tube 119–244 μm (mean 176 μm). Surface fusellar pattern of creeping tubes variable in vicinity of erect tubes and where branching of creeping tubes takes place; on long stretches of creeping tube fusellar strips are oblique and parallel, forming a longitudinal series of intersecting chevrons. Spacing between parallel strips 46–80 μm (mean 64 μm). Zigzag angles (point-to-point angle of divergence) 58–70° (mean 65°). Width of stolon within creeping tube 25–27 μm (mean 26 μm). Zooid individuals not seen within erect tubes.
-
Remarks
Although NHMO H2137 is from the same general area (Three Kings Shelf) as R. annulata, it appears not to be conspecific. Metric characters 1–4 (Table 2) are all non-overlapping and the zigzag suture of the former is confined to the middle part of the creeping tube. In comparing NHMO H2137 to the other specimens studied here, it appears closest to NIWA 90266 from Cavalli Seamount c. 200 km to the east. Although the zigzag pattern appears very similar, there is in fact no overlap in Char 4 (point-to-point angle of divergence) between the two morphologies, which suggest they might also not be conspecific.
B. Re-examination of holotype of Rhabdopleura annulata Norman, 1921 (NHMUK 1921.7.28.4, Three Kings Shelf, New Zealand (Fig. 3a, 6a, b; Table 2)
Partial description based on limited material available for SEM: Colony fragments with erect tubes of varying length (some broken) up to 1228 μm, the longest with ~34 irregular fusellae. Diameter of erect tubes 167–200 μm between fusellar collars (mean 180 μm), with tube openings subcircular to elongate-oval. Fusellar height irregular (38–54 μm, mean 45 μm). Fusellar collars irregular, projecting 5–23 μm (mean 16 μm) circumferentially from the tube. Creeping tubes of variable width, generally comprising a parallel-sided convex median linear portion flanked by a flat basal layer, of varying width and unpatterned, on either side. Median linear part of creeping tube c. 177 μm. Zigzag angles (point-to-point angle of divergence) 25–46° (mean 34°).
-
Remarks
Surface debris obscured the fusellar patterning of the creeping tubes in SEM images, but this pattern was able to be seen in a light micrograph (Fig. 3a, 6b). It shows very wide curved zigzag sutures that occupy the full width of the creeping tube, with the zigzag angles extending onto the flat basal layer. What is surprising is that cotype specimen NHMUK 1921.7.28.1 (Fig. 2a, 3d, 6c), from a different station on the Three Kings Shelf, differs from the holotype in some respects (compare data columns 1–4 in Table 2). Character 2 (number of fusellar collars in 500 μm) is non-overlapping. Furthermore, the zigzag pattern on the creeping tube of the cotype is not as wide as in the holotype. While the amount of material that could be separated from the holotype and cotype of R. annulata for SEM examination is admittedly limited, the metric data and difference in fusellar zigzag patterning suggest that the two type specimens might not be conspecific. NHMO H2137 from the Three Kings Shelf may represent a third species from the same general area.
Rhabdopleura from four different New Zealand localities showing variations in erect and creeping tubes, their width, the annuli of erect tubes and the fusellar patterning of creeping tubes. a, b NIWA 90264, Spirits Bay, northern New Zealand, 57 m; c, d NIWA 90266, Cavalli Seamount, 470–480 m; e, f NIWA 158791, Otago Shelf, 130–180 m; g, h NIWA 158156, Campbell Plateau, 432–531 m. Scale bars: a–c, e, f, h 100 μm; d, g 200 μm
a–h Rhabdopleura from another Spirits Bay locality (NIWA 158157, 59 m) showing additional variation, i.e. long stretches of erect tube (a) with parallel narrow annuli along most of erect-tube length, but more-developed and curled towards the proximal end. b The tube opening in a more magnified; c a section of the tube in a just distal to midlength; d a short tube (possibly broken), with curled annuli in the part near the substratum; e opening of another unbroken erect tube; f stolon emergent from a cellular mass, with some globular components, in a slightly swollen part of a creeping tube; g, h variations in creeping-tube width and fusellar patterning, the latter typically weakly and thinly developed throughout the whole colony. Scale bars: a 1 mm; b–f 100 μm, g, h 200 μm
C. Other Rhabdopleura specimens from the New Zealand region (Fig. 2c, e, 3b, e–h, j, 7a–h, 8a–h; Table 1)
Data pertaining to these specimens show a singular lack of conformity for all characters. While some characters clearly overlap or coincide, others do not. For example, NIWA 158518 from north of subtropical Raoul Island on the Kermadec Ridge (c. 29° S) is closest to NIWA158791 from the Otago Shelf (c. 46° S), and the two entities may be conspecific. They differ most in the average distance between fusellar collars (Char 3), but the ranges overlap. NIWA 90264 from Spirits Bay (SE part of Three Kings Shelf) differs from the material from near Raoul Island in character 2, which is non-overlapping, but it could be conspecific with NIWA 158517 from the Snares Plateau (c. 48° S); all characters overlap and the zigzag sutures are similar (cf. Fig. 3g, h (Snares) and Fig. 6b (Spirits Bay)). NIWA 90266 from Cavalli Seamount has the highest average number of fusellar collars (28) in 500 μm; it could be related to NHMO H2137 from the Three Kings Shelf but character 4 (point-to-point angle of divergence in fusellar zigzags) is non-overlapping. Finally, NIWA 158156 from the subantarctic Campbell Plateau differs in several significant respects, having the largest erect-tube diameter of all other samples in Table 2. It matches NHMO H2137 in having larger zigzag angles (Char 4), but the latter has a much narrower tube diameter.
-
Remarks
Comparison of metric and non-metric data in Table 2 for all eleven samples suggests either that morphological diversity in Rhabdopleura is extremely variable or that there might be many cryptic species. Clearly, a major constraint in studies of Rhabdopleura, including our own, is the availability of specimens from a wide range of localities amenable to molecular sequencing to compare with morphological results.
Discussion
This study compares Rhabdopleura specimens from a wide range of latitudes (c. 29–49° S) and depths (57–630 m) in the New Zealand region, plus Singapore, using six morphological characters. We consider four of these (2, 4–6) to be effectively new. Character 2, pertaining to the number of fusellar collars over a fixed distance of 500 μm in the distal part of an erect tube proximal to the aperture, is supported by the statement in Maletz et al. (2016) that early fuselli of an erect tube can be of a lesser height than distal or apertural fuselli but a lesser height may also be seen at thecal apertures. Concerning characters 4–6, although authors have historically noted the zigzag pattern within fusellar sutures on creeping tubes, they have not taken this further by measuring zigzag angles or describing the range of variation shown by the sutures (which can also vary in thickness). The overlooked potential usefulness of the pattern of creeping-tube sutures may be attributable to paucity of material in many instances, though it would be useful to apply it and some of the other new characters to populations of R. annulata along European coasts, for example. The new characters used here are suggested to be potentially useful for morphological discrimination within the genus, the results of which could be tested in parallel with molecular sequencing pending the discovery of fresh material. When we began our study, it was anticipated that examination of NIWA collection samples from a range of localities in the New Zealand region would suggest at least one widespread species. The sequence data and morphology indicate that there are at least two species on the Three Kings Shelf (assuming that Heron Island R. annulata is indeed this species—see below). We cannot conclusively say if there is one highly variable species or that there are instead several cryptic species. The latter hypothesis seems the more likely, but what is now needed is preserved new material of each of the variants for sequencing, which can be accompanied by detailed measurements of within-colony variation in the same samples.
Norman (1921) had suggested that New Zealand R. annulata was conspecific with Siboga Expedition material from Indonesia that was described by Harmer (1905) but left in open nomenclature. This conclusion seemed supported by the attribution of Heron Island material to R. annulata by Ramírez-Guerrero et al. (2020). In our study, molecular data for the Singapore specimen does not match that from Heron Island. Instead, it is sister to new material from northern New Zealand (NHMO H2137) in the mitochondrial genome phylogeny (Fig. 4b), and both are closely related to R. normani (Fig. 4a) in the ribosomal phylogeny.
Our study finds that the length of mitochondrial genomes of our material is similar to R. compacta but that gene order is slightly different. Specifically, for the two samples sequenced in this study, the position of COII was between ATP6 and NAD6, whereas in R. compacta it was between COIII and ATP8 (Fig. S1). While some invertebrates exhibit extensive mitochondrial gene rearrangements even between congenerics (Chen et al. 2018; Tempestini et al. 2020), others demonstrate remarkable conservation of gene order over millions of years (Ren et al. 2010; Quek et al. 2021). Mitochondrial gene arrangements may serve as important phylogenetic characters, particularly in deep phylogenies (Dowton et al. 2002; Zhang et al. 2018). The similarity in gene arrangement between the two mitochondrial genomes suggests that the two taxa diverged more recently when compared to the other members of Rhabdopleura, which was corroborated by our mitochondrial genome phylogeny (Fig. 4b). Furthermore, interspecific pairwise distances for both ribosomal genes (16S and 18S) were lower between New Zealand and Singapore samples (Tables S2 and S3), cementing the close evolutionary relationship between the two samples. Nevertheless, intraspecific distances for 16S rRNA in R. recondita were lower than 4%, whereas the smallest interspecific distance stood at 13.5% between Rhabdopleura species (Tables S2 and S3). In contrast, 18S rRNA was much less variable, with intraspecific distance below 0.5%, whereas interspecific distances never went above 2.5%. Taken together, this suggests that 16S might be a useful barcoding gene for this clade, although more samples are required in order to determine an appropriate barcoding gap distance, particularly from conspecifics (reviewed in DeSalle and Goldstein 2019).
If, as appears the case from the morphological data, there is more than one species on the Three Kings Shelf, that would be consistent with what is already known about the biodiversity of the Three Kings Shelf/Spirits Bay area, which is considered New Zealand’s marine-biodiversity hotspot (Cryer et al. 2000; Taylor and Gordon 2003). For example, Cryer et al. (2000) reported ~300 species of Bryozoa and ~215 species of sponges from an area of about 200 km2 north of Spirits Bay, including many multispecies genera. Indeed, five closely related species of the bryozoan genus Steginoporella, all sequenced (Orr et al. 2022), are found in the vicinity of the Three Kings Islands.
Simakov et al. (2015) estimated the origin of Rhabdopleura and the rest of the hemichordates to be at 559 Ma, placing it in deep evolutionary time. Fossils of putative Rhabdopleura and other pterobranchs have been discovered as early as Cambrian Series 3 (509 Ma) (Durman and Sennikov 1993; Maletz and Steiner 2015; Sennikov 2016). The small number of nominal species described so far could be an artefact of the difficulty in identifying them, given their unostentatious morphology (Chapman et al. 1995). Of course, the question remains—is Rhabdopleura as highly conserved genetically as it is morphologically? Based on our results, it appears that 16S rRNA could be a useful marker for identifying different species once sufficient sampling has been conducted for this enigmatic clade. If morphological variability suggests cryptic or otherwise unrecognized species, hinted at by differences in widths of erect- and creeping tube fusellar bands and the zigzag angles on the creeping tubes, then it is interesting to speculate on what might generate these differences. Inasmuch as fusellar strips are formed from secretions of the head shield of the zooids stretching from the erect-tube and creeping-tube apertures (Dilly 1986, 1993; Rigby and Dilly 1993; Maletz and Cameron 2016), then head-shield size, and individual zooid movements, may be correlated with fusellar-strip width. A study of tube-building in R. normani led Dilly (1986) to postulate that the “beautiful symmetry and regularity” of fusellar patterns may be the result of “a restricted and limited programme of tube-building available to the zooid”. Hence, size and behavioural differences among rhabdopleurid zooids may be species-specific. If fusellar zigzag patterns (inter alia) have taxonomic significance at the species level, then such characters will have applicability to fossil rhabdopleurids as well as living ones, as they can be preserved in thin organic films (Maletz and Steiner 2015; Sennikov 2016, pl. 1) and by bioimmuration (Taylor and Todd 2001, Fig. 3.2.5.2d).
It should be noted, however, that variations in zooid behaviour and tube morphometry might be caused by different environmental conditions (e.g. water currents, food availability, and sedimentation rates). For example, Stebbing (1970b) observed in R. compacta that fusellus height in erect tubes can reflect degeneration and regeneration of zooids that may in part be seasonally related. In this species, a number of fuselli of normal height may be succeeded distally by shorter fuselli, which then gradually increase in size towards the distal aperture. He attributed this increase in annulus height to increasing zooid size following regeneration. Degenerating zooids were commonly seen in autumn, but Stebbing (1970a) also noted that embryos can be produced throughout the year, presumably related to continuing food availability and/or or nutrient reserves in so-called dormant buds. Ecological studies and growth studies are desirable for the commonest Rhabdopleura species. For example, if R. normani is as widespread as reported, then it may be fairly eurythermal and less subject to seasonal degeneration than range-restricted R. compacta, which in any case may depend less on the physiological effects of seasonal temperature than the seasonal availability of planktonic food.
Larval traits indicate an important role in dispersal and offspring survival in Rhabdopleura. The larva is yolky, well-ciliated, non-planktonic and relatively short-lived (Stebbing 1970b, 1973; Dilly 1973; Lester 1988; Sato et al. 2008; Strano et al. 2019) and relatively few are produced per zooid. Stebbing reported that larvae of R. compacta are ready to settle up to 2 days after release, but they could delay metamorphosis for up to 10 days (Sato et al. 2008), after which they apparently have reduced ability to metamorphose. According to Lester (1988), swimming larvae of R. normani settle within 24 h after being released from the parent tube and metamorphosis takes place over 7–10 days. Strano et al. (2019) found that metamorphosis in R. recondita took 10 days in laboratory conditions at 21°C. It is clear that Rhabdopleura larvae are lecithotrophic and have only enough nutrient reserves to sustain a period of swimming for 1–2 days followed by a period of metamorphosis of up to 10 days. Accordingly, reproductive effort per offspring is higher, and larval mortality is assumed to be lower than it would be if planktotrophic. In reviewing what was known about larval ecology in marine invertebrates and the implications for paleobiology, Jablonski and Lutz (1983) observed that while species with lecithotrophic larvae appear to be typically geographically more restricted, geologically short-ranging and exhibit high speciation and extinction rates as a consequence of their characteristically low larval dispersal capabilities, there are exceptions, as subsequent studies have confirmed. For example, whereas all ascidians are lecithotrophic, those taxa with the highest speciation rates are those with the largest range sizes, while those with the least dispersive larval development have the lowest speciation rates and smallest geographic ranges (Maliska et al. 2013), contrary to findings from echinoids and gastropods in the fossil record. A comprehensive study of sacoglossan gastropods determined that lecithotrophy evolved at least 27 times in this group and that while lecithotrophic lineages arose often, they rarely persisted long enough to diversify (Krug et al. 2015). Rhabdopleuridae differs from such clades as Ascidiacea and Sacoglossa, and indeed most fossilizable clades, for being far less morphologically diverse over geological time, which means that lecithotrophy has likely been the sole reproductive mode throughout the Phanerozoic.
Remarkably, this low-diversity clade has persisted. It remains to be demonstrated if the clade is truly of such low diversity as appears at the present day. Our study suggests more morphological diversity exists than is generally appreciated but how this relates to genetic diversity remains to be seen. Previous studies sequencing the genomes of low-diversity, ancient lineages such as tuatara, coelacanth and horseshoe crab, found slow rates of evolution (Amemiya et al. 2013; Gemmell et al. 2020; Shingate et al. 2020). Nevertheless, mitochondrial rearrangements in pterobranchs were previously suggested in Li et al. (2019) to possibly correlate with higher evolutionary rates such as in ascidian tunicates. Even between congeners in Rhabdopleura, we found a minor rearrangement of mitochondrial gene order (Fig. S1). Taken together, we suggest that this clade is likely more speciose than is currently sampled, and more extensive sampling efforts of both morphology and genomics (i.e., whole genome sequencing) may reveal previously undiscovered species of this inconspicuous but highly distinctive creature.
References
Allman GJ (1869) On Rhabdopleura, a new form of Polyzoa, from deep-sea dredging in Shetland. Q J Microscopical Sci n.s. 4:95–109
Amemiya CT, Alföldi J, Lee AP, Fan S, Philippe H, MacCallum I, Braasch I, Manousaki T, Schneider I, Rohner N, Organ C, Chalopin D, Smith JI, Robinson M, Dorrington RA, Gerdol M, Aken B, Biscotti MA, Barucca M, Baurain D, Berlin AM, Blatch GL, Buonocore F, Burmester T, Campbell MS, Canapa A., Cannon JP, Christoffels A, De Moro G, Edkins AL, Fan L, Fausto AM, Feiner N, Forconi M, Gamieldien J, Gnerre S, Gnirke A, Goldstone JV, Haerty W, Hahn ME, Hesse U, Hoffman S, Johnson J, Karchner SI, Kuraku S, Lara M, Levin JZ, Litman GW, Mauceli E, Miyake T, Mueller MG, Nelson DR, Nitsche A, Olmo E, Ota T, Pallavicina A, Panji S, Picone B, Ponting CP, Prohaska SJ, Przybylski D, Saha NR, Ravi V, Ribeiro FJ, Sauka-Spengler T, Scapigliati G, Searle SMJ, Sharpe T, Simakov O, Stadler PF, Stegeman JJ, Sumiyama K, Tabbaa D, Tafer H, Turner-Maier J, van Heusden P, White S, Williams L, Yandell M, Brinkmann H, Volff J-N, Tabin CJ, Shubin N, Schartl M, Jaffe DB, Postlethwait JH, Venkatesh B, Di Palma F, Lander AM, Lindblad-Toh K (2013) The African coelacanth genome provides insights into tetrapod evolution. Nature 496:311–316. https://doi.org/10.1038/nature12027
Bankevich A, Nurk S, Antipov D, Gurevich AA, Dvorkin M, Kulikov AS, Lesin VM, Nikolenko SI, Pham S, Prjibelski AD, Pyshkin AV, Sirotkin AV, Vyahhi N, Tesler G, Alekseyev MA, Pevzner PA (2012) SPAdes: a new genome assembly algorithm and its applications to single-cell sequencing. J Computational Biol 19:455–477. https://doi.org/10.1089/cmb.2012.0021
Beli E, Aglieri G, Strano F, Maggioni D, Telford M, Piraino S, Cameron CB (2018) The zoogeography of extant rhabdopleurid hemichordates (Pterobranchia: Graptolithina), with a new species from the Mediterranean Sea. Invertebr Syst 32:100–110. https://doi.org/10.1071/IS17021
Burdon-Jones C (1954) The habitat and distribution of Rhabdopleura normani Allman. Universitetet i Bergen Årb Naturvitenskapelig Rekke 11:1–17
Capella-Gutiérrez S, Silla-Martínez JM, Gabaldón T (2009) trimAl: a tool for automated alignment trimming in large-scale phylogenetic analyses. Bioinformatics 25:1972–1973. https://doi.org/10.1093/bioinformatics/btp348
Chan PP, Lowe TM (2019) tRNAscan-SE: Searching for tRNA genes in genomic sequences. In: Kollmar M (ed) Gene prediction: methods and protocols. Humana, New York, pp 1–14
Chapman AJ, Durman PN, Rickards RB (1995) Rhabdopleuran hemichordates: new fossil forms and review. Proc Geol Assoc 106:293–303. https://doi.org/10.1016/S0016-7878(08)80240-4
Chen L, Chen P-Y, Xue X-F, Hua H-Q, Li Y-X, Zhang F, Wei S-J (2018) Extensive gene rearrangements in the mitochondrial genomes of two egg parasitoids, Trichogramma japonicum and Trichogramma ostriniae (Hymenoptera: Chalcidoidea: Trichogrammatidae). Sci Rep 8:7034. https://doi.org/10.1038/s41598-018-25338-3
Cryer M, O’Shea S, Gordon DP, Kelly M, Drury J, Morrison M, Hill A, Saunders H, Shankar U, Wilkinson M, Foster G (2000) Distribution and structure of benthic invertebrate communities between North Cape and Cape Reinga. Final Research Report for Ministry of Fisheries Research Project ENV9805 Objectives 1-4. NIWA, Wellington
Darriba D, Posada D, Kozlov AM, Stamatakis A, Morel B, Flouri T (2020) ModelTest-NG: a new and scalable tool for the selection of DNA and protein evolutionary models. Mol Biol Evol 37:291–294. https://doi.org/10.1093/molbev/msz189
DeSalle R, Goldstein P (2019) Review and interpretation of trends in DNA barcoding. Front Ecol Evol 7:302. https://doi.org/10.3389/fevo.2019.00302
Dierckxsens N, Mardulyn P, Smits G (2017) NOVOPlasty: de novo assembly of organelle genomes from whole genome data. Nucleic Acids Res 45:e18. https://doi.org/10.1093/nar/gkw955
Dilly PN (1973) The larva of Rhabdopleura compacta (Hemichordata). Mar Biol 18:69–86. https://doi.org/10.1007/bf00347923
Dilly PN (1985) A note on the status of Rhabdopleura from Bermuda. J Mar Biolog Assoc UK 65:987–991. https://doi.org/10.1017/S0025315400019469
Dilly PN (1986) Modern pterobranchs: observations on their behaviour and tube building. In: Hughes CP, Rickards RB (eds) Palaeoecology and biostratigraphy of graptolites. Geological Soc Spec Publ 20:261–269. https://doi.org/10.1144/GSL.SP.1986.020.01.27
Dilly PN (1993) Cephalodiscus graptolitoides sp. nov., a probable extant graptolite. J Zool 229:69–78. https://doi.org/10.1111/j.1469-7998.1993.tb02621.x
Dilly PN, Ryland JS (1985) An intertidal Rhabdopleura (Hemichordata, Pterobranchia) from Fiji. J Zool 205:611–623. https://doi.org/10.1111/j.1469-7998.1985.tb03548.x
Donath A, Jühling F, Al-Arab M, Bernhart SH, Reinhardt F, Stadler PF, Middendorf M, Bernt M (2019) Improved annotation of protein-coding genes boundaries in metazoan mitochondrial genomes. Nucleic Acids Res 47:10543–10552. https://doi.org/10.1093/nar/gkz833
Dowton M, Belshaw R, Austin AD, Quicke DLJ (2002) Simultaneous molecular and morphological analysis of braconid relationships (Insecta: Hymenoptera: Braconidae) indicates independent mt-tRNA gene inversions within a single wasp family. J Mol Evol 54:210–226. https://doi.org/10.1007/s00239-001-0003-3
Durman PN, Sennikov NV (1993) A new rhabdopleurid from the Middle Cambrian of Siberia. Palaeontology 36:283–296
Gasteiger E, Gattiker A, Hoogland C, Ivanyi I, Appel RD, Bairoch A (2003) ExPASy: the proteomics server for in-depth protein knowledge and analysis. Nucleic Acids Res 31:3784–3788. https://doi.org/10.1093/nar/gkg563
Gemmell NJ, Rutherford K, Prost S, Tollis M, Winter D, Macey JR, Adelson DL, Suh A, Bertozzi T, Grau JH, Organ C, Gardner PP, Muffalo M, Patricio M, Billis K, Martin FJ, Flicek P, Petersen B, Kang L et al (2020) The tuatara genome reveals ancient features of amniote evolution. Nature 584:403–409. https://doi.org/10.1038/s41586-020-2561-9
Gordon DP (2009) Baudina gen. nov., constituting the first record of Pasytheidae from Australia, and Sinoflustridae fam. nov., with a checklist of Bryozoa and Pterobranchia from Beagle Gulf. The Beagle: Records Museums Art Galleries Northern Territory 25:43–54
Harmer SF (1905) The Pterobranchia of the Siboga Expedition with an account of other species. Siboga-Expeditie 26:1–133
Hincks T (1880) A history of the British marine Polyzoa. Van Voorst, London
Jablonski D, Lutz RA (1983) Larval ecology of marine benthic invertebrates: paleobiological implications. Biol Rev 58:21–89. https://doi.org/10.1111/j.1469-185X.1983.tb00380.x
Jain SS, Gordon DP, Huang D, Kuklinski P, Liow LH (2022) Targeted collections reveal new species and records of Bryozoa and the discovery of Pterobranchia in Singapore. Raffles Bull Zool 70:257–274. https://doi.org/10.26107/RBZ-2022-0011
Johnston TH (1937) A note on the occurrence of Rhabdopleura annulata in South Australian waters. Rec South Aust Mus 6:105–107
Jullien J (1890) Description d'un Bryozoaire nouveau du genre Rhabdopleura. Bull Société Zoologique de France 15:180–183. https://doi.org/10.5962/bhl.part.18723
Jullien J, Calvet L (1903) Bryozoaires provenant des campagnes de l'Hirondelle. Résultats Campagnes scientifiques accompli par le Prince Albert I 23: 1–188, 18 pls
Katoh K, Standley DM (2013) MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol Biol Evol 30:772–780. https://doi.org/10.1093/molbev/mst010
Kozlov AM, Darriba D, Flouri T, Morel B, Stamatakis A (2019) RAxML-NG: a fast, scalable and user-friendly tool for maximum likelihood phylogenetic inference. Bioinformatics 35:4453–4455. https://doi.org/10.1093/bioinformatics/btz305
Krug PJ, Vendetti JE, Ellingson RA, Trowbridge CD, Hirano YM, Trathen DY, Rodriguez AK, Swennen C, Wilson NG, Valdés AA (2015) Species selection favors dispersive life histories in sea slugs, but higher per-offspring investment drives shifts to short-lived larvae. Systematic Zool 64:983–999. https://doi.org/10.1093/sysbio/syv046
Kumar S, Stecher G, Li M, Knyaz C, Tamura K (2018) MEGA X: Molecular Evolutionary Genetics Analysis across computing platforms. Mol Biol Evol 35:1547–1549. https://doi.org/10.1093/molbev/msy096
Lester SM (1988) Settlement and metamorphosis of Rhabdopleura normani (Hemichordata: Pterobranchia). Acta Zoologica 69:111–120. https://doi.org/10.1111/j.1463-6395.1988.tb00907.x
Li Y, Kocot KM, Tassia MG, Cannon JT, Bernt M, Halanych KM (2019) Mitogenomics reveals a novel genetic code in Hemichordata. Genome Biol Evol 11:29–40. https://doi.org/10.1093/gbe/evy254
López Gappa JJ (1987) Presencia del género Rhabdopleura (Hemichordata), Pterobranchia) en la plataforma continental Argentina Physis A, 45(108):33–36
Maletz J, Cameron CB (2016) Part V, second revision, chapter 3: Introduction to the class Pterobranchia Lankester, 1877. Treatise Online 82:1–15
Maletz J, Steiner M (2015) Graptolite (Hemichordata, Pterobranchia) preservation and identification in the Cambrian Series 3. Palaeontology 58:1073–1107. https://doi.org/10.1111/pala.12200
Maletz J, Bates DEB, Brussa ED, Cooper RA, Lenz AC, Riva JF, Toro BA, Zhang Y (2014) Glossary of the Hemichordata. Treatise Online 62:1–23
Maletz J, Lenz AC, Bates DEB (2016) Part V, second revision, chapter 4: Morphology of the pterobranch tubarium. Treatise Online 76:1–63
Maliska ME, Pennell MW, Swalla BJ (2013) Developmental mode influences diversification in ascidians. Biol Lett 9:20130098. https://doi.org/10.1098/rsbl.2013.0068
Martin M (2011) Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet.journal 17:10–12. https://doi.org/10.14806/ej.17.1.200
Mierzejewski P (1986) The enigma of graptolite ancestry: lesson from a phylogenetic debate. In: Hoffman A, Nitecki M (eds) Problematic fossil taxa. Oxford University Press, New York, pp 184–225
Mierzejewski P, Kulick C (2001) Graptolite-like fibril pattern in the fusellar tissue of Palaeozoic rhabdopleurid pterobranchs. Acta Palaeontol Pol 46:349–366
Mitchell CE, Melchin MJ, Cameron CB, Maletz J (2013) Phylogenetic analysis reveals that Rhabdopleura is an extant graptolite. Lethaia 46:34–56. https://doi.org/10.1111/j.1502-3931.2012.00319.x
Norman JR (1921) Rhabdopleura. British Antarctic “Terra Nova” Exped Nat Hist Rep, Zoology 4:95–102
Orr RJS, Di Martino E, Ramsfjell M, Gordon DP, Berning B, Chowdhury I, Craig S, Cumming RL, Figuerola B, Florence W, Harmelin J-G, Hirose M, Huang D, Jain SS, Jenkins HL, Kotenko ON, Kuklinski P, Lee HE, Madurell T, McCann L, Mello HL, Obst M, Ostrovsky AN, Paulay G, Shunatova NN, Smith AM, Souto-Derungs J, Vieira L, Voje KL, Waeschenbach A, Zágoršek K, Warnock R, Liow LH (2022) Paleozoic origins of cheilostome bryozoans and their parental care inferred by a new genome-skimmed phylogeny. Sci Adv 8:eabm7452. https://doi.org/10.1126/sciadv.abm7452
Perseke M, Hetmank J, Bernt M, Stadler PF, Schlegel M, Bernhard D (2011) The enigmatic mitochondrial genome of Rhabdopleura compacta (Pterobranchia) reveals insights into selection of an efficient tRNA system and supports monophyly of Ambulacraria. BMC Evol Biol 11:134. https://doi.org/10.1186/1471-2148-11-134
Quek ZBR, Chang JJM, Ip YCA, Chan YKS, Huang D (2021) Mitogenomes reveal alternative initiation codons and lineage-specific gene order conservation in echinoderms. Mol Biol Evol 38:981–985. https://doi.org/10.1093/molbev/msaa262
Rambaut A, Drummond AJ, Xie D, Baele G, Suchard MA (2018) Posterior summarization in Bayesian phylogenetics using Tracer 1.7. Syst Biol 67:901–904. https://doi.org/10.1093/sysbio/syy032
Ramírez-Guerrero GM, Kocot KM, Cameron CB (2020) Zooid morphology and molecular phylogeny of the graptolite Rhabdopleura annulata (Hemichordata, Pterobranchia) from Heron Island, Australia. Can J Zool 98:844–849. https://doi.org/10.1139/cjz-2020-0049
Ren J, Liu X, Jiang F, Guo X, Liu B (2010) Unusual conservation of mitochondrial gene order in Crassostrea oysters: evidence for recent speciation in Asia. BMC Ecol Evol 10:394 https://www.biomedcentral.com/1471-2148/10/394
Rigby S, Dilly PN (1993) Growth rates of pterobranchs and the lifespan of graptolites. Paleobiology 19:459–475. https://doi.org/10.1017/S00948373000114081
Ronquist F, Teslenko M, van der Mark P, Ayres DL, Darling A, Höhna S, Larget B, Lio L, Suchard MA, Huelsenbeck JP (2012) MrBayes 3.2: efficient Bayesian phylogenetic inference and model choice across a large model space. Syst Biol 61:539–542. https://doi.org/10.1093/sysbio/sys029
Sars GO (1872) On some remarkable forms of animal life from the great deeps off the Norwegian coast. I. Partly from posthumous manuscripts of the late Professor Dr. Michael Sars. Brøgger & Christie, Christiania
Sato A, Bishop JDD, Holland PWH (2008) Developmental biology of pterobranch hemichordates: history and perspectives. Genesis 46:587–591. https://doi.org/10.1002/dvg.20395
Schepotieff A (1909) Die Pterobranchier des Indischen Ozeans. Zool Jahrb (Syst) 28:429–448
Schindelin J, Arganda-Carreras I, Frise E, Kaynig V, Longair M, Pietzsch T, Preibisch S, Rueden C, Saalfeld S, Schmid B, Tinevez J-Y, White DJ, Hartenstein V, Eliceira K, Tomcak P, Cardona A (2012) Fiji: an open-source platform for biological-image analysis. Nat Methods 9:676–682. https://doi.org/10.1038/nmeth.2019
Sennikov NV (2016) Morphology of the exoskeleton and soft tissues of Cambrian rhabdopleurids. Paleontol J 50:1626–1636. https://doi.org/10.1134/S0031030116140070
Shepherd SA (1997) Acorn worms and a pterobranch (phylum Hemichordata). In: Shepherd SA, Davies M (eds) Marine invertebrates of southern Australia. Part III. South Australian Research and Development Institute & Flora and Fauna of South Australia Handbooks Committee, Adelaide, pp 1028–1039
Shingate P, Ravi V, Prasad A, Tay B-H, Garg KM, Chattopadhyay B, Yap L-M, Rheindt FE, Venkatesh B (2020) Chromosome-level assembly of the horseshoe crab genome provides insights into its genome evolution. Nat Commun 11:2322. https://doi.org/10.1038/s41467-020-16180-1
Simakov O, Kawashima T, Marlétaz F, Jenkins J, Koyanagi R, Mitros T, Hisata K, Bredeson J, Shoguchi E, Gyoja F, Yue J-X, Chen Y-C, Freeman RM, Sasaki A, Hikosaka-Katayama T, Sato A, Fujie M, Baughman KW, Levine J et al (2015) Hemichordate genomes and deuterostome origins. Nature 257:459–465. https://doi.org/10.1038/nature16150
Stebbing ARD (1970a) The status and ecology of Rhabdopleura compacta (Hemichordata) from Plymouth. J Mar Biol Assoc UK 50:209–221. https://doi.org/10.1017/S0025315400000722
Stebbing ARD (1970b) Aspects of the reproduction and life cycle of Rhabdopleura compacta (Hemichordata). Mar Biol 5:205–212. https://doi.org/10.1007/bf00346908
Stebbing ARD (1973) The larva of Rhabdopleura compacta (Hemichordata). Mar Biol 18:69–86. https://doi.org/10.1007/BF00347923
Strano F, Micaroni V, Beli E, Mercurio S, Scarì G, Pennati R, Piraino S (2019) On the larva and the zooid of the pterobranch Rhabdopleura recondita Beli, Cameron and Piraino, 2018 (Hemichordata, Graptolithina). Mar Biodivers 49:1657–1666. https://doi.org/10.1007/s12526-018-0933-2
Suyama M, Torrents D, Bork P (2006) PAL2NAL: robust conversion of protein sequence alignments into the corresponding codon alignments. Nucleic Acids Res 34 (Web Server Issue):W609–W612. https://doi.org/10.1093/nar/gkl315
Swalla BJ, van der Land J (2022) Hemichordata World Database. Rhabdopleura Allman, 1869. Accessed through: World Register of Marine Species at: https://www.marinespecies.org/aphia.php?p=taxdetails&id=137594 on 2022-05-06
Tassia MG, Cannon JT, Konifoff CE, Shenkar N, Halanych KM, Swalla BJ (2016) The global diversity of Hemichordata. PLoS ONE 11(10):e0162564. https://doi.org/10.1371/journal.pone.0162564
Taylor PD, Gordon DP (2003) Endemic new cyclostome bryozoans from Spirits Bay, a New Zealand marine-biodiversity “hotspot”. NZ J Mar Freshw Res 37:653–669. https://doi.org/10.1080/00288330.2003.9517196
Taylor PD, Todd JA (2001) Bioimmuration. In: Briggs DEG, Crowther PR (eds) Palaeobiology II. Blackwell Science, Oxford, pp 285–289
Tempestini A, Massamba-N’Siala G, Vermandele F, Beaudreau N, Mortz N, Dufresne F, Calosi P (2020) Extensive gene rearrangements in the mitogenomes of congeneric annelid species and insights on the evolutionary history of the genus Ophryotrocha. BMC Genomics 21:815. https://doi.org/10.1186/s12864-020-07176-8
Urbanek A, Dilly PN (2000) The stolon system in Rhabdopleura compacta (Hemichordata) and its phylogenetic implications. Acta Palaeontol Pol 45:201–226
Wheeler DL, Barrett T, Benson DA, Bryant SH, Canese K, Chetvernin V, Church DM, DiCuccio M, Edgar R, Federhen S, Geer LY, Kapustin Y, Khovayko O, Landsman D, Lipman DJ, Madden TL, Maglott DR, Ostell J, Miller V et al (2007) Database resources of the National Center for Biotechnology Information. Nucleic Acids Res 36(Suppl. 1):D13–D21. https://doi.org/10.1093/nar/gkl1031
Zhang S-Q, Che L-H, Li Y, Liang D, Pang H, Ślipiński A, Zhang P (2018) Evolutionary history of Coleoptera revealed by extensive sampling of genes and species. Nat Commun 9:205. https://doi.org/10.1038/s41467-017-02644-4
Acknowledgements
We thank the anonymous reviewers and the associate editor for comments that improved this manuscript. NIWA Invertebrate Collection staff, Wellington, are thanked for their assistance concerning specimen processing and data. Field collection in Singapore was authorized by the National Parks Board (permit number NP/RP19-006). We thank Sudanshi S. Jain (National University of Singapore) and Dr. Piotr Kuklinski (Institute of Oceanology, Polish Academy of Sciences) for field assistance in Singapore. Professor Abby Smith (University of Otago, New Zealand) organized two cruises (funded by the ERC grant to L.H. Liow) that supplied some New Zealand specimens for study (processed by Dr Hannah Mello). Principal Curator Miranda Lowe, Natural History Museum, London, is thanked for making available type and related material for study.
Funding
Open access funding provided by University of Oslo (incl Oslo University Hospital). This project received funding from the European Research Council (ERC) under the European Union’s Horizon 2020 research and innovation programme (grant agreement no. 724324 to L.H. Liow) and the National Research Foundation, Prime Minister’s Office, Singapore, under its Marine Science R&D Programme (grant number MSRDP-P03 to D. Huang). Some samples were collected during the following research programs: “Seamounts: their importance to fisheries and marine ecosystems”, undertaken by the NIWA and funded by the former New Zealand Foundation for Research, Science and Technology (C01X0028 (SFA023)) with additional funding from the former Ministry of Fisheries (project no. ZBD2001/10); and the Scientific Observer Program funded by the New Zealand Ministry for Primary Industries.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Ethical approval
All applicable international, national and/or institutional guidelines for animal testing, animal care and use of animals were followed by the authors.
Sampling and field studies
All necessary permits for sampling and observational field studies have been obtained by the authors from the competent authorities and are mentioned in the Acknowledgements, if applicable. The study is compliant with CBD and Nagoya protocols.
Data availability
Numerical data are tabulated in the article. Genetic data are published in GenBank and accession numbers are available in Table S1.
Author contribution
Conceptualization: Dennis Gordon, Randolph Quek, Lee Hsiang Liow. Methodology: Dennis Gordon, Randolph Quek, Russell Orr, Andrea Waeschenbach, Danwei Huang, Francesca Strano, Mali Ramsfjell. Formal analysis and investigation: Dennis Gordon, Randolph Quek, Russell Orr. The first draft of the manuscript was written by Dennis Gordon and Randolph Quek and all authors commented on previous versions of the manuscript. All authors read and approved the final manuscript.
Additional information
Communicated by K. Kocot
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
ESM 1
(XLSX 88 kb)
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Gordon, D.P., Quek, Z.B.R., Orr, R.J.S. et al. Morphological diversity and a ribosomal phylogeny of Rhabdopleura (Hemichordata: Graptolithina) from the Western Pacific (Singapore and New Zealand), with implications for a re-evaluation of rhabdopleurid species diversity. Mar. Biodivers. 53, 4 (2023). https://doi.org/10.1007/s12526-022-01310-3
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s12526-022-01310-3