Abstract
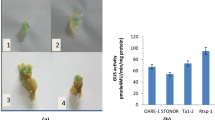
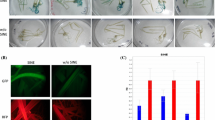
Retrotransposons are the most abundant mobile elements in the plant genome and seem to play an important role in genome reorganization induced by environmental challenges. Their success in this function depends on the ability of their promoters to regulate plant adaptation to biotic and abiotic stresses. In this study, the promoter region of FaRE1 was amplified in the strawberry genome, and promoter::GUS fusion was constructed. We produced transgenic strawberry plants carrying FaRE1 promoter::GUS-fusion genes, and monitored GUS reporter activity. Histochemical and fluorimetric GUS analysis these plants showed the characteristics of the FaRE1 promoter were activated by either hormones treatments with ABA, NAA, and 2,4-D or cold stress. In addition, we found the GUS reporter was activated in the leaves of transgenic strawberry plants using 5-azaC. These results suggest that the promoter of FaRE1 may act as different signal transduction pathways, allowing FaRE1 retrotransposon to be activated in response to multiples challenges.





Similar content being viewed by others
References
Anderson JP, Badurzaaufari E, Schenk PM, Manners JM, Desmond OJ, Ehlert C, Maclean DJ, Ebert PR, Kazan K (2004) Antagonistic interaction between abscisic acid and jasmonate-ethylene signaling pathways modulates defense gene expression and disease resistance in Arabidopsis. Plant Cell 16:3460–3479
Barkan A, Martienssen RA (1991) Inactivation of maize transposon Mu suppresses a mutant phenotype by activating an outward-reading promoter near the end of Mu1. Proc Natl Acad Sci USA 88:3502–3506
Beguiristain T, Grandbastien MA, Puigdomenech P, Grandbastien MA, Van Sluys MA (2001) retrolycl subfamilies by different U3 LTR regulatory regions in the Lycopersicon genus. Mol Gen Genet 266:35–41
Bradford MM (1976) A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem 72:248–254
Casacuberta JM, Santiago N (2003) Plant LTR-retrotransposons and MITEs: control of transposition and impact on the evolution of plant genes and genomes. Gene 311:1–11
Cheng C, Daigen M, Hirochika H (2006) Epigenetic regulation of the rice retrotransposon Tos17. Mol Genet Genomics 276(4):378–390
Dai H, Zhang Z, Guo X (2007) Adventitious bus regeneration from leaf and cotyledon explants of Chinese hawthorn (Crataegus pinnatifida Bge. Var. major N.E.Br.). In Vitro Cell Drv Biol Plant 43:2–8
Feschotte C, Jiang N, Wessler RS (2002) Plant retrotransposable elements: where genetics meets genomics. Nat Rev Genet 3:329–341
Han JS, Szak ST, Boeke JD (2004) Transcriptional disruption by the L1 retrotransposon and implications for mammalian transcriptomes. Nature 429:268–274
He P, Ma Y, Zhao G, Dai H, Li H, Chang L, Zhang Z (2010) FaRE1: a transcriptionally active Ty1-copia retrotransposon in strawberry. J Plant Res 123:707–714
Hirochika H, Okamoto H, Kakutani T (2000) Silencing of retrotransposons in Arabidopsis and reactivation by the ddm1 mutation. Plant Cell 12:537–369
Hung SH, Yu CW, Lin CH (2005) Hydrogen peroxide functions as a stress signal in plants. Bot Bull Acad Sin 41:1–10
Jefferson RA, Kavanagh TA, Bevan MW (1987) GUS fusions: β-glucuronidase as a sensitive and versatile gene fusion marker in higher plants. EMBO J 6:3901–3907
Kato M, Miura A, Bender J, Jacobsen SE, Kakutani T (2003) Role of CG and non-CG methylation in immobilization of transposons in Arabidopsis. Curr Boil 13:421–426
Kidwell MG, Lisch DR (2000) Transposable elements and host genome evolution. Trends Ecol Evol 15:95–99
Konishi M, Yanagiaswa S (2005) Signaling crosstalk between ethylene and other molecules. Plant Biotechnol 22:401–407
Kosugi S, Ohashi Y, Nakajima K, Arai Y (1990) An improved assay for β-glucuronidase in transformed cells: methanol almost completely suppresses a putative endogenous β-glucuronidase activity. Plant Sci 70:133–140
Kumar A, Bennetzen JL (1999) Plant retrotransposons. Annu Rev Genet 33:479–532
Lorenzo O, Chico JM, Sanchez-Sreeano JJ, Solano R (2004) Jasmonate-insensitives encodes a MYC transcription factor essential to discriminate between different jasmonate-regulated defense responses in Arabidopsis. Plant Cell 16:1938–1950
Ma Y, Sun H, Zhao G, Dai H, Gao X, Li H, Zhang Z (2008) Isolation and characterization of genomic retrotransposon sequences from octoploid strawberry (Fragaria × ananassa Duch.). Plant Cell Rep 27:499–507
Martienssen RA, Colot DV (2001) DNA methylation and epigenetic inheritance in plants and filamentous fungi. Science 293:1070–1074
McCarty EM, Liu J, Lizhi G, McDonald JF (2002) Long terminal repeat retrotransposons of Oryza sativa. Genome Biol 3: research 0053.1–0053.11
McInerney JM, Nawrocki JR, Lowrey CH (2000) Long-term silencing of retroviral vectors is resistant to reversal by trichostatin A and 5-azacytidine. Gene Ther 7:653–663
Melayah D, Bonnivard E, Chalhoub B, Audeon C, Grandbastien MA (2001) The mobility of the tobacco Tnt1 retrotransposon correlates with its transcriptional activation by fungal factors. Plant J 28:159–168
Meyers BC, Tingery SV, Organte MM (2001) Abundance, distribution, and transcriptional activity of repetitive elements in the maize genome. Genome Res 11:1660–1676
Miura A, Yonebayashi S, Watanabe K, Toyama T, Shimada H, Kakutain T (2001) Mobilization of trasposons by a mutation abolishing full DNA methylation in Arabidopsis. Nature 411:212–214
Nehra NS, Karta KK, Stushnlff C (1991) Nuclear DNA content and isozyme variation in relation to morphogenic potential of strawberry (Fragaria × ananassa) callus cultures. Can J Bot 69:239–244
Okamoto H, Hirochika H (2001) Silencing of transposable elements in plants. Trends Plant Sci 6:527–534
Prakash AP, Kumar PP (1997) Inhibition of shoot induction by 5-azacytidine and 5-aza-2′-deoxycytidine in Petunia involves DNA hypomethylation. Plant Cell Rep 16:719–724
Rabinowicz PD, Palmer LE, May BP, Hemann MT, Iowe SW, Mccombie WR, Martienssen RA (2003) Genes and transposons are differentially methylated in plants, but not in mammals. Genome Res 13:2658–2664
Suoniemi A, Anamthawat-Jonsson K, Arma T, Schulman AH (1996) Retrotransposon BARE-1 is major, dispersed component of the barley (Horderm vulgare L) genome. Plant Mol Biol 30:1321–1329
Tahara M, Aoki T, Suzuka S, Yamashita H, Tanaka M, Matsunaga S, Kokumai S (2004) Isolation of an active element from a high-copy-number family of rerotransposons in the sweet potato genome. Mol Gen Genomics 272:116–127
Takeda S, Sugimoto K, Otsuki H, Hirochika H (1999) A 13-pb cis-regulatory element in the LTR promoter of the tobacco retrotransposon Tto1 is involved in responsiveness to tissue culture, wounding, methyl jasmonate and fungal elicitors. Plant J 18:383–393
Tapoa G, Verdugo I, Yánez M, Ahumada I, Theoduloz C, Cordero C, Poblete F, González E, Ruiz-Lara S (2005) Involvement of ethylene in stress-induced expression of the TLC1.1 retrotransposon from Lycopersicon chilenes Dun. Plant Physiol 138(4):2075–2086
Terakami S, Matsuta N, Yamamoto T, Sugaya S, Genna H, Soejima J (2007) Agrobacterium-mediated transformation of the dwarf pomegranate (Punica granatum L. var. nana). Plant Cell Rep 26:1243–1251
Vance V, Vaucheret H (2002) RNA silencing in plants-defense and counter-defense. Science 292:2277–2280
Xiao B, Huang Y, Tang N, Xiong L (2007) Over-expression of a LEA gene in rice improves drought resistance under the field conditions. Theor Appl Genet 115(1):35–46
Zhang H, Huang Z, Xie B, Chen Q, Tian X, Zhang X, Zhang H, Lu X, Huang D, Huang R (2004) The ethylene-, jasmonate-, abscisic acid- and NaCl-responsive tomato transcription factor JERF1 modulates expression of GCC box-containing genes and salt tolerance in tobacco. Planta 220(2):262–270
Acknowledgements
This work was supported by the National Natural Science Foundation of China (No. 30871689) and Special Fund for Agro-scientific Research in the Public Interest (201003064–3).
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
He, P., Ma, Y., Dai, H. et al. Characterization of the Hormone and Stress-Induced Expression of FaRE1 Retrotransposon Promoter in Strawberry. J. Plant Biol. 55, 1–7 (2012). https://doi.org/10.1007/s12374-011-9180-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12374-011-9180-9