Abstract
Introduction
Pyrotinib is a newly developed tyrosine kinase inhibitor whose in vivo clearance relies heavily on cytochrome P450 3A4 (CYP3A4) activity. Clinical trials are ongoing to explore the effects of coadministration with CYP3A4 perpetrators on pyrotinib exposure. The present study aims to utilize physiologically based pharmacokinetic (PBPK) modeling to predict CYP3A4-based drug interactions of pyrotinib.
Methods
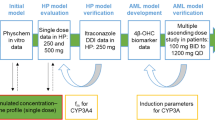
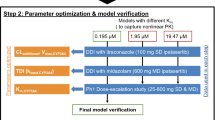
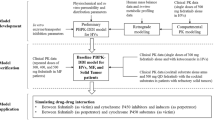
Pyrotinib PBPK model was developed in the PK-Sim® multicompartmental physiology structure. Physiochemical parameters were obtained from the literature, and clearance-related parameters were optimized by fitting clinical single-dose pharmacokinetic data. Pharmacokinetic parameters from the model output were compared with the observed data to validate the model predictive performance. Using validated CYP3A4 perpetrator models, we conducted PBPK simulations for drug interactions in a virtual population to explore the impacts of comedication with these perpetrators.
Results
The PBPK model accurately describes pyrotinib single- and multi-dose pharmacokinetics. The model also predicts dramatic exposure change of pyrotinib in the presence of itraconazole and rifampicin, though the impact of rifampicin is somewhat underestimated. According to model predictions, coadministration with typical potent or moderate CYP3A4 perpetrators increases pyrotinib concentration by over sixfold, extinguishing the possibility of dose adjustment for pyrotinib. A weak CYP3A4 inhibitor has minimal influence on pyrotinib pharmacokinetics.
Conclusion
PBPK modeling provides valuable information to avoid irrational medication when receiving pyrotinib chemotherapy.



Similar content being viewed by others
References
Ferlay J, Colombet M, Soerjomataram I, Parkin DM, Pineros M, Znaor A, et al. Cancer statistics for the year 2020: an overview. Int J Cancer. 2021;149:778–89.
Cao W, Chen HD, Yu YW, Li N, Chen WQ. Changing profiles of cancer burden worldwide and in China: a secondary analysis of the global cancer statistics 2020. Chin Med J (Engl). 2021;134(7):783–91.
Oh DY, Bang YJ. HER2-targeted therapies—a role beyond breast cancer. Nat Rev Clin Oncol. 2020;17(1):33–48.
Blair HA. Pyrotinib: first global approval. Drugs. 2018;78(16):1751–5.
Ma F, Li Q, Chen S, Zhu W, Fan Y, Wang J, et al. Phase I study and biomarker analysis of pyrotinib, a novel irreversible Pan-ErbB receptor tyrosine kinase inhibitor, in patients with human epidermal growth factor receptor 2-positive metastatic breast cancer. J Clin Oncol. 2017;35(27):3105–12.
Zhu Y, Li L, Zhang G, Wan H, Yang C, Diao X, et al. Metabolic characterization of pyrotinib in humans by ultra-performance liquid chromatography/quadrupole time-of-flight mass spectrometry. J Chromatogr B Analyt Technol Biomed Life Sci. 2016;1033–1034:117–27.
Meng J, Liu XY, Ma S, Zhang H, Yu SD, Zhang YF, et al. Metabolism and disposition of pyrotinib in healthy male volunteers: covalent binding with human plasma protein. Acta Pharmacol Sin. 2019;40(7):980–8.
Liu Y, Zhang Q, Lu C, Hu W. Multiple administrations of itraconazole increase plasma exposure to pyrotinib in chinese healthy adults. Drug Des Dev Ther. 2021;15:2485–93.
Cai MM, Dou T, Tang L, Sun QY, Zhai ZH, Wang HP, et al. Effects of rifampicin on antineoplastic drug pyrotinib maleate pharmacokinetics in healthy subjects. Invest New Drugs. 2022;40(4):756–61.
Zhang X, Yang Y, Grimstein M, Fan J, Grillo JA, Huang SM, et al. Application of PBPK modeling and simulation for regulatory decision making and its impact on US prescribing information: an update on the 2018–2019 submissions to the US FDA’s office of clinical pharmacology. J Clin Pharmacol. 2020;60(Suppl 1):S160–78.
Wendl T, Frechen S, Gerisch M, Heinig R, Eissing T. Physiologically-based pharmacokinetic modeling to predict CYP3A4-mediated drug-drug interactions of finerenone. CPT Pharmacometrics Syst Pharmacol. 2022;11(2):199–211.
Pilla Reddy V, Walker M, Sharma P, Ballard P, Vishwanathan K. Development, verification, and prediction of osimertinib drug-drug interactions using PBPK modeling approach to inform drug label. CPT Pharmacometrics Syst Pharmacol. 2018;7(5):321–30.
Hanke N, Frechen S, Moj D, Britz H, Eissing T, Wendl T, et al. PBPK models for CYP3A4 and P-gp DDI prediction: a modeling network of rifampicin, itraconazole, clarithromycin, midazolam, alfentanil, and digoxin. CPT Pharmacometrics Syst Pharmacol. 2018;7(10):647–59.
Kanacher T, Lindauer A, Mezzalana E, Michon I, Veau C, Mantilla JDG, et al. A physiologically-based pharmacokinetic (PBPK) model network for the prediction of CYP1A2 and CYP2C19 drug-drug-gene interactions with fluvoxamine, omeprazole, S-mephenytoin, moclobemide, tizanidine, mexiletine, ethinylestradiol, and caffeine. Pharmaceutics. 2020;12(12):1191.
Frechen S, Solodenko J, Wendl T, Dallmann A, Ince I, Lehr T, et al. A generic framework for the physiologically-based pharmacokinetic platform qualification of PK-Sim and its application to predicting cytochrome P450 3A4-mediated drug-drug interactions. CPT Pharmacometrics Syst Pharmacol. 2021;10(6):633–44.
Nishimura M, Yaguti H, Yoshitsugu H, Naito S, Satoh T. Tissue distribution of mRNA expression of human cytochrome P450 isoforms assessed by high-sensitivity real-time reverse transcription PCR. Yakugaku Zasshi. 2003;123(5):369–75.
Rodgers T, Rowland M. Physiologically based pharmacokinetic modelling 2: predicting the tissue distribution of acids, very weak bases, neutrals and zwitterions. J Pharm Sci. 2006;95(6):1238–57.
Britz H, Hanke N, Volz AK, Spigset O, Schwab M, Eissing T, et al. Physiologically-based pharmacokinetic models for CYP1A2 drug-drug interaction prediction: a modeling network of fluvoxamine, theophylline, caffeine, rifampicin, and midazolam. CPT Pharmacometrics Syst Pharmacol. 2019;8(5):296–307.
FDA U. Drug Interactions & Labeling last update 2020. https://www.fda.gov/drugs/development-resources/drug-interactions-labeling.
PRESCRIBING INFORMATION for INLYTA® (axitinib) tablets 2012. https://www.accessdata.fda.gov/drugsatfda_docs/label/2012/202324lbl.pdf.
Li X, Wang Y, Zhu X, Zheng L. Clinical study of drug-drug interaction between omeprazole and pyrotinib after meal. Br J Clin Pharmacol. 2022;88(5):2349–58.
Acknowledgements
The authors thank Jiangsu Hengrui Pharmaceuticals Co., Ltd., for providing clinical data of pyrotinib and Dr. Bin Ye from Shenzhen Institute of Advanced Technology for providing graphing support.
Funding
This study is funded by the Research Fund of Anhui Institute of Translational Medicine (project no. 2022zhyx-B10) and the Natural Science Foundation of Anhui Province (no. 2108085QH383). The Rapid Service Fee was funded by the authors.
Author Contributions
Liang Ni, Liang Zheng, Yueyue Liu, Wenwen Xu, Yingjie Zhao, Ling Wang, Qian Zhang, Wei Hu, and Xijing Chen contributed to the study concept and design, modeling, data analysis, drafting of the manuscript, and revision of the manuscript.
Disclosures
Liang Ni, Liang Zheng, Yueyue Liu, Wenwen Xu, Yingjie Zhao, Ling Wang, Qian Zhang, Wei Hu, and Xijing Chen have no conflicts of interest to declare in this work.
Compliance with Ethics Guidelines
This article is based on previously conducted studies and does not contain any new studies with human participants or animals performed by any of the authors.
Data Availability
The model file will be publicly available in the Open Systems Pharmacology Community (https://github.com/Open-Systems-Pharmacology).
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Ni, L., Zheng, L., Liu, Y. et al. Physiologically Based Pharmacokinetic Modeling to Simulate CYP3A4-Mediated Drug-Drug Interactions for Pyrotinib. Adv Ther 40, 4310–4320 (2023). https://doi.org/10.1007/s12325-023-02602-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12325-023-02602-1