Abstract
Species delimitation in sternaspid polychaetes is currently based on the morphology of a limited suite of characters, namely characters of the ventro-caudal shield—a unique feature of the family. Sternaspid species description has increased rapidly in recent years; however, the validity of the shield as a diagnostic character has not been assessed through molecular means. This study performs the largest molecular taxonomy of Sternaspidae to date, using the nuclear gene 18S, and the mitochondrial genes 16S and cytochrome oxidase subunit I (COI) to assess phylogenetic relationships within the family, to reassess the placement of Sternaspidae within the wider polychaete tree and to investigate the effectiveness of the shield as a diagnostic morphological character. This study includes many new records and reports Sternaspis affinis Stimpson, 1864 from USA Pacific coastline and genetic connectivity between specimens identified as Sternaspis cf. annenkovae Salazar-Vallejo & Buzhinskaja, 2013 from off southeastern Australia and specimens identified as Sternaspis cf. williamsae Salazar-Vallejo & Buzhinskaja, 2013 from the northwestern Pacific. In addition, we investigate material identified as Sternaspis cf. scutata (Ranzani, 1817) in the English Channel and compare with S. scutata through both molecular and morphological means. We further perform a detailed morphological and molecular investigation of new sternaspid material collected from the Southern Ocean and Antarctic Peninsula and regard Sternaspis monroi Salazar-Vallejo, 2014 syn. n. as a junior synonym of Sternaspis sendalli Salazar-Vallejo, 2014, two species recently described from the region, raising questions concerning the validity of current morphological delimitation.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Sternaspid polychaetes are widespread and often abundant in sedimented sea floors, reported globally from a variety of substrates, including gravelly muds (Hartman 1963), coarse sand, broken shells, soft mud (Treadwell 1914) and deep-sea clays and muds (Rouse and Pleijel 2001), at depths ranging from low intertidal to at least 6489 m (Sendall 2006; Salazar-Vallejo and Buzhinskaja 2013). Commonly known as mud owls (Sendall and Salazar-Vallejo 2013), these distinctive round-bodied or peanut-shaped worms are easily recognized by their characteristic and often colourful ventro-caudal shield. Currently, Sternaspidae is comprised of 43 species in three genera, with the largest genus, Sternaspis Otto, 1821 (Annelida: Sternaspidae Carus, 1863), containing 33 species (Salazar-Vallejo 2017). Both morphological assessments (Rouse and Fauchald 1997; Rouse and Pleijel 2001) and molecular phylogenetics (Rousset et al. 2007; Struck et al. 2007; Struck et al. 2008; Andrade et al. 2015) consistently place Sternaspidae as a sister taxon to Fauveliopsidae; however, the affinities of this sister pair to other families remains unclear (Osborn and Rouse 2011). Moreover, assessments of within-Sternaspidae relationships are scant, primarily limited to morphological cladistics (Sendall 2006), with only a single molecular investigation conducted thus far (Kobayashi et al. 2018).
Sternaspids can be abundant and even dominant in some ecosystems, with Sternaspis species reported to be amongst the most abundant benthic species in areas of southern Chile (Rozbaczylo et al. 2006); Jiaozhou Bay, China (Wang et al. 2006); the southwestern coast of India (Joydas and Damodaran 2009); shallow muddy bottoms in Bahia, Brazil (Pires-Vanin et al. 2011); Arctic waters (Balsom 2003); and the northwestern Mediterranean (Labrune et al. 2007; Harmelin-Vivien et al. 2009; Lorenti et al. 2011) and abundant year-round in seasonally hypoxia-stressed soft bottoms of Ariake Bay, Japan (Yoshino et al. 2010; Yoshino et al. 2014; Yoshino et al. 2016). Since the 1980s, Sternaspis scutata (Ranzani, 1817) has been reported to have greatly expanded its range into UK waters, where it is regarded as a non-native species (Townsend et al. 2006; Shelley et al. 2008).
Considered to be motile deposit feeders (Jumars et al. 2015), sternaspids have been observed as vigorous burrowers that rarely approach the sediment surface interface (Dorgan et al. 2006). Feeding and moving are facilitated by a completely eversible section of the body known as the introvert (see video of live specimen in Online Resource 1), which includes the head and thorax; the first three chaetigers are armed externally with curved hooks or spines (Fig. 1). Whole or partial invagination of this introvert often occurs during specimen fixation (Méndez and Yáñez-Rivera 2015) meaning that potential characters for species identification and delimitation, in addition to metrics such as body size, are often not observable. Other, more visible characters, such as features of the chaetae are highly conserved amongst sternaspids (Sendall and Salazar-Vallejo 2013), thus limiting the number of potential diagnostic morphological characters within the family.
In 2013, the family underwent a major revision (Sendall and Salazar-Vallejo 2013) that established a standardized method of identification and description based on the morphology of the ventro-caudal shield, where the shield is argued to be the best diagnostic morphological feature of the family. The shield, composed of mineralised iron (Bartolomaeus 1998), is located on the posterior-ventral surface of the worm and is made of two symmetrical sclerotinized plates (Fig. 1), divided along the mid-ventral axis and covered by a thick cuticle (Vejdovsky 1882). Features of the shield can vary between species and are used as diagnostic characters (Sendall and Salazar-Vallejo 2013); however, intraspecific plasticity and ontological variation in these characters have been observed across many species (Sendall and Salazar-Vallejo 2013).
In the years since this review, 26 new species have been described from locations around the globe using these morphological criteria; the current number of species is almost treble the number of species prior to 2013. Many of these new species have been described from relatively conservative geographic ranges. For example, three species have been described from waters around Vietnam (Zhadan et al. 2017), six from the South China Sea (Wu et al. 2015; Wu and Xu 2017), five from the tropical-subtropical Eastern Pacific (Salazar-Vallejo and Buzhinskaja 2013; Salazar-Vallejo 2017) and two from the Scotia Sea (Salazar-Vallejo 2014). For some species, materials have been scarce such that caudal shield features, particularly in terms of ontological and intraspecific variation, are poorly understood. Furthermore, there has been no genetic work to test whether variations in shield morphology fall within a consistent range so that morphological species delimitation remains practical and to affirm the usage of shield characteristics for delimitation, despite many new species displaying overlapping or sympatric geographic ranges.
A lack of reliable diagnostic morphological characters can lead to two essential problems: firstly, the “low morphology problem” (Van Oppen et al. 1996), whereby the diversity of morphologically similar yet genetically distinct species is greatly underestimated (Knowlton 1993) and secondly, the “cosmopolitan syndrome”, whereby the lumping together of such species inflates geographical range, leading to cosmopolitan distributions reported and hypothesised for morphologically simple species—a biogeographic artefact resulting from over-conservative taxonomy (Thorpe and Solé-Cava 1994; Klautau et al. 1999; Knowlton 2000; Dawson and Jacobs 2001; Gómez et al. 2002). This phenomenon is common amongst many marine invertebrate species and is particularly frequent amongst polychaetes (Westheide and Schmidt 2003; Bleidorn et al. 2006).
Building on morphological studies that began in the late 1980s, numerous molecular investigations have shown that many previously considered cosmopolitan polychaete species in fact consist of species complexes—groups of closely related species that are morphologically cryptic yet genetically distinct; each individual species often possesses a geographic range that is smaller than that of the complex as a whole (Nygren 2014, and references therein). However, following genetic investigation, not all widely dispersed polychaete species are necessarily composed of species complexes and may genuinely have broad and cosmopolitan distributions, even in species with short-lived or direct developing larvae (e.g. Westheide et al. 2003; Schulze 2006; Meyer et al. 2008; Ahrens et al. 2013; Georgieva et al. 2015; Eilertsen et al. 2018).
Sternaspid species are often reported as having global distributions. In particular, Sternaspis scutata has been recorded worldwide from every ocean (Sendall and Salazar-Vallejo 2013, and references therein) and is the most represented sternaspid species in the literature (Sendall 2006). Sendall and Salazar-Vallejo (2013) drastically reduced the known distribution of S. scutata to encompass the Mediterranean Sea and the English Channel, suggesting that records from its previous range belong to other described and possibly undescribed sternaspids. This alludes to the phenomenon of discovering many distinct species within the range of a previously assumed cosmopolitan polychaete. However, at present only one genetic investigation of sternaspids and their distribution has been undertaken (Kobayashi et al. 2018). This study found evidence of multiple unidentified species amongst GenBank specimens recorded as S. scutata from non-type localities, though also revealed intraspecific connectivity over a large geographic range in a case study of deep-water specimens from the northwestern Pacific Ocean identified as Sternaspis cf. williamsae Salazar-Vallejo & Buzhinskaja, 2013.
These results emphasize the need for further genetic investigations of sternaspid distributions—morphologically similar species do not necessarily share the same biological characteristics (Nygren 2014), and it is therefore imperative that an accurate taxonomy and distribution of species is determined, as misinterpretation can have wider implications for ecological and conservational assessments and evaluations.
This study aims to conduct a molecular phylogeny of Sternaspidae using all available data, in addition to re-assessing the placement of Sternaspidae within the wider polychaete tree. Within-family phylogenetic analyses include new material from the western Pacific, from deep waters off southeastern Australia and from the English Channel, the Mediterranean Sea and the Southern Ocean, in addition to newly published sequence data from the South China, East China, Bohai and Yellow seas and the northwestern Pacific from Japan and the Kuril Islands. We also assess the diagnostic power of the ventro-caudal shield through morphological and molecular investigations of shield variation in English Channel and Southern Ocean material, leading to the synonymy of two Southern Ocean species recently described from the region, previously differentiated by features of the ventro-caudal shield (Salazar-Vallejo 2014), Sternaspis sendalli Salazar-Vallejo, 2014 and Sternaspis monroi Salazar-Vallejo, 2014 syn. n.
Materials and methods
Sampling localities and specimens
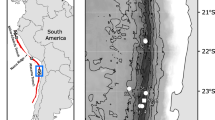
Sternaspid material was obtained from a variety of localities and sources (Fig. 2; Table 1; see Online Resource 2 for detailed material table): whole specimens from the Natural History Museum London (NHMUK) collections, specimen tissue vouchers from other institutions and sequence data, either from other institutions or downloaded from NCBI GenBank (https://www.ncbi.nlm.nih.gov/genbank/) (see Online Resource 3 for accession numbers). Type material of Sternaspis monroi and S. sendalli were available from NHMUK collections for morphological analyses.
The largest dataset examined in this study consisted of material collected from the Southern Ocean (Fig. 3) primarily from the British Antarctic Survey SO-AntEco (South Orkneys-State of the Antarctic Ecosystem) expedition on board the Royal Research Ship (RRS) James Clark Ross cruise JR15005, between February and March 2016. Material from localities further along the Antarctic Peninsula collected from RRS James Clark Ross cruises JR308 and JR144 was also available for examination (see Online Resource 2 for cruise and collection information). Sternaspids from these cruises were collected from a range of depths (455–1500 m) using either an Agassiz Trawl, Rauschert dredge, or Epi-benthic sledge and fixed onboard in 80% ethanol.
Distribution of sampling localities around the Southern Ocean and Antarctic Peninsula for all new Antarctic sternaspid material examined. SO, South Orkneys. Coastline data obtained from http://openstreetmapdata.com/, edited in QGIS
The next largest sample consisted of sternaspid material obtained from the UK during field sampling conducted between the 25th and 28th of June 2017 within Plymouth harbour. Benthic material was collected onboard the Research Vessel (RV) Callista, owned by the School of Ocean and Earth Science, University of Southampton. Sediment was collected using a 50-L Van Veen Grab sample and sorted for benthic invertebrates through a 1.0-mm mesh sieve. Live invertebrates were sorted, imaged, identified and fixed in 80% ethanol onboard.
Darwin Core (Wieczorek et al. 2012) and the GGBN data standard (Droege et al. 2016) were used in the management and transfer of specimen and derived sample data between the central museum collections database, a molecular collections database and external repositories (e.g. GenBank, WoRMS, OBIS, GBIF, GGBN, ZooBank).
Molecular taxonomy and analysis
Molecular analyses were conducted using data from two mitochondrial genes, cytochrome c oxidase subunit I (COI) and 16S, and one nuclear gene, 18S. The use of COI in single-gene analyses as per the Barcode of Life initiative (Hebert et al. 2003; Ratnasingham and Hebert 2007) has been proposed as an efficient method of detecting cryptic diversity in polychaetes (Carr et al. 2011) and in other studies can often (but not always) identify cryptic polychaete species without the need for other genes (Nygren 2014). However, COI can be difficult to obtain in some taxa (Brasier et al. 2016), and alternate mitochondrial genes such as 16S have also been used effectively in analyses of polychaete genetic diversity, either in addition to or replacing COI (e.g. Nygren et al. 2009; Wiklund et al. 2009a, 2009b; Brasier et al. 2016), and has been shown to fulfil multiple barcoding gene criteria despite a slower evolutionary rate than COI (Brasier et al. 2016). The longer and more conservative nuclear gene, 18S, has often been utilized in conjunction with mitochondrial genes for assessing deeper taxonomic relationships (Meißner and Blank 2009; Nygren et al. 2009; Wiklund et al. 2009b).
In total, 58 taxa were included in the combined three-gene phylogenetic analyses: 9 from Sternaspidae and 49 from 14 other polychaete families. Often regarded as a basal family in Polychaeta (e.g. Struck et al. 2007; Weigert et al. 2014), the amphinomid Amphinome rostrata (Pallas, 1766) was selected as the outgroup to root the tree (accession numbers for new sequence data can be found in Table 1; accession numbers of previously published sequences can be found in Online Resource 3). Separate 16S and COI datasets of all sternaspid sequences were examined in order to assess relationships within Sternaspidae, using taxa associated closely with Sternaspidae in the above phylogenetic analyses to root the tree.
Where whole specimens were available, the selection of specimens to dissect for DNA extraction was non-random; specimens were chosen based on biogeographic distribution and morphological variation in order to sample the widest range of both, respectively. Tissue was dissected from the dorsal or lateral abdomen body wall, leaving more character-rich body regions such as the ventro-caudal shield and posterior and anterior regions undamaged and available for potential morphological re-examination following DNA analyses.
The following protocol for DNA extraction, amplification and sequencing, and subsequent analyses followed the Laboratory Pipeline established by Glover et al. (2016). DNA was extracted with DNeasy Blood and Tissue Kit (Qiagen) using a Hamilton Microlab STAR Robotic Workstation. Approximately 1800 base pairs (bp) of 18S, 450 bp of 16S and 650 bp of COI were amplified using the primers listed in (Table 2). Polymerase chain reaction (PCR) mixtures consisted of 1 μl of each primer, 2 μl of template DNA and 20 μl of Red Taq DNA Polymerase 1.1X MasterMix (VWR), giving a mixture total of 24 μl. PCR programs were carried out in a Thermal Cycler with the following temperature profile: initial denaturation at 95 °C/5 min, 35 cycles of denaturation at 94 °C/45 s, annealing at 55 °C/45 s, extension at 72 °C for 2 min and a final extension at 72 °C for 10 min. Purification of PCR products was done using a Millipore Multiscreen 96-well PCR Purification System, and sequencing was carried out on an ABI 3730XL DNA Analyser (Applied Biosystems) at the Natural History Museum Sequencing Facility, using the same primers as in the PCR reactions (Table 2).
All newly examined material was sequenced for COI and 16S. Following identification of distinct clades from both genes, 18S was sequenced for representative specimens from each distinct genetic clade.
Fragments of overlapping sequences were merged into consensus sequences using Geneious v10.2.3 (Kearse et al. 2012) and aligned using the following Geneious plugins with default settings: MAFFT (Katoh and Standley 2013) for 16S and 18S, and MUSCLE (Edgar 2004) for COI.
MrBayes v3.2.6 (Ronquist et al. 2012) was used to conduct all Bayesian phylogenetic analyses. All analyses were run three times for 10,000,000 generations, with 2,500,000 generations discarded as burn-in. The software jModelTest v2.1.4 (Darriba et al. 2012) was used to estimate the best fitting nucleotide substitution model for each analysis using Akaike and Bayesian information criteria (AIC and BIC respectively). The model GTR + I + G was selected for each gene partition of the combined analysis and for the 16S within-sternaspid analysis; different models were obtained for each codon partition of the COI within-sternaspid analysis, with GTR + I selected for the first and second codon partitions and GTR + G for the third codon partition.
IQ-TREE (Nguyen et al. 2015) was also used to perform maximum likelihood (ML) analyses. Best fitting models for these analyses were found using the IQ-TREE ModelFinder function (Kalyaanamoorthy et al. 2017) based on the Bayesian information criterion (BIC)—in the combined analysis, the model TNe + R4 was found to be the best fit for 18S while the model GTR + F + I + G4 was found to be the best fit for both 16S and COI; for the within-sternaspid analyses, the best fitting model for 16S was TIM2 + F + G4, while the models TIMe + I, TPM3 + F + I and TIM2 + F + G4 were obtained for the first, second and third codon partitions of the COI analysis, respectively. All ML analyses were conducted with 1000 bootstrap pseudoreplicates using the ultrafast bootstrap approximation algorithm (Minh et al. 2013; Hoang et al. 2018). All trees were edited using FigTree v1.4 (Rambaut 2012) and Adobe Photoshop CS6.
Monophyletic clades determined from 16S and COI within-Sternaspidae tree topologies were assessed for both within and between-clade variation by a pairwise comparison of sequence divergence based on the proportion of nucleotide sites at which sequences differ, using the uncorrected p distance (Nei and Kumar 2000), which was calculated using Mesquite v3.2 (Maddison and Maddison 2017).
A separate COI dataset of all Southern Ocean sternaspids in which COI was successfully sequenced (28, out of 32 specimens) was generated for parsimony haplotype network analysis using TCS v1.21 (Clement et al. 2002) through the software PopART (http://popart.otago.ac.nz).
Morphological analysis
Specimens were examined and imaged using a Leica M216 stereomicroscope fitted with a Canon EOS600D camera, with a 1-mm scale bar placed alongside each specimen. Sediment particles adhering to the body and shield of the specimens were carefully removed using a fine brush, and the branchiae of some specimens were removed using fine forceps in order to examine branchial plates. The relative size of the shield can be used to extrapolate estimates of body size (Lim and Hong 1996; Méndez and Yáñez-Rivera 2015), which is otherwise difficult to obtain, as the introvert is often invaginated upon specimen fixation. The length and width of one ventral plate and the maximum length and width of the abdomen were measured from images taken for each specimen using the ImageJ v1.48 (Schneider et al. 2012) software. Abdomen length, as opposed to total body length was recorded to standardize measurements, as the introvert was not exposed in the majority of specimens examined. Scanning electron microscopy using a SEM FEI Quanta 650 was conducted on several whole specimens, following a preparation of graded ethanol dehydration, critical point drying and gold coating. Specimen morphology was described following approaches and terminology proposed by Sendall and Salazar-Vallejo (2013). In several taxon assignments, the open nomenclature ‘cf’ was used as a precautionary approach where morphological resemblance to known taxa was ambiguous.
Results
Placement of Sternaspidae within Annelida
Combined analyses of 18S, 16S and COI found Sternaspidae as sister to Scalibregmatidae with weak support and this sister pair in turn sister to Cirratuliformia (Acrocirridae, Flabelligeridae and Cirratulidae) with moderate support (Fig. 4). The position of Fauveliopsidae is often reported as sister to Sternaspidae in phylogenetic analyses (e.g. Rousset et al. 2007; Struck et al. 2007; Struck et al. 2008) and more recently in a phylogenomic analysis that utilized thousands of genes, again finding strong support for this sister pair (Andrade et al. 2015). The position of Fauveliopsidae in this study was unresolved—however, this is likely due to a lack of data, as the analysis was limited to three genes and only three fauveliopsid sequences were available for use, two 18S sequences and one 16S sequence (Online Resource 3), with one of the 18S sequences (Fauveliopsis sp.) shown to be particularly long branched in relation to other sequences in the dataset. Furthermore, basal support values across the tree were poorly resolved, limiting interpretation of inter-familial relationships.
Phylogeny of combined Bayesian analyses of three genes, 18S RNA, 16S RNA and cytochrome oxidase subunit I (COI) with 9 sternaspid taxa, 48 taxa from 12 Sedentaria polychaete families and one amphinomid as the outgroup. Vertical lines and text indicate different polychaete families, and a box highlights Sternaspidae. Support values are presented as maximum likelihood (ML) bootstrap values/Bayesian posterior probability values. Asterisks denote nodes where both bootstrap support and Bayesian posterior probabilities were ≥ 95 and 0.99 respectively. A hyphen indicates instances of no ML support
Systematics
Sternaspidae Carus, 1863
Sternaspis Otto, 1820
Sternaspis affinisStimpson, 1864
Material examined: All examined material loaned from the Scripps Institution of Oceanography Benthic Invertebrate Collection (SIO-BIC). Specimens SIO-BIC A5918 and SIO-BIC A6281, collected from East Sound, WA, USA (depth 25 m), and off Santa Barbara, CA, USA (depth 100 m), respectively (see Table 1; Online Resource 2). See also Online Resource 1 for footage of live specimen SIO-BIC-A5918.
Description: Preserved specimen SIO-BIC A5918 with a completely inverted introvert (Fig. 5a); SIO-BIC A6281 with introvert fully everted (Fig. 5c); Full body length of SIO-BIC A6281 11.7 mm, width 5.1 mm. Body light tan to beige in colour and covered with fine papillae; papillae are densest and largest on segments 7–9, becoming more widely spaced on the preceding six segments, and more fine on subsequent segments; body papillae are often encrusted with a fine sandy sediment.
Sternaspis affinis Stimpson, 1864. a Specimen SIO-BIC A5918; b live images of specimen SIO-BIC A5918, ventral and dorsal view (image credit: Greg Rouse); c specimen SIO-BIC A6281 (p = papillae); d dorsal view of introvert highlighting chaetal bundles in specimen SIO-BIC A6281; e dorsal view of lateral marginal fascicles highlighting oval arrangement of chaetae in specimen SIO-BIC A6281. All scale bars = 1 mm
Prostomium hemispherical, browner in colour than surrounding tissue and slightly opalescent; eyespots not visible; peristomium rounded, without obvious papillae; mouth region damaged in specimen SIO-BIC A6281 (Fig. 5c), but long papillae apparent on sections of mouth still visible.
First three chaetigers bearing bundles of 12–15 thick, bronze-coloured slightly falcate introvert hooks that become darker at the tips (Fig. 5c, d). One pair of long, digitate gonopodial lobes present on the ventral side between segments 7 and 8 (Fig. 5c). Abdomen consists of seven segments, with fine papillae that become denser, longer and more filament-like on the dorsum than on the ventral body wall, particularly towards the shield region.
Ventro-caudal shield ranging from brick red to orange in colour, with distinct concentric lines, particularly towards the shield margins; ribbing present throughout shield of SIO-BIC A6281 (Fig. 5c) but less distinct in the shield interior of SIO-BIC A5918 (Fig. 5a); anterior margins rounded, anterior keels slightly visible; anterior depression ranging from deep and triangular (Fig. 5c) to relatively more shallow and rounded (Fig. 5a). Suture visible throughout shield. Lateral margins slightly rounded, relatively straight. Posterio-lateral corners well developed in both specimens, demarked by a particularly large diagonal rib; posterior fan slightly expanded beyond corners; posterior margin distinctly crenulated in SIO-BIC A6281 (Fig. 5c), but more smooth in SIO-BIC A5918 (Fig. 5a); both specimens with distinct median and lateral notches; notches notably deep in SIO-BIC A6281 (Fig. 5c).
Marginal chaetal fascicles surround the shield, with 10 ovally arranged fascicles on either side of lateral margins (Fig. 5c) and five linearly arranged posterior fascicles per shield plate. Numerous thick, coiled branchiae protrude from branchial plates, interspersed with fine, long and filamentous papillae.
Remarks: These specimens match the neotype description of Sternaspis affinis in Sendall and Salazar-Vallejo (2013), with variation between the two specimens within the intraspecific variation reported in a case study of S. affinis morphology (also Sendall and Salazar-Vallejo 2013) and within general patterns of ontological variation reported for sternaspids. Furthermore, SIO-BIC A5918 was collected from East Sound, WA, USA—roughly 60 km from the type locality of S. affinis, off Vancouver, BC, Canada.
Genetic data (see sections on within-Sternaspidae Phylogenetics and Population genetics) revealed relatively low intraspecific variation between SIO-BIC A5918 and SIO-BIC A6281 in both 16S and COI analyses, despite a geographic distance of approximately 1800 km between Washington and California collection sites. Furthermore, these specimens fell into a clade with GenBank COI sequence data identified as Sternaspis fossor Stimpson, 1854 collected from off Bamfield, BC, Canada, approximately 250 km from the SIO-BIC A5918 collection site (see Fig. 17 in later results section). The geographical location of the 16S S. fossor GenBank sequence is unknown.
Sternaspis affinis and S. fossor are morphologically similar species; until its neotype description and reinstatement in 2013, S. affinis was considered to be a junior synonym of S. fossor (Sendall and Salazar-Vallejo 2013). Sternaspis fossor is now known to differ from S. affinis primarily in that posterior corners and ribbing of the ventro-caudal shield are poorly developed and that it is reported from the northwestern Atlantic from along the East Coast of Canada and the USA, whereas S. affinis is reported from the eastern Pacific, from Alaska to California (Sendall and Salazar-Vallejo 2013). The GenBank S. fossor sequences were published in 2011, i.e. before S. affinis was reinstated as a species in 2013; it is thus likely that these sequences are currently misidentified and are in fact S. affinis, based both on geographic proximity to the type locality of S. affinis and the phylogenetic and population genetic relationships revealed in this study (see Figs. 17 and 18a).
Sternaspiscf.annenkovaeSalazar-Vallejo & Buzhinskaja, 2013
Material examined: Specimens IN2017_V03-040-138 and IN2017_V03-040-139, collected off southeastern Australia, 2500 m (see Table 1; Online Resource 2).
Description: Two specimens, both with introvert everted (Fig. 6a, b)—the introvert is fully inflated and bulbous in specimen IN2017_V03-040-139, with segments between the introvert and the rest of the body appearing cinched (Fig. 6b). Body slightly bi-coloured in both specimens, with abdomen opaque and light tan in colour, and the introvert slightly darker and orange. Body finely papillose, with a fine silty sediment adhering to papillae in both specimens and with papillae largest and densest on segments 7 and 8 and; body papillae more eroded on IN2017_V03-040-139. Both specimens similar in size, with a body length and width of 11.7 mm and 5.2 mm for IN2017_V03-040-138 and 11.4 mm and 4.6 mm for IN2017_V03-040-139.
Prostomium projected, rounded and slightly conical, similar in colour to surrounding tissue; eyespots not observed; peristomium rounded; mouth rounded, heavily papillose and particularly dark in specimen IN2017_V03-040-138 (Fig. 6c)—it is unclear if this is due to dark coloured sediment adhering to the mouth, in contrast to the pale sediment that encrusts the body, or whether this is true pigmentation.
Introvert chaetigers bearing bundles of 10–12 brassy, slightly falcate hooks that are darker at the distal tips (Fig. 6c, d). Gonopodial lobes not clearly observable and could either be eroded or hidden between segmental folds of segments 7 and 8.
Abdomen with around seven segments, papillose, with denser bands of papillae somewhat observable on segments close to the shield (Fig. 6a).
Ventro-caudal shield orange to brick red, with ribbing and fine concentric rings; inner rings at the anterior-most end of the shield form a plate that appears to be somewhat raised relative to the rest of the shield; suture visible throughout shield; anterior depression relatively shallow; anterior margins somewhat angular; lateral margins gently rounded; posterio-lateral corners not distinct; posterior margin straight, faintly crenulated; fan barely projected past posterior corners; lateral notches not distinct; fans are somewhat divergent, forming a distinct, deep and triangular median notch, displays some medial fusion between; posterior margins slightly narrower than anterior margins.
Marginal chaetal fascicles with 9–10 lateral bundles on either side of the shield, with chaetae arranged ovally, and 5–6 posterior fascicles per shield plate, with linearly arranged chaetae.
Branchiae mostly eroded, but where present range from thick to slender, and coiled to relatively straight, interspersed with dense filamentous branchial papillae; branchial plates narrow and rounded, slightly wider anteriorly and curved around the anus.
Remarks: These specimens somewhat resemble the deep-water Pacific species, Sternaspis annenkovae, in that they bear shields that are wider anteriorly, with divergent fans that form a deep median notch, and with indistinct lateral notches, poorly developed posterior corners, a straight posterior margin, ribbing and fine concentric lines. It is possible that these specimens also display some bi-coloration, a feature that is unique to S. annenkovae within the genus. However, the bi-colouration is subtle and not particularly distinct in these specimens when compared with S. annenkovae. Further, the anterior shield margins of these specimens are only slightly wider than posterior margins, whereas anterior margins are distinctly wider in S. annenkovae.
These specimens also bear similarities to two other deep-water pacific species, S. maureri Salazar-Vallejo & Buzhinskaja, 2013 and S. williamsae Salazar-Vallejo & Buzhinskaja, 2013 including bearing anteriorly expanded shields with truncate posterior margins. However, the shields of these specimens have more distinct concentric lines and are less anteriorly expanded than in S. maureri and S. williamsae. They also lack the lateral notches present in S. maureri and have more developed median notches than in S. williamsae—the median notch is barely developed in S. maureri. Further, the prostomium is not notably smaller than the mouth, as in S. maureri and S. williamsae.
It has been previously noted that S. annenkovae, S. maureri and S. willimasae all bear similarities (Salazar-Vallejo and Buzhinskaja 2013). Our Australian specimens mostly resemble S. annenkovae, closely followed by S. williamsae; however, it is difficult to definitively place the Australian specimens as one over the other, or as a morphologically distinct but separate species, particularly considering the fact that, with only two similar sized specimens available, both general and ontological intraspecific variation cannot be assessed. Due to this ambiguity, we identify these specimens as S. cf. annenkovae.
Interestingly, in terms of genetic data, these specimens appear to be closely related to other deep-water specimens identified in Kobayashi et al. (2018) as S. cf. williamsae from the northwestern Pacific, with relatively low interspecific variation between two clades (see section on within-Sternaspidae Phylogenetics), despite a geographic distance of over 9000 km. Without further sampling from sites between these disparate localities, it is again difficult to definitively identify these specimens. It is worth noting that the type locality of S. annenkovae off the northern Kuril islands is closer to S. cf. williamsae collection sites than S. cf. williamsae is to the type locality of S. williamsae, off Oregon, USA; conversely, the Australian specimens are closer to the type locality of S. williamsae than to S. annenkovae. Furthermore, Kobayashi et al. (2018) found considerable morphological variation within the S. cf. williamsae sample population, with some specimens resembling S. annenkovae more than S. williamsae.
Sternaspiscf.scutata(Ranzani, 1817)
Material examined: Twenty specimens (Figs. 7 and 8a–h) (identifier prefix P17_145) collected from the English Channel (depth 18 m), ranging in size from 5 to 20 mm in length, all with everted introverts (Fig. 7a–b).
Sternaspis cf. scutata (Ranzani, 1817) specimen P17_145_O; a whole body, ventral view; b SEM detail of anterior region (pr = prostomium; m = mouth); c SEM detail of ventral shield region and associated chaetal groups (cc = capillary chaetae; pc = peg chaetae; poch = posterior chaetal fascicles; lch = lateral chaetal fascicles); d SEM detail of shield surface, highlighting filamentous surface papillae (sp). Scale bars 1.0 mm (a–c) and 300 μm (d)
Sternaspis cf. scutata (Ranzani, 1817); a–h shield variation in English Channel (P17_145) specimens ranging in size from large to small; a specimen P17_145_A; b specimen P17_145_C; c specimen P17_145_F; d specimen P17_145_G; e Specimen P17_145_I; f specimen P17_145_M; g specimen P17_145_Q; h specimen P17_145_U; iSternaspis scutata specimen SIO-BIC A5986, whole animal. All scale bars = 1 mm
One Mediterranean specimen SIO-BIC A5986, collected off Rovinij, Croatia (depth 25 m), small (2.1 mm long abdomen), introvert fully inverted (Fig. 8i). Specimen loaned from the Scripps Institution of Oceanography Benthic Invertebrate Collection (see Table 1; Online Resource 2).
Description: Body leathery, beige and opaque in large specimens, becoming thinner, paler and more translucent in smaller specimens. Introvert somewhat paler than abdomen in specimens of all sizes. Body finely papillated, more eroded on introvert than on abdomen; papillae more visible in smaller specimens when contrasted against pale, transluscent integument (Fig. 8i).
Prostomium hemispherical, opalescent; eyespots not observed; peristomium rounded; mouth circular, densely papillated (Fig. 7b). First three chaetigers bearing bundles of 8–10 widely separated, slightly falcate hooks (Fig. 7b); hooks range in colour from bronze in larger specimens to pale gold in smaller specimens, but tend to have darker subdistal tips in specimens of all sizes. Gonopodial lobes often eroded, but when present long and somewhat cirriform, projecting lateroventrally between segments 7 and 8. Abdomen with seven segments; the abdomen begins to bear longitudinal wrinkles in larger specimens, particularly as the integument becomes thicker and more leathery.
Ventro-caudal shield ranging in colour from orange and red in small specimens to blue-black in the largest specimens; shield covered by integument bearing minute filamentous papillae (Fig. 7c, d), with the integument thickest in the smallest specimens (Fig. 8i) and becoming heavily eroded in the largest specimens (Fig. 8a, b). All shields with concentric lines and ribbing, though the latter less pronounced in smaller specimens; in the smallest specimen, SIO-BIC_A5986, concentric lines are barely visible through thick, papillated integument (Fig. 8i); thin suture somewhat visible throughout all shields, tending to be more distinct in the anterior-most regions and becoming less defined posteriorly; anterior depression deep, triangular and becoming slightly shallower and rounded in smaller specimens; anterior margins truncate (Fig. 8c, e) to angular (Fig. 8d, f); introvert not exposed; lateral margins relatively straight across specimens of all sizes, though appearing slightly more rounded in smaller specimens (Fig. 8h, i); posterior corners well developed, demarked by distinct diagonal rib; posterior margins ranging from smooth (Fig. 8e) to slightly crenulated (Fig. 8f); posterior fan ranging from projecting beyond posterior corners with a shallow median notch (Fig. 8a, b, e, g) to relatively truncate, not projecting past corners, with an indistinct median notch (Fig. 8d, f, h).
Marginal chaetal fascicles (Fig. 7c) with 9–10 lateral bundles of ovally arranged chaetae on either side of the shield, and six posterior bundles per plate, with chaetae in a slightly curved arrangement and with bundles slightly offset and parallel to each other (see Sendall 2006). Peg chaetae often broken or hidden by chaetae and branchiae, but when present, appear to be as long as the first lateral chaetal fascicle (Fig. 7c); peg-associated capillary chaetae sometimes present (Fig. 7c).
Branchiae abundant, thick and coiled; interbranchial papillae long, filamentous and abundant; branchial plates diverge from anus, thick and ovoid in shape and densely papillated.
Remarks: Specimens from the English Channel more or less agreed with the neotype description of Sternaspis scutata in Sendall and Salazar-Vallejo (2013). Interestingly, there was some variation in a key diagnostic character, the projection of the posterior fan, which ranged from projected beyond posterior corners and notched medially, to truncate, not projecting past corners, and with no distinct median notch.
Notably, a truncate posterior fan not projecting past posterio-lateral corners is a diagnostic character of S. thalassemoides Otto, 1821, another Mediterranean sternaspid, sympatric with S. scutata. Sternaspis thalassemoides and S. scutata are two morphologically similar species; the former had previously been regarded as a junior synonym of the latter, until 2013 when S. thalassemoides was reinstated as a species, based on the character of a truncate fan not expanding past posterio-lateral corners (Sendall and Salazar-Vallejo 2013). A third species, S. assimilis (Malmgren, 1867), originally described from the northeastern Atlantic, off Île de Ré, France, was initially considered to be a junior synonym of S. scutata (Sendall, 2006), but was then found to be more similar to, and thus suggested to be a junior synonym of, S. thalassemoides, as the fan is only slightly projected in this species (Sendall and Salazar-Vallejo 2013).
Our English Channel specimens included individuals of similar sizes with both typical S. scutata posterior margins (Fig. 8c, e) and more truncate S. thalassemoides-like margins (Fig. 8d, f), in addition to specimens with more intermediate, slightly projected fans, perhaps similar to S. assimilis. Some of this variation may be due to ontogenic variation, with shield features such as the posterior fan and posterior corners poorly developed in smaller specimens. However, both types of diagnostic shield margin were observed in moderately sized adults (Fig. 8c–f), and there was variation in the degree of fan projection and depth of median notch even amongst the largest of specimens (Fig. 8a, b). Despite the morphological variation observed, molecular analyses (see section on within-Sternaspidae phylogenetics and population genetics) found very little genetic structure within the English Channel samples.
The single Mediterranean specimen, SIO-BIC A5986, collected off Croatia in the Adriatic Sea was one of the smallest specimens examined, possibly juvenile/sub-juvenile. Thus, full ontogenic development of shield characters could not be assessed. However, no character of SIO-BIC A5986 (Fig. 8i) strongly conflicted with the diagnostic characters of S. scutata, with slight median notch and fan projection evident. Notably, the specimen closely resembled English Channel specimens of a similar size.
In molecular analyses, these specimens, in addition to sequence data from specimens identified as S. scutata from the Bay of Biscay, northeastern Atlantic and another Mediterranean specimen identified as S. scutata, SIO-BIC A1012 (Fig. 9) collected off Banyuls France, formed a monophyletic clade (see Figs. 16 and 17). However, while SIO-BIC A5986 was found to be almost identical to SIO-BIC A1012 and the northeastern Atlantic sequence, differing by only one nucleotide mutation in COI analyses (see Fig. 18b), English Channel specimens displayed moderate genetic isolation from both northeastern Atlantic and Mediterranean sequences (see Figs. 16 and 17, Table 5). The type locality of S. scutata is within Aegean Sea, in the Eastern Mediterranean; the recent neotype description of the species included specimens from the original type locality, though it is worth noting that this description also incorporated a large quantity of additional material from other localities, including individuals collected from off Rovinij, Croatia (the same locality as SIO-BIC A5986) and others from across the Mediterranean, along with several northeastern Atlantic specimens collected off Portugal and France (Sendall and Salazar-Vallejo 2013).
Both morphology and molecular data support the identification of SIO-BIC A5986 as Sternaspis scutata. However, as the English Channel specimens exhibited variable morphology and were relatively genetically isolated from specimens close to the type locality of the species, we cannot with confidence identify them as S. scutata and instead tentatively identify them as S. cf. scutata for the time being. Wider sampling and further molecular and morphological investigations of sternaspids from the English Channel to the Mediterranean are necessary to definitively confirm presence of S. scutata in the English Channel—such investigations could potentially lead to the reinstatement of S. assimilis if the English Channel population is found to be a cryptic species. Further studies within the Mediterranean would also be useful in re-assessing the species status of S. thalassimoides.
Sternaspis sendalliSalazar-Vallejo, 2014
Sternaspis monroi Salazar-Vallejo, 2014 syn. n.
Material examined: Sixty-five individuals from new Southern Ocean material were examined, with a partial or wholly everted introvert observed in only 12 specimens (Table 1; Online Resource 2; see Table 3 for locality information and body size metrics).
Type materials from Natural History Museum London collections were also examined: S. sendalli holotype (NHM 1930.10.8.2372–2400), paratypes (NHM 1930.10.8.2372–2400p), S. monroi holotype (NHM 1930.10.8.2372–2400R), paratypes (NHM 1930.10.8.2372–2400Rp).
Description: Body opaque, yellow to pale beige in colour (Figs. 10a, 11a, 12 and 13), with paler and more translucent body walls observed in very small specimens (Fig. 13); body papillae are significantly or partially eroded in many specimens, though when present are fine and evenly spaced, often encrusted with sand particles, and tending to become longer and denser on and around segments 7 and 8, and then sparser on the introvert; gonopodial lobes are often eroded, but observed as short, small and digitate if present (Fig. 11c).
Sternaspis sendalli Salazar-Vallejo, 2014, Specimen JR16_227_A from off the NW South Orkney Islands, site 227; a whole body, ventral view; b SEM anterior, ventral view, showing exposed introvert (pr = prostomium; m = mouth); c SEM posterior, ventral view, showing the ventro-caudal shield (lch = lateral shield chaetae; poch = posterior shield chaetae); d close up of the ventro-caudal shield, ventral view (sp = shield papillae); e, f close up images of posterior shield chaetae, showing “furry” fibrous outer sheath and smooth inner chaetal filament where outer sheath is broken (e). Scale bars 2 mm (a–c), 500 μm (d) and 20 μm (e–f)
Sternaspis sendalli Salazar-Vallejo, 2014, specimen JR16_210_N from off the NW South Orkney Islands, Site 210. a Whole body, ventral view; b posterior, dorsal view, showing branchial plates, with branchiae and interbranchial papillae mostly removed; c SEM anterior, ventral view, showing exposed introvert (m = mouth; gl = gonopodial lobes) (note: a broken branchial filament from the posterior end is tangled between the anterior chaetal hooks surrounding the mouth); d SEM anterior chaetal hooks, lateral view; e SEM posterior end, ventral view, showing the ventro-caudal shield (lch = lateral shield chaetae; poch = posterior shield chaetae). Scale bars 2.5 mm (a), 1 mm (b–c), 200 μm (d), 2 mm (e)
Variation in the ventro-caudal shield in Sternaspis sendalli Salazar-Vallejo, 2014 from new Southern Ocean material. Horizontal pairs represent specimens of similar sizes. Though few specimens possessed all diagnostic characters, left-hand column contains specimens with more S. sendalli-like characters, defined by a posterior fan that extends beyond the posterior-lateral corners, a wide median notch, and the presence of lateral notches, and the right-hand column contains specimens with more S. monroi-like characters (S. monroi Salazar-Vallejo, 2014), such as a truncate posterior fan, a narrow median notch, and no lateral notches; a specimen LI_EBS_1; b specimen JR16_210_L; c specimen JR16_316_A; d specimen JR16_38_A; e specimen JR16_63_A; f specimen JR16_38B JR16_210_L; g specimen JR16_227_A; h specimen JR16_210_S. All scale bars = 1 mm
Where visible, the prostomium is observed as pale, hemispherical and projected, with no discernable eyespots (Fig. 10a, b); peristomium round, with sparse fine papillae; mouth round, densely papillated (Figs. 10b and 11c).
The first three chaetigers bearing bundles of 6–7 introvert hooks (Figs. 10a, b and 11a, c); hooks slightly falcate, range in colour from brassy to dark brown in colour, though darker at the tips in all cases; hook tips often broken or damaged, particularly in larger specimens, however when intact are observed to have tapered tips (Figs. 10a, b and 11a, c, d).
Ventral shield colour varies, ranging from orange (Figs. 12a and 14) and brick red (Figs. 10a and 12b–d, f, g) to dark maroon (Fig. 12e), with grey to grey-brown shields (Figs. 11a and 12h) observed in 12 individuals; an integument bearing fine filamentous papillae covers most shield surfaces (Fig. 10d), though this varies in thickness, and is eroded in some specimens (Fig. 12c); diagonal ribs often visible though occasionally indistinct where integument is thick and intact (Fig. 12h); concentric lines are primarily fine and poorly defined to not visible at all, but are moderately defined in some specimens (Fig. 12e); suture consistently distinct, wide and visible throughout shield, tending to expand posteriorly in most specimens; anterior margins rounded to sub-angular with a shallow and wide anterior depression; anterior keels vary in terms of exposure (e.g. Fig. 12b, not visible; Fig. 12g, visible); lateral margins rounded, tending to expand posteriorly in large specimens; posterior corners well developed; posterior fan ranges from projecting past posterior corners, notched medially and laterally (Figs. 10a and 12a, c, e, g), to truncate, not significantly expanding beyond posterior corners, and with poorly defined notches (Figs. 11a and 12b, d, f), in additional to specimens with intermediate characters (e.g. Fig. 12h, wide median notch, slightly projected fan, poorly developed lateral notches); posterior margins slightly crenulated (Fig. 12f, g) to smooth (Fig. 12c, d).
Three sub-juvenile/juvenile Sternaspis sendalli Salazar-Vallejo, 2014 specimens from new Southern Ocean material, highlighting rounded orange shields, expanded anteriorly and increasingly translucent body walls with decreasing size; a specimen JR16_41_A; b specimen LI_EBS_3_A; c specimen LI_EBS_3_B. All scale bars = 1 mm
Marginal chaetal fascicles as nine lateral and five to six posterior chaetal bundles (Figs. 10 c and 11e), with all chaetae arranged in oblique rows; posterior chaetae often damaged; peg chaetae not observed, assumed to be damaged and/or hidden by sediment or interbranchial papillae; peg-associated capillary chaetae often present, though delicate and easily broken. SEM reveals that marginal chaetae are covered by a thick fibrous sheath (Fig. 10e, f) likely similar to the sheath and fibrous matrix that bind chaetal bundles to form peg chaetae, as observed using SEM by Zhadan et al. (2017).
Branchiae are often eroded and lost, though where present are abundant, long and coiled. Interbranchial papillae are long, dense and fine, often coated by sediment. Where branchiae are removed, branchial plates bordering the anal peduncle are observed to vary from narrow, curved and somewhat tapered anteriorly to anteriorly expanded and rounded (Figs. 11b and 13).
Juveniles: In small specimens and juveniles, the shield is rounded and smooth, with lateral margins expanded anteriorly and posterior corners indistinct (Fig. 14). With increasing body size, the lateral margins tend to expand posteriorly and the plates become somewhat less rounded, with corners, ribbing and margin crenulation tending to become more distinct and shield colour tending to darken (Figs. 12 and 14).
Type material: Type materials of Sternaspis monroi and S. sendalli were examined (Fig. 15) and found to agree with corresponding species descriptions outlined in Salazar-Vallejo (2014). Both species were described from material originally collected from the Scotia Sea off the South Orkney Islands, identified as S. scutata by Monro (1930). Both species are similar morphologically, differentiated primarily by characters of the ventro-caudal shield. According to the original description (Salazar-Vallejo 2014) barely defined concentric lines, fan projected past posterior corners, notched medially and laterally, and with a barely crenulated margin, characterize the shield of S. sendalli (Fig. 15a, c, e, g), whereas the shield of S. monroi is characterized by an absence of concentric lines, a truncate fan not projecting past posterior corners, a short narrow median notch without lateral notches and a crenulated posterior margin (Fig. 15b, d, f, h).
Left-hand column (a, c, e, g) Sternaspis sendalli Salazar-Vallejo, 2014 type specimen ventro-caudal shields; a holotype (NHM 1930.10.8.2372–2400); c, e, g paratypes (NHM 1930.10.8.2372–2400p), right-hand column (b, d, f, h) Sternaspis monroi Salazar-Vallejo, 2014 type specimen ventro-caudal shields; b holotype (NHM 1930.10.8.2372–2400R); d, f, h paratypes (NHM 1930.10.8.2372–2400Rp); h paratype shield with ontogenic deformity. All scale bars = 1 mm
Remarks: The morphological variation observed amongst the Antarctic specimens examined included specimens that agreed with the morphological descriptions of either Sternaspis sendalli (Figs. 10a and 12a, c, e) or S. monroi (Fig. 11a and 12b, d, f), as well as specimens with intermediate morphologies (e.g. Fig. 12h), that could not be with confidence allocated to one species over the other. The size of the animal did not appear to influence the presence of key diagnostic characters. For example, extended and truncate posterior fans were observed both in large and small specimens (Fig. 12).
Despite morphological variation, molecular analyses revealed very little genetic variation (see Figs. 16, 17 and 19; Tables 4 and 5) with no genetic structure based on geography or shield morphology observed. We therefore consider S. monroi Salazar-Vallejo, 2014 a new synonym of S. sendalli Salazar-Vallejo, 2014. As both species were first described in the same paper (Salazar-Vallejo 2014), we prefer to use the name of a current researcher, Kelly Sendall, who participated in the revision of the family in 2013.
Phylogeny obtained from Bayesian analysis of the 16S RNA gene showing the relationships between all Sternaspis spp. material obtained during the study (see Table 1; Online Resource 2), in addition to a GenBank sequence indicated by bold text (see Online Resource 3). Quotation marks are used where it is likely that the given species names of GenBank sequences are misidentifications. Vertical lines and text indicate different geographic localities. Support values are presented as maximum likelihood (ML) bootstrap values/Bayesian posterior probability values. Asterisks denote nodes where both bootstrap support and Bayesian posterior probabilities were ≥ 95 and 0.99 respectively. A hyphen indicates instances of no ML support
Phylogeny obtained from Bayesian analysis of the cytochrome oxidase subunit I (COI) gene showing the relationships between all Sternaspis spp. material obtained during the study (see Table 1; Online Resource 2), in addition to GenBank sequences indicated by bold text (see Online Resource 3). Quotation marks are used where it is likely that the given species names of GenBank sequences are misidentifications. Vertical lines indicate different geographic localities. Support values are presented as maximum likelihood (ML) bootstrap values/Bayesian posterior probability values. Asterisks denote nodes where both bootstrap support and Bayesian posterior probabilities were ≥ 95 and 0.99 respectively
Phylogenetic relationships within Sternaspis
Of the three genera that constitute Sternaspidae, there was no obtainable molecular data for Petersenaspis Sendall & Salazar-Vallejo, 2013, and only two sequences from a single species, Caulleryaspis nuda Salazar-Vallejo & Buzhinskaja, 2013, were available for Caulleryaspis Sendall & Salazar-Vallejo, 2013. Furthermore, the Caulleryaspis sequences in question were previously found to be nested within Sternaspis (Kobayashi et al. 2018), which either raises questions surrounding the validity of C. nuda and generic diagnostic characters or suggests that these specimens may have been misidentified; addressing either of these is beyond the scope of this study. Therefore, the following analyses were primarily based on Sternaspis spp.
The taxa included differed between 16S and COI analyses due to a variable availability of sequences, a poorer sequencing success rate for COI and fewer 16S sequences on GenBank relative to COI (Table 1; Online Resource 3). Different taxa and low sampling relative to the current number of valid species mean that phylogenetic and biogeographic interpretations are limited. Furthermore, deeper branches were often poorly resolved with low support values. However, several consistent patterns of note are observed for both phylogenies (Figs. 16 and 17).
S. affinis and S. fossor—West Coast North American clade
Whole specimen vouchers A5198 and A6281, collected from off the coast of Washington and California states respectively, and which matched the morphological description of Sternaspis affinis (see the ‘Systematics’ section) fell into a clade with specimens identified on GenBank as Sternaspis fossor in both 16S and COI analyses (maximum within-clade uncorrected p distances of 0.005 and 0.009 for 16S and COI analyses respectively (Tables 4 and 5)) (Figs. 16 and 17). As discussed earlier (see the Remarks section for S. affinis in the ‘Systematics’ section), it is likely that GenBank S. fossor sequences are misidentified S. affinis specimens—the entire West Coast North American clade therefore can be seen to represent a single species, S. affinis.
Sternaspis cf. annenkovae, S. cf. williamsae and other Pacific clades
As with Kobayashi et al. (2018), we also found specimens identified as Caulleryaspis cf. nuda (Sea of Okhotsk) to be nested within Sternaspis rather than as a separate genus, close to S. cf. williamsae (off eastern Japan and the Kuril Islands), with a minimum uncorrected p distance of 0.005 and 0.019 between C. cf. nuda and S. cf. williamsae clades in the 16S and COI analyses respectively (Figs. 16 and 17; Tables 4 and 5). This, and considering that S. cf. williamsae displayed relatively high intra-clade variability (maximum uncorrected p distance of 0.01 in COI analysis (Table 5)) highlights a close affinity between these northwestern Pacific species, as discussed in Kobayashi et al. (2018).
In both analyses, deep-water specimens from off the southeastern coast of Australia identified as S. cf. annenkovae (see the ‘Systematics’ section) fell close to the C. cf. nuda + S. cf. williamsae clade (minimum uncorrected p distance between this clade and S. cf. annenkovae clade of 0.007 for 16S and 0.035–0.038 for COI (Tables 4 and 5)) (Figs. 16 and 17). This is despite a geographic distance of up to ~ 9600 km between specimens in these clades.
Other closely related clades despite large geographic distances include GenBank S. scutata sequences from the Bay of Bengal and the Bering Sea (minimum uncorrected p distance of 0.021 between clades in COI analysis (Table 5)) and specimens from the East Coast of Japan and a GenBank S. scutata sequence from Southern Chile (minimum uncorrected p distance of 0.023 in COI analysis (Table 5)) (Fig. 17).
Conversely, a number of genetically discrete specimens were also observed from within relatively small geographic ranges, for example from around the islands of Japan (several unidentified species off the East Coast of Japan and S. costata Marenzeller, 1879 from the Ariake Sea, southwestern Japan) (Figs. 16 and 17) and within the South China Sea (S. radiata Wu & Xu, 2017 and S. spinosa Sluiter, 1882) (Fig. 16). In the COI analysis, Sternaspis sp. 7 (GK612), one of the unidentified Japanese specimens, was nested within the S. costata clade (maximum uncorrected p value of 0.005 (Table 5)) (Fig. 17). The reported range of S. costata spans from Sakhalin Island, Russia, along the eastern and southwestern coastlines of the Japanese archipelago, to the Philippines, with the neotype described from the Boso Peninsula, off Chiba, eastern Japan (Sendall and Salazar-Vallejo 2013)—Sternaspis sp. 7 (GK612) was collected close to the neotype locality of S. costata (Table 1; Online Resource 2), and based on both molecular and geographic data, we suggest identifying it as such. No 16S sequences were available for other S. costata specimens; however, in the 16S analysis, Sternaspis sp. 7 (GK612) was found to be closely associated with S. chinensis Wu, Salazar-Vallejo, & Xu, 2015 (minimum uncorrected p distance of 0.012 between clades (Table 4)) (Fig. 16), in turn sister to the S. radiata clade, forming a broader clade of shallow water East Asian species, excluding S. spinosa.
Sternaspis scutata
The two Sternaspis scutata specimens collected from Mediterranean formed a monophyletic clade in both analyses (maximum uncorrected p value of 0.002 for both 16S and COI analyses (Tables 4 and 5)). In the COI analysis (Fig. 17; Table 5), only one out of five GenBank sequences identified as S. scutata fell close to the Mediterranean S. scutata clade. This sequence was collected from the North Eastern Atlantic coast (the Bay of Biscay), with a minimum uncorrected p distance between this clade and the Mediterranean clade of 0.002, the same as the within-clade uncorrected p distance for Mediterranean specimens. Notably, several other sequences recorded on GenBank as S. scutata from non-Atlantic localities did not fall near this clade.
Sternaspids identified as S. cf. scutata from the English Channel formed a clade with low intra-clade variation in both analyses (maximum uncorrected p values of 0.002 and 0.0 for 16S and COI analyses respectively) and fell close to the Mediterranean S. scutata clade both in the 16S analysis (minimum uncorrected p distance between clades 0.002) and the COI analysis (minimum uncorrected p value between English Channel clade and both Mediterranean and North East Atlantic clades of 0.026) (Figs. 16 and 17; Tables 4 and 5).
In the COI analysis, a phenomenon in which nodes were supported by high bootstrap but low posterior probability values was observed between the two Mediterranean specimens within the larger S. scutata clade and within a handful of other clades, such as the West Coast North America clade (Fig. 17). The phenomenon of recovering high posterior probability but moderate bootstrap values is widespread (see Lewis et al. 2005 and references therein), though the opposite can also occur, depending on the data (Cummings et al. 2003). However, Bayesian posterior probability and maximum likelihood bootstrap are not equivalent measures of support (Alfaro et al. 2003), and the exact relationship between the two for any given dataset is complex (Cummings et al. 2003). The incongruence between support values in this analysis was only observed at internal, intraspecific nodes within often well-supported clades. Within these clades, low variation between sequences and low sampling coverage relative to the population may have possibly resulted in too little phylogenetic information to resolve internal topologies more accurately.
Intra-clade variation in new Southern Ocean material
In both analyses, Antarctic specimens identified as S. sendalli formed a monophyletic clade with low intra-clade variation, with a maximum uncorrected p distance of 0.003 in both 16S and COI analyses (Figs. 16 and 17; Tables 4 and 5).
Population genetic analyses
Haplotype network analyses were performed for several clades highlighted in COI phylogenetic analyses. A haplotype network (Fig. 18a) constructed for the clade of North American West Coast specimens (see Fig. 17) revealed interesting patterns of connectivity. The site of collection for sequences identified as Sternaspis fossor on GenBank, collected off Bamfield, British Columbia, Canada (Online Resource 3), is located approximately 250 km from the site of collection for S. affinis SIO-BIC A5918 from East Sound, Washington, USA; both Washington and British Columbia specimens are in turn roughly 1800 km from the site of collection for S. affinis SIO-BIC A6281, off Santa Barbara, CA, USA. However, a single British Columbia sequence, ‘S. fossor’ 1 (accession number: HM473681), displayed fewer nucleotide mutations between it and S. affinis SIO-BIC A5918 (nucleotide substitutions, n = 4) and with S. affinis SIO-BIC A6281 (n = 7) than with the rest of the S. fossor sequences collected from the same site (n = 8). Likewise, S. affinis SIO-BIC A5918 is closer in terms of genetic structure to S. affinis SIO-BIC A6281 (n = 7) than to the majority of the British Columbia specimens (n = 8), despite being much closer geographically. As discussed earlier (see relevant sections in ‘Systematics’ and ‘Phylogenetic relationships within Sternaspis’), it is likely that these GenBank sequences have been misidentified and are in fact S. affinis. Though moderate intraspecific variation is present in this clade, for example relative to the Southern Ocean clade (see the next section), these results show genetic structure over a large geographic range that is not necessarily influenced by geographic proximity.
Haplotype networks based on cytochrome oxidase subunit I (COI). Each circle represents a sampled haplotype, with circle size proportional to the frequency of that haplotype, indicated by the key. Each bar represents one mutation. Colour indicates collection locality, as listed in the key. Bold text indicates sequences sourced from GenBank. Quotation marks are used where it is likely that the given species names of GenBank sequences are misidentifications. a A network of specimens identified as Sternaspis affinis Stimpson, 1864 (Table 1; Online Resource 2) and sequences identified on GenBank as Sternaspis fossor Stimpson, 1854 (Online Resource 3) constructed using COI (585 bp); b a network of specimens identified as Sternaspis scutata (Ranzani, 1817) and S. cf. scutata (Table 1; Online Resources 2 and 3) constructed using COI (545 bp)
Conversely, in a COI haplotype network analysis of the clade consisting of the English Channel specimens identified as S. cf. scutata, and the North East Atlantic (Bay of Biscay) sequence and two Mediterranean sequences identified as S. scutata (Fig. 18b), the English Channel haplotype had a distance of n = 15 nucleotide mutations from the Bay of Biscay sequence, which in turn was separated by n = 1 nucleotide mutation from S. scutata SIO-BIC A5986 (collected off Rovinj, Croatia), itself n = 1 mutation away from S. scutata SIO-BIC A1012 (collected off Banyuls, France). This highlights a degree of genetic isolation between English Channel specimens and more southern Atlantic and Mediterranean specimens, which was also reflected in the uncorrected p values (Table 5) and appears to confirm the presence of non-Mediterranean S. scutata in the northeast Atlantic, as previously reported based on morphology (e.g. Sendall and Salazar-Vallejo 2013), while questioning the presence of the species further north in the English Channel.
It is worth noting that in this case, the Straits of Gibraltar may act as a geographic barrier to dispersal, in addition to the relatively large distances between the English Channel and Mediterranean collection sites (between roughly 3500–4500 km apart). However, the Bay of Biscay sequence also shares the same major geographic barrier and similar geographic distances to the Mediterranean sites, despite very low genetic distance from the Mediterranean specimens. Furthermore, the distance between English Channel S. cf. scutata and the Bay of Biscay sequences (roughly 800 km apart) is less than half the distance between those of S. affinis specimens from Washington and California, despite there being considerably less genetic distance between the North American specimens.
A 582-bp COI haplotype network analysis (figure not shown) of S. cf. williamsae and southwestern Pacific specimens identified as S. cf. annenkovae revealed the latter to be n = 20 nucleotide mutations from the network of S. cf. williamsae haplotypes, over a much greater distance of roughly 9000 km than in the above two analyses (see Kobayashi et al. 2018 for a COI Haplotype network analysis of S. cf. williamsae sequences from the northwestern Pacific).
Genetic structure in terms of geographic distribution and morphological variation
In total, 28 COI sequences were successfully obtained from the new Southern Ocean specimens, belonging to three haplotypes (haplotype a = 10 individuals, b = 17, c = 1) (Fig. 19). The three haplotypes differed by only two mutations, and no clear patterns in morphological characters, such as shield colour or posterior fan projection, were apparent for the different haplotypes. No pattern in relation to geographic distribution was observed either, with the two largest haplotypes (a = 10, b = 17) both including specimens from all major South Orkney Island localities, in addition to Adelaide Island localities—sites approximately 1400 km apart. The n = 10 haplotype is seen to be nested within the rest of the Antarctic clade observed in Fig. 17, though total intra-clade variation did not exceed 0.003 in terms of uncorrelated p value (Table 5). COI is a relatively fast-evolving gene (Hebert et al. 2003), and the lack of genetic variation over a large geographic area suggests a high degree of gene flow within the population and a lack of reproductive barriers.
Sternaspis sendalli Salazar-Vallejo, 2014 parsimony haplotype network of cytochrome c oxidase subunit I (COI) (572 bp) for new sternaspid material from the Southern Ocean. Different colours represent the distribution of haplotypes in relation to the geographic locality of specimens, indicated by the above legend, whereby SO is an abbreviation for the South Orkney Islands (see Fig. 3 for map of localities). Each circle represents a sampled haplotype, with circle size proportional to the frequency of that haplotype, indicated by the key. Haplotype a = 10 individuals, b = 17, c = 1. Each bar represents one mutation. The ventro-caudal shields from all individuals included in the network are shown, whereby specimens of the same haplotype and of the same geographic locality are blocked together. White scale bar represents 1 mm in all images
Discussion
Placement of Sternaspidae within Annelida
This study provides the largest molecular taxonomy of Sternaspis to date, in addition to the first focused molecular attempt at resolving the position of Sternaspidae in the wider polychaete tree. In the late 1990s, Sternaspidae was initially found to be part of a clade later named Cirratuliformia, alongside families such as Acrocirridae, Flabelligeridae, Cirratulidae and Fauveliopsidae (Rouse and Fauchald 1997). More recent molecular studies found Sternaspidae as sister to Fauveliopsidae, with the pair not closely related to other cirratuliformids (Rousset et al. 2007; Struck et al. 2007). However, though only included as outgroup taxa, a recent phylogenomic study of archiannelids using Illumina-based data found strong support for a sister pair of Sternaspidae and Fauveliopsidae that was in turn part of Cirratuliformia (Andrade et al. 2015). A number of morphological features also support the pairing of Sternaspidae with Fauveliopsidae (e.g. Rouse and Fauchald 1997)—some fauveliopsids have even been observed to bear a ventral cuticular shield on posterior segments (Petersen 2000), which may be homologous to the sternaspid ventral-caudal shield. While limited to three genes, the results of our study also suggested a close association of Sternaspidae to Cirratuliformia, though direct sister relationships to Sternaspidae were unclear, and basal relationships were poorly resolved overall. More recent still, a molecular phylogeny of Paraonidae, a family not included in this study, found Sternaspidae as sister to Paraonidae, though only one sternaspid species was included in the analysis, and no fauveliopsids (Langeneck et al. 2019). Greater taxonomic and genetic coverage, in addition to the use of omics techniques such as transcriptomic data, should be undertaken in future studies in order to resolve the position and affiliations of Sternaspidae within the wider polychaete tree.
Within-Sternaspis relationships and phylogeography
The phylogenetic analyses in this study revealed some notable results within the available material. For example, COI analyses found the GenBank sternaspid sequence collected from the Bay of Biscay to be the first record of Sternaspis scutata outside the Mediterranean confirmed by molecular data. However, other sternaspids identified as S. scutata on GenBank from the Bering Sea, the Bay of Bengal and Southern Chile were shown to be different species to each other and to Mediterranean S. scutata, supporting similar findings in Kobayashi et al. (2018) and further rejecting the reported cosmopolitan distribution of S. scutata. This also highlights the issue of misidentification of data deposited at public online databases such as GenBank. Species misidentification is anecdotally widespread within these databases (Morton 2018), with specific cases of marine invertebrate misidentification often reported (e.g. Lima et al. 2017; Morton 2018). In this case, the Chilean sequence likely belongs to S. chilensis Díaz-Díaz & Rozbaczylo, 2017, described from some of the same source material as the sequence (Maturana et al. 2011; Díaz-Díaz and Rozbaczylo 2017); however, the relatively low intra-clade variation between specimens from the Bering Sea and the Bay of Bengal is interesting as it suggests gene flow over a massive geographic area, as well as sandwiching South-East Asia, an apparent hotspot for sternaspid diversity based on current taxonomy (Fig. 2).
Kobayashi et al. (2018) also found connectivity and low genetic structure over a large geographic range in a case study of S. cf. williamsae from deep (3000–4550 m) North West Pacific waters around eastern Japan and the Kuril islands, with distances between specimens up to ~ 1500 km, similar to the 1400 km range of the Antarctic specimens investigated in the current study. Likewise, the authors also found high variability in shield morphology despite relatively low genetic structure, with some specimens morphologically resembling S. annenkovae, another deep-water Pacific species, originally described from waters around the Kuril Islands.
Our study, which utilized the sequence data submitted by Kobayashi et al. (2018) in addition to unpublished sequence data from the same source material, corroborated the results of connectivity within the S. cf. williamsae population and of the close relationship between specimens identified as Caulleryaspis cf. nuda and S. cf. williamsae, which formed a distinct clade in both studies. Remarkably, we found a close association between this clade and two deep-water (2500 m) specimens identified as S. cf. annenkovae collected from off the southeastern coast of Australia—a distance of just under 10,000 km away, highlighting connectivity across a massive geographic range (see Remarks section for S. cf. annenkovae in the ‘Systematics’ section). Connectivity over large geographic ranges in marine taxa can be determined by a number of factors, including oceanographic currents, the physiology and behaviour of larvae, and connectivity cycles that span multiple generations, as found for the ornate spiny lobster Panulirus ornatus (Fabricius, 1798), which similarly displays genetic homogeneity across a distributional range of up to 10,000 km from Australia to Vietnam (Dao et al. 2015). Anthropogenic factors may also play a role—dispersal by ships’ ballast water is thought to account for the connectivity between northwest Atlantic and northwest Pacific populations of the capitellid polychaete Capitella teleta Blake, Grassle & Eckelbarger, 2009 (Tomioka et al. 2016).
Considering the moderate intra-clade variation within S. cf. williamsae, the inter-clade distance between the northwestern Pacific clade and the Australian specimens is remarkably low. Sampling from across a distribution range is important for establishing species boundaries as geographically separate populations can be connected through several intermediate populations (Nygren 2014), and the small number of Australian specimens available and lack of specimens from geographic intermediaries makes it difficult to assess intraspecific rates of variation and whether genetic variation is continuous or discrete between the localities of the two clades. Notably, in terms of morphology the Australian specimens resemble not only S. annenkovae but also S. williamsae and S. maureri too—three species that were recently described from deep-water pacific localities (Salazar-Vallejo and Buzhinskaja 2013; see Fig. 2 for type localities).
Low genetic diversity over great horizontal distances has been reported for a number of deep-sea invertebrate taxa (e.g. Etter et al. 2011; Georgieva et al. 2015; Zhang et al. 2015; Havermans 2016; Kobayashi et al. 2018; Kobayashi and Araya 2018). Cold environments are known to slow larval metabolism (Shilling and Manahan 1994) and Kobayashi et al. (2018) suggest that the colder temperatures of deep-sea waters may arrest larval sternaspid metabolism and development, which, in addition to deep-water countercurrents that circulate along central Japan to the Aleutian Peninsula (Kawabe and Fujio 2010), allows for greater dispersion S. cf. williamsae over the northwestern Pacific. Furthermore, sternaspids appear to have lecithotrophic larvae (see Rouse and Pleijel 2001)—the large yolk reserves of lecithotrophic larvae are thought to further prolong the larval period and therefore larval dispersal ability at low temperatures (Jollivet et al. 1998). Perhaps it is these conditions, combined with a complex system of deep-water Pacific currents, that allow for even greater connectivity across the Pacific Ocean. Further molecular investigations will be needed to clarify the patterns of Pacific sternaspid connectivity and to assess the prospect of a cosmopolitan deep-water Pacific species or a pan-Pacific species complex.
The Antarctic specimens were also collected from relatively deep waters (~ 200–1500 m) and showed high connectivity along the western Antarctic Peninsula, reflecting the widespread distributional patterns recently found for many cryptic polychaete species collected from a similar depth range from across the West Antarctic (Brasier et al. 2017). However, Brasier et al. (2017) also found species from this region with highly restricted ranges, concluding that differences in Southern Ocean species distributions are more likely to rely on a complex of factors, rather than just a single factor such as oceanographic currents.
In contrast, our results also show instances of high genetic diversity within relatively small geographic ranges. For example, sequence data from two sympatric shallow water species from the South China Sea, S. radiata and S. spinosa were not closely related in 16S analyses (see Fig. 16; Table 4) despite the proximity of their collection sites (Online Resource 2). It has been suggested that, while cold deep-sea waters may slow larval development, larvae may develop much more rapidly in warmer, shallower waters, which could reduce dispersal ability, increasing barriers to gene flow and leading to higher diversity in the shallows (Kobayashi et al. 2018). Waters around the Japanese archipelago appear to host several genetically discrete sternaspid species from a range of depths, such as S. costata from shallow waters off eastern and southwestern Japan, and at least three unidentified species off eastern Japan, ranging in depth from 120 to 1682 m (see Table 1; Online Resource 2). These species did not tend to cluster together in either the 16S or COI analyses, but often with clades from other, often distant localities, perhaps suggesting bathymetric rather than geographic segregation in terms of species boundaries. In the 16S analysis, the close relationship between S. costata from Japan and S. chinensis from the East China, Yellow and Bohai Seas may reflect a recent isolation event; during the last glacial maximum 20–15 ka, the Japanese archipelago was connected to the Korean Peninsula and the continental coastline of China, with rising sea levels around 10 ka isolating Japan from the mainland and creating the modern day East China, Yellow and Bohai Seas (Kaizuka 1980). In general however, our capacity to make phylogeographic inferences in this study was restricted, as only a very limited coverage of total sternaspid diversity was accessible through sequence data, and many of the taxa available were singletons, or known from just one or two sequences (Table 1; Online Resources 2 and 3). Information on interpopulation connectivity and standard rates of intraspecific variation was therefore difficult to obtain due to low geographic and bathymetric coverage. Regardless, it is evident from our analyses that an interesting and complex picture in terms of phylogeographic relationships is present within this family and is worth further investigation, with the potential for both ocean-wide connectivity and cryptic species within small geographic ranges. Particular questions of interest to address include patterns of connectivity and isolation along the western North American coastline in the case of S. affinis and between the English Channel and the Mediterranean Sea concerning S. cf. scutata (see systematic remarks and population genetics sections on respective species) and, more generally, the effects of depth zonation on species boundaries.
Synonymy of S. sendalli with S. monroi syn. n.
Specimens from the Southern Ocean and Antarctic Peninsula collected from depths ranging from ~ 200 to 1500 m displayed considerable variation in terms of shield morphology, with shield morphotypes of both Sternaspis species described from the region (S. sendalli and S. monroi) present in the sample. However, molecular analyses of COI and 16S barcode regions revealed that all specimens formed a monophyletic clade with very low intra-clade variation despite a relatively large geographic range and no discernable patterns of genetic structure based on shield morphology or geography evident from haplotype network analyses of COI.
Interspecific variation ten times the mean intraspecific variation between clades has been suggested as a rule of thumb for delimiting cryptic species based on COI barcodes (Hebert et al. 2003) and has previously been used to assess polychaete cryptic diversity (e.g. Carr et al. 2011). Brasier et al. (2016) found that interspecific variation between cryptic clades in Antarctic polychaetes to be on average 20 times greater than the mean intraspecific variation for COI and ranging from 9 to 28 times greater than mean intraspecific variation for 16S, suggesting that a universal threshold for cryptic polychaete delimitation by barcode is questionable. Regardless, the intra-clade variation within the Antarctic clade was consistently low for both COI and 16S in terms of uncorrected p distance, and variation between the two largest COI haplotypes did not approach suggested threshold levels. Thus, we propose that S. monroi should be regarded as a junior synonym of S. sendalli due to no convincing molecular support for either different clades or species found within the sampled specimens, despite considerable morphological variation observed in the diagnostic shield characters currently used to delimit S. sendalli and S. monroi—this result raises questions surrounding the usage of shield morphology in Sternaspis species delimitation.
Variation in the ventro-caudal shield—challenges to morphological delimitation
Similar to the Southern Ocean material, English Channel specimens identified as Sternaspis cf. scutata also exhibited considerable morphological variation, particularly in terms of shield margin, with shield morphotypes resembling either S. thalassemoides or S. scutata, two sympatric species described from the Mediterranean Sea that are delimited based on shield margin characteristics. As with the Southern Ocean material, morphological variation in these characters in the English Channel specimens was not reflected in genetic structure.
The shield variation observed in both S. sendalli and S. cf. scutata has serious implications for sternaspid taxonomy as it stands today, as the majority of the 33 currently valid Sternaspis species have been described in roughly the past 5 years alone, primarily based on shield morphology and without corresponding molecular data. It is also worth noting however that several previously identified species did tend to fall into reciprocally monophyletic clades in our analyses, such as S. radiata and S. spinosa (Fig. 16), two relatively similar, sympatric shallow water species, suggesting that morphological delimitation may be valid for some species. It is vital that further molecular studies in conjunction with morphological analyses are carried out in order to fully test the validity of the ventro-caudal shield as a diagnostic character. Features of the shield are also amongst the diagnostic characters that delimit the two other sternaspid genera, Caulleryaspis and Petersenaspis, though there has been no molecular work to support the erection of either genus. Both this study and Kobayashi et al. (2018) found sequences identified as Caulleryaspis cf. nuda to be nested within Sternaspis as an immediate sister to S. cf. williamsae. Caulleryaspis nuda has previously been noted to resemble S. williamsae, particularly in terms of overall shield outline and the presence of a thin, loosely adhered layer of sediment on the shield, but with C. nuda differing by a more convex, delicate and pliable shield margin (Salazar-Vallejo and Buzhinskaja 2013). A soft, flexible ventro-caudal shield with a firmly attached layer of sediment was an original diagnostic feature of the genus Caulleryaspis (Sendall and Salazar-Vallejo 2013), which was then later amended to include sternaspids with soft shields but only loosely adhered sediment that could be easily brushed off, such as C. nuda (Salazar-Vallejo and Buzhinskaja 2013). Results of both Kobayashi et al. (2018) and the current study question the diagnostic power of a soft shield margin at the generic level; it is possible that firmly adhered sediment is a more informative feature, but this is unclear without more thorough investigations of these characters. The placement of C. nuda is also uncertain without an assessment of representatives from the type locality and greater sampling coverage of congeners in general, as is any potential revision of Caulleryaspis without inclusion of the generic type. Regardless, these results highlight the need for a rigorous molecular examination of the generic relationships within Sternaspidae and of the validity of generic diagnostic characters.
It is interesting to note the degree of ontogenic deformity present in type material of S. monroi syn. n.; additional studies examining the growth pattern of the shield would be useful in assessing any environmental influence on shield development and characters such as concentric growth lines. The degree to which epigenetic factors may influence the development of shield characters is unknown, but this knowledge could be vital in order to assess whether certain shield characters are determined genetically and therefore could remain important diagnostic characters.
A lack of reliable diagnostic morphological characters is not uncommon within Annelida. For example, diagnostic characters in the family Capitellidae such as chaetal arrangement have been shown to vary ontogenically within individuals (Ewing 1982; Fredette 1982; George 1984), and phylogenetic analyses have found several morphological characters traditionally used to delineate capitellids at the generic level be uninformative for many genera (e.g. Tomioka et al. 2018).
If attributes of the ventro-caudal shield are found to be unreliable diagnostic characters in future assessments, there is a possibility that many currently valid species may be composed of morphologically similar yet genetically distinct species. Conversely, multiple species may be in fact be part of a single species with considerable intraspecific morphological variability, as was found in S. sendalli. Where robust diagnostic morphological characters are lacking, a combined approach using as much data as possible been suggested in order to define species boundaries, where ecological, physiological and reproductive data may be utilized in addition to morphology and molecular data (see Nygren 2014 and references therein); future sternaspid taxonomy should utilize this more holistic approach, incorporating geographic and bathymetric data where possible, in addition to a robust molecular dataset, consisting of both nuclear and mitochondrial data. An accurate taxonomy and effective methods of identification are vital for documenting diversity, ecosystem management and conservation, particularly for vulnerable ecosystems experiencing rapid environmental change.
References
Ahrens JB, Borda E, Barroso R, Paiva PC, Campbell AM, Wolf A, Nugues MM, Rouse GW, Schulze A (2013) The curious case of Hermodice carunculata (Annelida: Amphinomidae): evidence for genetic homogeneity throughout the Atlantic Ocean and adjacent basins. Mol Ecol 22(8):2280–2291. https://doi.org/10.1111/mec.12263
Alfaro ME, Zoller S, Lutzoni F (2003) Bayes or bootstrap? A simulation study comparing the performance of Bayesian Markov chain Monte Carlo sampling and bootstrapping in assessing phylogenetic confidence. Mol Biol Evol 20(2):255–266. https://doi.org/10.1093/molbev/msg028
Andrade SCS, Novo M, Kawauchi GY, Worsaae K, Pleijel F, Giribet G, Rouse GW (2015) Articulating “archiannelids”: phylogenomics and annelid relationships, with emphasis on meiofaunal taxa. Mol Biol Evol 32(11):2860–2875. https://doi.org/10.1093/molbev/msv157
Balsom AL (2003) Macroinfaunal community composition and biomass, and bacterial and viral abundances from the Gulf of Alaska to the Canadian archipelago: a biodiversity study. Master’s Thesis, University of Tennessee
Bartolomaeus T (1998) Chaetogenesis in polychaetous Annelida—significance for annelid systematics and the position of the Pogonophora. Zoology 100(4):348–364
Blake JA, Grassle JP, Eckelbarger KJ (2009) Capitella teleta, a new species designation for the opportunistic and experimental Capitella sp. I, with a review of the literature for confirmed records. Zoosymposia 2:25–53
Bleidorn C, Podialowski L, Bartolomaeus T (2006) The complete mitochondrial genome of the orbiniid polychaete Orbinia latreilli (Annelida, Orbiniidae)—a novel gene order for Annelida and implications for annelid phylogeny. Gene 370:96–103. https://doi.org/10.1016/j.gene.2005.11.018
Brasier MJ, Wiklund H, Neal L, Jeffreys R, Linse K, Ruhl H, Glover AG (2016) DNA barcoding uncovers cryptic diversity in 50% of deep-sea Antarctic polychaetes. R Soc Open Sci 3(11):160432. https://doi.org/10.1098/rsos.160432
Brasier MJ, Harle J, Wiklund H, Jeffreys RM, Linse K, Ruhl HA, Glover AG (2017) Distributional patterns of polychaetes across the West Antarctic based on DNA barcoding and particle tracking analyses. Front Mar Sci 16(4):356. https://doi.org/10.3389/fmars.2017.00356
Carr CM, Hardy SM, Brown TM, Macdonald TA, Hebert PDN (2011) A tri-oceanic perspective: DNA barcoding reveals geographic structure and cryptic diversity in Canadian polychaetes. PLoS One 6(7):e22232. https://doi.org/10.1371/journal.pone.0022232
Carus JV (1863) Vermes. In: Peters WCH, Carus JV, Gerstäcker CEA (eds) Handbuch der zoologie. Wilhelm Engelmann, Leipzig, pp 422–484
Clement M, Snell Q, Walker P, Posada D, Crandall K (2002) TCS: estimating gene genealogies. IEEE IPDPS 0184 9(10):1657–1659. https://doi.org/10.1109/IPDPS.2002.1016585
Cohen BL, Gawthrop A, Cavalier-Smith T (1998) Molecular phylogeny of brachiopods and phoronids based on nuclear–encoded small sub-unit ribosomal RNA gene sequences. Philos Trans R Soc B 1524 353(1378):2039. https://doi.org/10.1098/rstb.1998.0351
Cummings MP, Handley SA, Myers DS, Reed DL, Rokas A, Winka K (2003) Comparing bootstrap and posterior probability values in the four-taxon case. Syst Biol 52(4):477–487
Dao HT, Smith-Keune C, Wolanski E, Jones CM, Jerry DR (2015) Oceanographic currents and local ecological knowledge indicate, and genetics does not refute, a contemporary pattern of larval dispersal for the ornate spiny lobster, Panulirus ornatus in the South-East Asian archipelago. PLoS One 10(5):e0124568. https://doi.org/10.1371/journal.pone.0124568
Darriba D, Taboada GL, Doallo R, Posada D (2012) jModelTest 2: more models, new heuristics and parallel computing. Nat Methods 9(8):772–772. https://doi.org/10.1038/nmeth.2109
Dawson MN, Jacobs DK (2001) Molecular evidence for cryptic species of Aurelia aurita (Cnidaria, Scyphozoa). Biol Bull 200(1):92–96. https://doi.org/10.2307/1543089
Díaz-Díaz O, Rozbaczylo N (2017) Sternaspis chilensis n. sp., a new species from austral Chilean channels and fjords (Annelida, Sternaspidae). Zootaxa 4254(2):269–276. https://doi.org/10.11646/zootaxa.4254.2.7
Dorgan K, Jumars P, Johnson B, Boudreau B (2006) Macrofaunal burrowing: the medium is the message. Oceanogr Mar Biol Annu Rev 44:85–121
Droege G, Barker K, Seberg O, Coddington J, Benson E, Berendsohn WG, Bunk B, Butler C, Cawsey EM, Deck J, Döring M (2016) The global genome biodiversity network (GGBN) data standard specification. Database 2016:1–11. https://doi.org/10.1093/database/baw125
Edgar RC (2004) MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res 32(5):1792–1797. https://doi.org/10.1093/nar/gkh340
Eilertsen MH, Georgieva MN, Kongsrud JA, Linse K, Wiklund H, Glover AG, Rapp HT (2018) Genetic connectivity from the Arctic to the Antarctic: Sclerolinum contortum and Nicomache lokii (Annelida) are both widespread in reducing environments. Sci Rep 8(1):4810
Etter RJ, Boyle EE, Glazier A, Jennings RM, Dutra E, Chase MR (2011) Phylogeography of a pan-Atlantic abyssal protobranch bivalve: implications for evolution in the deep Atlantic. Mol Ecol 20:829–843. https://doi.org/10.1111/j.1365-294X.2010.04978.x
Ewing RM (1982) A partial revision of the genus Notomastus (Polychaeta: Capitellidae) with a description of a new species from the Gulf of Mexico. Proc Biol Soc Wash 95:232–237
Fabricius JC (1798) Entomologia Systematica emendata et aucta, secundum classes, ordines, genera, species adjectis synonimis locis observationibus descriptionibus. Hafniae. I–IV. Supplementum Entomologiae Systematicae Copenhagen 1–572
Folmer O, Black M, Hoeh W, Lutz R, Vrijenhoek R (1994) DNA primers for amplification of mitochondrial cytochrome c oxidase subunit I from diverse metazoan invertebrates. Mol Mar Biol Biotechnol 3(5):294–299
Fredette TJ (1982) Evidence of ontogenetic setal changes in Heteromastus filiformis (Polychaeta: Capitellidae). Proc Biol Soc Wash 95:194–197
George JD (1984) The behavior and life history of a mangrove dwelling capitellid (Polychaeta). In: Hutchings PA (ed) Proceedings of the First International Polychaeta Conference, Sydney. The Linnean Society of New South Wales, Sydney, pp 323–337
Georgieva MN, Wiklund H, Bell JB, Eilertsen MH, Mills RA, Little CTS, Glover AG (2015) A chemosynthetic weed; the tubeworm Sclerolinum contortum is a bipolar cosmopolitan species. BMC Evol Biol 15(1):280. https://doi.org/10.1186/s12862-015-0559-y
Glover GA, Dahlgren GT, Wiklund H, Mohrbeck I, Smith RC (2016) An end-to-end DNA taxonomy methodology for benthic biodiversity survey in the Clarion-Clipperton Zone, central Pacific abyss. J Mar Sci Eng 4(1):2. https://doi.org/10.3390/jmse4010002
Gómez A, Serra M, Carvalho GR, Lunt DH, Baum D (2002) Speciation in ancient cryptic species complexes: evidence from the molecular phylogeny of Branchionus plicatilis (Rotifera). Evolution 56(7):1431–1444. https://doi.org/10.1554/0014-3820(2002)056[1431:SIACSC]2.0.CO;2
Harmelin-Vivien ML, Banaru D, Dierking J, Hermand R, Letourneur Y, Salen-Picard C (2009) Linking benthic biodiversity to the functioning of coastal ecosystems subjected to river runoff (NW Mediterranean). Anim Biodivers Conserv 32(2):135–145
Hartman O (1963) Submarine canyons of Southern California, 3. Systematics: polychaetes. Allan Hancock Pacific Exped 27:1–93
Havermans C (2016) Have we so far only seen the tip of the iceberg? Exploring species diversity and distribution of the giant amphipod Eurythenes. Biodiversity 17:12–25. https://doi.org/10.1080/14888386.2016.1172257
Hebert PDN, Cywinska A, Ball SL (2003) Biological identifications through DNA barcodes. Proc R Soc B 270(1512):313–321. https://doi.org/10.1098/rspb.2002.2218
Hoang DT, Chernomor O, von Haeseler A, Minh BQ, Vinh LS (2018) UFBoot2: improving the ultrafast bootstrap approximation. Mol Biol Evol 35:518–522. https://doi.org/10.1093/molbev/msx281
Jollivet D, Comtet T, Chevaldonné P, Hourdez S, Desbruyeres D, Dixon DR (1998) Unexpected relationship between dispersal strategies and speciation within the association Bathymodiolus (Bivalvia)-Branchipolynoe (Polychaeta) inferred from the rDNA neutral ITS2 marker. Cah Biol Mar 39:359–362
Joydas TV, Damodaran R (2009) Infaunal macrobenthos along the shelf waters of the west coast of India, Arabian Sea. Indian J Mar Sci 38(2):191–204
Jumars PA, Dorgan KM, Lindsay SM (2015) Diet of worms emended: an update of polychaete feeding guilds. Annu Rev Mar Sci 7(1):497–520. https://doi.org/10.1146/annurev-marine-010814-020007
Kaizuka S (1980) Late cenozoic palaeogeography of Japan. GeoJournal 4(2):101–109
Kalyaanamoorthy S, Minh BQ, Wong TKF, von Haeseler A, Jermiin LS (2017) ModelFinder: fast model selection for accurate phylogenetic estimates. Nat Methods 14:587–589. https://doi.org/10.1038/nmeth.4285
Katoh K, Standley DM (2013) MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol Biol Evol 30(4):772–780. https://doi.org/10.1093/molbev/mst010
Kawabe M, Fujio S (2010) Pacific Ocean circulation based on observation. J Oceanogr 66(3):389–403. https://doi.org/10.1007/s10872-010-0034-8
Kearse M, Moir R, Wilson A, Stones-Havas S, Cheung M, Sturrock S, Buxton S, Cooper A, Markowitz S, Duran C, Thierer T, Ashton B, Meintjes P, Drummond A (2012) Geneious basic: an integrated and extendable desktop software platform for the organization and analysis of sequence data. Bioinformatics 28(12):1647–1649. https://doi.org/10.1093/bioinformatics/bts199
Klautau M, Russo C, Lazoski C, Boury-Esnault N, Thorpe JP, Solé-Cava AM (1999) Does cosmopolitanism in morphologically simple species result from overconservative systematics? A case study using the marine sponge Chondrilla nucula. Evolution 53:1414–1422
Knowlton N (1993) Sibling species in the sea. Annu Rev Ecol Syst 24:189–216
Knowlton N (2000) Molecular genetic analyses of species boundaries in the sea. Hydrobiologia 420(1):73–90. https://doi.org/10.1023/A:1003933603879
Kobayashi G, Araya JF (2018) Southernmost records of Escarpia spicata and Lamellibrachia barhami (Annelida: Siboglinidae) confirmed with DNA obtained from dried tubes collected from undiscovered reducing environments in northern Chile. PLoS One 13(10):e0204959. https://doi.org/10.1371/journal.pone.0204959
Kobayashi G, Mukai R, Alalykina I, Miura T, Kojima S (2018) Phylogeography of benthic invertebrates in deep waters: a case study of Sternaspis cf. williamsae (Annelida: Sternaspidae) from the northwestern Pacific Ocean. Deep Sea Res Part II Top Stud Oceanogr 154:159–166. https://doi.org/10.1016/j.dsr2.2017.12.016
Labrune C, Grémare A, Amouroux JM, Sardá R, Gil J, Taboada S (2007) Assessment of soft-bottom polychaete assemblages in the Gulf of Lions (NW Mediterranean) based on a mesoscale survey. Estuar Coast Shelf Sci 71(1–2):133–147. https://doi.org/10.1016/j.ecss.2006.07.007
Langeneck J, Barbieri M, Maltagliati F, Castelli A (2019) Molecular phylogeny of Paraonidae (Annelida). Mol Phylogenet Evol 136:1–13. https://doi.org/10.1016/j.ympev.2019.03.023
Lewis PO, Holder MT, Holsinger KE (2005) Polytomies and Bayesian phylogenetic inference. Syst Biol 54(2):241–253
Lim HS, Hong JS (1996) Distribution and growth pattern of Sternaspis scutata (Polychaeta: Sternaspidae) in Chinhae Bay, Korea. Korean J Fish Aquat Sci 29(4):537–545
Lima FD, Berbel-Filho WM, Leite TS, Rosas C, Lima SM (2017) Occurrence of Octopus insularis Leite and Haimovici, 2008 in the tropical northwestern Atlantic and implications of species misidentification to octopus fisheries management. Mar Biodivers 47(3):723–734. https://doi.org/10.1007/s12526-017-0638-y
Lorenti M, Gambi MC, Guglielmo R, Patti FP, Scipione MB, Zupo V, Buia MC (2011) Soft-bottom macrofaunal assemblages in the Gulf of Salerno, Tyrrhenian Sea, Italy, an area affected by the invasion of the seaweed Caulerpa racemosa var. cylindracea. Mar Ecol 32(3):320–334. https://doi.org/10.1016/j.ecss.2006.07.00710.1111/j.1439-0485.2011.00472.x
Maddison WP, Maddison DR (2017) Mesquite: a modular system for evolutionary analysis. Version 3.2 http://mesquiteproject.org. Accessed 14 Aug 2017
Malmgren AJ (1867) Annulata Polychaeta Spetsbergiæ, Grœnlandiæ, Islandiæ et Scandinaviæ. Hactenus Cognita. Ex Officina Frenckelliana, Helsingforslæ pp 127, plate 14
Marenzeller EV (1879) Südjapanische Anneliden. I. (Amphinomea, Aphroditea, Lycoridea, Phyllodocea, Hesionea, Syllidea, Eunicea, Glycerea, Sternaspidea, Chaetopterea, Cirratulea, Amphictenea.). Denkschriften der Kaiserlichen Akademie der Wissenschaften, Mathematisch-naturwissenschaftliche Classe, Wien 41(2):109–154, plates 1–6
Maturana CS, Moreno RA, Labra FA, González-Wevar CA, Rozbaczylo N, Carrasco FD, Poulin E (2011) DNA barcoding of marine polychaetes species of southern Patagonian fjords. Rev Bol Mar Oceanogr 46(1):35–42. https://doi.org/10.4067/S0718-19572011000100005
Medlin L, Elwood HJ, Stickel S, Sogin ML (1988) The characterization of enzymatically amplified eukaryotic 16S-like rRNA-coding regions. Gene 71(2):491–499. https://doi.org/10.1016/0378-1119(88)90066-2
Meißner K, Blank M (2009) Spiophanes norrisi sp. nov. (Polychaeta: Spionidae)—a new species from the NE Pacific coast, separated from the Spiophanes bombyx complex based on both morphological and genetic studies. Zootaxa 2278:1–25. https://doi.org/10.11646/zootaxa.2278.1.1
Méndez N, Yáñez-Rivera B (2015) Distribution and morphometry of the deep-sea sternaspids, Sternaspis maior, Sternaspis uschakovi, and Caulleryaspis fauchaldi (Polychaeta), in Mexican Pacific waters. Bull Mar Sci 91(4):457–467. https://doi.org/10.5343/bms.2015.1046
Meyer A, Bleidorn C, Rouse GW, Hausen H (2008) Morphological and molecular data suggest a cosmopolitan distribution of the polychaete Proscoloplos cygnochaetus Day, 1954 (Annelida, Orbiniidae). Mar Biol 153(5):879–889. https://doi.org/10.1007/s00227-007-0860-4
Minh BQ, Nguyen MAT, von Haeseler A (2013) Ultrafast approximation for phylogenetic bootstrap. Mol Biol Evol 30(5):1188–1195. https://doi.org/10.1093/molbev/mst024
Monro CCA (1930) Polychaete worms, 2nd edn. Discovery Reports, Cambridge, pp 1–222
Morton B (2018) Fake news. Mar Pollut Bull 128:396–397. https://doi.org/10.1016/j.marpolbul.2018.01.042
Nei M, Kumar S (2000) Molecular evolution and phylogenetics: Oxford University Press
Nguyen LT, Schmidt HA, von Haeseler A, Minh BQ (2015) IQ-TREE: a fast and effective stochastic algorithm for estimating maximum likelihood phylogenies. Mol Biol Evol 32:268–274. https://doi.org/10.1093/molbev/msu300
Nygren A (2014) Cryptic polychaete diversity: a review. Zool Scr 43(2):172–183. https://doi.org/10.1111/zsc.12044
Nygren A, Sundberg P (2003) Phylogeny and evolution of reproductive modes in Autolytinae (Syllidae, Annelida). Mol Phylogenet Evol 29(2):235–249. https://doi.org/10.1016/S1055-7903(03)00095-2
Nygren A, Eklöf J, Pleijel F (2009) Arctic-boreal sibling species of Paranaitis (Polychaeta, Phyllodocidae). Mar Biol Res 5(4):315–327. https://doi.org/10.1080/17451000802441301
Osborn KJ, Rouse GW (2011) Phylogenetics of Acrocirridae and Flabelligeridae (Cirratuliformia, Annelida). Zool Scr 40(2):204–219. https://doi.org/10.1111/j.1463-6409.2010.00460.x
Otto AG (1821) Animalium maritimorum nondum editorum genera duo. Verhandlungen der Kaiserlichen Leopoldinisch-Carolinischen Akademie der Naturforscher 10:618–634 plates 50–51
Pallas PS (1766) Miscellanea zoologica. Quibus novae imprimis atque obscurae animalium species describuntur et observationibus iconibusque illustrantur. Petrum van Cleef, Hagí Comitum pp xii + 224, plate 14
Palumbi SR (1996) Nucleic acids II: the polymerase chain reaction. Mol Syst 2(1):205–247
Palumbi S, Martin A, Romano S, McMillan WO, Stice L, Grabowski G (1991) The simple fool’s guide to PCR. University of Hawaii, Department of Zoology and Kewalo Marine Laboratory, Honolulu, HI
Petersen ME (2000) A new genus of Fauveliopsidae (Annelida: Polychaeta), with a review of its species and redescription of some described taxa. Bull Mar Sci 67(1):491–515
Pires-Vanin AMS, Muniz P, De Léo FC (2011) Benthic macrofauna structure in the northeast area of Todos os Santos Bay, Bahia State, Brazil: patterns of spatial and seasonal distribution. Braz J Oceanogr 59:27–42. https://doi.org/10.1590/S1679-87592011000100003
Rambaut A (2012) FigTree v1. 4. Molecular evolution, phylogenetics and epidemiology. University of Edinburgh, Institute of Evolutionary Biology, Edinburgh
Ranzani C (1817) Descrizione di una nuova specie del genere Thalassema. Opuscoli Scientifici, Bologna 1:112–116 plate 114
Ratnasingham S, Hebert PDN (2007) BOLD: The Barcode of Life Data System (http://www.barcodinglife.org). Mol Ecol Notes 7(3):355–364. https://doi.org/10.1111/j.1471-8286.2007.01678.x. Accessed 14 Aug 2017
Ronquist F, Teslenko M, van der Mark P, Ayres DL, Darling A, Höhna S, Larget B, Liang L, Suchard MA, Huelsenbeck JP (2012) MrBayes 3.2: efficient Bayesian phylogenetic inference and model choice across a large model space. Syst Biol 61(3):539–542. https://doi.org/10.1093/sysbio/sys029
Rouse GW, Fauchald K (1997) Cladistics and polychaetes. Zool Scr 26(2). https://doi.org/10.1111/j.1463-6409.1997.tb00412.x
Rouse GW, Pleijel F (2001) Polychaetes. Oxford University Press, Oxford
Rousset V, Pleijel F, Rouse GW, Erséus C, Siddall ME (2007) A molecular phylogeny of annelids. Cladistics 23(1):41–63. https://doi.org/10.1111/j.1096-0031.2006.00128.x
Rozbaczylo N, Moreno R, Díaz-Díaz O, Martínez S (2006) Poliquetos bentónicos submareales de fondos blandos de la región de Aysén, Chile: Clado Terebellida (Annelida, Polychaeta). Ciencia y Tecnología del Mar 29(1)
Salazar-Vallejo SI (2014) Three new polar species of Sternaspis Otto, 1821 (Polychaeta: Sternaspidae). Zootaxa 3861(4):333–344. https://doi.org/10.11646/zootaxa.3861.4.3
Salazar-Vallejo SI (2017) Six new tropical sternaspid species (Annelida, Sternaspidae) with keys to identify genera and species. Zool Stud 56(32):1–27. https://doi.org/10.6620/ZS.2017.56-32
Salazar-Vallejo SI, Buzhinskaja G (2013) Six new deep-water sternaspid species (Annelida, Sternaspidae) from the Pacific Ocean. ZooKeys 348:1–27. https://doi.org/10.3897/zookeys.348.5449
Schneider CA, Rasband WS, Eliceiri KW (2012) NIH Image to ImageJ: 25 years of image analysis. Nat Methods 9(7):671–675
Schulze A (2006) Phylogeny and genetic diversity of Palolo worms (Palola, Eunicidae) from the tropical North Pacific and the Caribbean. Biol Bull 210(1):25–37. https://doi.org/10.2307/4134534
Sendall K (2006) Review and revision of the genus Sternaspis (Polychaeta: Sternaspidae) using cladistics on morphological characters. Master of Science Thesis, University of Victoria, Canada. Retrieved from http://dspace.library.uvic.ca:8080/handle/1828/2189. Accessed 21 April 2017
Sendall K, Salazar-Vallejo SI (2013) Revision of Sternaspis Otto, 1821 (Polychaeta, Sternaspidae). ZooKeys 286:1–74. https://doi.org/10.3897/zookeys.286.4438
Shelley R, Widdicombe S, Woodward M, Stevens T, McNeill CL, Kendall MA (2008) An investigation of the impacts on biodiversity and ecosystem functioning of soft sediments by the non-native polychaete Sternaspis scutata (Polychaeta: Sternaspidae). J Exp Mar Biol Ecol 366(1–2):146–150. https://doi.org/10.1016/j.jembe.2008.07.018
Shilling FM, Manahan DT (1994) Energy metabolism and amino acid transport during early development of Antarctic and temperate echinoderms. Biol Bull 187(3):398–407. https://doi.org/10.2307/1542296
Sjölin E, Erséus C, Källersjö M (2005) Phylogeny of Tubificidae (Annelida, Clitellata) based on mitochondrial and nuclear sequence data. Mol Phylogenet Evol 35(2):431–441. https://doi.org/10.1016/j.ympev.2004.12.018
Sluiter CP (1882) Ueber einen indischen Sternaspis und seine Verwandschaft zu den Echiuren. Natuurkundig tijdschrift voor Nederlandsch Indië uitgegeven door de Natuurkundige Vereeniging. Batavia, 41:235–287, plates 1–3
Stimpson W (1854) Synopsis of the marine Invertebrata of Grand Manan: or the region about the mouth of the Bay of Fundy, New Brunswick. Smithsonian Contributions to Knowledge 6:1–66 plates 1–3
Stimpson W (1864) Descriptions of new species of marine Invertebrata from Puget Sound, collected by the naturalists of the North-West Boundary Commission, A.H. Campbell, Esq., Commissioner. Proc Acad Nat Sci Philadelphia 153–161
Struck TH, Schult N, Kusen T, Hickman E, Bleidorn C, McHugh D, Halanych KM (2007) Annelid phylogeny and the status of Sipuncula and Echiura. BMC Evol Biol 7(1):57. https://doi.org/10.1186/1471-2148-7-57
Struck TH, Nesnidal MP, Purschke G, Halanych KM (2008) Detecting possibly saturated positions in 18S and 28S sequences and their influence on phylogenetic reconstruction of Annelida (Lophotrochozoa). Mol Phylogenet Evol 48(2):628–645. https://doi.org/10.1016/j.ympev.2008.05.015
Thorpe JP, Solé-Cava AM (1994) The use of allozyme electrophoresis in invertebrate systematics. Zool Scr 23(1):3–18. https://doi.org/10.1111/j.1463-6409.1994.tb00368.x
Tomioka S, Kondoh T, Sato-Okoshi W, Ito K, Kakui K, Kajihara H (2016) Cosmopolitan or cryptic species? A case study of Capitella teleta (Annelida: Capitellidae). Zool Sci 33(5):545–555. https://doi.org/10.2108/zs160059
Tomioka S, Kakui K, Kajihara H (2018) Molecular phylogeny of the family Capitellidae (Annelida). Zool Sci 35(5):436–445. https://doi.org/10.2108/zs180009
Townsend M, Worsfold TM, Smith PRJ, Martina LJ, McNeill CL, Kendall MA (2006) Occurrence of Sternaspis scutata (Polychaeta: Sternaspidae) in the English Channel. Cah Biol Mar 47(3):281
Treadwell AL (1914) Polychaetous annelids of the Pacific coast: in the collections of the zoological museum of the University of California. Univ Calif Publ Zool 13:175–234
Van Oppen MJH, Klerk H, Olsen JL, Stam WT (1996) Hidden diversity in marine algae: some examples of genetic variation below the species level. J Mar Biol Assoc UK 76(1):239–242. https://doi.org/10.1017/S0025315400029192
Vejdovsky F (1882) Untersuchungen über die Anatomie, Physiologie und Entwicklung von Sternaspis. Denkschriften der Mathematisch-Naturwissenschaftlichen Classe der kaiserlichen Akademie der Wissenschaften 43:33–90 plates l–10
Wang J, Lin X, Wang H (2006) Ecological characteristics of dominant polychaete species from the Jiaozhou Bay. Dong wu xue bao. Acta Zool Sin 52(1):63–69
Weigert A, Helm C, Meyer M, Nickel B, Arendt D, Hausdorf B, Santos SR, Halanych KM, Purschke G, Bleidorn C, Struck TH (2014) Illuminating the base of the annelid tree using transcriptomics. Mol Biol Evol 31(6):1391–1401. https://doi.org/10.1093/molbev/msu080
Westheide W, Schmidt H (2003) Cosmopolitan versus cryptic meiofaunal polychaete species: an approach to a molecular taxonomy. Helgol Mar Res 57(1):1–6. https://doi.org/10.1007/s10152-002-0114-2
Westheide W, Haß-Cordes E, Krabusch M, Müller M (2003) Ctenodrilus serratus (Polychaeta: Ctenodrilidae) is a truly amphi-Atlantic meiofauna species—evidence from molecular data. Mar Biol 142(4):637–642. https://doi.org/10.1007/s00227-002-0960-0
Wieczorek J, Bloom D, Guralnick R, Blum S, Döring M, Giovanni R, Robertson T, Vieglais D (2012) Darwin Core: an evolving community-developed biodiversity data standard. PLoS One 7(1):e29715. https://doi.org/10.1371/journal.pone.0029715
Wiklund H, Glover AG, Dahlgren TG (2009a) Three new species of Ophryotrocha (Annelida: Dorvilleidae) from a whale-fall in the North-East Atlantic. Zootaxa 2228:43–56
Wiklund H, Glover AG, Johannessen PJ, Dahlgren TG (2009b) Cryptic speciation at organic-rich marine habitats: a new bacteriovore annelid from whale-fall and fish farms in the North-East Atlantic. Zool J Linnean Soc 155(4):774–785
Wu X, Xu K (2017) Diversity of Sternaspidae (Annelida: Terebellida) in the South China Sea, with descriptions of four new species. Zootaxa 4244(3):403–415. https://doi.org/10.11646/zootaxa.4244.3.8
Wu X, Salazar-Vallejo SI, Xu K (2015) Two new species of Sternaspis Otto, 1821 (Polychaeta: Sternaspidae) from China seas. Zootaxa 4052(3):373–382. https://doi.org/10.11646/zootaxa.4052.3.7
Yoshino K, Hamada T, Yamamoto K, Hayami Y, Yamaguchi S, Ohgushi K (2010) Effects of hypoxia and organic enrichment on estuarine macrofauna in the inner part of Ariake Bay. Hydrobiologia 652(1):23–38. https://doi.org/10.1007/s10750-010-0313-9
Yoshino K, Katano T, Ito Y, Hamada T, Fujii N, Hayami Y (2014) Community assembly by limited recruitment in a hypoxia-stressed soft bottom: a case study of macrobenthos in Ariake Bay. Plankton Benthos Res 9(1):57–66. https://doi.org/10.3800/pbr.9.57
Yoshino K, Nagayoshi M, Sato M, Katano T, Ito Y, Fujii N, Hamada T, Hayami Y (2016) Life history of Sternaspis costata (Sternaspidae: Polychaeta) in Ariake Bay, Japan. J Mar Biol Assoc UK 96(3):647–655. https://doi.org/10.1017/S0025315415000880
Zhadan AE, Tzetlin AB, Salazar-Vallejo SI (2017) Sternaspidae (Annelida, Sedentaria) from Vietnam with description of three new species and clarification of some morphological features. Zootaxa 4226(1):75–92. https://doi.org/10.11646/zootaxa.4226.1.3
Zhang H, Johnson SB, Flores VR, Vrijenhoek RC (2015) Intergradation between discrete lineages of Tevnia jerichonana, a deep-sea hydrothermal vent tubeworm. Deep Sea Res Part II Top Stud Oceanogr 121:53–61. https://doi.org/10.1016/j.dsr2.2015.04.028
Acknowledgements
The authors would like to thank the British Antarctic Survey and all crew and scientists who participated in the three research cruises that provided Antarctic material, in particular thanking Huw Griffiths for supplying material from the SO-AntEco (JR15005) cruise, Katrin Linse for material from the BIOPEARL I (JR144) cruise and Peter Enderlein for material from the Benthic Biology of the Cold Hole (JR308) cruise. The authors also wish to thank the CSIRO Marine National Facility (MNF) for its support in the form of sea time on RV Investigator, support personnel, scientific equipment and data management for voyage IN2017_V03. Thanks also to Muriel Rabone for her great help with database and molecular collections management and to Emma Sherlock for her assistance with collections material and specimen curation. A further thank you to Charlotte Seid for her help in coordinating specimen loans from the Scripps Institution of Oceanography Benthic Invertebrates Collections. RD would like to thank Kuidong Xu and Sergio Salazar-Vallejo for their advice and correspondence. GK would like to thank Koji Seike for providing a specimen; Hajime Itoh, Mizuki Ohta, Tajima Nagisa and the crew and researchers R/V Shinsei Maru (KS-17-1 and KS-18-2) for their support in samplings; and Toshi Nagata and Hiroshi Ogawa for inviting GK to the research cruises. The cruises KS-17-1 and KS-18-2 were conducted as parts of a project supported by the Tohoku Ecosystem–Associated Marine Sciences (TEAMS) grant from the Ministry of Education, Culture, Sports, Science, and Technology (MEXT). KY would like to thank Kei Kimura for supervising DNA analyses and Ryo Orita for their collection of sternaspid samples.
Funding
This work was primarily conducted at the Natural History Museum London as part of the MSc course in Taxonomy and Biodiversity at Imperial College London. Work of XW was funded by the National Natural Science Foundation of China (No. 31601842).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
No animal testing was performed during this study.
Sampling and field studies
All necessary permits for sampling and observational field studies have been obtained by the authors from the competent authorities and are mentioned in the acknowledgements, if applicable. The study is compliant with CBD and Nagoya protocols.
Data availability
Genetic sequence data generated in this study were deposited in GenBank with Accession Numbers MK809970–MK810090. All other data generated or analyzed during this study are included in this published article and its supplementary information files.
Additional information
Communicated by P. Lana
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Video of live Sternaspis affinis Stimpson 1864 (specimen voucher number SIO-BIC A5918), highlighting the extrusion and intrusion of the introvert. Video credit: Greg Rouse. (MP4 23,293 kb)
Online Resource 2
Table of all new sternaspid material examined, presented in a Darwin Core archive format. Table includes collection, taxonomic, and occurrence data. (XLSX 64 kb)
Online Resource 3
Table of taxa with previously published sequence data used in phylogenetic analyses, with list of associated NCBI GenBank accession numbers. (XLSX 45 kb)
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Drennan, R., Wiklund, H., Rouse, G.W. et al. Taxonomy and phylogeny of mud owls (Annelida: Sternaspidae), including a new synonymy and new records from the Southern Ocean, North East Atlantic Ocean and Pacific Ocean: challenges in morphological delimitation. Mar. Biodivers. 49, 2659–2697 (2019). https://doi.org/10.1007/s12526-019-00998-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12526-019-00998-0